Bacteriology and Oral Diseases; pptx 9
1/93
Earn XP
Description and Tags
Dr. cugini slides
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
94 Terms
who coined the term microbiome?
Joshua Lederberg in 2001
microbiome, as Joshua Lederberg explains it
term to signify the ecological community of commensal, symbiotic, and pathogenic microorganisms that literally share our body space and have been all but ignored as determinants of health and disease
how was the microbiome being to be understood and by who?
David Relman’s group published in 2005 the prokaryotic signature of the human intestinal microbiota, they brute forced analysis of the 16S rRNA from mucosal tissue and fecal samples from 3 healthy adults and this allowed them to sequence the bacteria and get a sense of what was present
The human oral microbiome paper was written and preformed by who?
Flyd E. Dewhirst
The human oral microbiome paper
collected 16S rRNA gene sequences into a web database(HOMD) and analyzed 26,043 16S rRNA gene clones and 1000+ isolates from studies of the oral microbiota to determine the relative abundance of taxa and identify novel candidate taxa
before the genome age what was the method for identification?
plating
why did microbiologists already know about the presence of microinches?
many were trained as microbial ecologists
what institution was the human oral microbiome paper/tests done in and are they still doing research on the human oral microbiome?
Forsyth institute and yes they are still preforming research
what did the human oral microbiome paper give the idea for?
it gave the idea of a full blown humna microbiome project to understand the entire human microbiome and microorganism that live in it. It was very well thought out and curated methodology to have many different researchers come together and work on the project and put the idea of the microbiome to the world
what sites on a human are heavily colonized?
Nasal
Oral
Skin
Gastro-intestinal
urogenital
commensals
normal protective flora; what is on us that can help prevent pathogens to go in from the outside and keep us healthy
what are examples of commensals of humans?
Streptococcus species
e.g strep gordonii
in high levels can help prevent others from coming in, e.g S. mutans
Staphylococcus epidermidis
is only a pathogen if it goes into the wrong place, put while on skin actually helps
pathobionts
residents of the normal flora that have the potential to cause disease; basically commensals that COULD cause disease
examples of pathobionts
porphyromonas gingivalis
streptococcus mutans
these bacteria are normally present in low levels but a change in the microbiome that makes it dysbiotic causing outgrowth, growing in number in that area which can cause disease
dysbiotic / dysbiosis
a change in the microbiota, from a healthy pattern to one associated with disease
T/F everything about pathobionts is understood
False, we are still trying to understand this class; there is the potential for strong genetic susceptibility, or host genetics might play a role, etc
what were previously characterized pathogens that are not considered pathobionts
Neisseria meningitidis
usually found in nose but can cross the BBB and cause disease in some people
Haemophilus influenzae
can be found in ear or nasal cavity but if it grows a lot can cause different infections
*luckily we have vaccines for these bacteria now
Pathogens
organisms that come in from the outside and make us sick
Examples of Pathogens
Bacillus anthracis
Pseudomonas aeruginosa
Borrelia burgdorferi
what was the initial study of NIH HMP(human microbiome project)
first taking the samples from the subjects and tested the microbial communities that were present. from there they did metagenomics through either looking at the 16S rRNA or they did WGS(whole genome sequencing) :
16S rRNA
compared it to a database of 16S rRNA sequences
this information is freely available
WGS
were allowed to make reference sequences
they did this while also sequencing the sources of strains to be able to understand the reference sequences and know what they were looking at
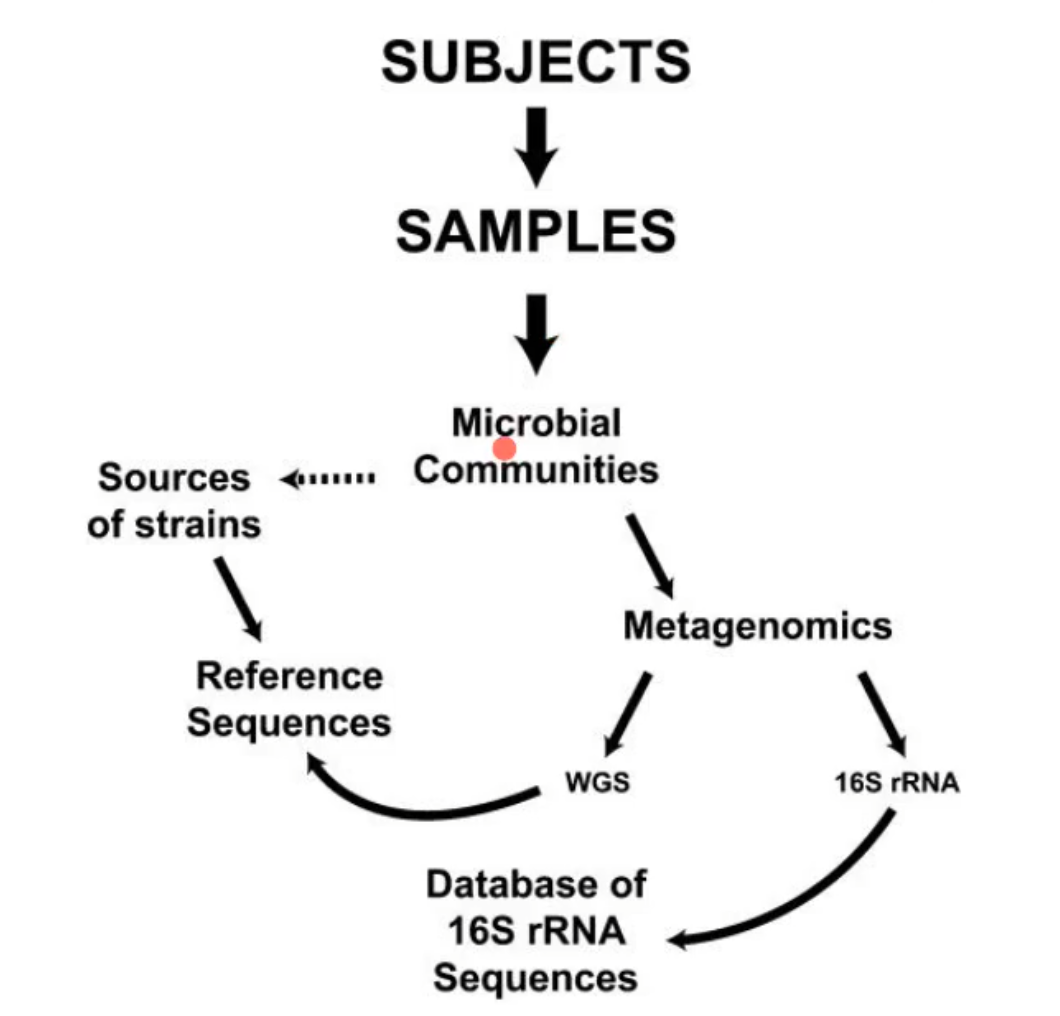
metagenomics
the study of genetic material from an entire community of organisms, often microbes, directly from an environmental sample
what were some problems and questions raised due to the HMP?
Ethical, legal, sand social implications, basically there was a lot of privacy concerns at the time
can ur microbiome be tracked back to u?
is it like a fingerprint?
if u have a disease that will normally be reported, will u get reported or is it still anonymous, etc
who owns this data?
*more specific concerns in picture

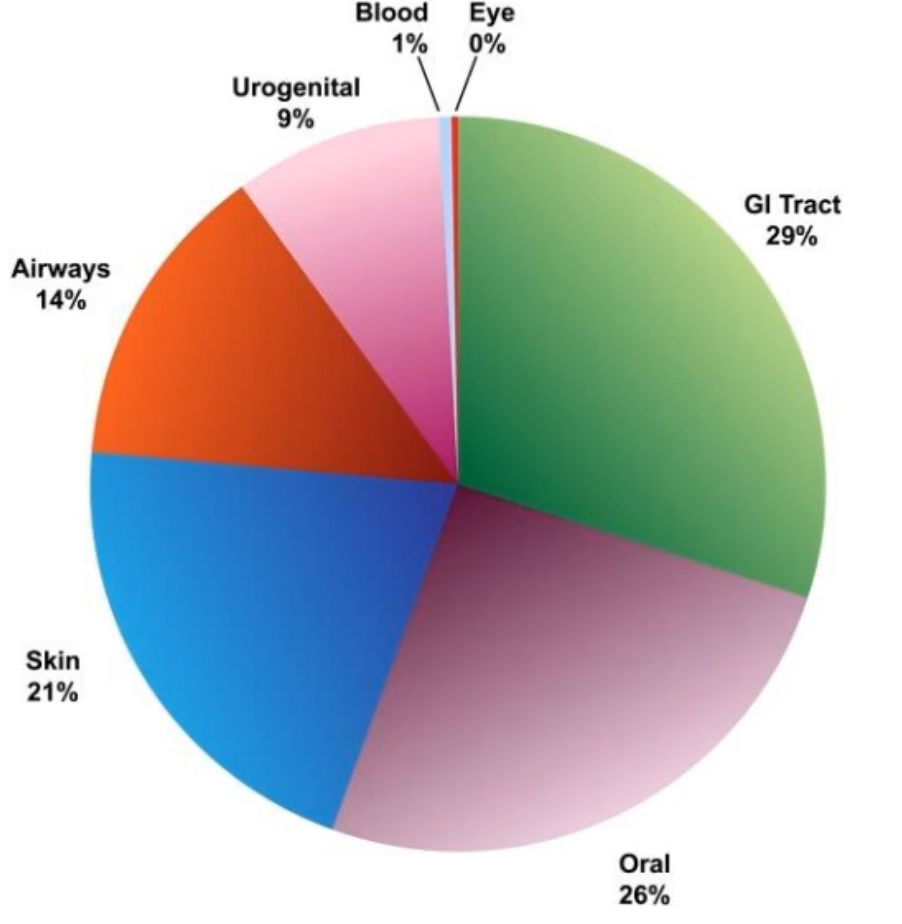
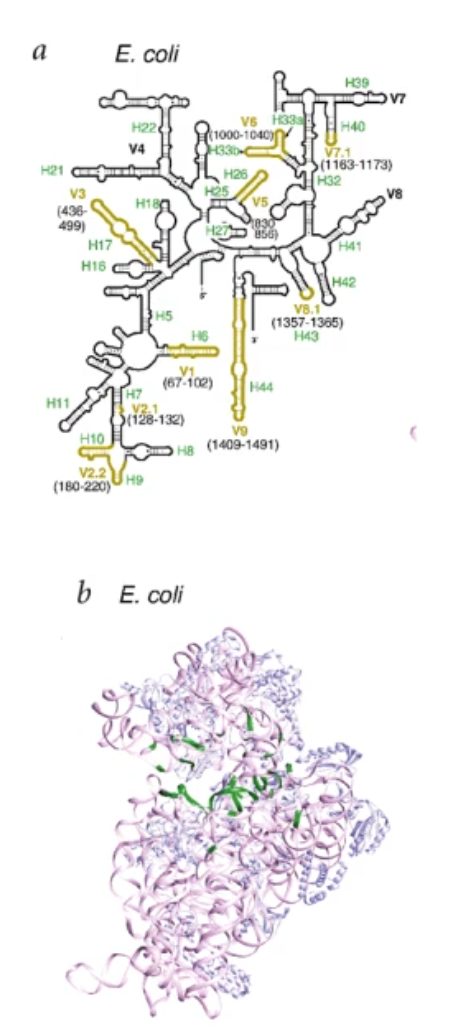
what does this figure show?
