Cell Bio Final Exam
1/66
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
67 Terms
What is Cell Culture?
Growth of Cells outside their natural environment
What are the two main types of Cell Cultures?
Primary Cells
Directly isolated from an organism, limited lifespan
Immortalized Cells:
Divide indefinitely, easier to maintain (often derived from cancer, or induced to behave cancer-like)
Adherent vs Suspension growing cells in culture
Adherent Cells
Grow attached to flask surfaces (most cells do this)
Suspension Cells
Grow Freely in growth media
What are some essential cell culture equipment?
Biosafety Cabinet (Laminar flow hood)
CO2 Incubator (37 C at 5% CO2)
Inverted Microscope
Centrifuge
37 C water Bath
Sterile Pipettes and Tissue Culture Flasks
When should cells ideally be passaged (Subcultured)? what do you have to worry about with too many passages?
1) Should be passaged during exponential growth phase at around 70-90% confluence
2) High passage numbers risk genetic drift
What is the role of Trypsin, Glucose, and DMEM in cell culture?
Trypsin is used to detach cells from flask surface
Glucose is used for energy
DMEM is growth medium (provides nutrients, amino acids, ect…)
What P-values determines whether results are statistically significant?
values below 0.05
What did Dr. O’Toole show BRAF to interact with, even though they were originally thought to be mutually exclusive?
TP53 interacts with mutated BRAF (BRAFV600E)
What did Dr. O’Toole use in order to identify neighboring biotinylated proteins?
TurboID
How many alleles do proto-oncogenes require to be mutated in order to be cancerous? Does this show that this is a dominant or recessive mutation? Is this a loss of function or gain of function?
1) 1 allele
2) Dominant
3) Gain of function
How many alleles do tumor suppressor genes require to be mutated in order to be cancerous? Does this show that this is a dominant or recessive mutation? Is this a loss of function or gain of function?
1) 2 alleles (both alleles)
2) Recessive
3) Loss of function
Protein Production can be regulated at many points, but what is the most important regulation point?
Transcription Initiation is the most important control point.
No unnecessary intermediates, most energy efficient.
What does the Chromatin-Remodeling Complex do?
Binds to Histone tails, and moves histones to expose DNA and allow for transcription.
(Especially helps if a promoter region in euchromatin is wrapped around a histone protein)
What are Histone AcetylTransferases (HAT) and Histone DeAcetylase (HDAC)? What do they do?
