Exam 3 - Molecular Genetics
1/159
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
160 Terms
location of DNA in prokaryotes
nucleoid (no membrane-bound organelles)
location of DNA in eukaryotes
nucleus (separates transcription and translation)
location of ribosomes in prokaryotes
cytoplasm
location of ribosomes in eukaryotes
endoplasmic reticulum (ER) and cytoplasm
how are genes organized in prokaryotes?
in operons - close together
how are genes organized in eukaryotes?
can be far apart - no operons
protein coding content on chromosome in prokaryotes
entire gene is coding
protein coding content on chromosome in eukaryotes
introns are removed and exons are coding
pre-translation mRNA processing in prokaryotes
minimal - some trimming and phosphates may be added to 3’ end
pre-translation mRNA processing in eukaryotes
splicing of exons
5’-cap (methyl group added to guanine)
3’-poly(A) tail
are transcription and translation coupled in prokaryotes?
yes they are
are transcription and translation coupled in eukaryotes?
no they are not
bacterial operon
clusters of co-regulated (all on or off) genes with related functions
why is there a correlation between transcription and translation in prokaryotes?
ribosomes start translating mRNA before transcription even finishes, so the rate of one process directly influences the other
What happens to transcription when translation is quick?
This ribosome movement blocks Rho-dependent termination, allowing transcription to continue (positive correlation)
What happens to transcription when translation is stalled or slow?
Naked mRNA is exposed, so Rho binds and prematurely terminates transcription (negative correlation)
In bacteria, what molecule causes premature termination when translation slows?
Rho factor
promoter
DNA sequence (100-1000 bp long) upstream of structural genes - where RNA polymerase will bind to initiate transcription
operator
region of DNA between promoter and structural genes where a repressor (regulatory protein) will bind (stops transcription)
regulatory genes
encode transcription factor proteins that influence transcription rate of the operon
repressor
turns off transcription in response to external stimuli
activator
increases transcription in response to external stimuli
What 3 regions of the promoter does RNAP recognize and bind to?
-35 region, -10 region (TATA/Pribnow box), +1 region (start site)
Which RNAP subunit interacts with the -10 and -35 promoter regions?
sigma
What happens to the sigma subunit of RNAP once transcription begins?
It falls off the core RNAP.
Beta subunit of RNAP
Has the main catalytic domain - separates DNA strands and polymerizes RNA synthesis
Beta’ subunit of RNAP
helps stabilize the DNA-RNA hybrid during transcription
Alpha subunits of RNAP
involved in assembly and stability of RNAP transcription complex
Omega subunit of RNAP
stabilizes core subunits, essential for holoenzyme assembly
Sigma subunit of RNAP
essential for promoter recognition and binding (not part of core)
Transcription - Step 1 - Initiation
sigma subunit binds to core RNA polymerase subunits
sigma subunit binds to promoter
beta subunit separates DNA strands at start site, creating a transcription bubble
attaches first nucleotide of RNA (A or G)
Transcription - Step 2 - Elongation
Beta subunit adds nucleotides to growing mRNA strand
sigma subunit is displaced by RNA strand and falls off.