the distribution by body site of bacteria that have been sequences under the HMP r are in the sequencing pipelines
break down of bacterial distribution by body site
Eye
0%
Blood
1%
Urogenital
9%
Airways
14%
Skin
21%
Oral
26%
GI tract
26%
this fata was collected in 2009 and research is still being done but data seems to be holding true
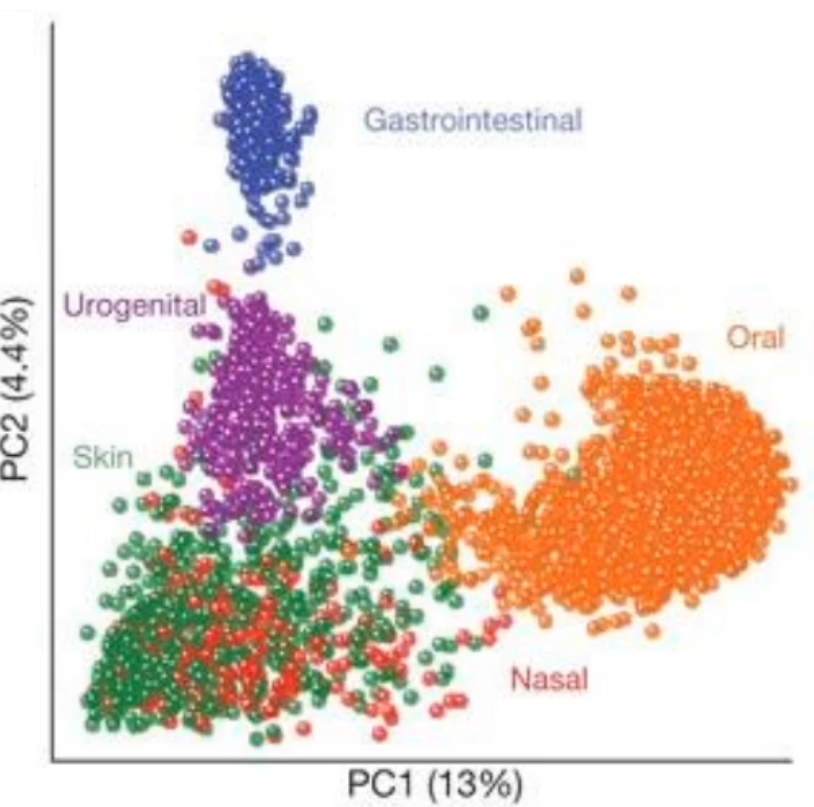
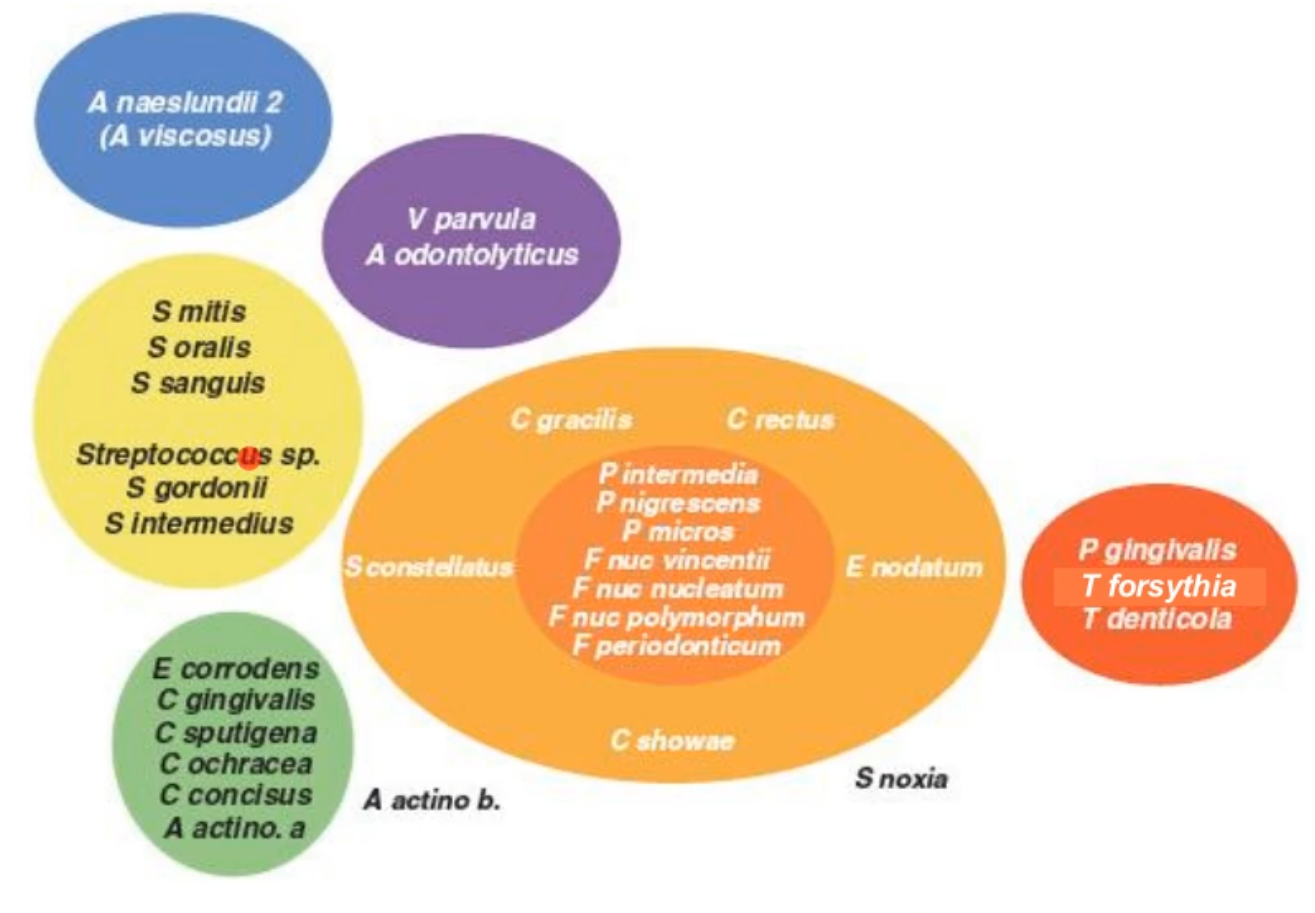
what does this diagram show/tell you?
it shows how each of these bacteria like to clump up together depending on where they inhabit. This tells us that the microbiome is unique to each induvial location and strongly determined by microbial habitat
this also means that there is not as much confusion.; of if a bacteria is (for example) a oral bacteria, or GI bacteria
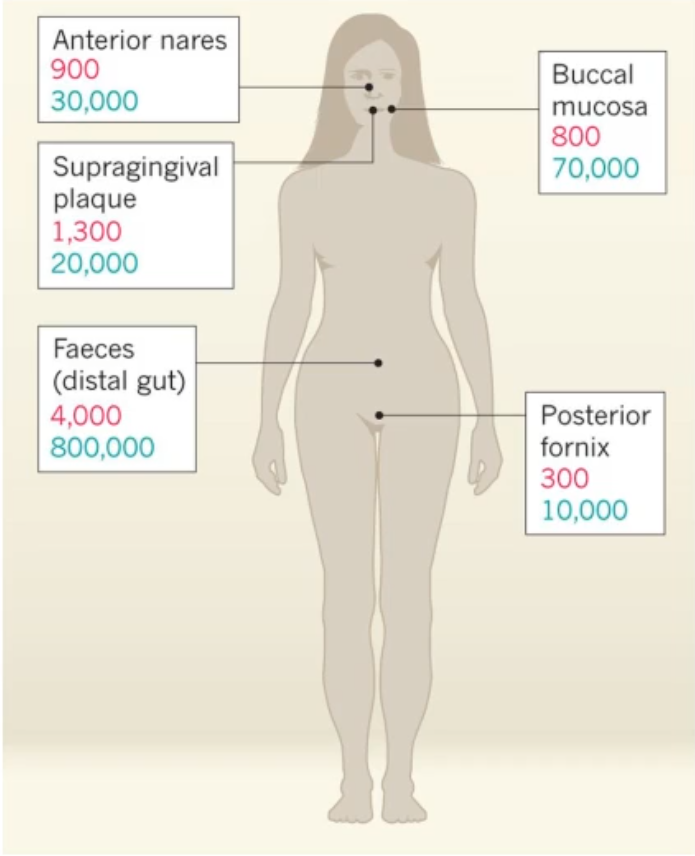
what does this show tell you?
