Epigenetics Theory
1/41
Earn XP
Description and Tags
Quiz 2
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
42 Terms
How are all the different cells in your body produced from a single genome?
Zygote → 200+ types of cells in human body
Cells vary in phenotype due to unique gene (and thus protein) expression profiles
How are all the different cells in your body produced from a single genome?
Gene expression is regulated by both extrinsic and intrinsic cues
Genes may be turned ‘on’ or ‘off’, or transcript levels can be increased/decreased
extrinsic ←> intrinsic (bidirectional relationship)
gene expression is regulated by
both extrinsic and intrinsic cues
Genes may be turned ‘on’ or ‘off’, or transcript levels can be increased/decreased
extrinsic
from the cell’s environment
other cells
organism's environment
• E.g., growth factors trigger intracellular signaling cascades → changes in transcription
intrinsic
DNA modification
Cell’s own machinery chemically modifies DNA in a way that affects gene expression
What is epigenetics?
Study of changes in gene expression (and thus phenotype) that occur due to chemical modifications of the genome, rather than a change to the DNA sequence itself
Cells have the mechanisms to copy epigenetic modifications during divisions
These modifications are however, reversible
What is epigenetics? - human twins
Lack of concordance in patterns of disease
30 – 60% concordance rate for vast array of diseases such as: schizophrenia, AD, MS, Crohn disease, asthma, diabetes, prostrate cancer
• Only 10% concordance rate for breast cancer
What is epigenetics? - other
Phenotypic variations can arise in absence of changes to the nucleotide sequence of genes
Experience and the environment can promote or inhibit gene expression
A lifetime of experience can alter behaviour, via the epigenome
What is gene expression?
Process by which cells convert the information encoded in our DNA into a functional product
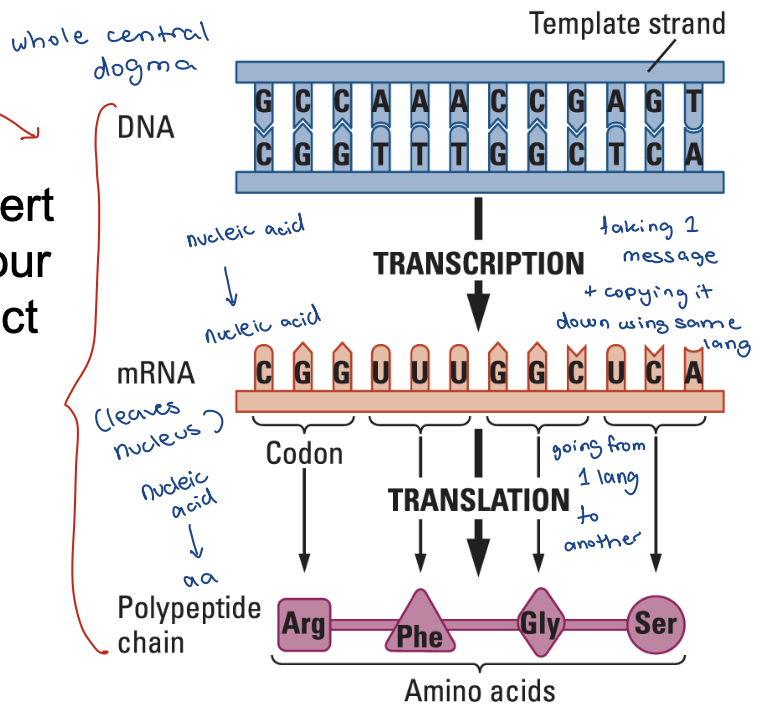
Gene structure and expression
Genes are made of DNA, and most genes code for protein products
Consist of coding regions and noncoding regions
Transcription initiated at the promoter region
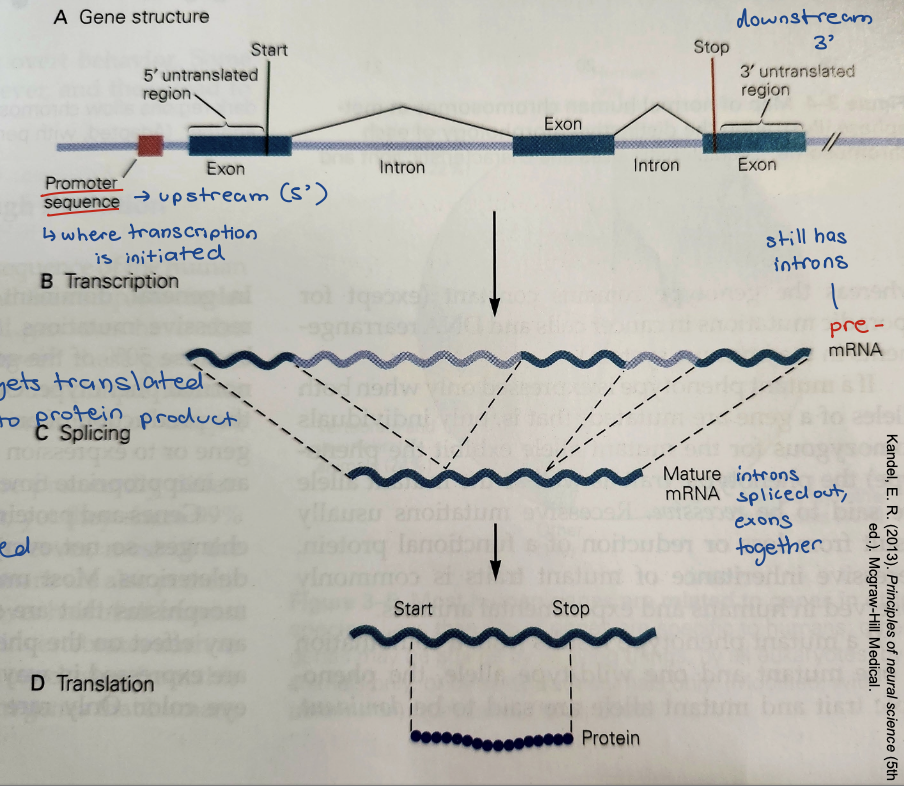
DNA consists of two
polynucleotide chains
template strand
coding strand
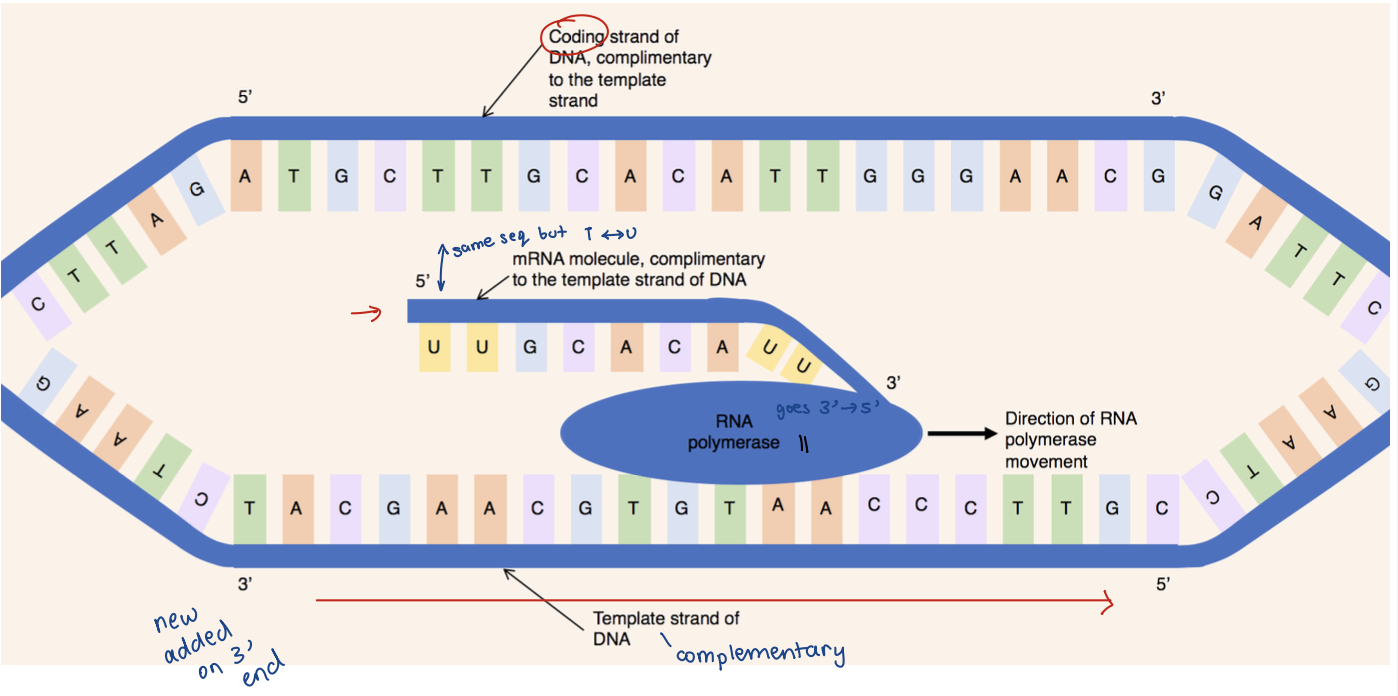
Template (-) strand (aka antisense strand)
Read by RNA polymerase (RNA pol II)
aka noncoding strand
Coding (+) strand (aka sense strand)
Sequence of mRNA is identical to this strand
aka complimentary
Control of transcription in eukaryocytes
Each cell in the body contains the DNA for every gene, but only expresses a subset of genes as RNAs
The brain expresses more genes than any other organ