Core RNAP subunits move in 5’-3’ direction
beta’ stabilizes DNA/RNA hybrid
alphas bind to upstream region of DNA (UP element at -50) to help stabilize complex
Transcription elongation complex (TEC)
DNA enters pincers of beta and beta’
Free nucleotides enter through beta secondary channel
beta active site = site of polymerization
RNA exits through beta exit channel
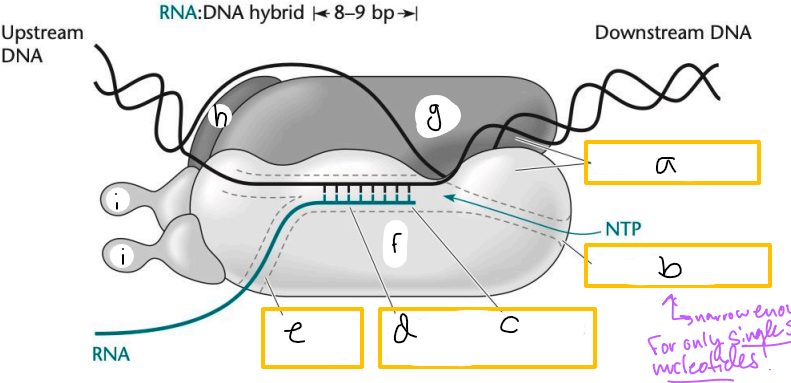
a? (TEC)
downstream jaws
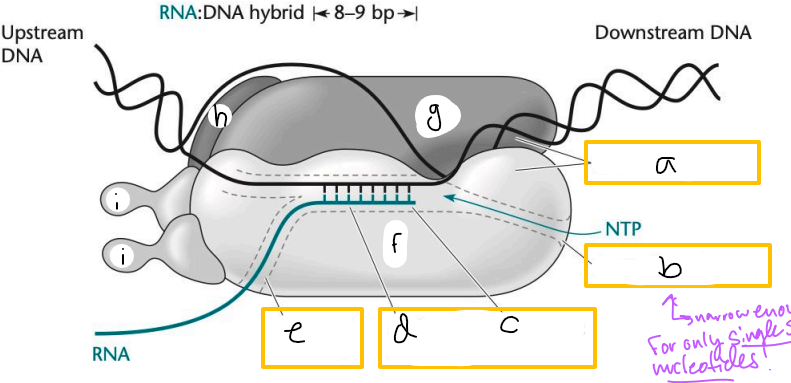
b? (TEC)
secondary channel
