Mole Bio Exam 3
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
What is homologous Recombination
the “crossing over” event during meiosis
essential for diversity, segregation, and repair during mitosis
What is Somatic Recombination
recombination in non-sex cells
mainly in Immune cells
Double Strand Break Repair Pathway
Recruitment of MRN and RAD proteins
Resection by exonucleases
Strand invasion
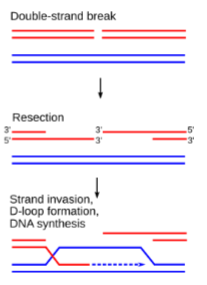
Types of Double Stranded Break Repair
Double Holliday Junction
Synthesis-dependent Single-strand Annealing
What types of Strand-Transfer proteins are there
Generic : promote exchange between DNA strands
Controlled : sequence specific DNA transfers (site-specific recombinases)
How does Phage lambda integrate into plasmids
Recombination between attP(phage plasmide) and attB(bacterial plasmid) sites
What do RAD and MRN do in DNA repair
RAD: recombination-repair
MRN single-stranded overhang in DNA ends
Non-Homologous End Joining
joining of double stranded breaks when homologous sequence is not available
Ligates blunt DNA ends
Break Induced Replication
break in chromosome at end
Uses non-homologous chromosome
Can cause Non-Reciprocal translocation (unbalance translocations of chromosome segments)
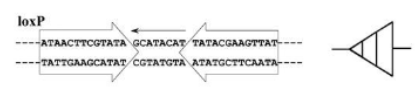
LoxP site and Cre recombinase
LoxP site is two inverted 13bp repeats sandwiching an 8bp core
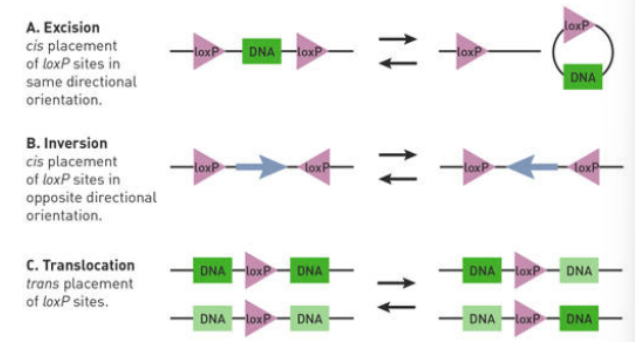
Cre recombinase can
Excise LoxP + DNA outside of site
Inverts DNA outside of LoxP
Translocate DNA outside of LoxP
FLP recombinase and FRT sites
FLP: Flippase that recognizes FRT site
FRT: like LoxP
Mechanism: FLP excises DNA between two FLP sites facing same direction
What is Recombination signal sequence (RSS)
The V+D+J recombination machinery uses sequences of 7bp separated by nonamers of 12-23bp (RSS)
signals VDJ recombination
VDJ recombination events in blood cells
Rag 1 or 2 recognizes RSS flanking VDJ
Rag complexes with HMGB1 and hairpin the DNA
Double stranded break
non-homologous repair and end joining (NHEJ)
random nucleotides are added at junctions
Transcription direction
moves 3’→5’ on DNA
Upstream vs Downstream of RNA polymerase
Up: polymerase has already passed over it
Down: yet to be transcribed
Primary Transcript
the newly synthesized RNA
Prokaryotic RNA Polymerase holoenzyme
5 subunits + sigma factor
C Terminal Domain contacts regulatory proteins
Prokaryote Transcription steps
Polymerase binds to promoter forming closed complex
Transcription is initiated forming bubble (open complex)
elongation moving 3’→5’ on DNA
termination
what does Sigma Factor do
Required for initiation
increases affinity for DNA promoter in RNA pol.
recognizes consensus sequences like TATA
can compete with other sigmas to bind at different promoters
displaced when transcription starts
Ternary Complex in Prokaryotic Transcription
Formed at initiation
RNA pol. + DNA + Dinucleotide primer
UP element
sequence upstream to promoter that enhances transcription
binds to CTD on RNA pol
What is promoter strength
frequency of RNA pol. transcription initiation
Down mutation vs up mutation in promoters
down : decrease promoter efficiency specificity to consensus sequence
up : opposite
Footprinting
DNA is amplified with PCR
radio labeled at ends
reacted with DNA binding proteins
treated with DNAase1 (cant endonuclease where protein is bound)
Creates different lengths of protein
analyzed with gel electrophoresis
no band is part of DNA with protein bound to it
Termination of Transcription
intrinsic : RNA polymerase recognizes terminator sequence in DNA→ makes hairpin in new RNA → cleavage
rho-dependent: needs rho proteins that bind to new RNA and cleaves RNA at rut site sequence on RNA