Diverse populations of neurons have different gene expression profiles
‘Upstream’ regulatory regions
Unique complement of transcription factors interacts with promoters and enhancer
‘upstream’ regulatory regions
Promoters, enhancers, and silencers ensure that the right gene is expressed in the right cells, at the right time
Unique complement of transcription factors interacts with
promoters and enhancer and silencers
Control of transcription in eukaryocytes
Proximal regulatory region: Promoter
Distal regulatory region: enhancers and silencers
Promoter consists of a ‘core’ rate promoter region and ‘promoter proximal’ regions
Core promoter contains the TATA box
Distal regulatory regions are specific to particular genes and particular tissues
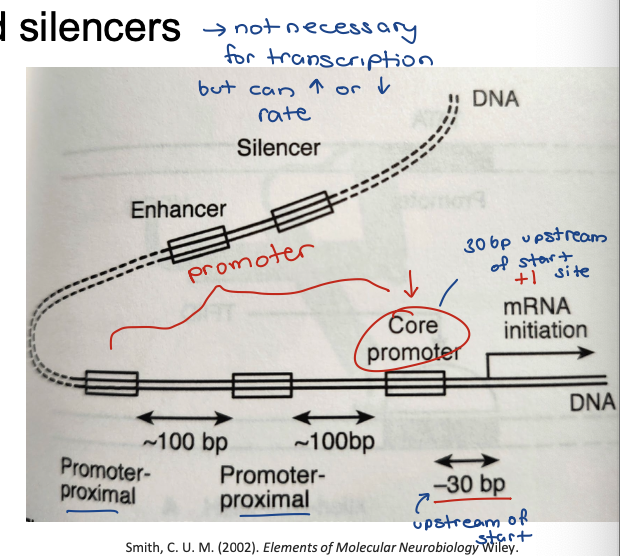
Transcription initation in eukaryocytes
Regulatory regions are recognized and bound by protein complexes
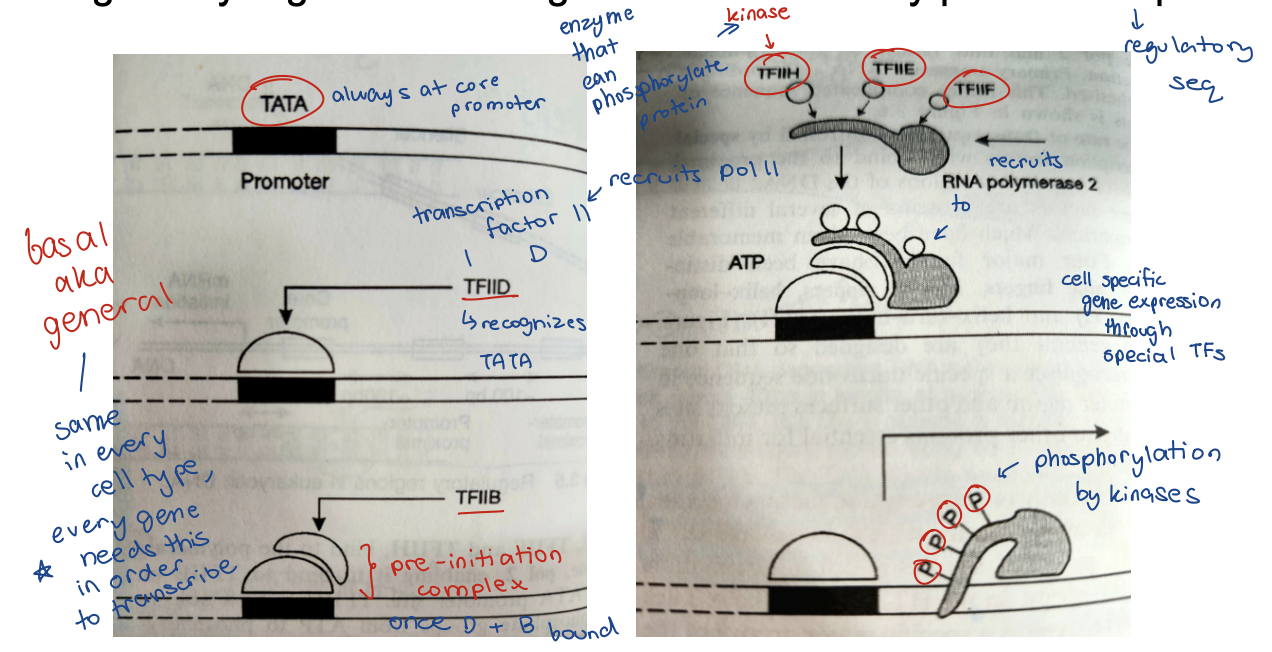
How can special transcription factors attached to distal regulatory regions affect the core promoter and its complex of basal transcription factors?
Special transcription factors called activators and
repressors, bind thousands of bp upstream of transcription
*control expression of specific genes
Activators – help basal TFs and/or RNA polymerase bind to core promoter
Repressors – may impede basal TFs or RNA polymerase such that they cannot bind to core promoter to initiate transcription
