BIOC 302 - DNA Polymerase and Replication
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
describe the process of protein purification
cell is lysed in test tube
centrifugation to separate soluble proteins (top) from other membrane components (bottom)
Proteins seperated using fractionation techniques: e.g., salt added and look for precipitation, chromatography (size-exclusion chromatography, hydrophobic interaction chromatography)
After each step, fractions tested for desired activity (e.g., DNA synthesis)
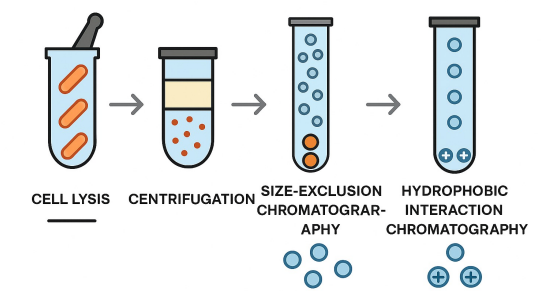
explain the steps of fractionation experiments
start with a cell or organism that contains your protein of interest
break it open under conditions that preserve the proteins (cold, proteases)
separate it based on: size, solubility, hydrophobicity, charge
repeat fractionation with another biophysical property until protein is homogenous
confirm activity via an activity assay
How was protein purification applied to the discovery of DNA polymerase?
After fractionation, each fraction was tested using the kornberg assay for DNA synthesis. Fractions that had the radioactive nucleotides in the DNA contained DNA polymerase. Then, repeated purification steps were used until DNA polymerase was isolated.
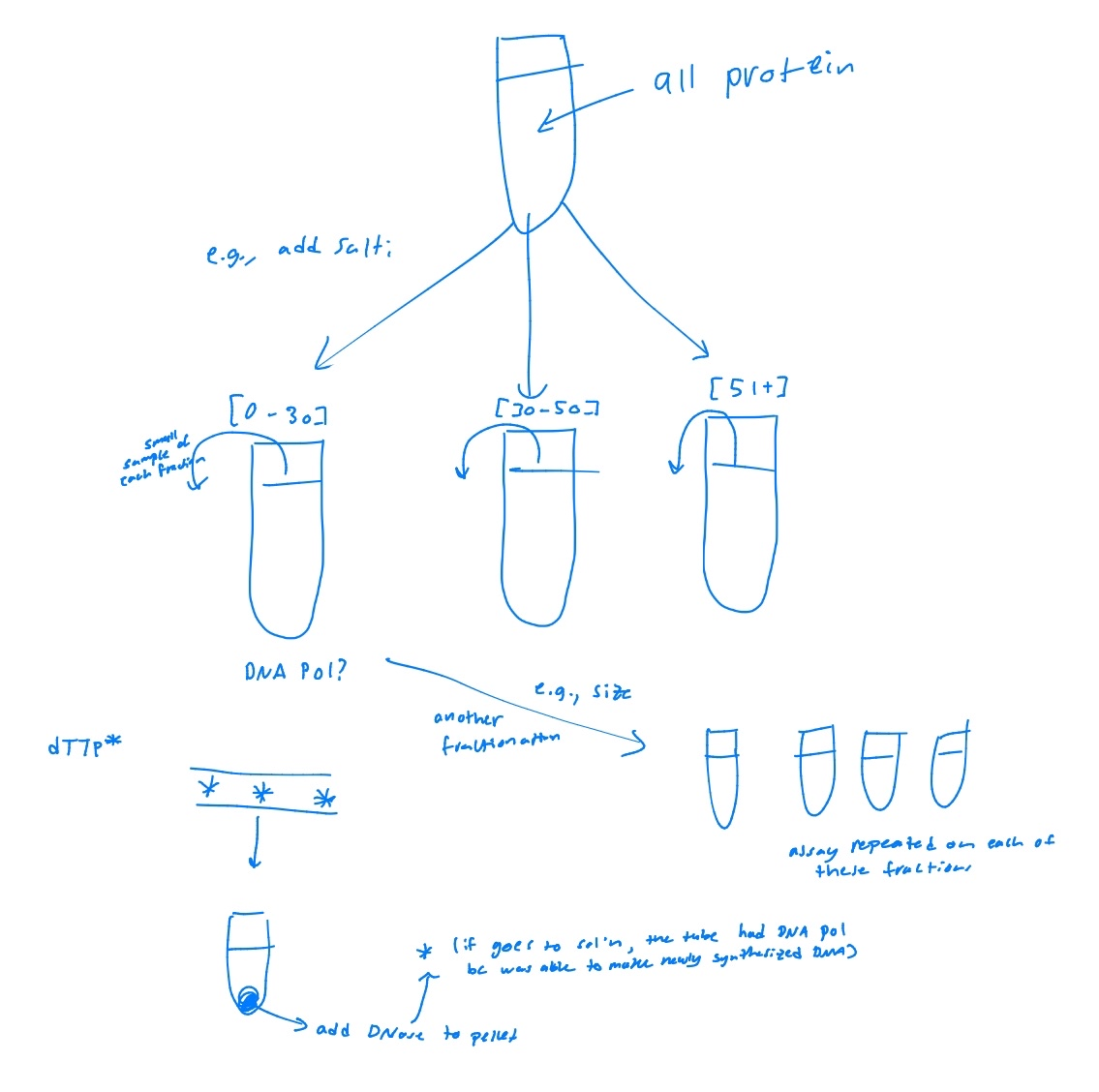
Explain the steps of the Kornberg experiment
mixed radio-labeled dT (deoxythymidine - nucleoside) (method 1) or radio-labeled dTTP (deoxythymidine triphosphate - nucleotide) (method 2) with isolated fraction
added acid to the reaction (after some time) and separated reaction mixture into 2 fractions: a) acid soluble and b) acid insoluble by centrifugation
acid insoluble mixture could be polymerized by DNA, indicating polymerase was present in the mixtyre