Histone AcetylTransferases
Adds acetyl group to histone tail
This allows chromatin relaxation
Histone DeAcetylase
Removes acetyl from histone tails
Promotes chromatin condensation (gene repression)
What does methylation of histone tails cause?
Methylation of histone tails condenses chromatin
This “hides” DNA sequence and blocks transcription
What do Transcription Regulators do?
Transcription regulators bind to regulatory DNA sequences & assist in RNA polymerase binding.
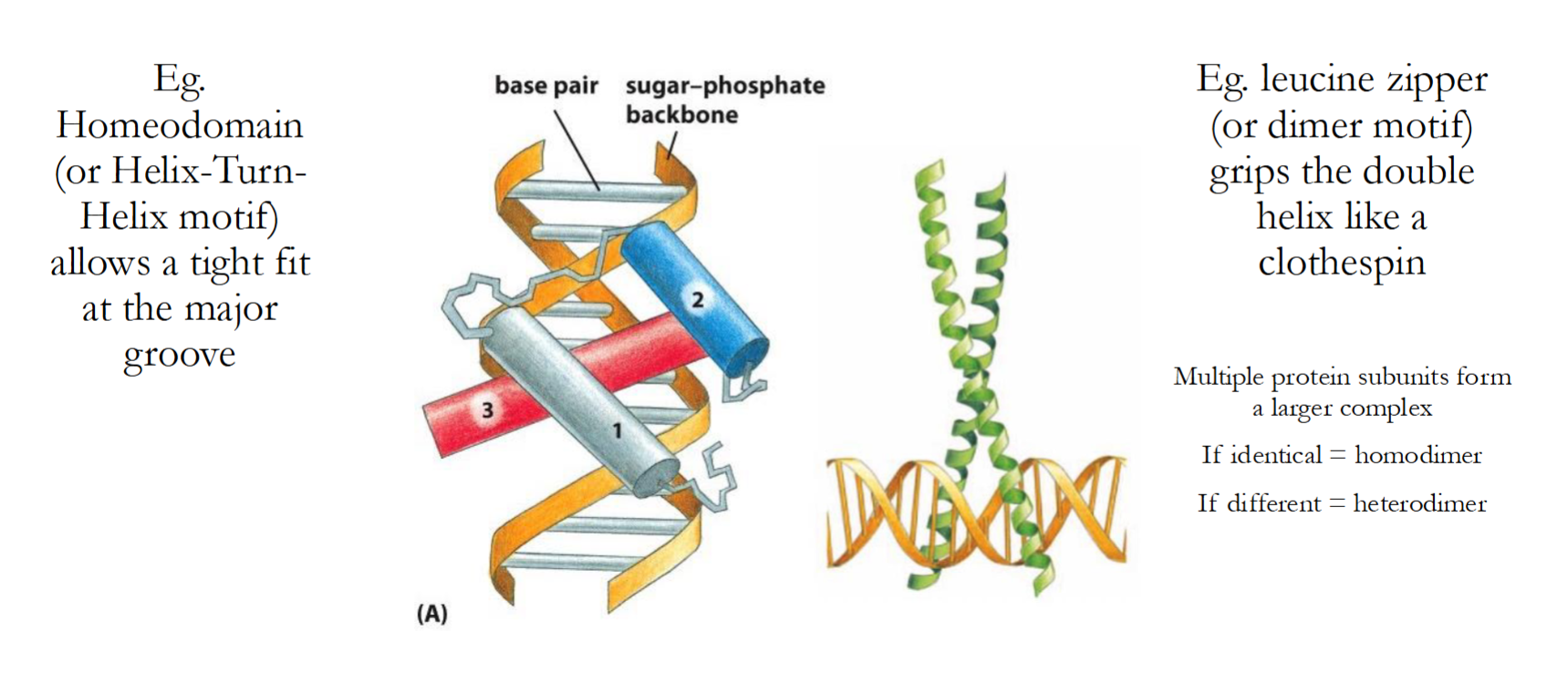
Transcription regulators bind to DNA using conserved ‘motifs’, what are these motifs?
Homeodomain/Helix-Turn-Helix
Allows a tight fit at the major groove
Leucine Zipper/Dimer Motif
Grips double helix like a clothespin
Multiple protein subunits form this complex, and:
-If identical = Homodimer
-If different = Heterodimer
Genes can be regulated at a distance with Enhancers/Repressors. How is this so?
Through the DNA looping around and meeting at a Mediator. The mediator is large and can touch both the core promoter and the Enhancer/Repressor.
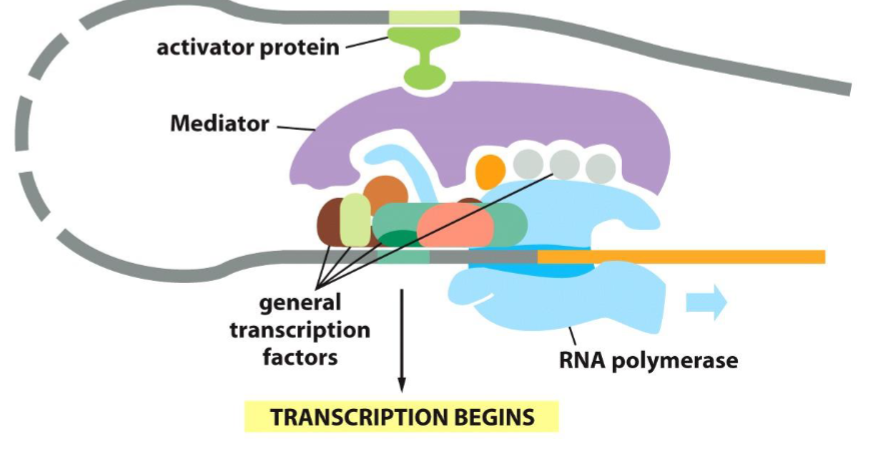
What are Enhancers and Repressors?
Enhancers and Repressors are regulator non-coding DNA sites located upstream/downstream from the promoter and allow regulatory proteins to bind that either promote or repress transcription.
What is a Mediator in Transcription?
A Mediator is a multi-subunit (>20 subunits) complex that links TFs (Transcription Factors) to RNA polymerase II
does not contain DNA-binding components itself (relies on TFs for specificity)
What is combinatorial control, and what dose it accomplish?
1) Combinatorial control is how genes can be regulated through a combination of multiple Transcription Factors, regulatory sequences, and other regulatory proteins.
2) allows complex organisms to be built from a manageable “toolkit” of regulatory proteins, and therefore, allows cell differentiation
A mutation in an exon of a gene may deactivate that gene everywhere in an organisms body. What would happen if a mutation occurred in one of the enhancer regions?
A mutation at an enhancer region would only deactivate this gene in cells where that enhancer is needed to express it.
(Ex: mutation in eye enhancer, eye no longer produces that gene)
Stickleback fish in freshwater lost their pelvic spines, the mutation that caused this occurred where and effected what gene?
1) Mutation deactivated the pelvic regulatory switch.
2) effected the Pitx1 gene only in the pelvic region.
In Bacteria, at what point does gene regulation almost exclusively occur? Why is this?