It shows the Estimations of number of microbial species(red) and number of microbial genes(blue)
*the genes are what make the mRNA and proteins that actually do things in those sites
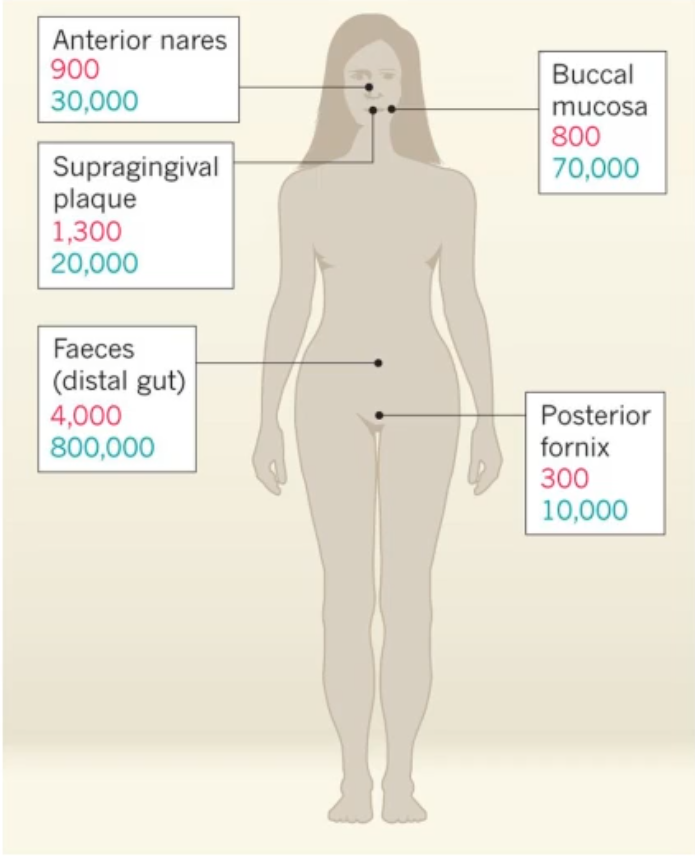
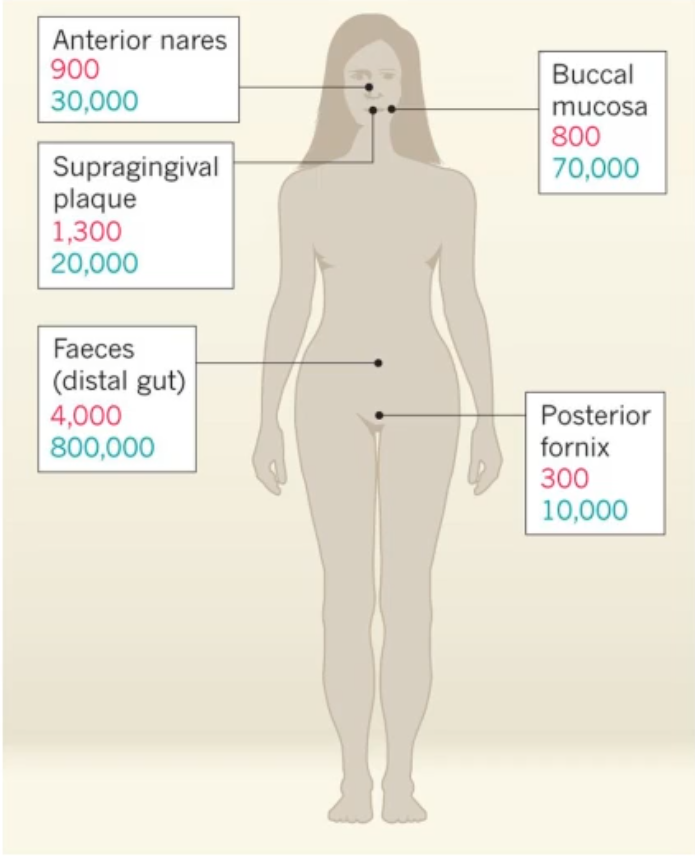
how was the data collected for this diagram and what additional details?
242 healthy adults were sampled
the highest diversity is in the gastrointestinal tract and supragingival plaque
Intermediate diversity in nose and buccal mucosa
lowest diversity in posterior fornix
*depending on how many and what genes are currently present, there are different metabolic environments present
what is something to keep in mind about the microbiomes in different locations?
you can have the carrying of varying microbial taxa(different bacteria) while metabolic pathways remain stable within a healthy population
*basically that the bacteria can change but the metabolic pathways stays stable, there are a lot of different bacteria that gives the same metabolic result; it doesn’t matter who provides the metabolic function, and the microbial niche and network of species is maintained by cooperative metabolism as the different metabolic byproducts from one organism provides nutrients for neighbors (common goods)
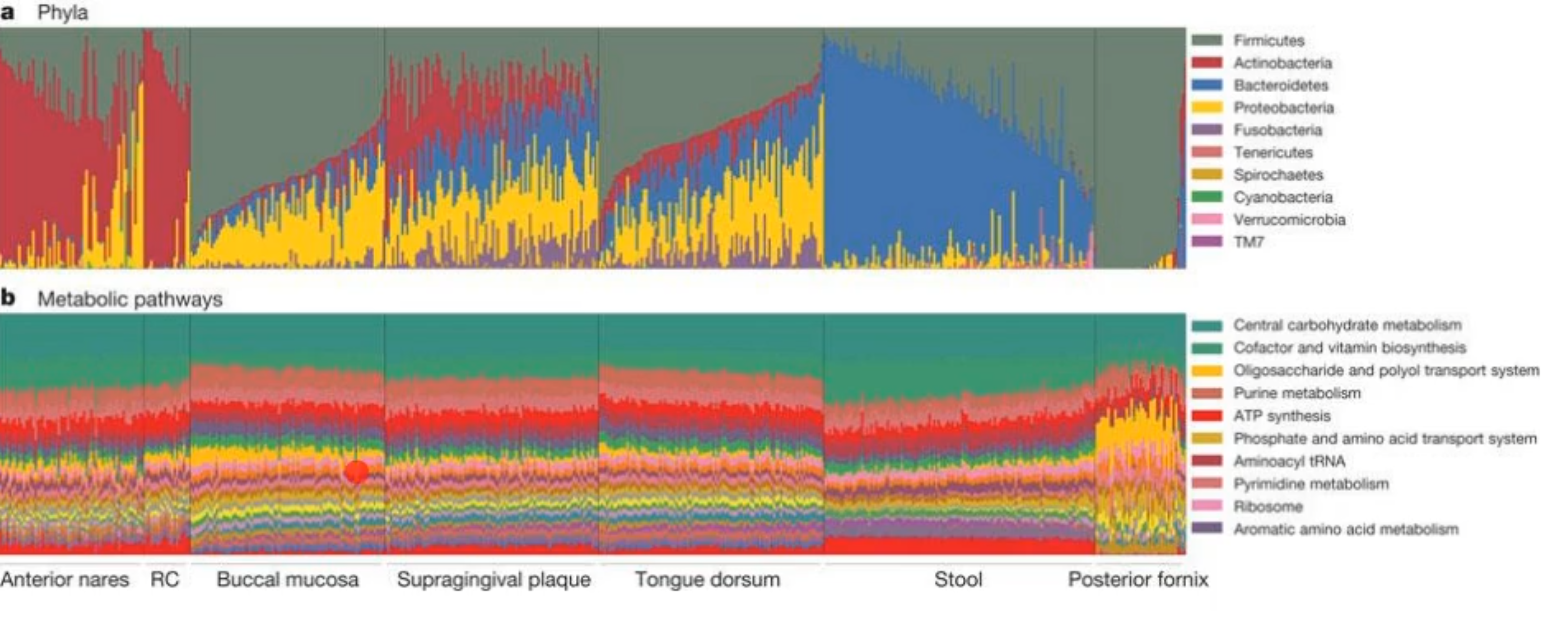
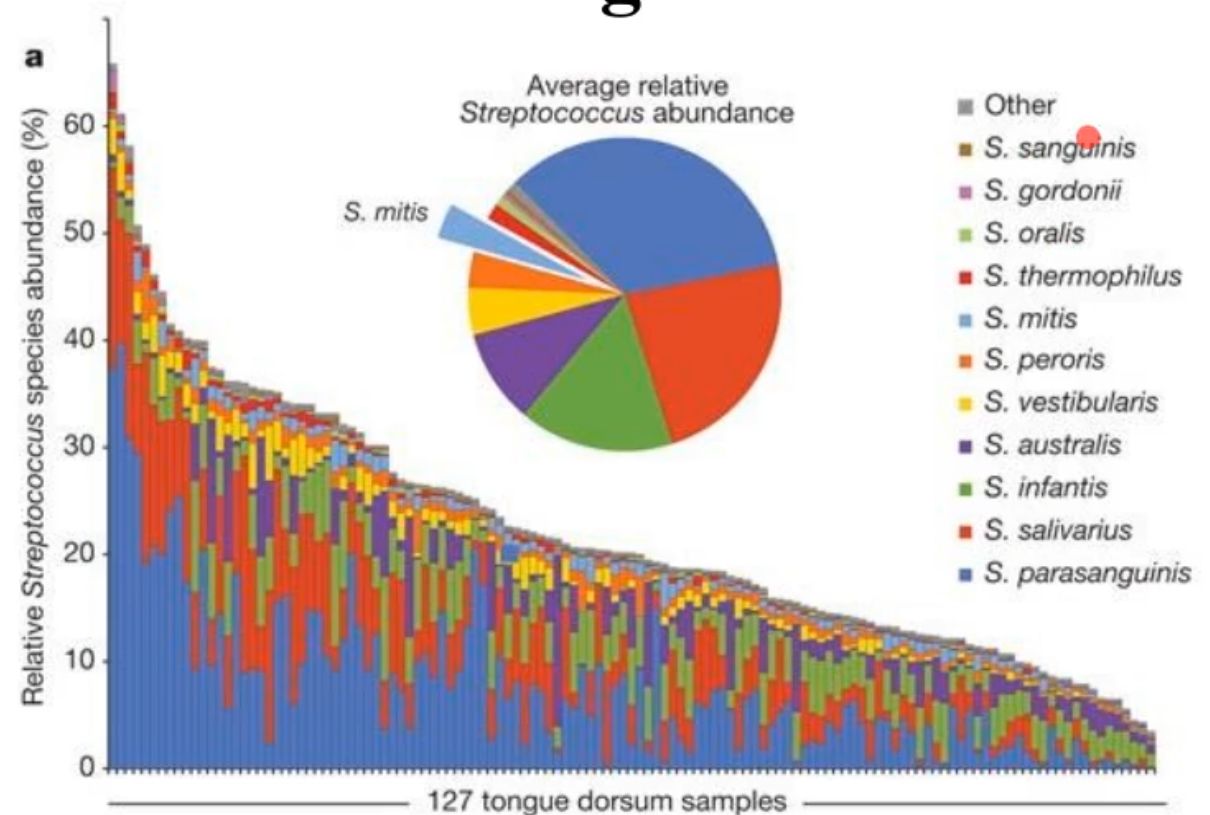
what species predominate on the tongue?