verified acid insoluble pellet contained bonafide polymerized DNA by treating with DNase - real nucleotide polymers would be broken up and radioactive would end up in the supernatant
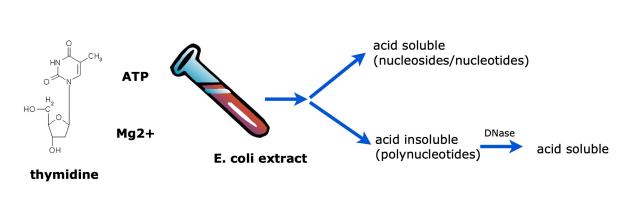
why is acid used in the Kornberg experiment?
nucleosides/nucleotides are acid soluble while polynucleotides are acid insoluble. Therefore, when acid is added, polynucleotides are gonna be able to be observed in the test tube as they are insoluble in acid.
why did the experiment stop working after the first mixture when dT was used instead of dTTP?
The nucleoside dT must be converted into a nucleotide (dTTP) to be incorporated into DNA. In the first mixture, any contaminating kinase could phosphorylate dT, allowing DNA polymerase to use it. During fractionation, DNA polymerase was separated from the kinase (they have different properties), so the kinase was no longer present. Without kinase, dT could not be converted to dTTP, and the polymerase could not add it to the growing DNA chain → no radioactive pellet formed.

What is the purpose of fractionation?
to isolate different proteins from a complex mixture
what is the role of DNase? why would we expect free nucleotides in solution and DNA polymers in the pellet?
DNase breaks the phosphodiester bonds. Thus DNase breaks down DNA polymer, releasing the radioactive nucleotide (now acid soluble) so that we know the nucleotide was actually in the polymer and not just trapped somewhere (like isolated protein that binds to nucleic acid). Free nucleotides in sol’n bc acid soluble. DNA polymers in pellet bc acid insoluble.
How would we adapt this experiment (Kornberg) if our goal was to instead demonstrate the presence and function of RNA polymerase?