1) Gene regulation in bacteria occurs almost exclusively at the level of transcription
2) This is because RNA is often translated while it is till being transcribed (no processing, less opportunities for regulation)
What is an Operon?
A group of genes that are all under the control of the same promoter, thus are transcribed as a unit.
Most commonly found in Bacteria
The entire operon can be regulated by the one promoter
What is polycistronic mRNA?
One mRNA transcript containing multiple genes.
Results from the transcription of an Operon
Activator and Repressor proteins can bind to promoters to do what?
Can bind to promoter sequences to increase or decrease transcription
In Bacteria, What is the Trp Operon? How is it regulated?
1) an Operon of 5 genes that are involved in the production of the amino acid Tryptophan.
2) When Tryptophan levels in the cell are high, the Tryptophan repressor (a homodimer) binds to the Trp regulatory region in the promoter, and physically blocks RNA Polymerase. When Tryptophan levels are low, the repressor does not bind.
In Bacteria, what is the Lac Operon? How is it regulated?
1) An Operon of 3 genes that are specific for the uptake and metabolism of lactose (needed to digest lactose).
2) An Activator AND a Repressor control the Lac Operon. Activator binds when Glucose levels are low. Repressor disassociates when Lactose levels are high. (Therefore, in order to transcribe, need both low glucose levels and high lactose levels).
Repressor: A tetramer that forms a loop in DNA that physically blocks RNA Pol.
Activator: Catabolite Activator Protein (CAP) (a homodimer) needs cAMP to bind in order to induce a slight allosteric shift, causing bound DNA to bend slightly, which helps to stabilize RNA Pol on the promoter
What determines the amount of mRNA and protein in a cell?
mRNA: level of transcription - amount of decay by ribonucleases = amount of mRNA
Protein: level of translation - amount of decay by proteases = Amount of proteins
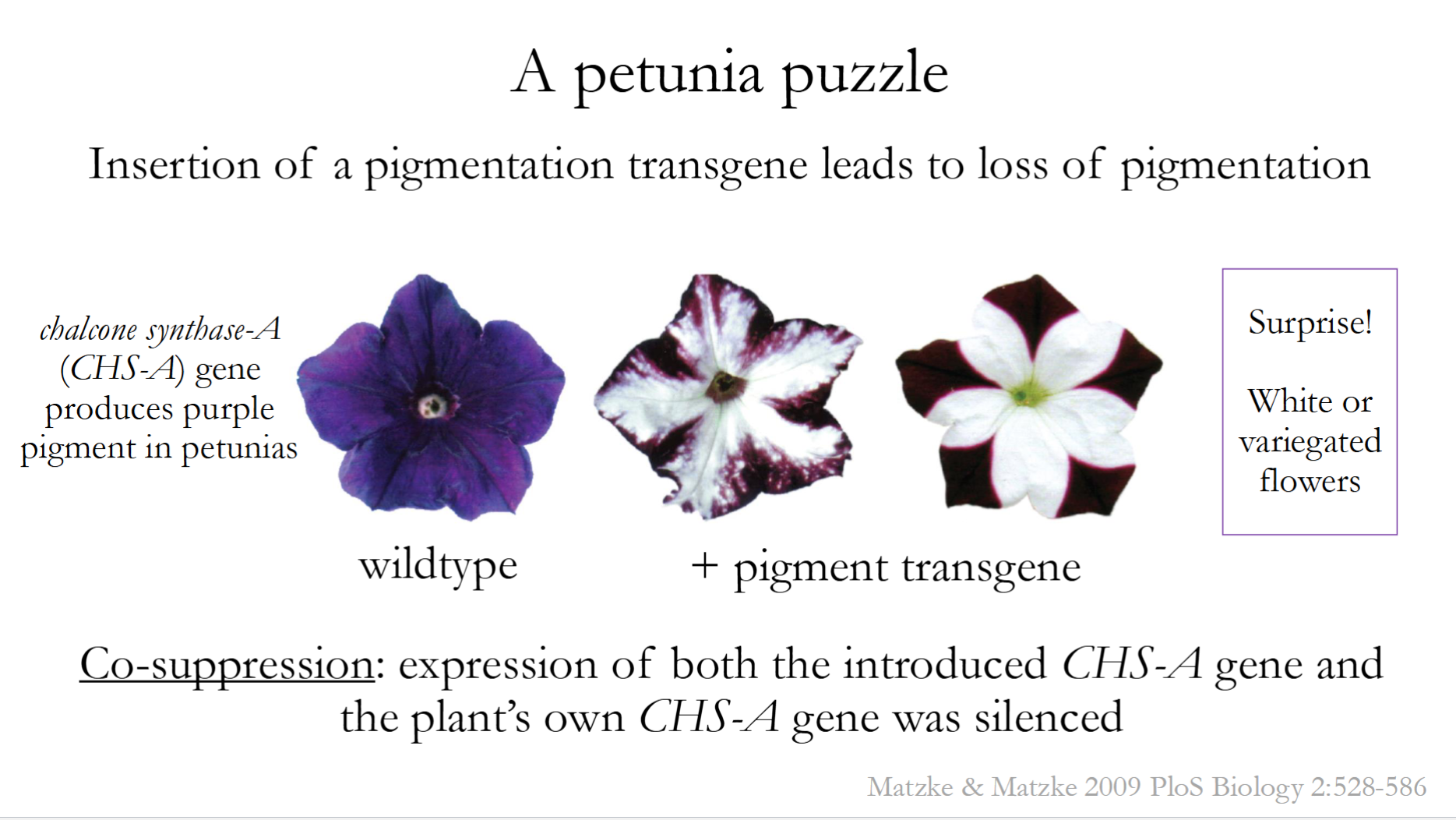
What is Co-Suppression?