Streptococcus species
*this confirmed what we already had known among oral microbiologists in that strep species matter in health
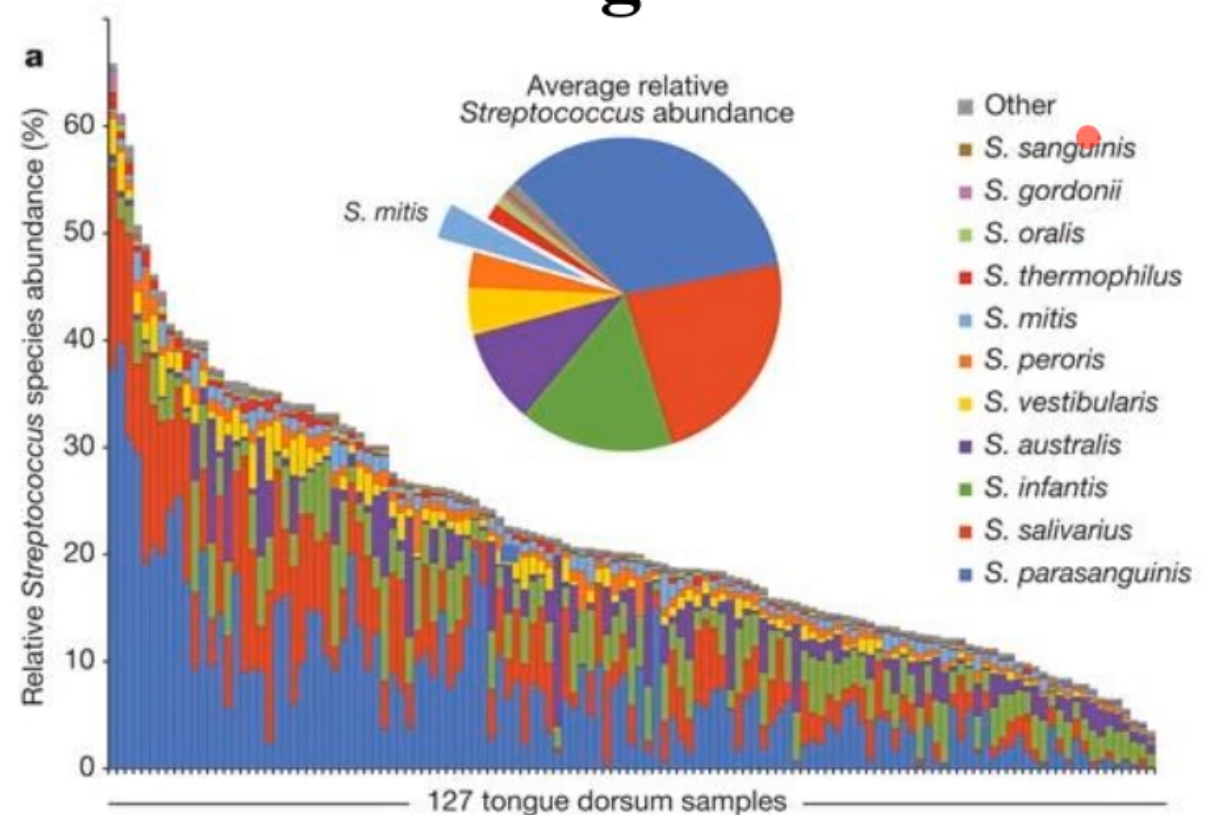
how many species are there in the oral cavity in 1 person?
there can be ~300 species present out of a possible 1000+; these all combine to form the dental plaque biofilm
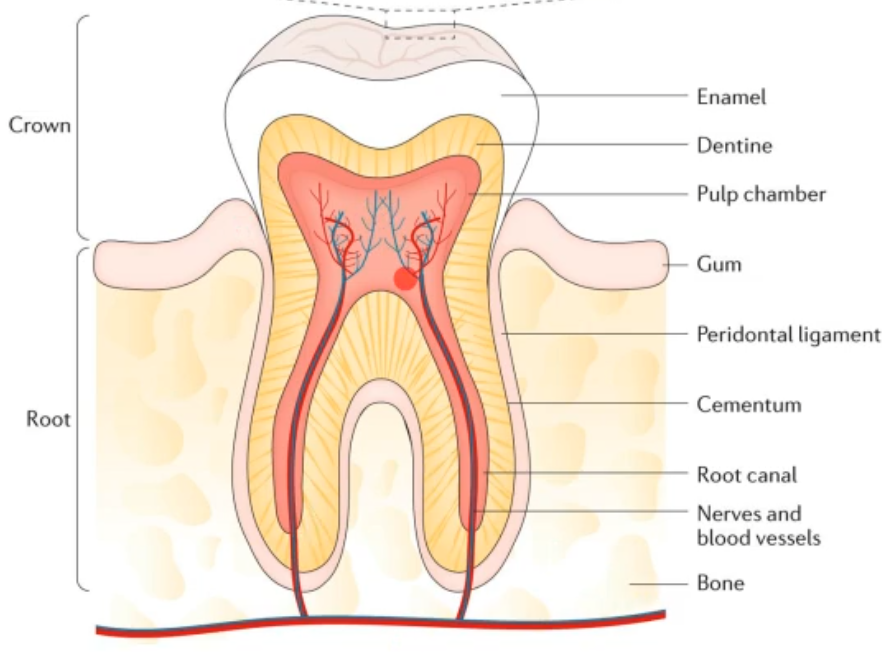
what does this diagram show?
the normal tooth anatomy
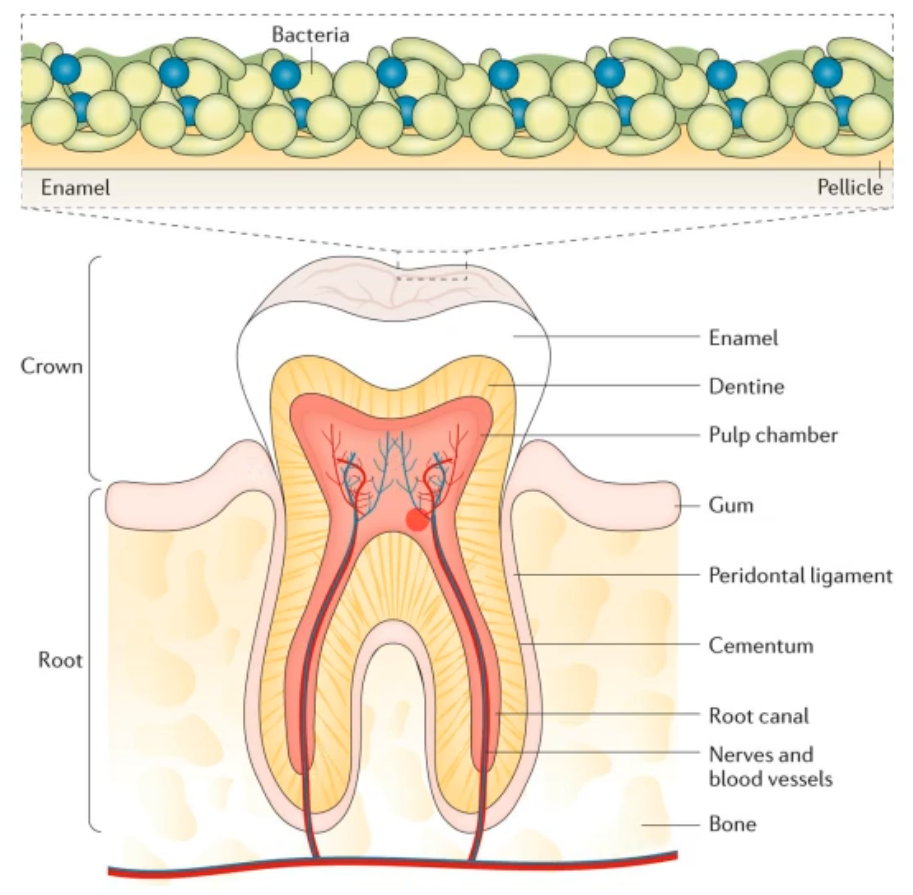
what does this diagram show?
the normal tooth anatomy and developing dental biofilm
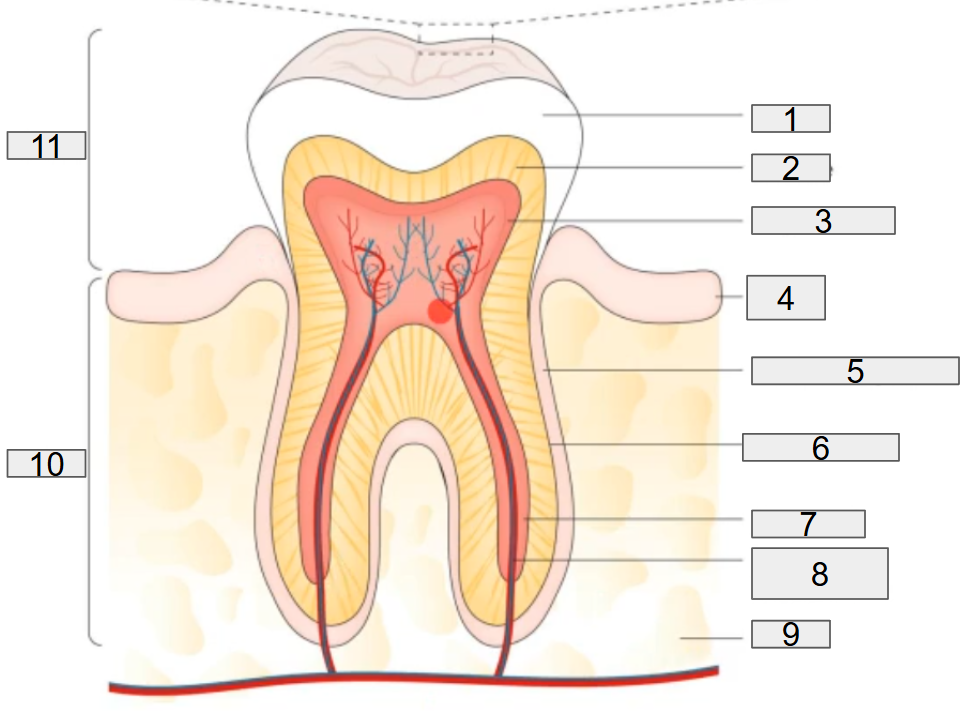
what does # 1 on the diagram representing?
enamel
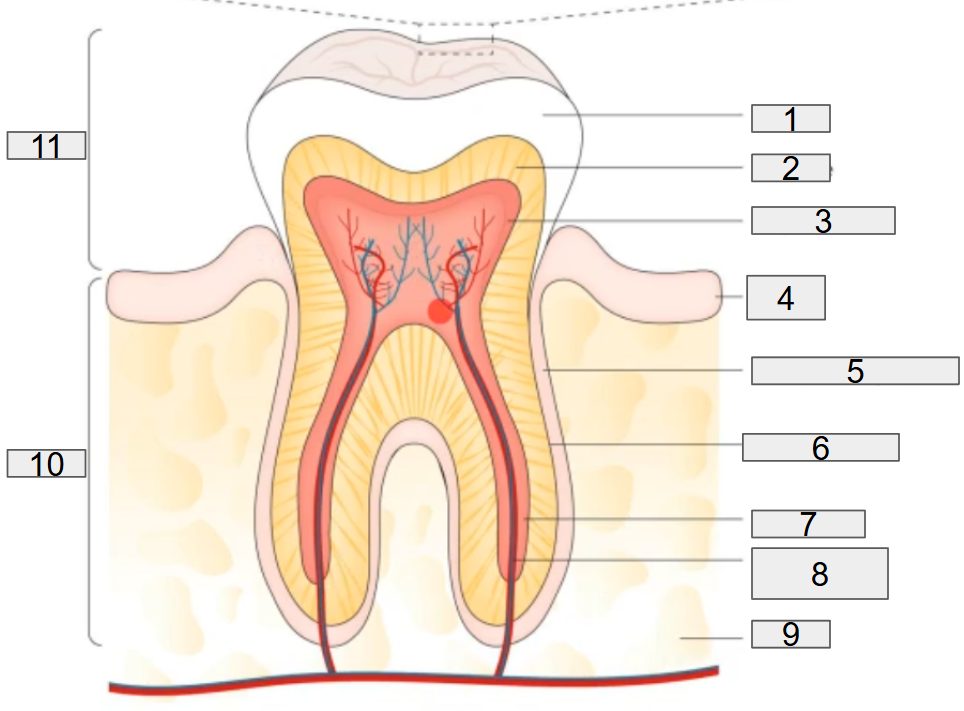
what does # 2 on the diagram representing?