Could use UTP and Rnase
what is the primary function of DNA polymerase 1?
cleanup during replication, recombination, and repair
what is the function of DNA pol. II?
involved in DNA repair (to fix DNA, u gotta break it first)
what is the function of DNA polymerase III?
its the principal replication enzyme in E. coli
what is the function of DNA pol. IV and V?
involved in an unusual form of DNA repair (they’re translesion polymerases that repair damage from UV radiation)
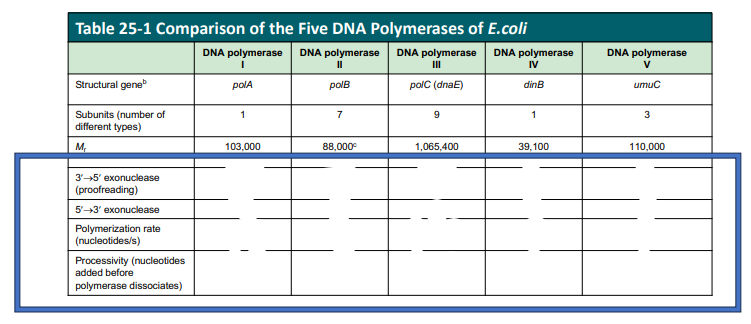
DNA Pol II is the processive polymerase
Pol I is the only one with the 5’→3’ exonuclease
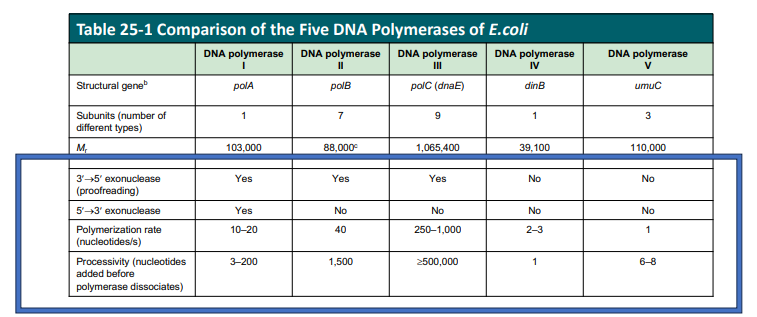
How were other DNA polymerases discovered?
by looking at mutant E.coli with no DNA pol activity. Other DNA polymerases were isolated after an E. coli mutant without functional DNA Pol I was identified. It turns out that Pol III is the replicative polymerase, and the others are involved in DNA repair. DNA pol I is the prototype for all DNA polymerases (enzymatically and structurally)
What is the 3’ → 5’ exonuclease activity of DNA Pol I?
it is used for proofreading and removes incorrectly paired nucleotides behind the polymerase, ensuring high replication fidelity
what is the DNA Pol I. 5’ → 3’ exonuclease activity used for?
Nick translation (not actually translation). The 3’→ 5’ domain is in front of the enzyme and performs nick translation. Nick translation is when a break or nick in the DNA is moved along the enzyme (DNA Pol removes nucleotides one by one after the initial nick). The active site of DNA Pol I comes along behind the 5’ → 3’ exonuclease and adds nucleotides. The purpose of this is for DNA repair and the removal of RNA primers during replication. (btw for nick translation to occur, there has to be a nick present already in the backbone of the DNA).
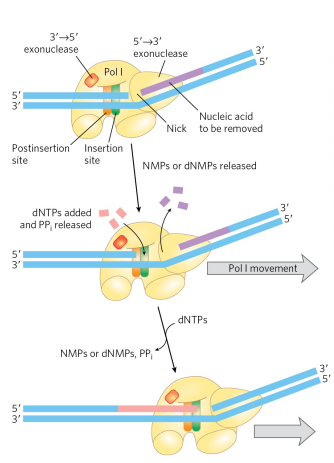
What is the Klenow fragment of DNA Pol I and what activities does it retain?
The Klenow fragment is produced by mild protease treatment of DNA Pol I and retains the polymerase activity and 3′→5′ proofreading exonuclease activity, but it loses the 5′→3′ exonuclease activity responsible for nick translation.
what are the 4 requirements of DNA-templated DNA polymerase the synthesize new DNA?