A gene-silencing phenomenon where introducing an extra transgenic copy of a gene causes both the transgene and the endogenous (native) gene to be silenced.
Caused by RNA Interference (RNAi)
Famously seen in petunias (Insertion of a pigmentation transgene leads to loss of pigmentation)
What is RNA Interference (RNAi)?
RNAi is a way to directly target RNA for degradation via the RNA-Induced Silencing Complex (RISC)
What does the RNA-Induced Silencing Complex (RISC) do?
RISC is able to bind to a specific RNA sequence that is complementary to its own single stranded RNA fragment (either miRNA or siRNA) and then use its Argonaute unit (Ago) to cleaves the target mRNA, which is then degraded.
How are siRNAs made? How do they interact with RISC?
siRNAs are produced from double-stranded foreign (typically virus) RNA
Endonuclease DICER cuts up foreign RNA into fragments to generate siRNA (19-28bp)
siRNAs are incorporated into RISC
Ago 2 unit discards the passenger strand and keeps the Guide Strand which is methylated for stability
RISC then cleaves target RNA that is complementary to Guide RNA (this helps to protect the cell from infections)
binding to target RNA must be tight
What is the DICER enzyme?
DICER cuts up double stranded RNA.
Made of several subunits:
RNaseIIIa and RNaseIIIb make up the catalytic domain (do the cutting)
“Ruler” helix & PAZ domain (which binds the 3’ end of foreign RNA), the ruler determines length of the fragment (19-28bp)
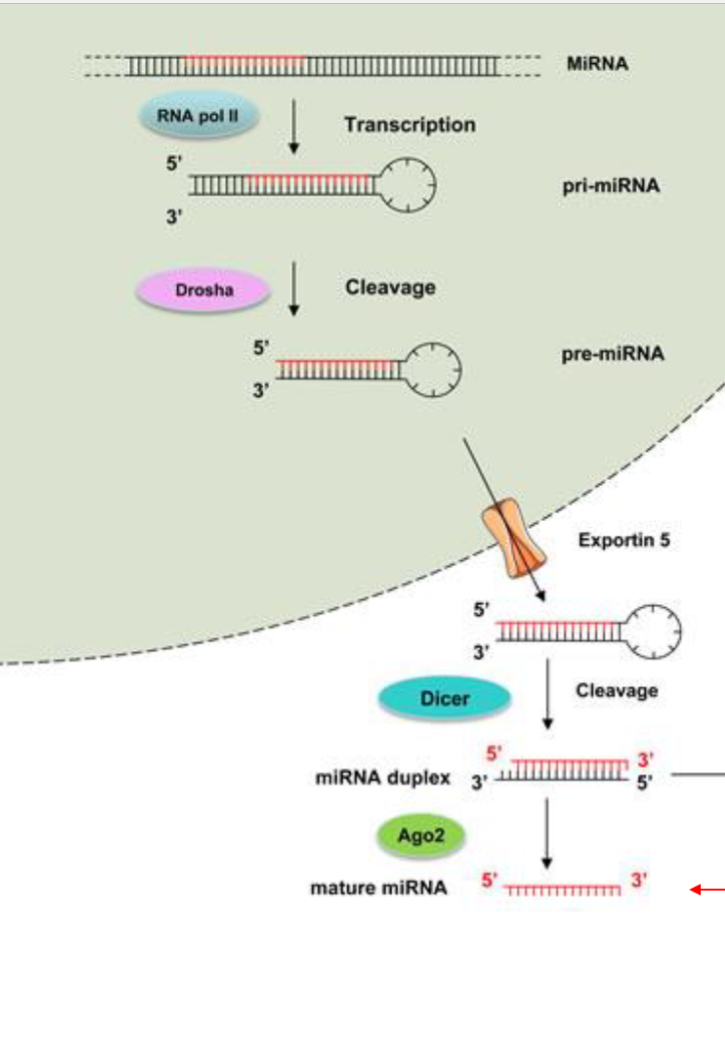
How are miRNAs made? How do they interact with RISC?
miRNAs are synthesized in the nucleus and processed before they can be apart of the RISC complex.