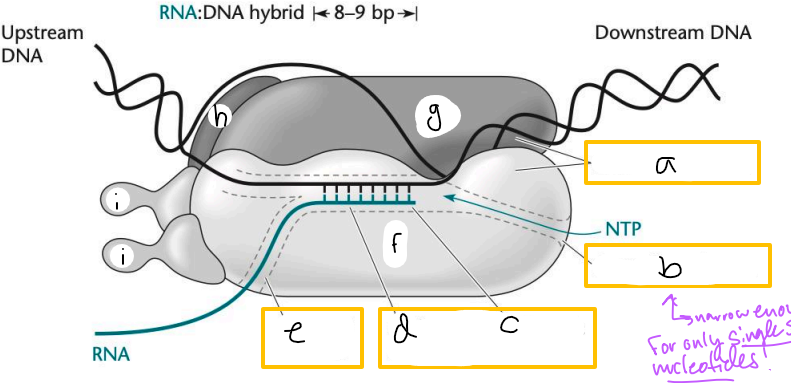
c? (TEC)
active site
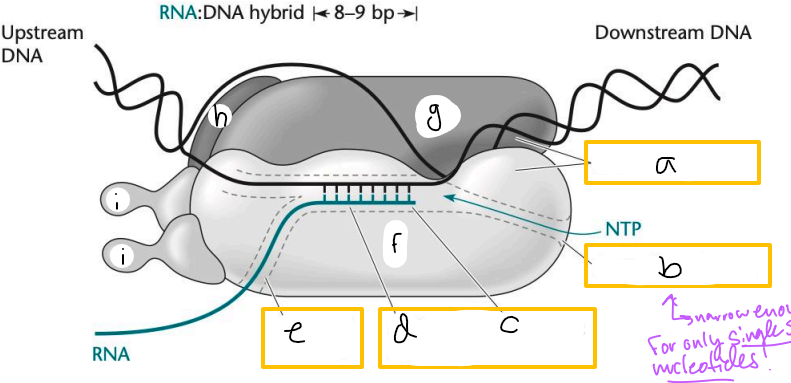
d? (TEC)
active site channel
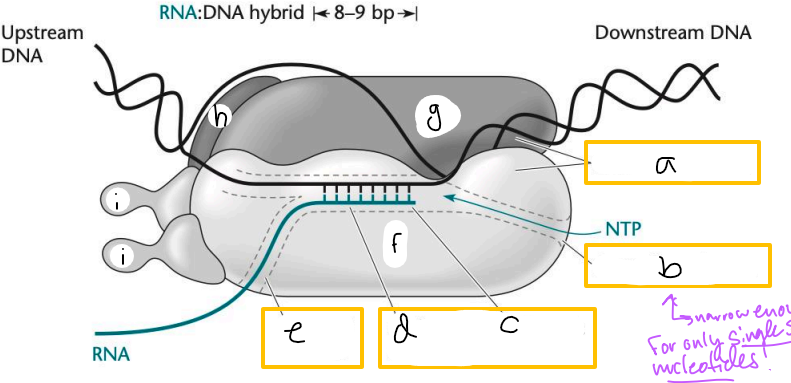
e? (TEC)
RNA exit channel
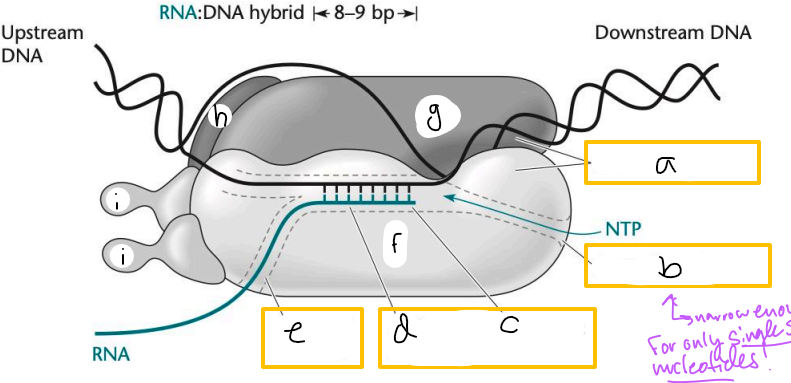
f? (TEC)
beta subunit
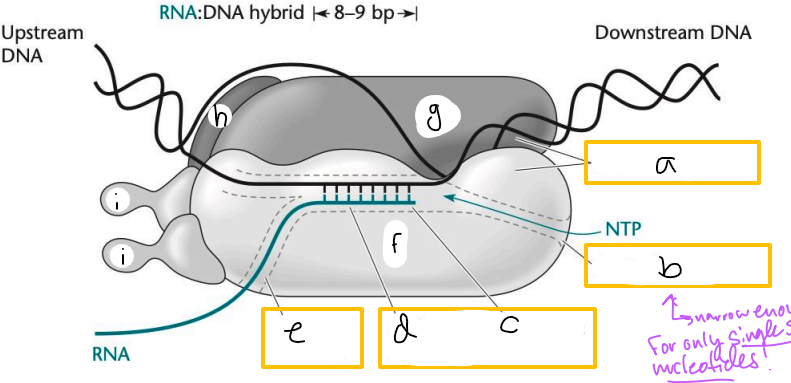
g? (TEC)
beta’ subunt
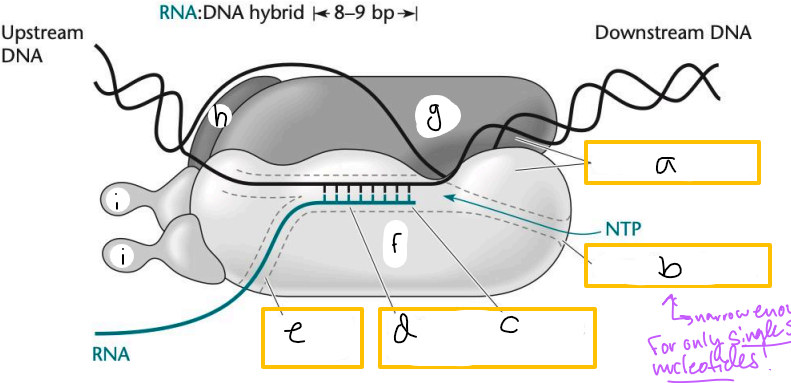
h? (TEC)
omega subunit
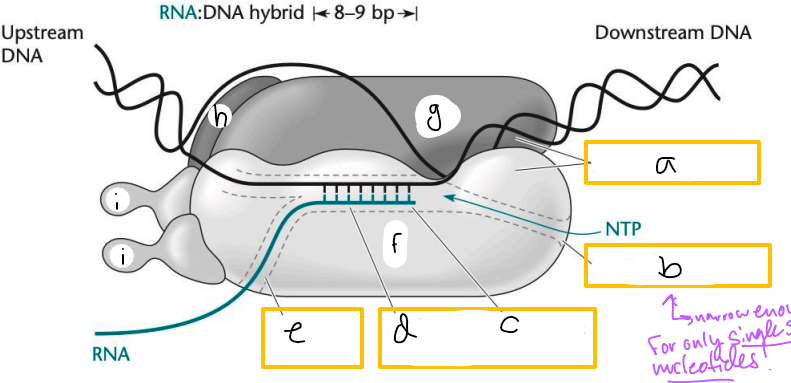
i’s? (TEC)
alpha subunits
Which strand is the template strand that RNAP creates a compliment of?