Single-stranded template
deoxyribonucleotide with a 5’ triphosphate (dNTPs)
Magnesium ions (essential cofactor for the polymerase)
annealed primer (often RNA) with a free 3’ OH
how does DNA Pol. synthesize new DNA polymers and in what direction?
it synthesizes new DNA in a template driven process, and synthesis only occurs in the 5’→3’ direction
explain the steps of the DNA polymerase reaction mechanism
incoming triphosphate nucleotide forms W-C interaction with the template strand
Mg2+ stabilizes the 3’OH and alpha phosphate
bond forms between alpha phosphate and 3’ OH
beta and gamma phosphates released as PPi, and new phosphodiester bond is formed
as long as template strand is still available, a new substrate is generated for another round of polymerization
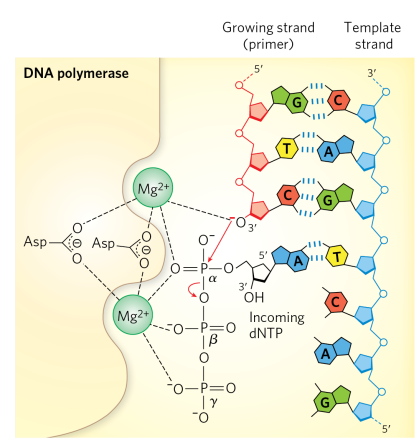
Why does DNA polymerase require a single-stranded template and a free 3′ hydroxyl, and how is a single-stranded template generated?
DNA polymerase requires a single-stranded template and a free 3′ hydroxyl to synthesize DNA in a template-driven 5′→3′ manner. Double-stranded DNA is too stable for polymerase to work, so a helicase, which is ATP-dependent and acts like a zipper, separates the two strands to create single-stranded templates for the polymerase to extend from the primer. Without a single-stranded template, polymerase cannot function.
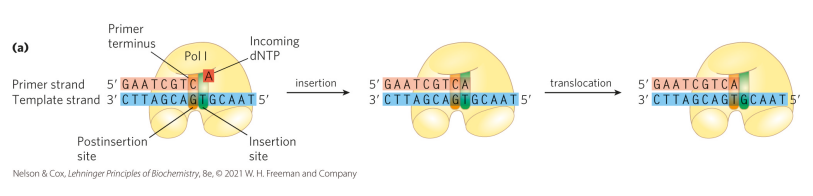
Why is single-stranded DNA unstable after helicase unwinds it, and what problem does this cause?
Once DNA is unwound by helicase, the single-stranded DNA is unstable because exposing the nitrogen-rich bases is energetically costly, and the strands could re-anneal (get back together). To prevent wasting this energy and to keep the template accessible for replication (so stay single-stranded), single-stranded DNA binding proteins stabilize the unwound strands so they don’t collapse back together. SSB proteins make non-specific interactions with the single stranded DNA. The motion of the protein (has lots of movement) allows for it to be quickly slid out of the way when needed so that replication can occur.
what mechanisms ensure fidelity of DNA replication?
W-C H-bonds
active site geometry - shape discrimination
proof-reading in the 3’→5’ exonuclease domain
mismatch repair
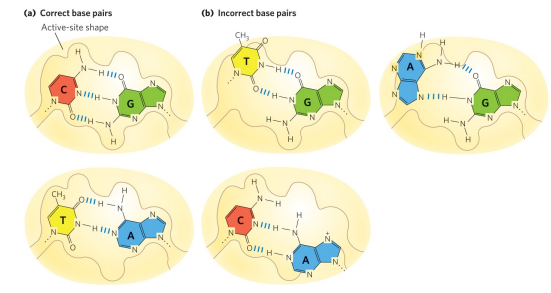
How does DNA polymerase use base-pair geometry (shape discrimination) to ensure fidelity of replicated DNA?
The active site of DNA polymerase has specific shape to perfectly fit the correct W-C base pairs, allowing for efficient catalysis of properly matched bases. Incorrect base pairs distort the geometry of the active site and do not fit properly, making them energetically unfavorable and less likely to be incorporated.