1) miRNAs are synthesized by RNA Pol II (Genomicaly, miRNA genes can be found in many places, in introns, between mRNA genes, in clusters, ect…) and form a hairpin loop structure (miRNA loops in on itself), this is pri-miRNA
2) Drosha (an endonuclease) trims pri-mRNA into pre-miRNA (just makes it shorter)
3) pre-miRNA is exported out of the nuclease by Exportin-5
4) DICER then cleaves of the loop (now we just have a double stranded miRNA segment)
5) Ago2 discards the passenger strand and keeps the Guide Strand (which is methylated for stability)
Ago2, miRNA segment (and a few other proteins) now make the RISC complex
RISC complex now searches for complementary target mRNA
RISC will either cleave mRNA or bind to it to block transcription (will eventually be degraded either way)
binding to complementary mRNA can be lose
Why do RNAi pathways exist?
RNAi likely evolved as an ancient antiviral defense system
note, Viruses have diverse strategies for dealing with RNAi (this suggests it is a major selective force in host-pathogen evolution)
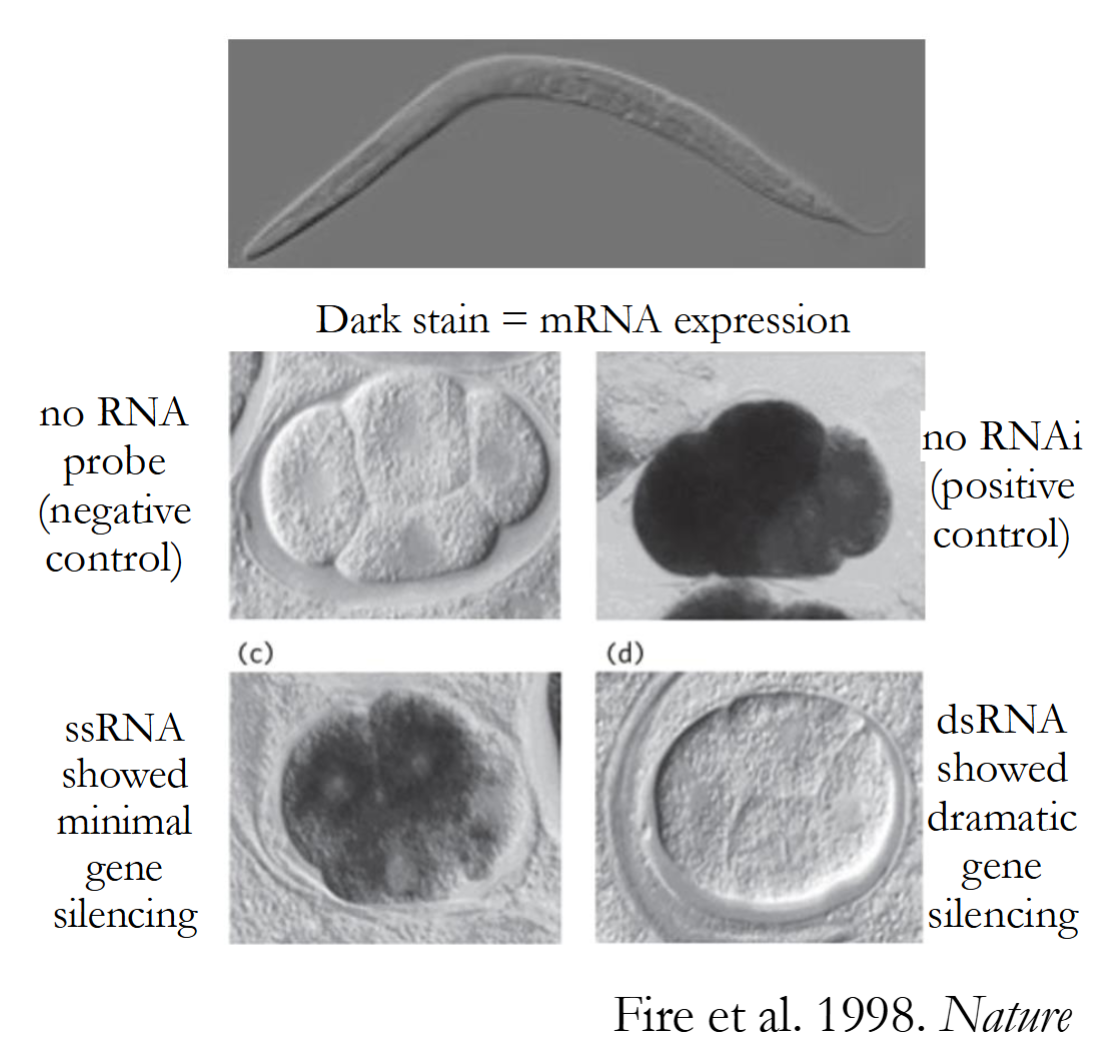
What was the Mello and Fire experiment?