dentine
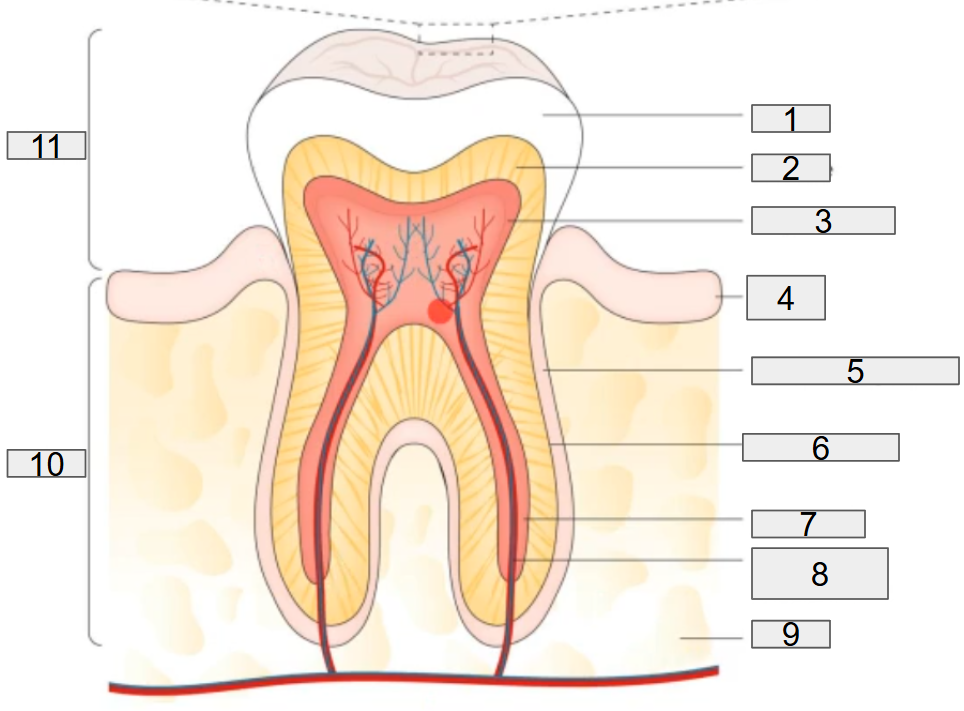
what does # 3 on the diagram representing?
pulp chamber
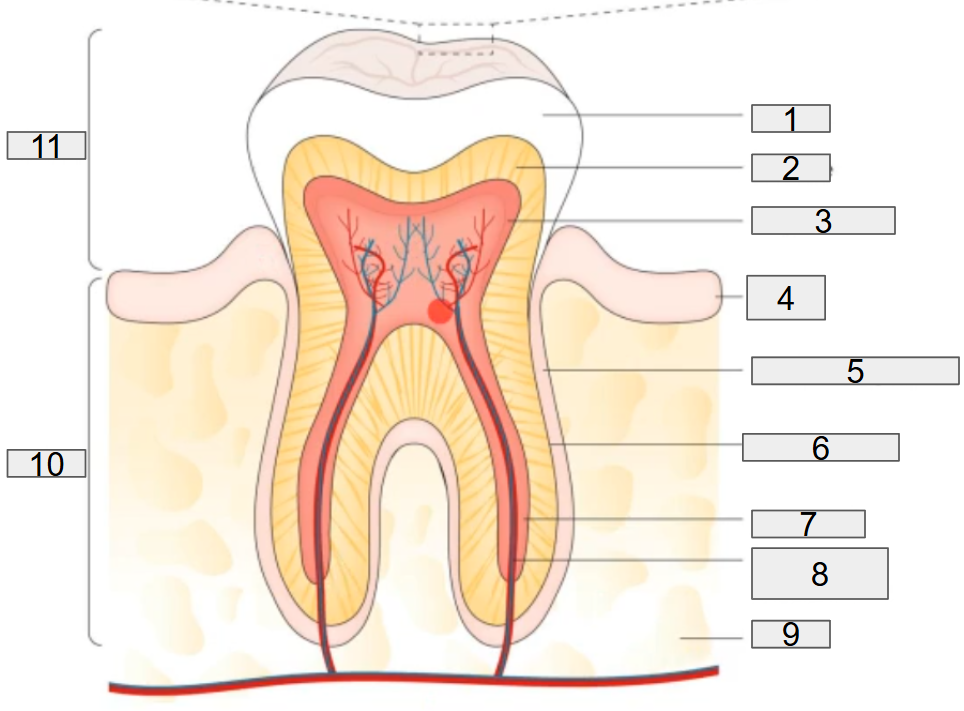
what does # 4 on the diagram representing?
gum
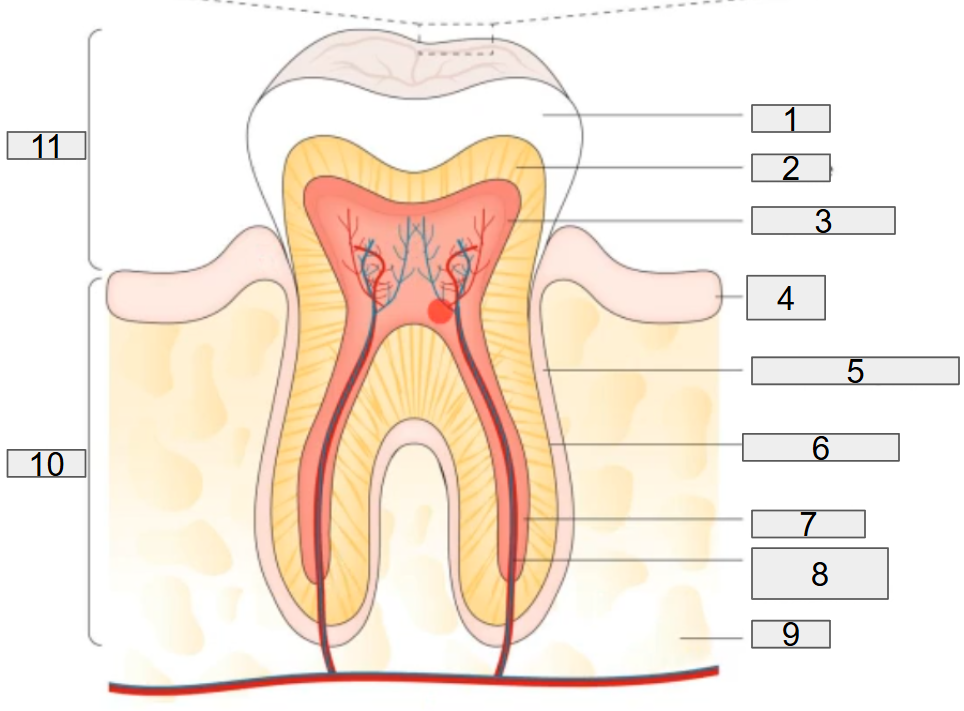
what does # 5 on the diagram representing?
deridontal ligament
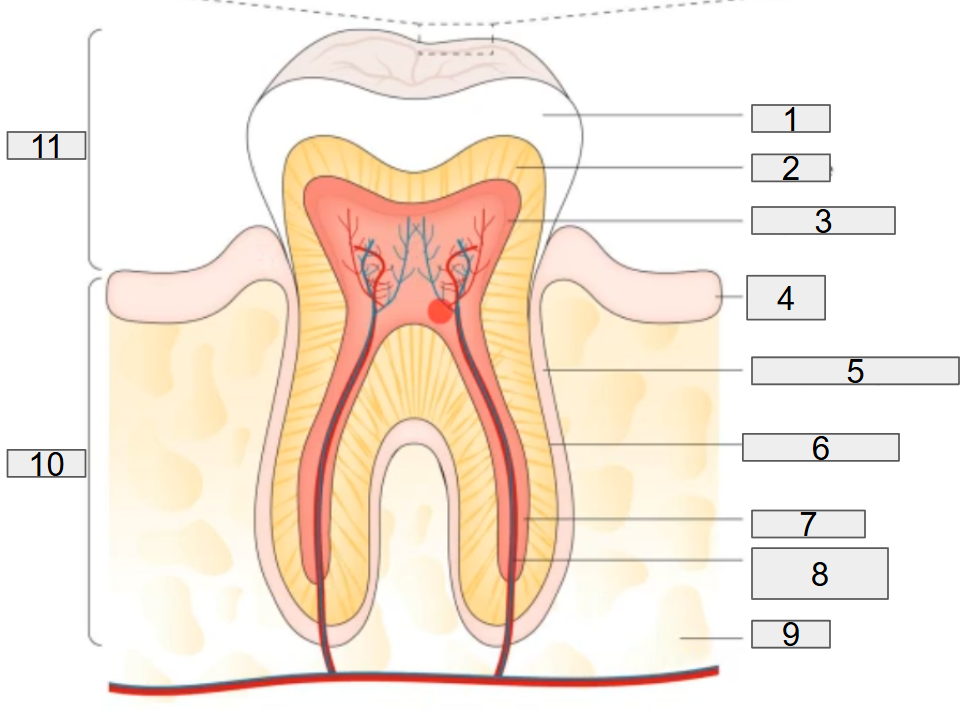
what does # 6 on the diagram representing?
cementum
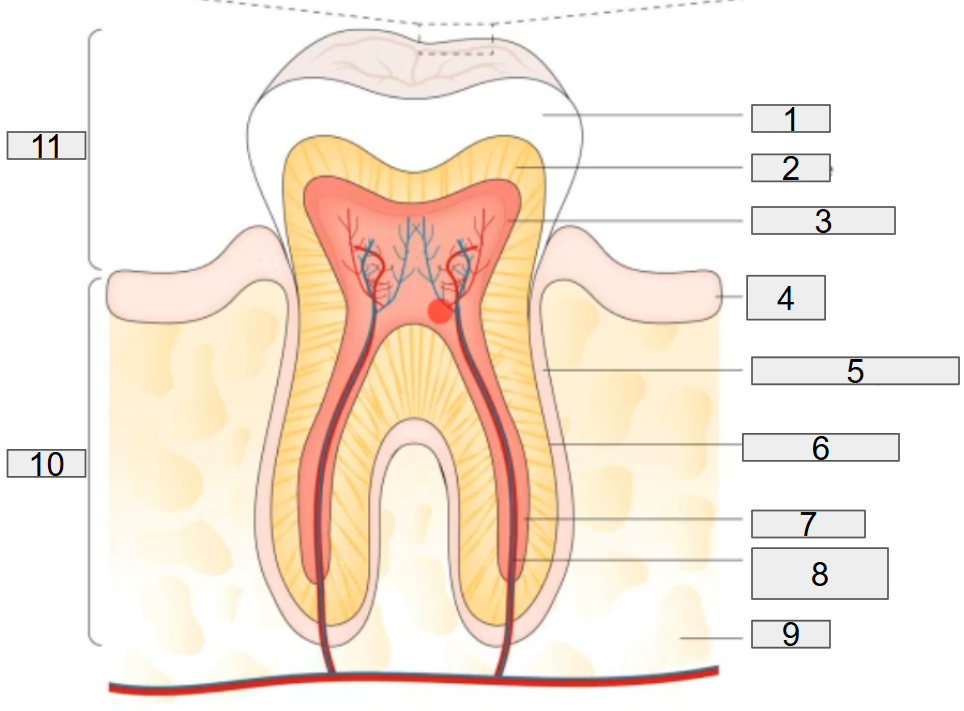
what does # 7 on the diagram representing?