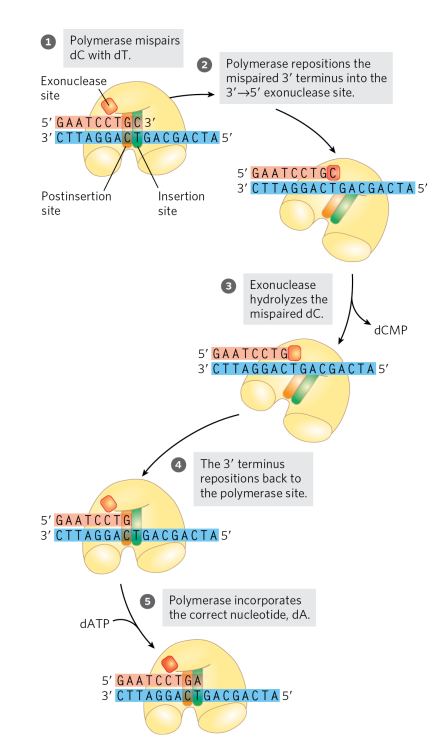
how does the proof-reading in the 3’→5’ exonuclease domain work? Explain the steps.
its used for error correction to ensure fidelity of the replicated DNA. When the wrong nucleotide is added, the DNA polymerase is unable to translocate and continue to add nucleotides. The 3’→5’ exonuclease permits the enzyme to remove a newly added nucleotide so that it can add the correct nucleotide (structure of the editing site favours non-canonical base pairs). (look at image for steps)
True or false: DNA polymerase is replicating a region of the genome and makes a mistake. After adding only one or two more nucleotides to the polymer, it is possible to remove the incorrect nucleotides using the 3’->5’ exonuclease domain
False bc if there is a mismatch that is not at the end, there is no opportunity to alter that region of the genome bc the DNA pol is not gonna go back and fix it.
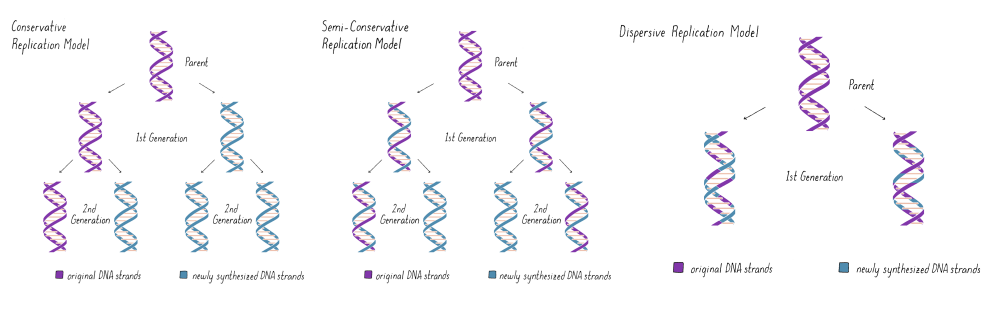
what are the 3 models of DNA replication?
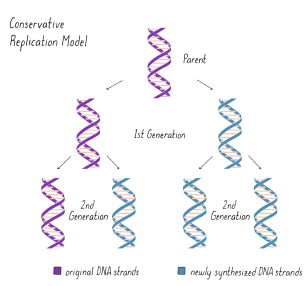
conservative (parent strands stay together and daughter strands stay together
semi-conservative (one parent strand, one daughter strand)
dispersive (done in pieces and put back together)
what is the Meselson-Stahl experiment?
it was an experiment done to determine the mechanism of DNA replication (conservative vs semi-conservative vs dispersive)
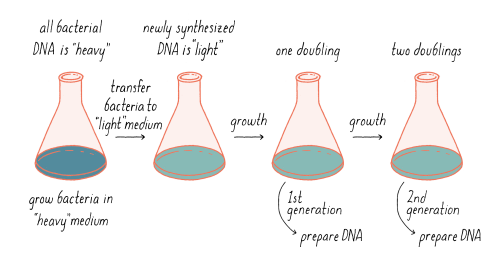
Explain how the Meselson-Stahl experiment was conducted
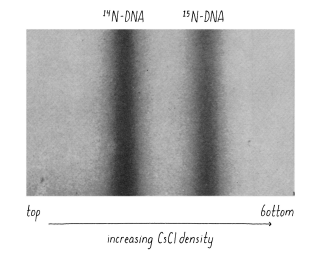
They grew E. coli in the presence of heavy nitrogen (15N) to label the bacteria’s DNA (since bacteria copy their entire complement of DNA, or genome, before every cell division). Then, they moved the bacteria to a normal 14N-containing medium (allowed the cells to divide once, separated the DNA by density). The results supported semiconservative replication. The results showed that at the onset of the experiment, all the DNA is heavy - as the E.coli would’ve been grown for many generations in the heavy nitrogen. Once the E. coli is transferred into light media, any newly synthesized DNA will be made from light nitrogen. By monitoring the heaviness of DNA after generations, a model replication could be supported (based on the weight of DNA).