Craig Mello and Andrew Fire studied RNAi in C. elegans nematode worms.
1) They introduced the dsRNA (double stranded RNA) and the ssRNA (Single Stranded RNA) into the worms, these were the treatments.
2) They found that dsRNA was better at triggering gene silencing (this is because DICER cuts up double stranded RNA to make siRNA, which is then incorporated into RISC), while ssRNA did not show much gene silencing
How do RNAi-based therapeutics generally work? What is an example of RNAi based drug?
Use the cell's natural gene silencing machinery to selectively "turn off" disease-causing genes.
Typically use Lipid Nanoparticles as vesicles to transfer desired material (Ex: synthetic siRNA) into cells.
Patisiran is an RNAi based drug to treat Hereditary Transthyretin Amyloidosis. (mutations in TTR gene cause protein misfolds and aggregates)
uses siRNA lipid nanoparticles
siRNA incorporated into RISC and then targets 3’ UTR of mutant TTR mRNA to prevent translation
What are the four key components of a Lipid Nanoparticle membrane?
1) Ionizable Lipid: facilitates endosomal escape
2) Cholesterol: Enhances stability
3) Helper Lipids: Promotes stability and fusogenicity (promotion of the fusion of two membranes)
4) PEG-Lipids: synthesized, provides stealth coating
What is the Xist non-coding long RNA? What does it do?
Xist Long RNA is a long transcript (>17kb) that remains in the nucleus and is not translated. It is involved with X chromosome inactivation in female mammals.
in mammals, one of two X chromosomes in female cells is epigenetically silenced.
This RNA molecule coats that X chromosome, a scaffold to recruit silencing proteins.
What is Sanger Sequencing? How is it similar to PCR?
1) Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides (ddNTPs) by DNA polymerase during in vitro DNA replication.
method requires a single-stranded DNA template, a DNA primer, a DNA polymerase, normal dNTPs, and modified ddNTPs, the latter of which terminate DNA strand elongation.
2) Similar to PCR in that it uses a DNA template stand which is heated to separate, primers, which are annealed to the strand, and synthesis by DNA Polymerase.
ddNTPs vs. dNTPs
ddNTPs: Dideoxynucleotide triphosphate, missing 3’ OH group (only has H)
prevents elongation
dNTPS: Deoxynucleotide triphosphate, have normal 3’ OH group
How does Automated Sanger Sequencing work? Who invented it?
1) uses fluorescent labeled ddNTPs to allow combined reactions and visualization by a computer.
2) Lee Hood
How do we most commonly work with DNA in vitro and in vivo
In Vitro: Amplification by PCR
In Vivo: Synthesis plasmid, put plasmid in bacteria, grow bacteria and isolate DNA.
What is a plasmid? What does a plasmid vector need? How do we use it?
1) Naturally occurring (in bacteria, ect..), circular, extrachromosomal DNA molecule that is a non-essential gene carrier.
2) Good synthetic plasmid vectors have three things:
Replication origin
Indicator tag on gene of interest (Ex: Green florescent protein)
Positive selection trait (usually antibiotic resistance. Makes sure only the bacteria that successfully uptake the plasmid survive)
3) Use plasmid vectors to create transgenic organisms and create DNA libraries. (among other things)
How could you sequence C. elegans DNA using plasmids?
1. grind up worms to get their DNA
2. make plasmids with worm DNA
3. sequence plasmids (shotgun sequencing)
What are two notable second generation DNA sequencing techniques? How do they work?
454 Roche Pyrosequencing
Uses Luciferase (from fireflies) to produce light each time a nucleotide is added.
Occurs in wells on a fiber optic chip so computer can see and measure the flashes (take a picture, did it flash? wash).
Solexa/ Illumina
DNA fragments are bound to a slide and amplified in situ.
Their “flow cell” allows lots of fragments to be sequenced in parallel.
Extended with blocked fluorescent nucleotides (each base is a different color), so only one base can be added at a time.
Image the slide, the color of each colony reports the base that was added last
What are Transposons?
Transposons are “jumping genes”, DNA sequences that can change their location in the genome
Transposons and other repetitive DNA elements can pose a problem to short-read DNA sequencing techniques
What did Barbara McClintock discover?