root canal
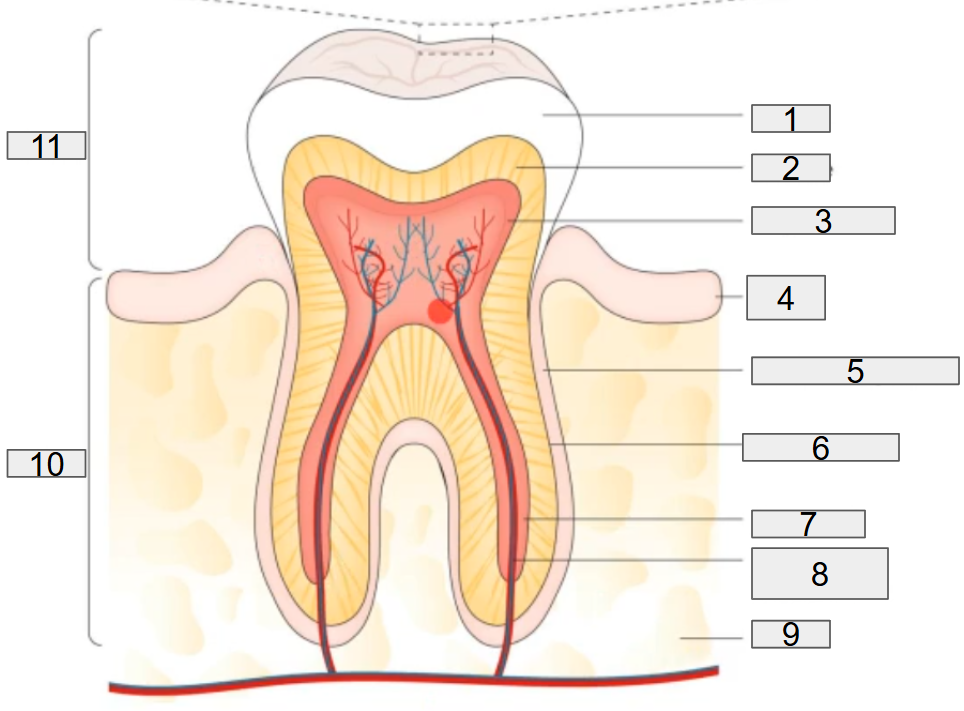
what does # 8 on the diagram representing?
nerves and blood vessels
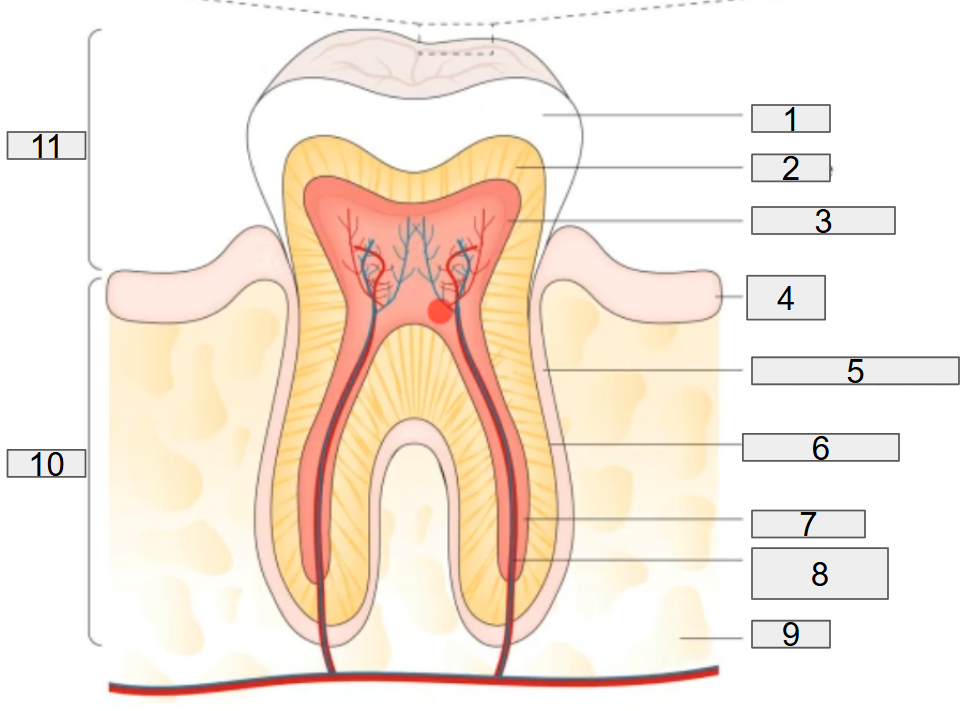
what does # 9 on the diagram representing?
blood
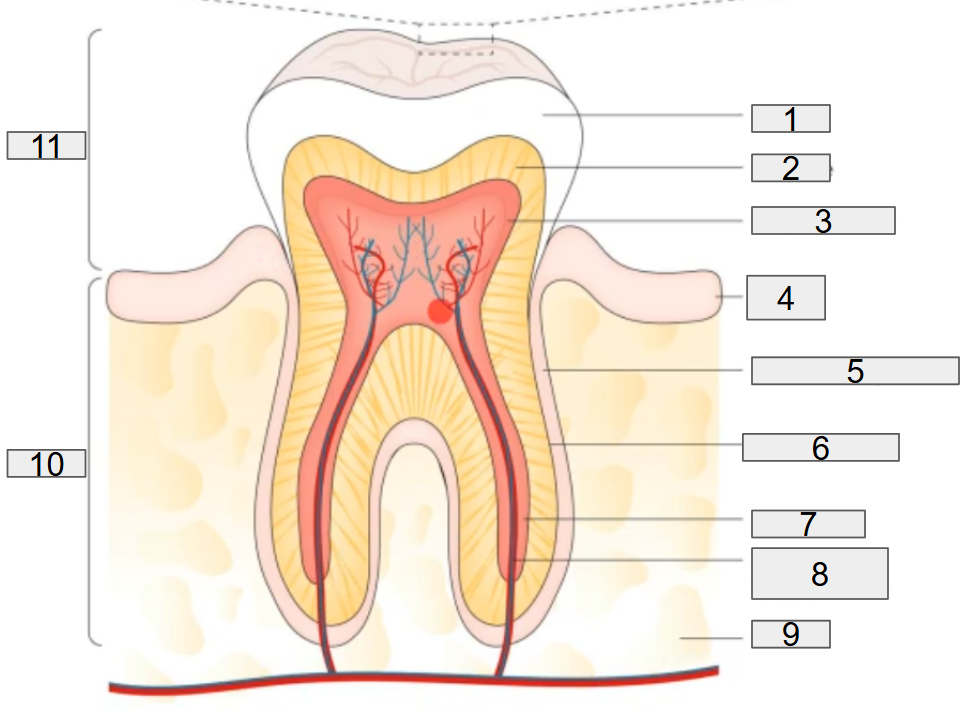
what does # 10 on the diagram representing?
Root
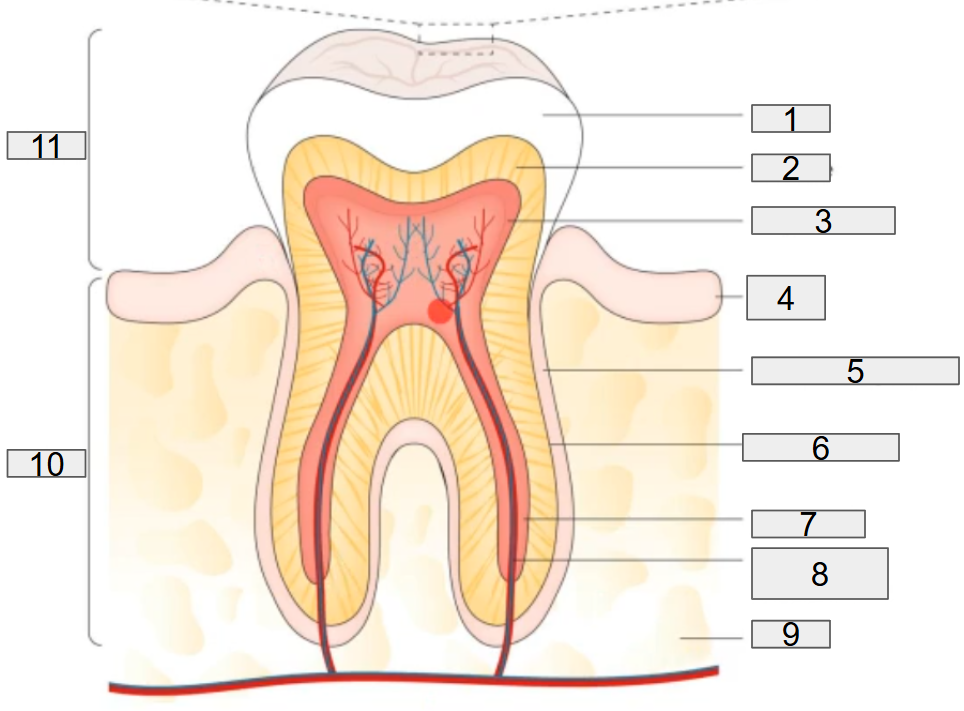
what does # 11 on the diagram representing?
Crown
what is the pellicle on tope of the enamel
proteins from the host (host proteins) that sit on the tooth forming a coat on the enamel, this allows for attachment of bacteria, more specifically for strep species and actinons species
other than the enamel, what other niches can bacteria colonize on?