In the Meselson-Stahl experiment, how is heavy and light DNA distinguished?
Distinguished on a cesium chloride density gradient - heavier DNA travels further
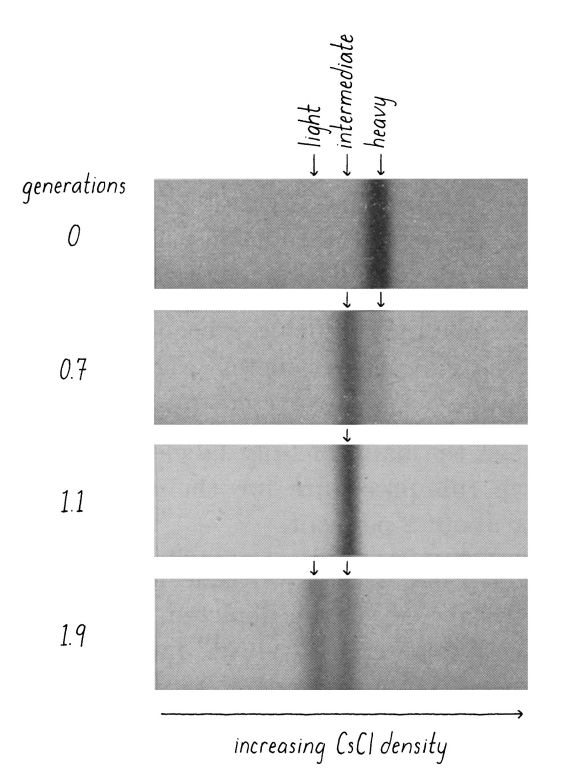
what was the result of the Meselson-Stahl experiment?
Semi-conservative replication. After 1 generation (1 doubling of the E. coli - means all the DNA has been duplicated) the DNA from these samples is a mixture of heavy and light - this rules out the conservative replication model. After 2 generations, the DNA is either all light, or a mixture of heavy and light which supports the semi-conservative model. (0 generation = all DNA is heavy. After one generation, DNA is gonna be composed of 1 daughter strand (light) and one parent strand (heavy) - DNA weight intermediate . After 2 generation, one DNA mlc will have 1 parent strand and 1 daughter strand - intermediate weight; and one DNA mlc with 2 daughter strands (light DNA) which is why we see 2 bands with 2 diff weights at 2 generations).

B. One heavy and one light
how many origins of replication in a circular genome?
1
How does DNA replication initiate and proceed in bacterial chromosomes?
Replication begins at a specific origin where the DNA strands separate. Replication forks form at both ends of the origin, allowing bidirectional replication. Both DNA strands are replicated simultaneously, and replication loops extend from the origin.
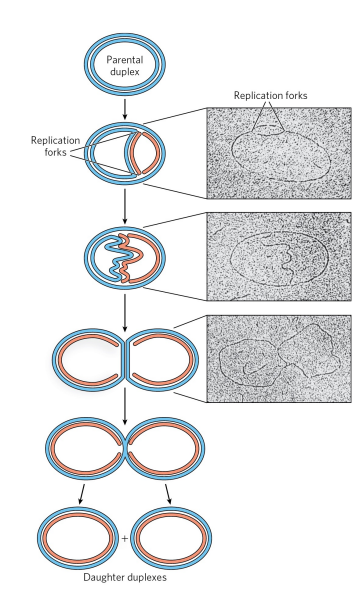
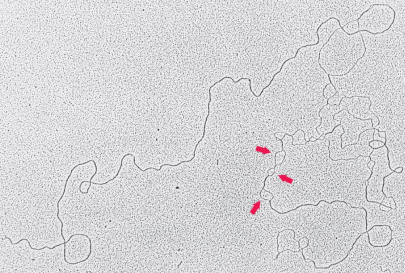
what is denaturation mapping?
selective denaturation of sequences unusually rich in A=T base pairs to provide landmarks along the DNA mlc. Generates a reproducible pattern of single-strand bubbles. At the bubbles, replication occurs bidirectionally, speeding up replication.