She discovered Transposons while doing experiments with maize (1950)
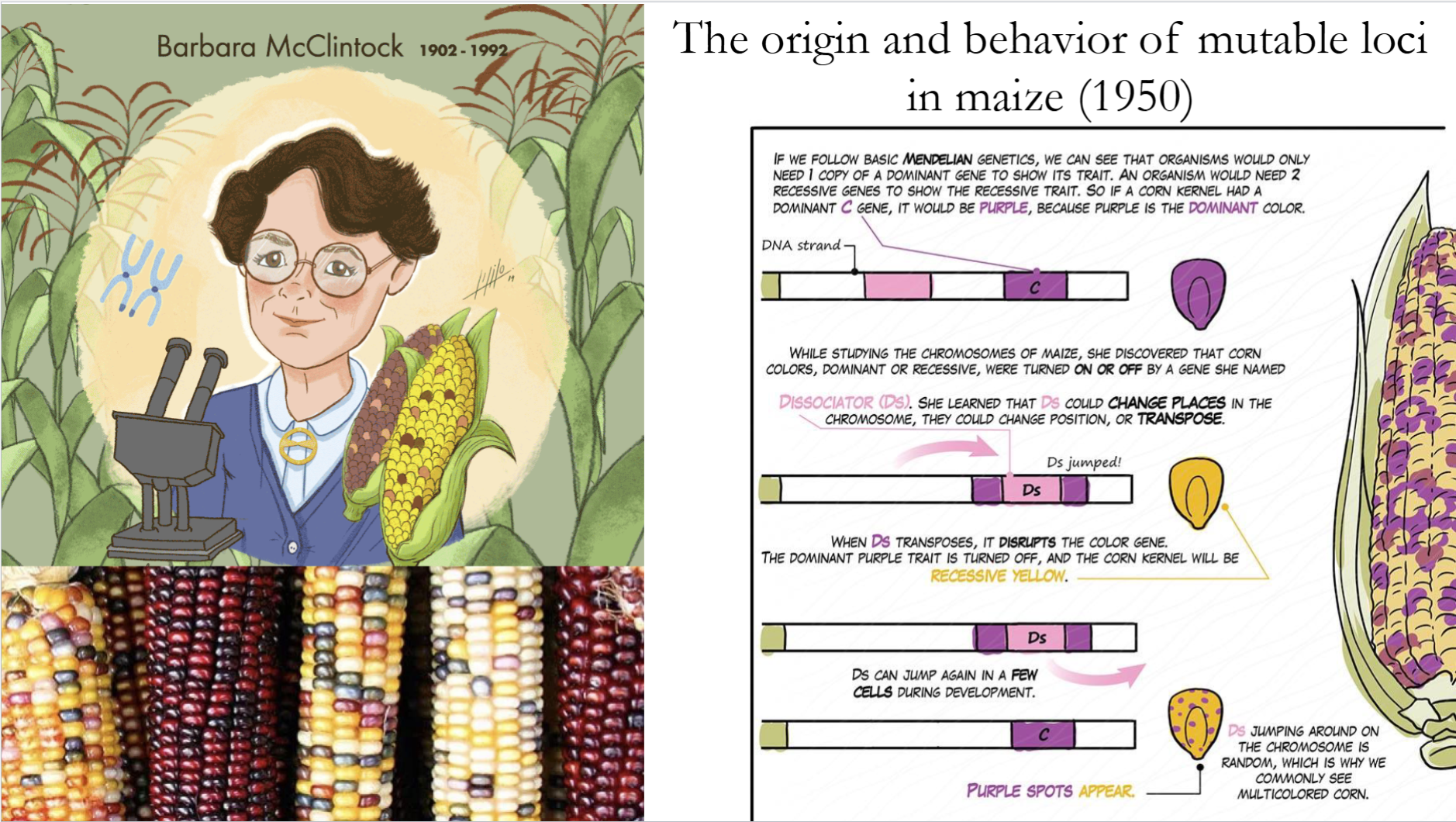

What is the likely evolutionary origin of Transposons?
Transposons are descendants of ancient viruses
What are two notable third generation (long-read) DNA sequencing techniques?
Pacific Biosciences (PacBio)
A single immobilized DNA molecule, 1 anchored polymerase
As polymerase incorporates labeled bases (4 different colors), light is emitted
Light is measured in real time (directly detect DNA elongation during sequencing)
Oxford Nanopore Technologies (Nanopore)
Tiny protein pores embedded in a synthetic memebrane
As nucleotides pass through pore they disrupt an electrical current
Each nucleotide leaves a specific electrical signature that is read by a computer.
What is a Transgenic Organism?
An organism who’s DNA has been altered by the introduction of genes from another organism
What are the two main functions of Transgenic genes?
1) As reporter genes (Ex: GFP)
2) To introduce new functions (Ex: Herbicide resistance for crops)
How can you make Transgenes heritable?
Germ line cells must be affected
Genes must be inserted into chromosomes
How do you get a Transgene into a cell in Prokaryotes and Eukaryotes?