Buccal mucosa
Saliva
Palate
tongue
*all depends on on the bacteria and what attachment sites they attach to
what is a biofilm?
a structured community of microorganisms encapsulated within a self-developed polymetric matrix and attached to a living or inert surface, they can be found everywhere and they function or facilitate nutrient acquisition and provides a stable, shelter environment
why do prokaryotes form biofilms?
surfaces provide a space to be occupied
Biofilms provide a degree of stability in the growth environment
Catalytic functions (metabolic) through localizing cells in close procimity
protection from a wide range of environmental challenges: metal toxicity, acid exposure, dehydration and salinity, phagocytosis, and antibiotics and antimicrobial agents
*basically protection and nutrients
how do biofilm formation form on surfaces?
bacteria will bind attach to a surfaces by expressing their adhesion molecules this can also allow for the bacteria to aggregate together
the aggregated bacteria’s will begin to grow on the surface, expressing more of the adhesion molecules and they begin to form an extracellular matrix (polysaccharides , DNA, lipids, proteins)
other bacteria will also being to attach to the pioneering bacteria
finally there will become detachment which allows for repeating of this cycle
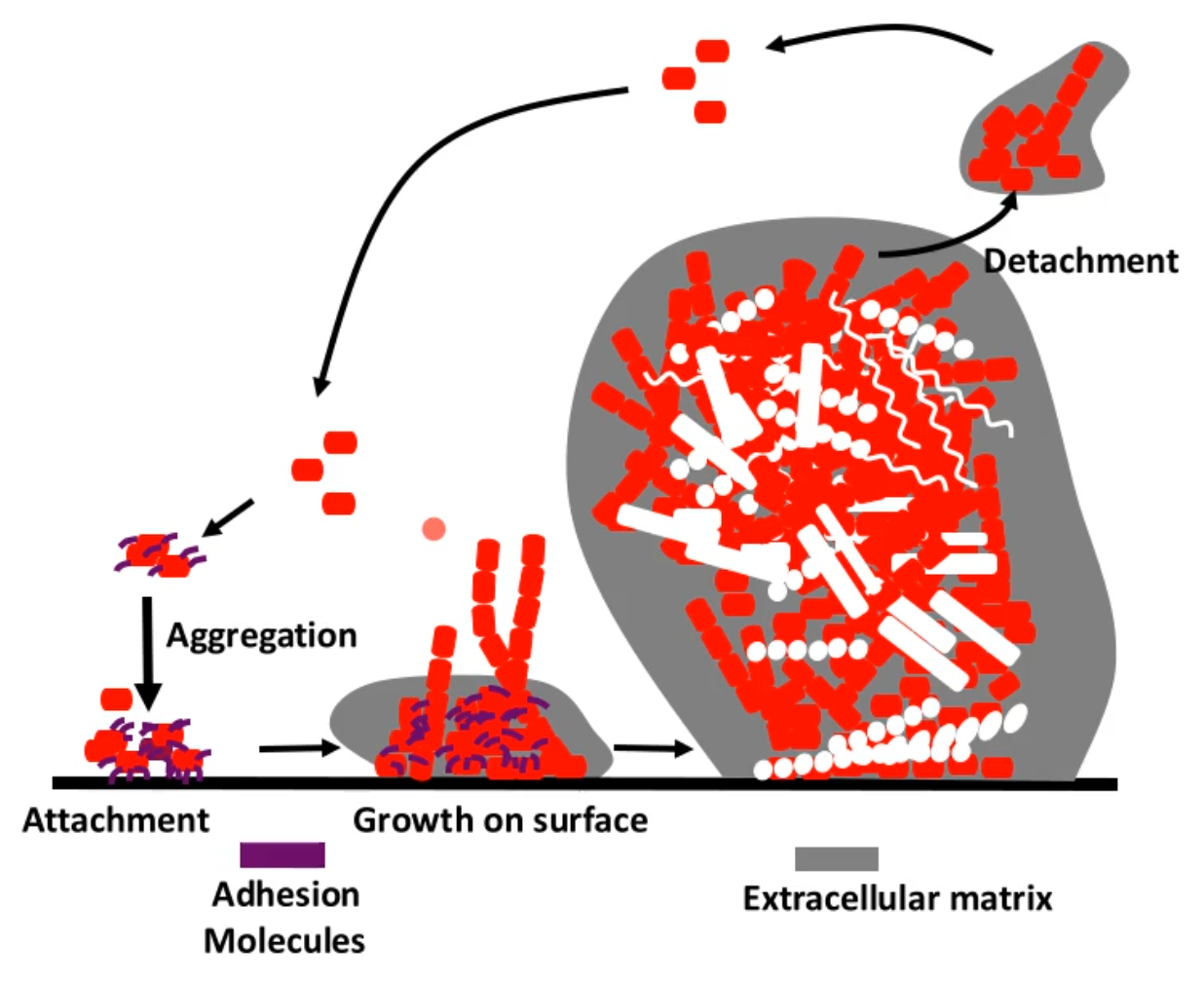
T/F dental plaque = biofilm
True
what makes oral biofilms unique?
in the oral cavity, if u cant attach and form a biofilm, u cant hang around (will get washed away). Attachment factors allow for attachment to surfaces or other bacteria and are thought of a virulence factors
why are attachment factors that important for bacteria in the oral cavity?
factors in the oral cavity such as saliva flow or nutrient rich GCF can increase during inflammation (chronic periodontitis) which results in washing away of the none attached bacteria and even the bacterial aggregates, so there is no second chance
there are still a number of pressures on the bacteria that are in biofilms, what are they?
this thing to note is that there is a mix of G-, G+, commences, etc in the biofilm to begin with
Nutrient availability
anoxic environments
no oxygen, not all organisms can withstand this
viruses
mammalian cell infiltration
DNA exchange
Hemin
blood that is available
chemical and physical microbe-microbe interactions
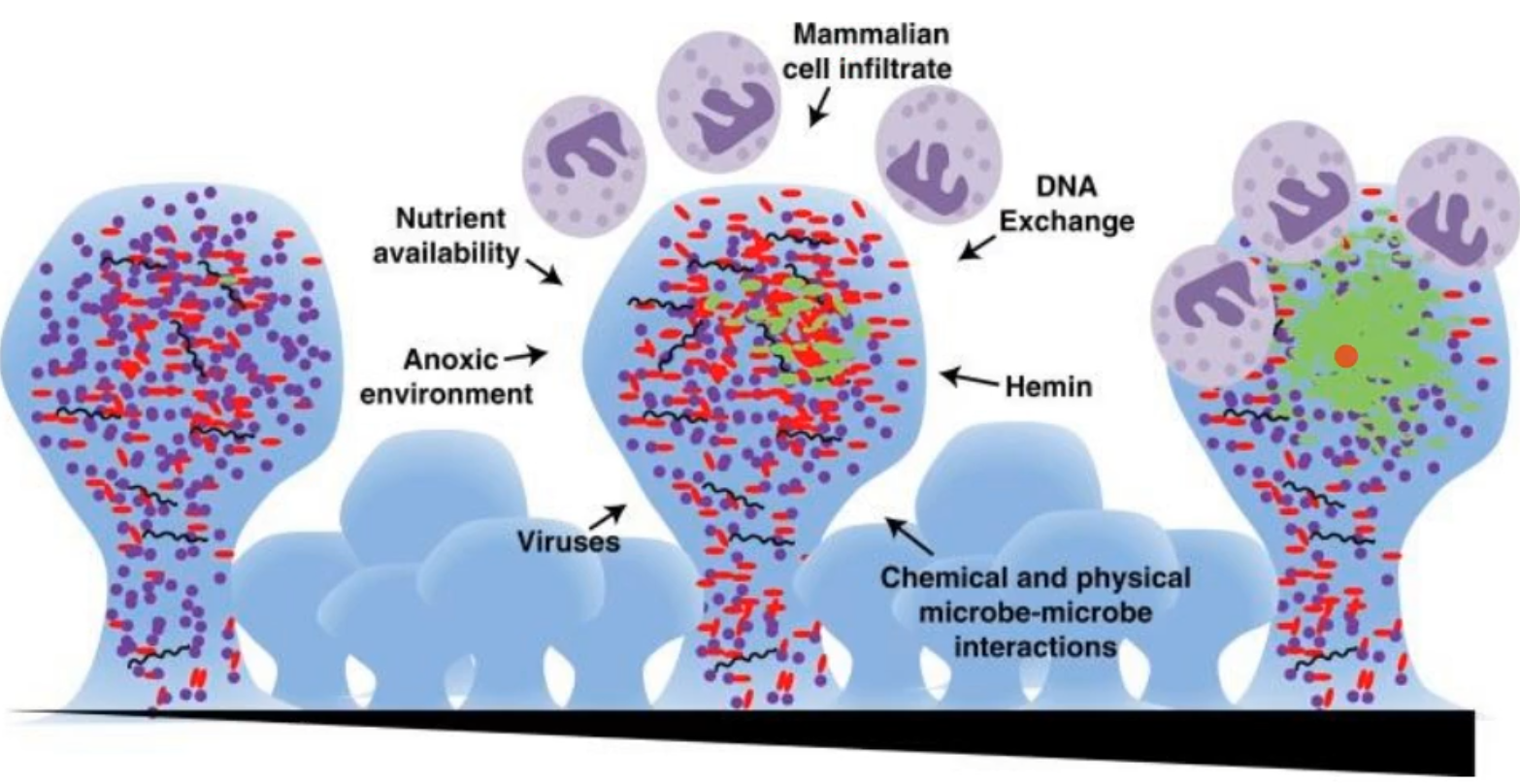
what facilitates proper biofilm development?
microbial co-aggregation; first primary/pioneer colonizers and then second colonizers
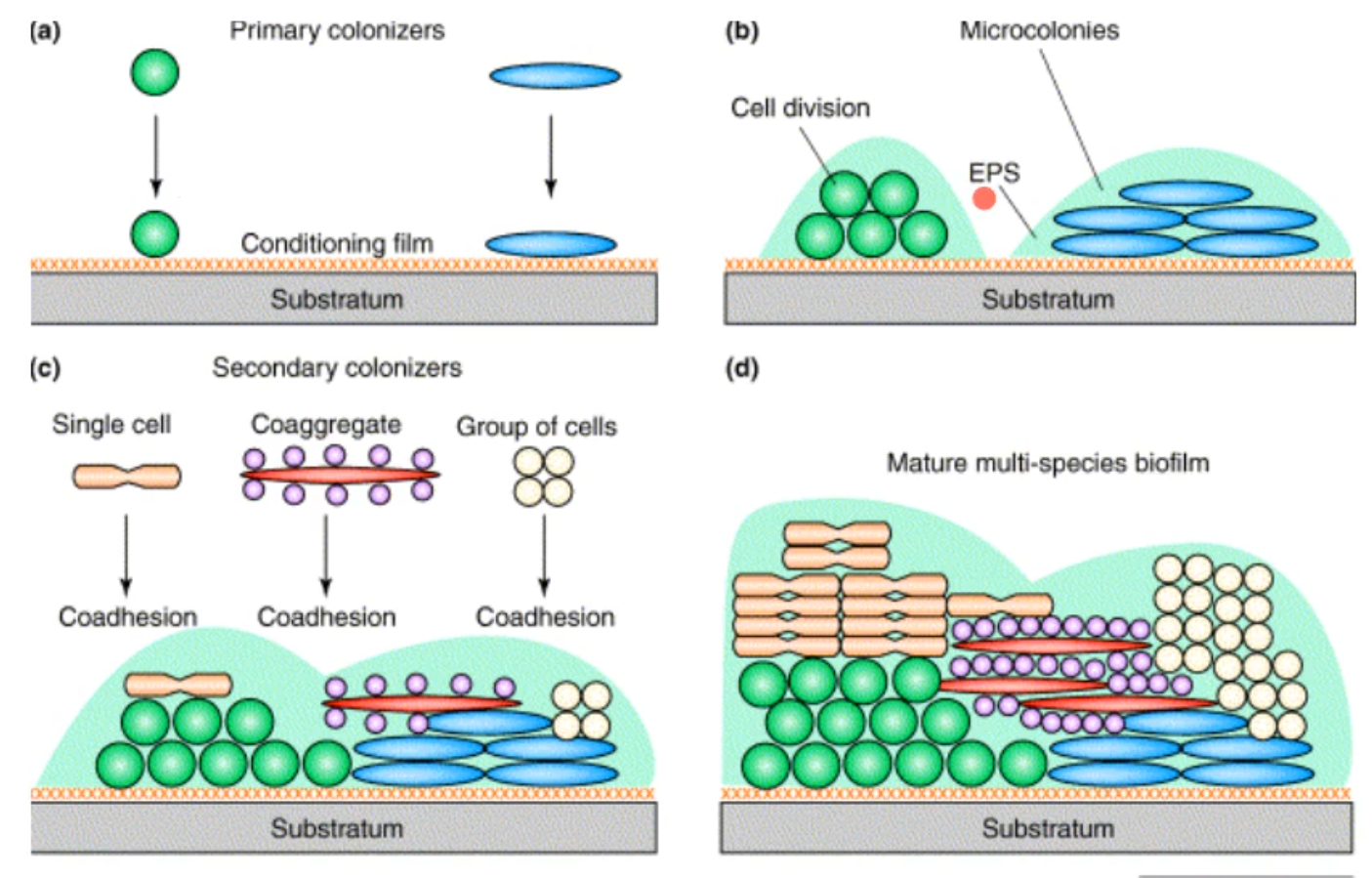
what is salivary pellicle defined as?