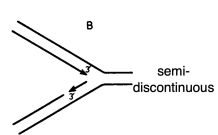
in semi-discontinuous DNA replication, the leading strand is going ___ the replication fork, and the lagging strand is going ___ the replication fork
toward
away from
What polymerase replicates the genome in E. coli and how does this differ in mammals?
In E. coli, DNA Polymerase III replicates the entire genome. In mammals, two separate polymerases are used: one replicates the leading strand, and the other replicates the lagging strand.
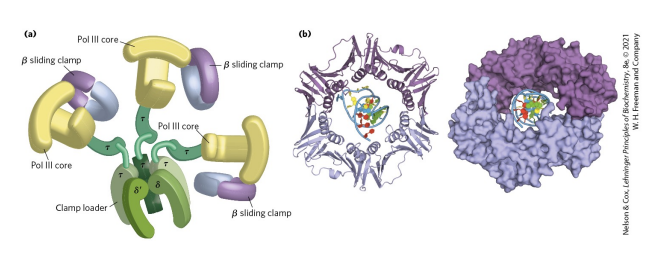
What is the point of the clamp loader and clamp in DNA polymerase III?
Clamp is needed for DNA polymerase III to be processive. Without it, there would be low fidelity since DNA polymerase on its down can fall off DNA easily. The clamp loader is needed to load the clamp onto the DNA-RNA hybrid (single-stranded DNA with the small amount of RNA - RNA primer on it). Clamp loading requires ATP binding and hydrolysis to add the clamp to the substrate.
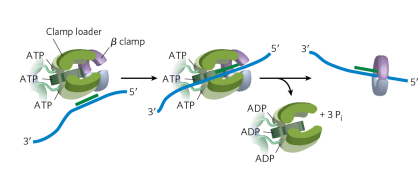
True or False: A mutation that renders the protein non-function in either the clamp or the clamp loader would have the same effect – it would reduce the processivity of DNA polymerase
True (we wouldn’t actually see a variant like this occurring in a life form bc you have to replicate ur DNA to survive)
what does primase do and why is it needed?
Primase synthesizes short RNA primers on single-stranded DNA, providing a free 3’ hydroxyl (-OH) group. This 3’ OH is essential because DNA polymerase can only form a phosphodiester bond by adding a nucleotide to an existing strand—it cannot start from scratch. The RNA primer allows DNA polymerase to attach to the 3’ OH and begin elongating the DNA strand.
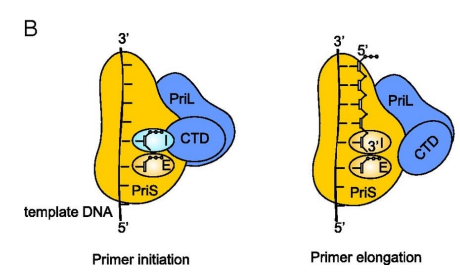
what is the problem with primase?