Prokaryotes:
Electroporation (Electric shock)
Calcium Chloride Chemical Transformation (Heat shock)
Eukaryotes
Viral Transduction (Uses a virus shell to deliver desired transgene)
Lipid Nanoparticles/ Liposomes
Sometimes diet (nematodes)
DNA microinjections
What is signal transduction?
Extracellular signals leading to changes inside the cell.
The same cell signal can result in different responses from different cell types. How is this possible?
Response is based on the target cells receptor/effector proteins (different cell types have different type of receptors and responders)
Combinations of cell signals regulate the cell’s response
How can some signaling molecules permeate the cell membrane while others cannot?
To permeate the cell membrane (which is typically done by facilitated transport or just diffusion), molecule must be:
Must be relatively small
Must be nonpolar (lipid soluble)
(Ex: Steroid hormones (derived from cholesterol), CO2, ect…)
Molecules that can NOT permeate the membrane:
Large molecules
ions
polar molecules
What kinds of signaling molecules will bind to Nuclear Receptors?
Signaling molecules that can pass through the cell membrane, may travel directly to the nucleus and bind to nuclear receptors. (Ex: Steroid hormones)
Once a nuclear receptor is activated, it will now act as a Transcription factor to regulate gene expression.
Theses signals are relatively slow but can have big affects.
How can intracellular signaling pathways be regulated?
1) Feedback regulation:
Positive feedback
negative feedback
2) Switch Proteins:
Signals can turn proteins in the cell on/off
Two major classes of switch proteins (Signaling by Phosphorylation, and Signaling by GTP-Binding Proteins)
What are G-Protein Coupled Receptors (GPCRs)?
Single polypeptide chain that crosses the cell membrane 7 times
Largest family of cell-surface receptors
mediate mot external cell signaling (smell, sight, sound, ect..)
Mediate immune cell migration.
A variety of signal factors act on GPCRs (hormones, Neurotransmitters, local mediators; half of our drugs work through GPCRs and their pathways)
How do GPCRs work?
GPCRs use G-proteins to relay signals into cell interior
G-proteins are mobile, activated by GTP (Guanosine Triphosphate) (GDP is replaced by GTP)
Active G-protein alpha subunit (has GTP) then activates some effector protein
What are the three G-protein subunits? Which one holds GDP/GTP?
1) Alpha subunit holds GDP/GTP (GTP is the active form)
2) Beta subunit
3) Gamma subunit
What are two toxins that can effect the activity of G-proteins?
Cholera toxin
modifies the G-protein α subunit, prevents G-protein inactivation (GTP can’t un-bind, G-protein always on).
Leads to dehydration and death via diarrhea
toxin is caused by Vibrio cholerae
Pertussis toxin
modifies the G-protein α subunit, prevents G-protein activation (GDP can’t un-bind, G-protein is always off)
leads to Whooping Cough (caused by whooping cough bacteria producing toxin in the lungs)
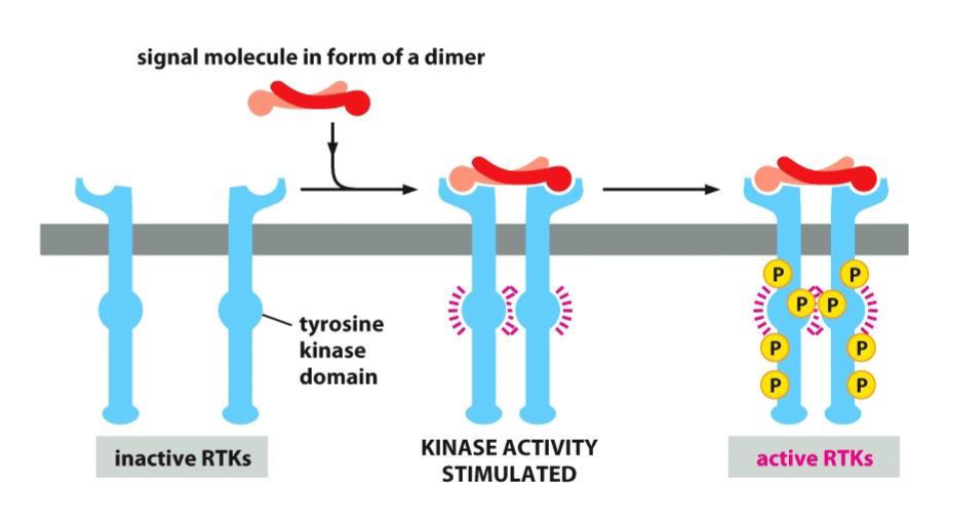
How do Receptor Tyrosine Kinases (RTK) work?
Inactivated two tailed RTKs are activated and brought together by a ligand binding and causing dimerization.
When the tails are associated with each other, tyrosines get phosphorylated by kinases.
Now act as “docking sites” for signaling proteins and set off signaling pathways.
Can GPCR and RTK signaling pathways interact with each other?
yes