Ex) CRE binding proteins (CREBs) bind to the CRE sequence
Special transcription factors often recognise short lengths of palindromic DNA
cAMP response element (CRE site) = cAMP responsive element
• 5’TGACGTCA3’ → ————AGT3’ usually palindrome sequence
Phosphorylation of CREB allows CREB binding protein (CBP) to attach to CREB
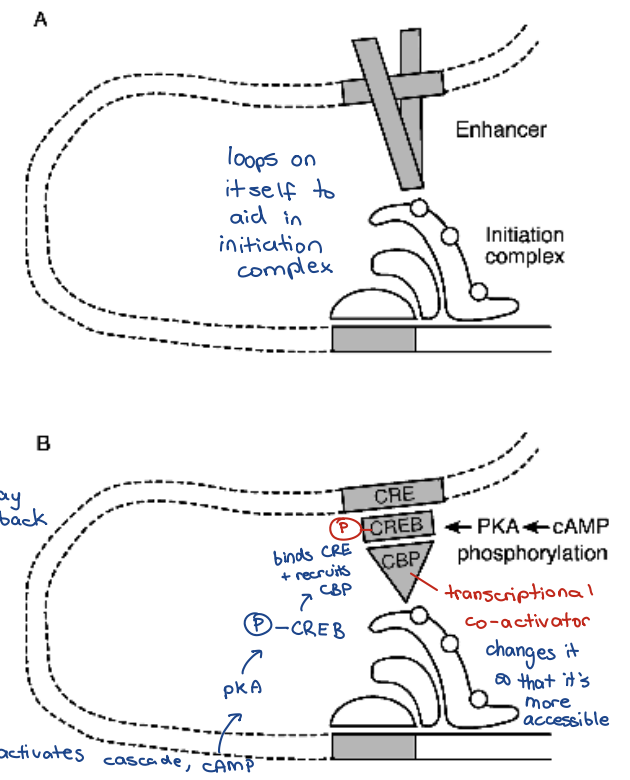
activator/repressor or enhancer/silencer
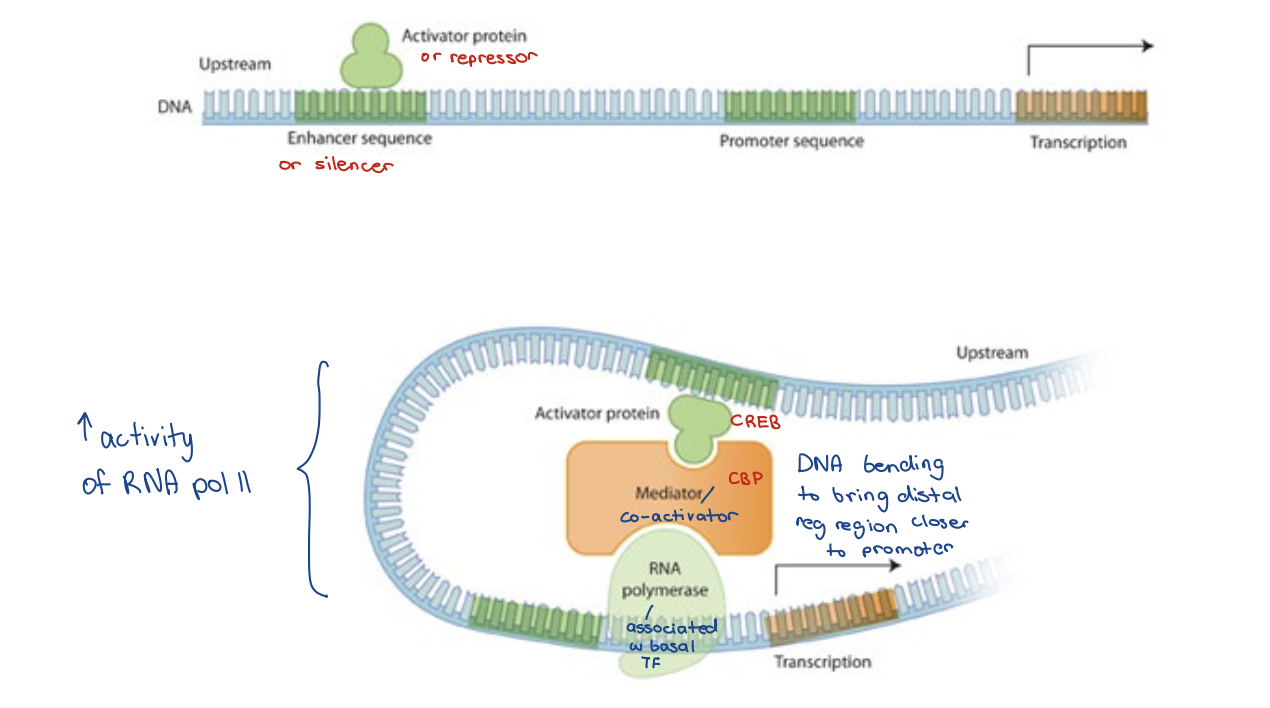
Combinatorial regulation
Genes may be controlled by several different transcription factors
Certain combinations of activators may need to be present (in the absence of certain repressors) to induce gene expression
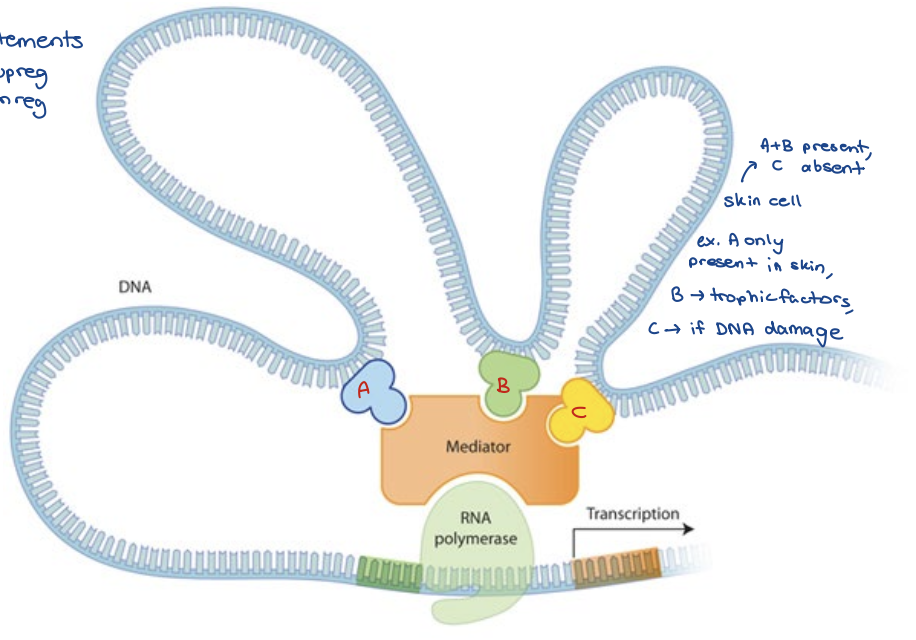
Cell-type specific gene expression profiles
Different cell types express characteristic set of transcription factors
Epigenetic modifications also determine whether a gene is turned ‘on’ or ‘off’
Important during development/cell differentiation
• Genome-wide patterns of epigenetic modifications are established in early development
Over an organism’s lifetime, the environment influences the epigenome
How is DNA organized within the cell?
60 trillion cells, 3.2 billion base pairs (haploid genome); ~20 000 protein coding genes (Amaral et al., 2023)
• ~2 m of DNA!
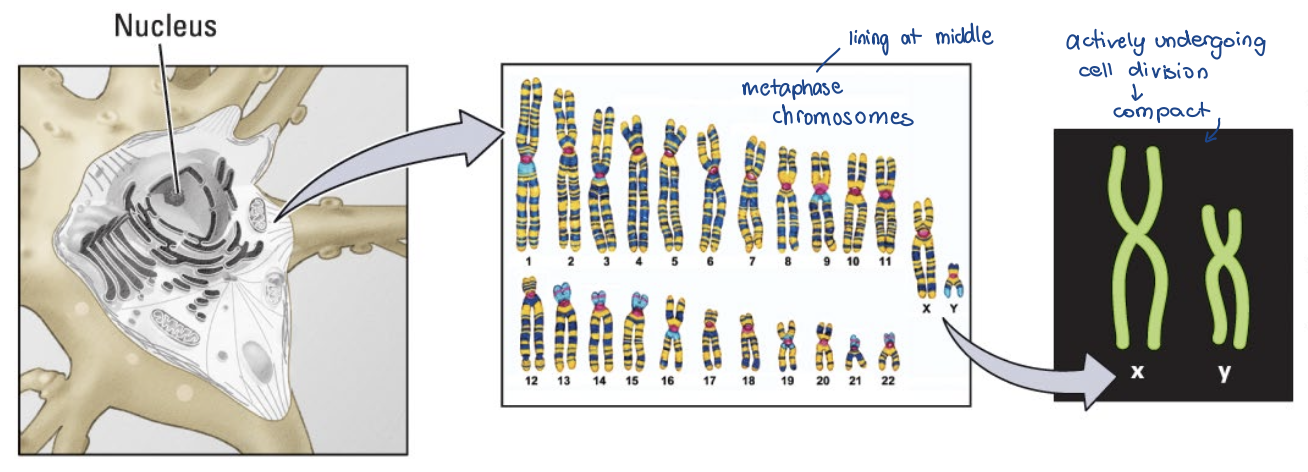
How is DNA organized within the cell?