primase is error prone (not very good at adding W-C interactions and making sure that the sequence is perfect)
Explain the steps of leading and lagging strand synthesis
1. Helicase (DnaB) unwinds DNA at the replication fork
2. Leading strand is synthesized in one piece
3. Lagging strand is synthesized in fragments – Okazaki fragments
4. Primase adds a primer to generate a free 3’OH
5. Clamp loader loads on a clamp to ensure synthesis is processive
6. Synthesis of lagging strand progresses until primer on older Okazaki fragment
7. Clamp loader (green structures in cartoon) re-loads clamps on new primer/template RNA-DNA hybrids until synthesis is complete
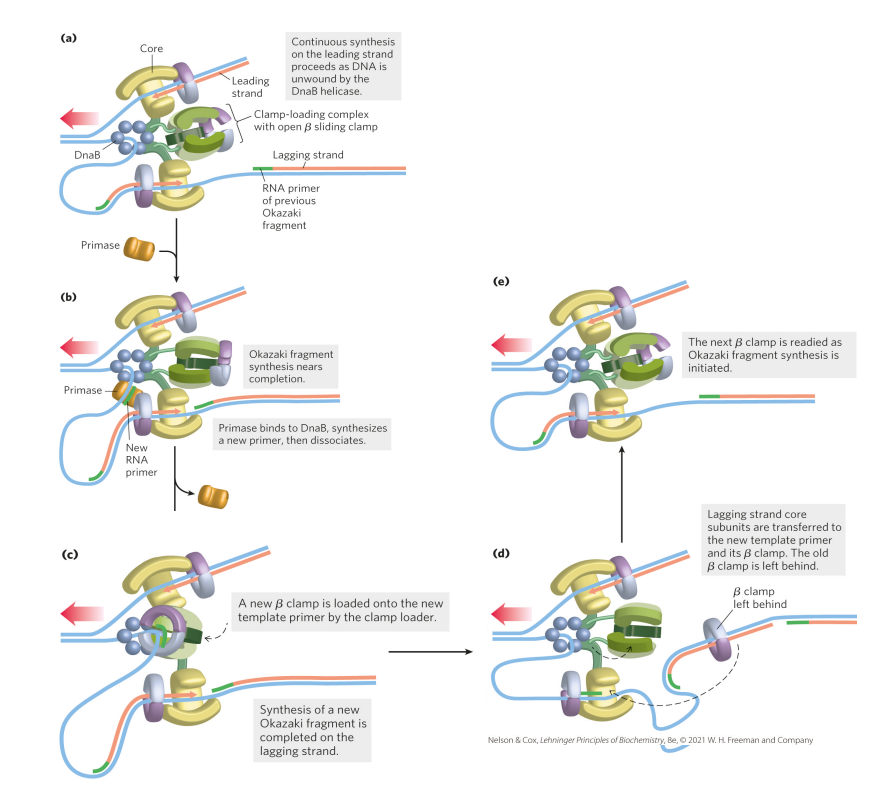
what problems does the synthesis of short fragments of DNA generate? How are these problems solved?
RNA - generating Okazaki fragments involve short regions of RNA-DNA hybrid complexes in the lagging strand. DNA pol I removes the RNA and replaces it with DNA through the specialized exonuclease activity “nick translation.”
Gaps - DNA ligase seals the remaining nick
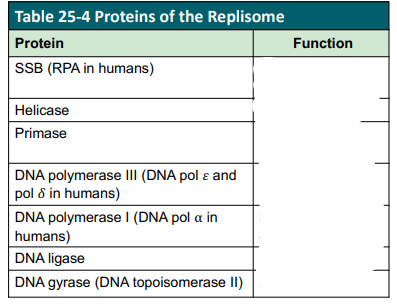
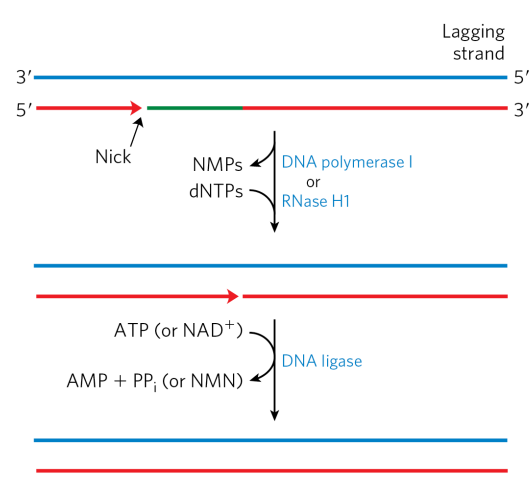

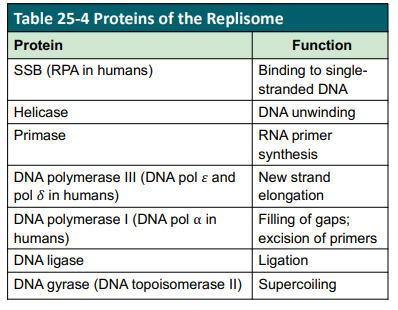

D
You want to generate a fragment of DNA with short concentrated fluorescent labels. Which polymerase would you choose and why?
Pol I bc adds a good amount of nucleotides (not too much, not too little) therefore good for seeing short fluorescent labels on DNA