the 3’-5’ strand
Which strand is the RNA product identical to with U’s replacing T’s?
the coding strand (5’-3’)
Transcription - Step 3 - Termination
polymerization continues until RNAP encounters terminator region of DNA
RNAP pauses after transcribing string of Us
Inverted G-C repeats bind to each other and form hairpin
Hairpin causes RNAP to dissociate
what site does the antibiotic rifampin bind to?
the beta subunit of RNAP, just before the active site
how does rifampin inhibit transcription?
only allows 2-3 nucleotides to be incorporated into nascent mRNA before physically blocking further transformation
what is the effect of a sigma factor mutation?
RNAP may not bind to the promoter correctly, decreasing transcription initiation
what is the effect of a mutation in the beta subunit of RNAP?
can reduce transcription elongation efficiency or cause resistances to antibiotics like rifampin
how do promoter mutations impact transcription?
change how strongly RNAP can bind, altering transcription levels (strong vs weak).
RNAP moving from left to right uses the ______ strand as template
bottom
RNAP moving from right to left uses the ___ strand as template
top
why are transcription errors 3-4 orders of magnitude higher than DNA replication errors?
RNAP does not have proofreading abilities, unlike DNA pol.
polycistronic
many genes in one mRNA (prokaryotic)
monocistronic
one gene per mRNA/transcript (eukaryotic)

Messenger RNA (mRNA)
coding; encodes the amino acid sequence of a protein
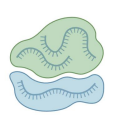
Ribosomal RNA (rRNA)
non-coding; ribozyme; make up the ribosome, translates mRNA to proteins
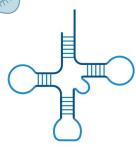
Transfer RNA (tRNA)
non-coding; ribozyme; read mRNA sequence to deliver amino acids during translation

Micro RNA (miRNA)
non-coding; regulatory RNA; inhibits mRNA translation as a form of regulating gene expression

Small interfering RNA (siRNA)
non-coding; regulatory RNA; selectively degrades mRNA as a form of regulating gene expression (RNAi pathway)
ribozyme
RNA molecules with catalytic domains that act similar to enzymatic protein domains
RNA primary structure
single stranded chain of ribonucleotides
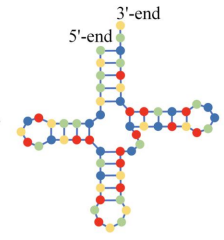
which RNA structure is this?
secondary (nearby areas of single RNA bind together) (hairpin)
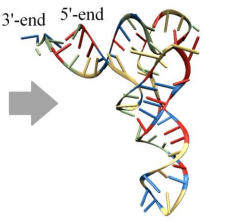
which RNA structure is this?
tertiary (more distant areas of an RNA strand interact)
What type(s) of bonds do DNA and RNA share?
hydrogen bonds and phosphodiester bonds
structure of DNA?
double helix, antiparallel, supercoiling
structure of RNA?
varied complex tertiary structures
information/function of DNA
coding/non-coding, carries genetic information
information/function of RNA
coding = mRNA, also has varied non-coding such as ribozymes and regulatory RNA
________ are essential for transcription termination
hairpins
In RNA, what can guanine non-canonically pair with?
uracil
What is a pseudoknot?
an RNA tertiary structure; creates 3D structure of RNA and is especially important for function of non-coding RNAs
what are the types of prokaryotic RNA modifications?
methylation
trimming
tRNA modifications
methylation of prokaryotic RNA
adds methyl (CH3) groups
increases RNA stability
can impact translation efficiency
can induce structural changes
trimming of prokaryotic RNA
ribonucleases (RNAses) make cuts in mRNA
can influence final RNA structure
cuts non-coding RNAs out of longer RNA transcripts
which bacterial RNA is most heavily modified?
tRNA
prokaryotic tRNA modifications
tRNA is cut from a longer mRNA transcript
2-40 modifications made depending on bacterium
what types of base modifications occur in tRNAs and why?
bases can be modified to dihydrouridine or pseudouracil to help tRNA fold correctly and function in translation
eukaryotic 5’ cap mRNA modification
capping enzyme adds 5’ 7-methylguanosine cap to prevent RNA degradation
eukaryotic RNA splicing
spliceosome proteins bind to 5’ splice site, cleaves intron-exon junction, and ligates exons.