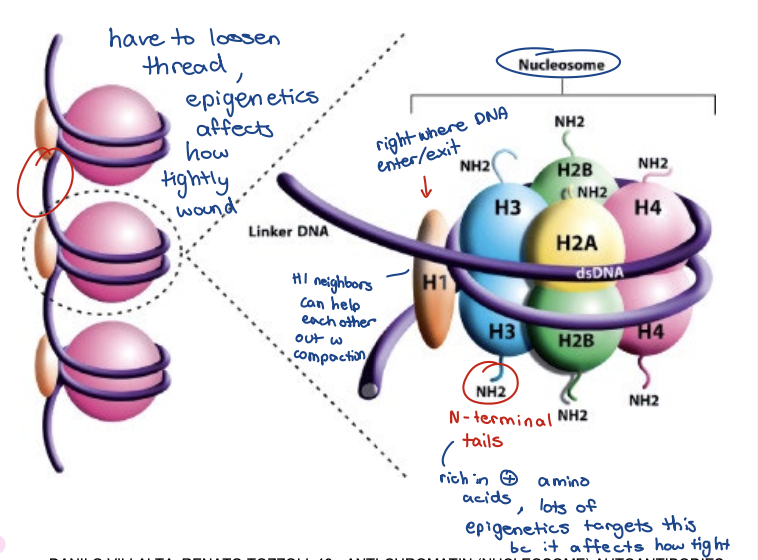
DNA packaging step 1
Nucleosomes are the unit of chromatin
DNA has a negatively charged phosphate backbone
DNA wrapped around core histone octamer
H2A, H2B, H3, H4 proteins
H1 linker
147 bp, wrapped 1.7 times
Primary function of histone proteins is transcriptional control
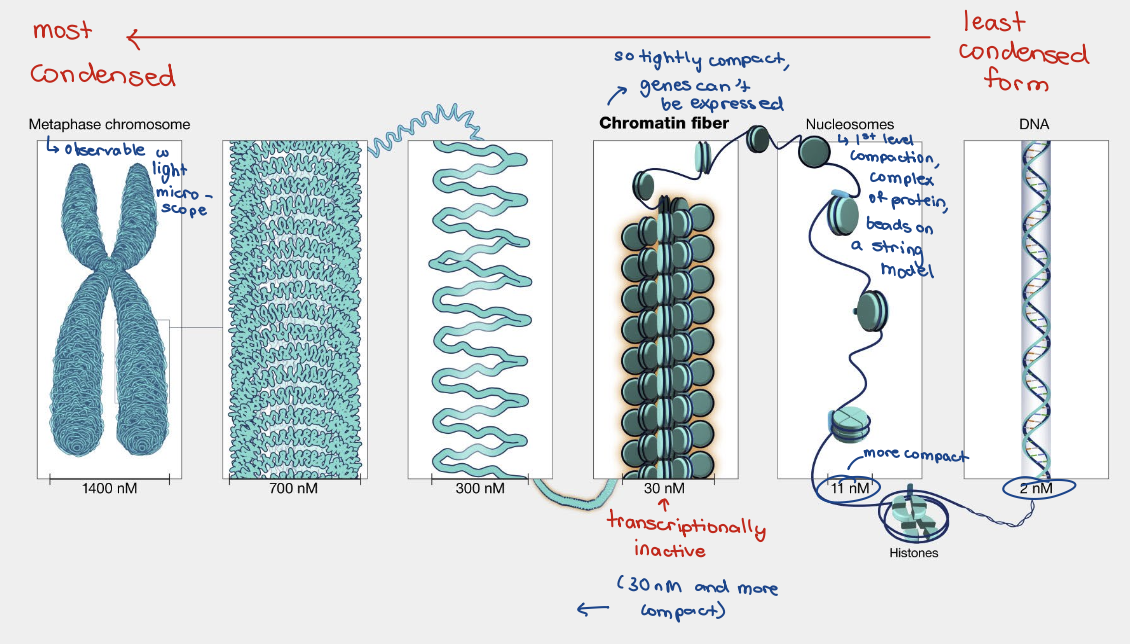
DNA packaging step 2
30 nm chromatin fibers
Transcriptionally dormant
Linker histones and N-terminal tails facilitate interactions between nucleosomes into higher order structures
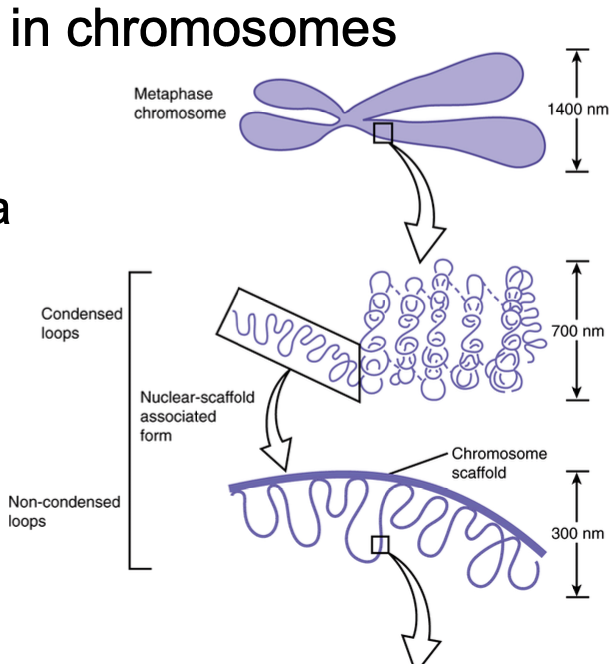
DNA packaging steps 3 and 4
300 nm fiber compaction in chromosomes
30 nm fiber forms loops anchored to a protein scaffold, establishing a 300 nm fiber
300 nm fiber further folds in on itself, forming a chromosome
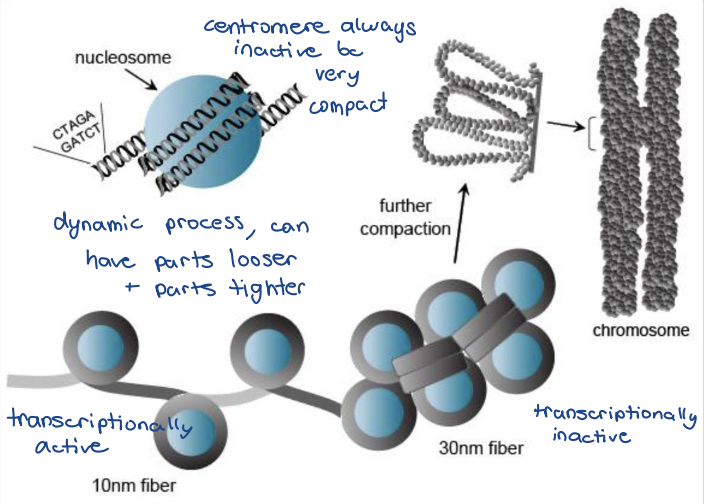
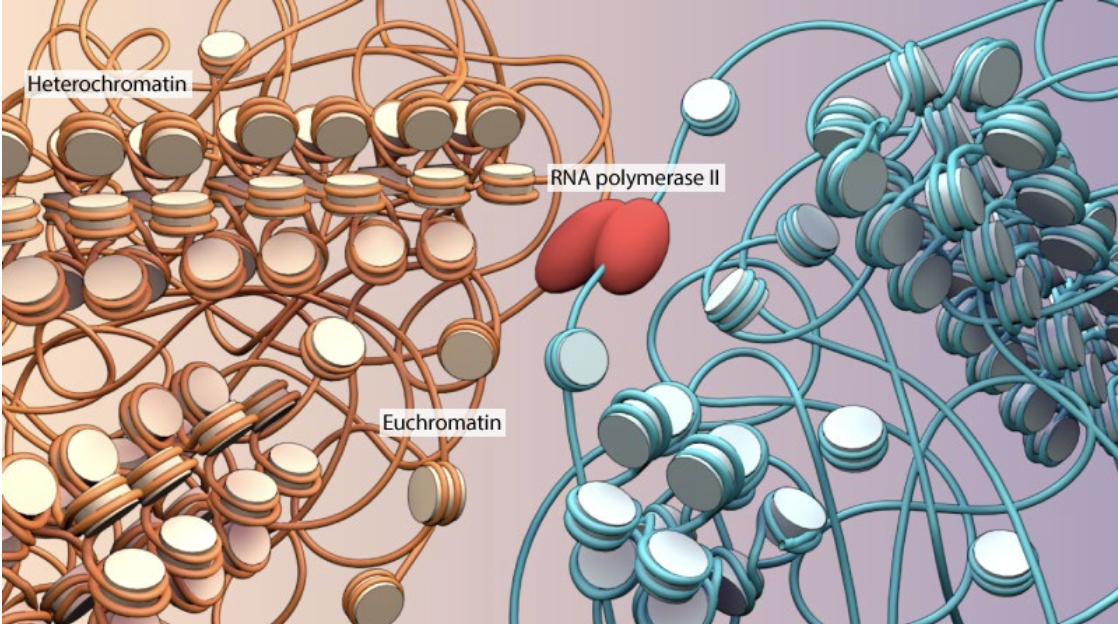
Euchromatin
looser/uncoiled, “open” state
transcriptionally active
~10 nm fiber
Function: permissive to transcription
Interphase chromosomes take up entire nuclear space
Heterochromatin
condensed, or “closed” state
transcriptionally inactive
predominates in non-coding regions
30 nm+
Functions: transcriptional repression, genome stability
-any given chromosome can have
tighter or loosen regions
Chromatin remodeling complexes
Nucleosomes are dynamic
Large protein complexes (ATPases)
Cooperate with specific DNA-binding proteins to
repress/activate gene expression
transcription
DNA replication
DNA repair
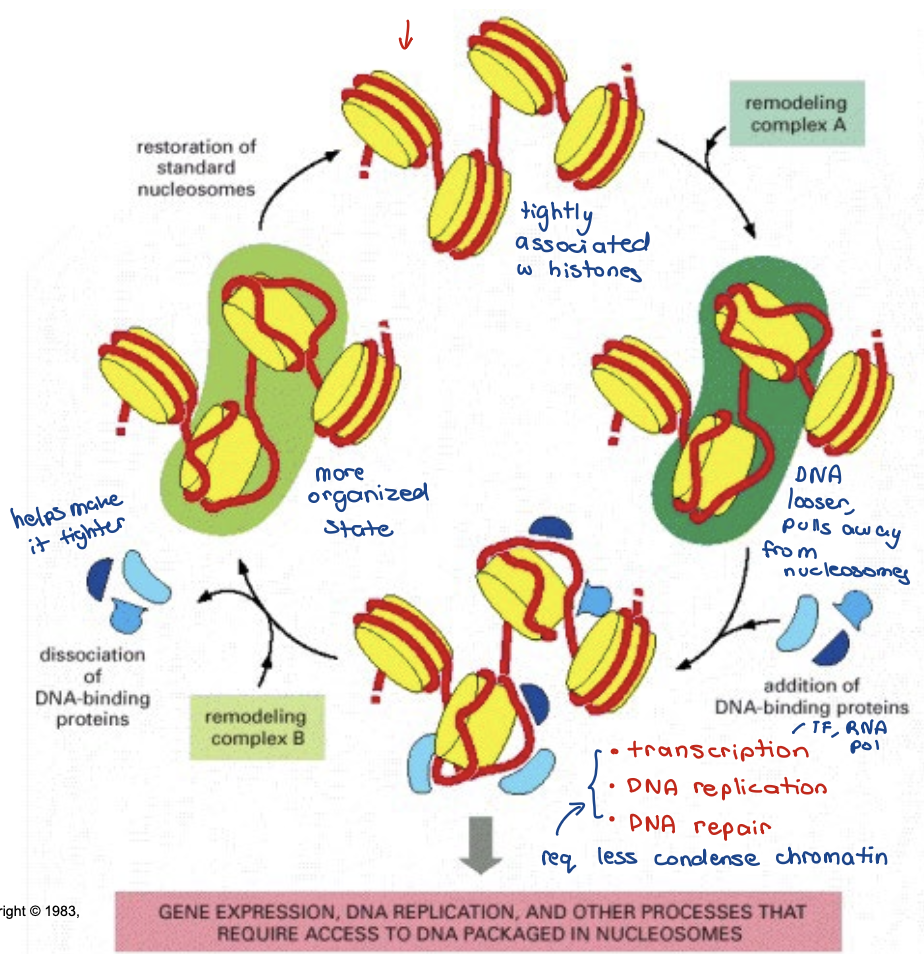
Chromatin remodeling complexes
Nucleosome assembly — promote gene silencing
Chromatin access — exposes binding sites for DNA binding protein
Nucleosome editing — replacing histone with a different variant
Chromatin remodeling complexes can render chromatin more
accessible to DNA-binding proteins
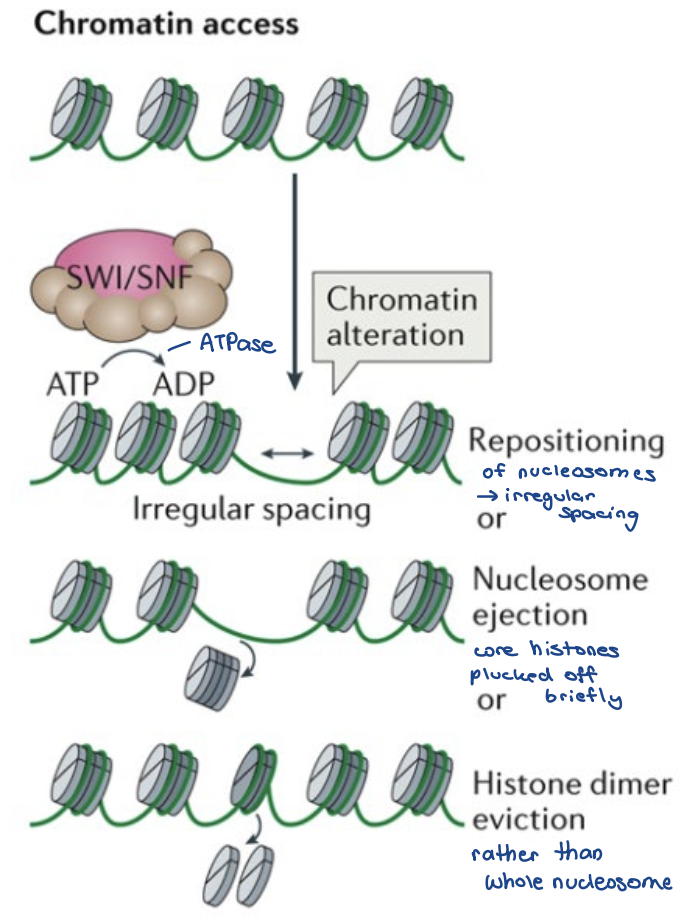
access remodellers
switch/sucrose non-fermentable (SWI/SNF) subfamily of remodellers
Recognize histone modifications
• Mobilize/unwrap or eject nucleosome/histones
• Transcriptional apparatus is granted access
Chemical modifications of the nucleosome dynamically regulate transcription
Posttranslational modification of histones