can create isoforms of the same gene leading to different functions
eukaryotic addition of 3’ poly(A) tail
polyadenylation factor syntheses the 3’ poly(A) tail on mRNA to increase stability
mRNA life span in prokaryotes
1-5 min
mRNA lifespan in eukaryotes
minutes to hours
Why does RNA degradation happen?
the cellular concentration of mRNA helps govern the level of gene expression
degradative pathways ensure mRNAs do not build up in the cell and make proteins that are not needed
ribonucleases (RNases) in RNA degradation
cleave sugar/phosphate backbone of RNA
endoribonucleases in RNA degradation
cleave backbone within an RNA strand (remove larger pieces, interior cleavage)
exoribonucleases in RNA degradation
cleave off one nucleotide at a time from ends of RNA strand (removes ends)
prokaryotic pathway of mRNA degradation
endoribonuclease cleaves mRNA into piece with hairpin structure at 3’ end and a piece without the hairpin structure (open 3’ end)
mRNA piece with open 3’ end gets degraded rapidly by exoribonucleases
mRNA piece with hairpin (blocks exoribonuclease) either:
gets cut again by endoribonuclease
degraded by 5’-3’ exoribonuclease
undergoes polyadenylation then is degraded by 3’-5’ exoribonucleases
What is a direct method for studying RNA?
when RNA is visualized or sequenced in its native state
What are the direct methods for studying RNA?
Fluorescence in-situ hybridization (FISH)
Northern Blot
Direct RNA sequencing (long-read; Oxford Nanopore / PacBio)
What is an indirect method for studying RNA?
RNA is converted to complimentary DNA (cDNA) and often requires amplification (reverse transcription)
What are the indirect methods for studying RNA?
Reverse Transcription PCR (RT-PCR)
Quantitative RT-PCR (qRT-PCR)
RNA-seq (short-read cDNA sequencing; Illumina)
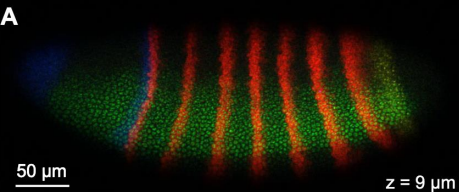

RNA Fluorescence in-situ hybridization (FISH)
Tissue is embedded in paraffin in sectioned into very thin slices that can be adhered to a slide
additionally cells can be fixed and applied to slide
Fluorescent probes are hybridized to tissue/cell sections
probes can adhere to DNA or RNA
Slides are imaged under fluorescent microscope
What can be learned from visualizing the location of RNA?
which cells express certain genes
body plan
RNA localization is key in development
Northern Blot
RNA is rn on a gel
Size-separated RNA is transferred to nylon membrane
Gene-specific radioactive probes are applied to membrane
Membrane is exposed to film. If the gene is expressed, radioactive probes hybridize to the membrane and expose the film.
What does reverse transcription do?
Creates double stranded DNA from an RNA template using a reverse transcriptase enzyme
Binds to 3’ end of mRNA and synthesizes single strand of cDNA in 5’-3’ direction
Degrades RNA strand
Synthesizes complimentary cDNA strand to form double-stranded DNA
How does RT-PCR work?
RNA is extracted and treated with DNase (to prevent DNA amplification) (semi-quantitative)
cDNA synthesis - mix:
oligo d(t) primers (eukaryotic) or random primers (prokaryotic)
reverse transcriptase
RNA
regular PCR proceeds with cDNA and gene-specific primers
PCR product is visualized on agarose gel. Can estimate abundance of RNA by comparing band brightness to a control, highly expressed gene.
What is quantitative RT-PCR?
A tool that uses fluorescence to ‘count’ how many copies of RNA are amplified during each round of PCR
How does quantitative RT-PCR (qRT-PCR) work?
cDNA is mixed with PCR reagents (SYBR or gene-specific fluorescent probes (more expensive))
qPCR cycler takes a fluorescent image after each cycle
Fluorescence is graphed from the images as an amplification curve
Ct or Cq value = cycle at which fluorescence raises above the background
Expression should always be normalized against a housekeeping gene
Can RNA be used as a template in PCR?
No, would need to make cDNA first. DNA polymerase cannot use RNA.