Acetylation of lysines (K)
Methylation of lysines (K) and
arginines (R)
Phosphorylation of serines (S) and threonines (T)
Ubiquitination of lysines (K)
DNA modification
• Cytosine methylation
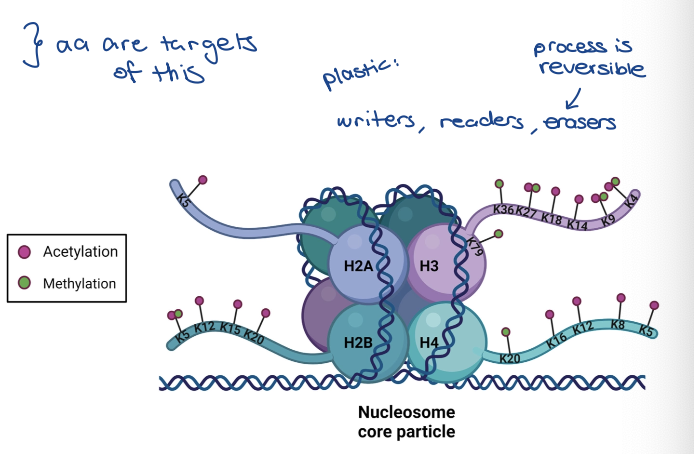
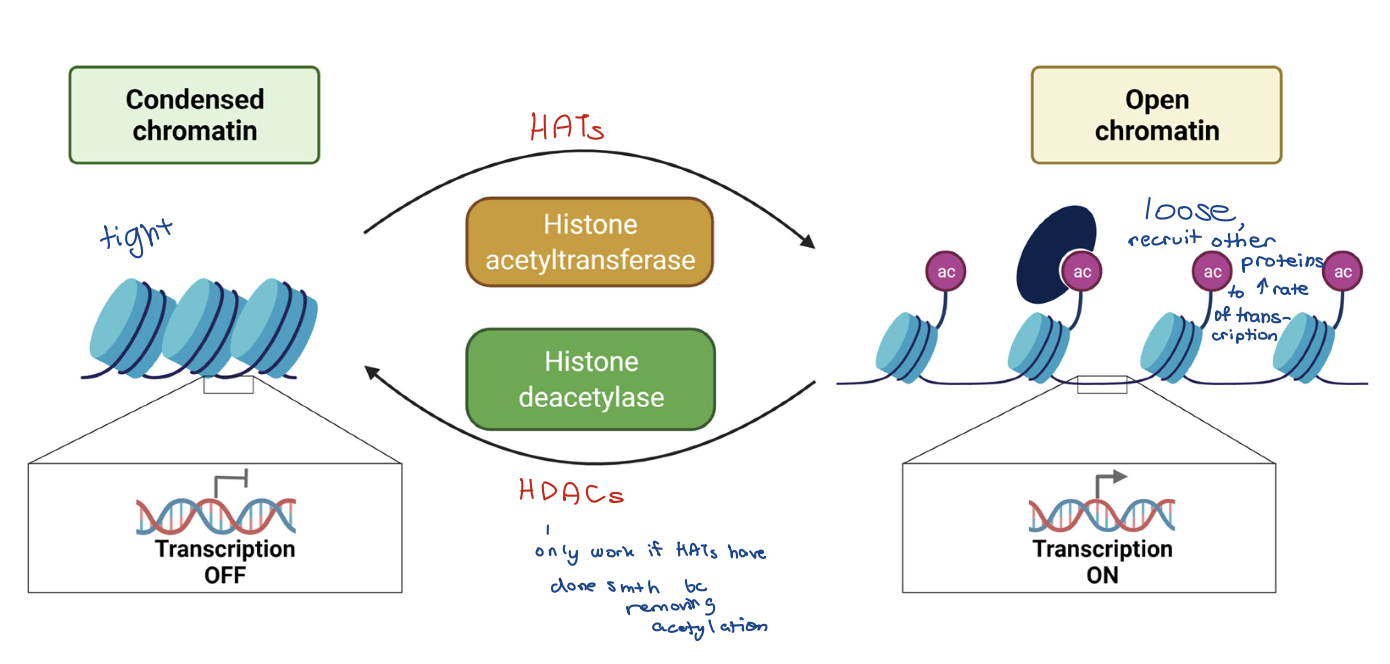
Histone acetylation and deacetylation
Histone acetyltransferases (HATs) –acetyl group added to lysine (+) (HATs → gene activation)
• N-terminal tails (preferred targets) for loosening up
• Internal globular domains
Acetylation destabilizes chromatin structure AND recruits chromatin remodelling enzymes
Reversed by histone deacetylases (HDACs)
Acetyl group: -Ac
HATs vs HDACs
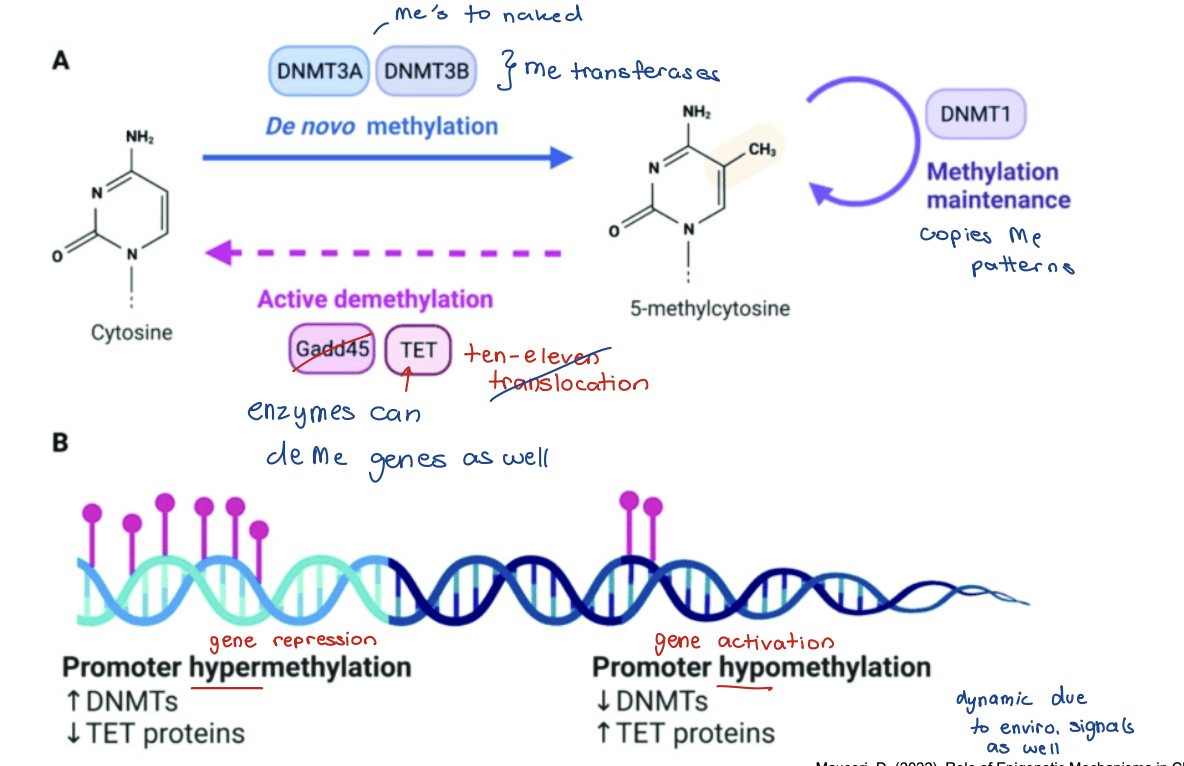
Methylation and demethylation of DNA
Methyl groups added to cytosine nucleotides by DNA methyltransferases (DNMTs)
• Especially targeted to CpG dinucleotides
Represses transcription
• Prevents binding of transcription factors be its bulky
• recruitment of methyl-CpG binding domain (MBD) proteins (repressors)
CpG sites are spread throughout genome and are usually methylated
• Exception is CpG islands long stretches of DNA
• Majority of gene promoters reside within CpG islands → usually not methylated
methylation indicated by: -Me
de novo methylation, methylation maintenance, active demethylation
Chromatin Remodelling – Summary
Condensed chromatin inaccessible to transcriptional machinery
Modulation of chromatin structure – 2 general mechanisms
1. Posttranslational modification (PTM) of histone tails and DNA methylation
2. ATP-dependent chromatin remodelling enzymes
Histone code hypothesis: PTMs occur in complex patterns which are “read” by cellular machineries
• Different combos at various amino acid residues may lead to either activation or repression of transcription
Modulation of chromatin structure – 2 general mechanisms
Posttranslational modification (PTM) of histone tails and DNA methylation
ATP-dependent chromatin remodelling enzymes
Histone code hypothesis
PTMs occur in complex patterns which are “read” by cellular machineries
Different combos at various amino acid residues may lead to either activation or repression of transcription