The thin layer of salivary proteins and glycoproteins that quickly adhere to the tooth surface after the tooth has been cleaned; this amorphous, bacteria-free layer may serve as an attachment medium for bacteria, which in turn form plaque
when the slivbary pellicle, what primary colonizers will use is as an attachment site?
gram positive aerobes
Streptococcus species and Actinomyces
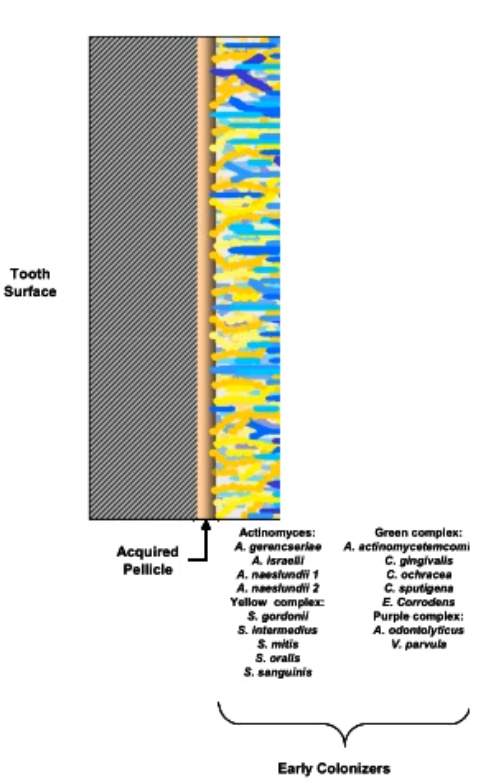
T/F members of the early colonizers can contribute to caries formation
True, Streptococcus mutans is a notorious acid producer that is a early colonizer and when the conditions are met for acids to be produces, the acids will begin to eat away at the enamel (demineralization of the enamel)
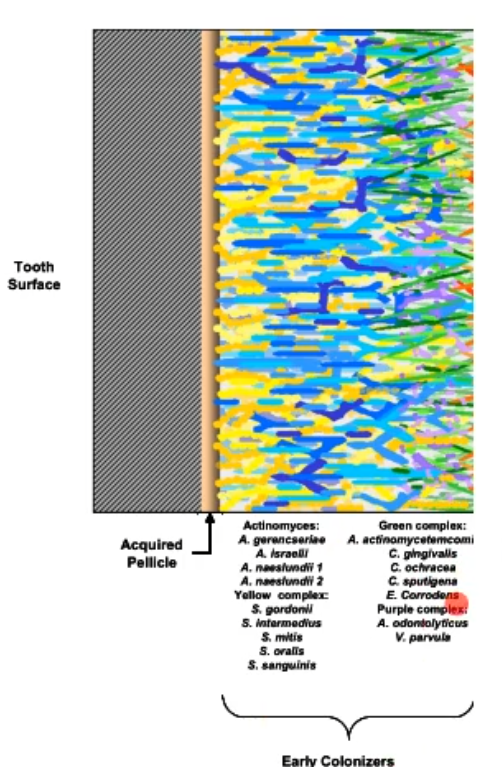
after the early colonizers attach, they provide (they become) an attachment site for the later colonizers, what are usually later colonizers?
gram negative anaerobes
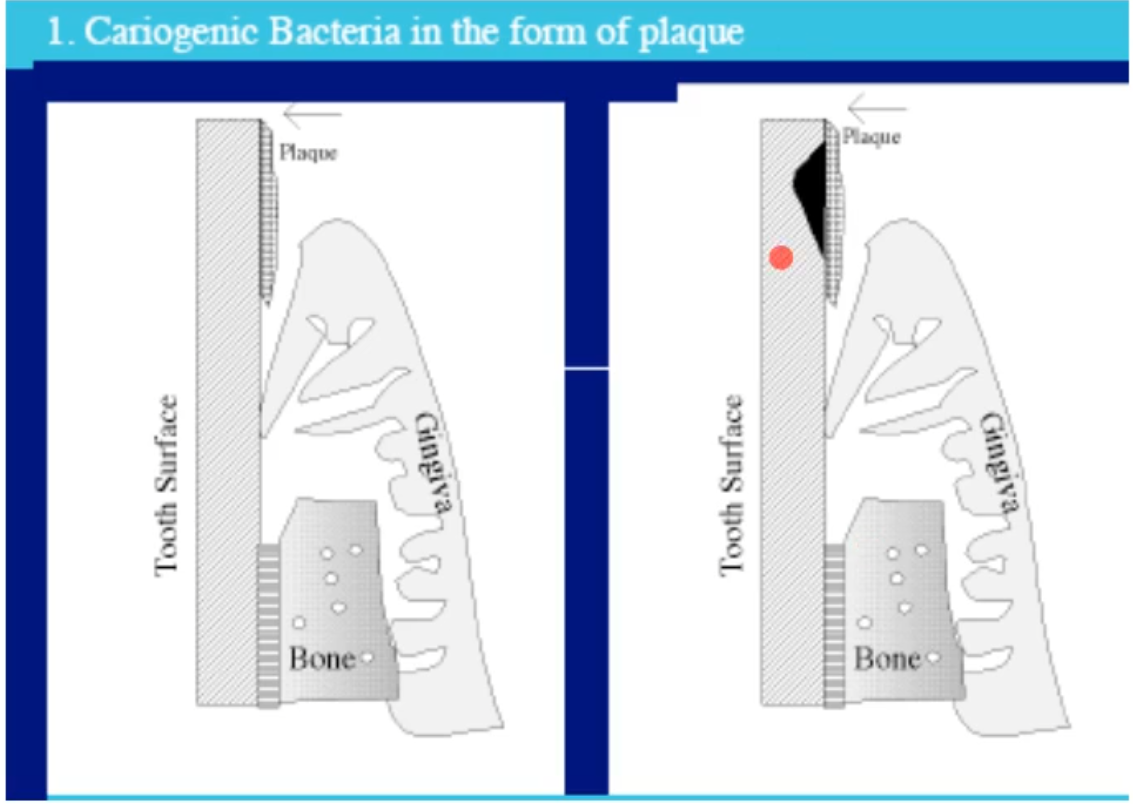
as the oral biofilm is forming, what is making up the subgingival plaque that borders the epithelial cells in the resulting cervices?
The later colonizers, usually what we talk about as pathogenetic (e.g pariopathogens); these can result in periodontal disease(tissue degradation) as they continue growing and going under the gingiva; late colonizers are highly and inflammophilic
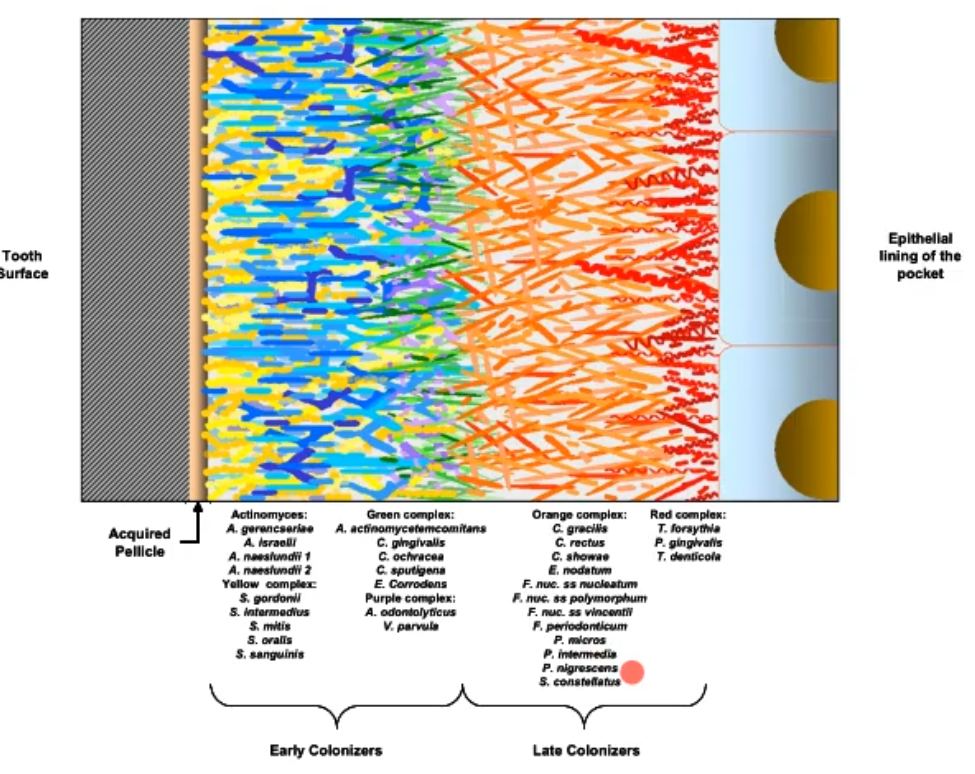
proteolytic bacteria
have a lot of proteins that they secrete or on their surface that allow for the degradation of other proteins
inflammophilic bacteria
will inflame local tissue which can result which is beneficial to the bacteria as it provides a nutrition source of the bacteria; however this results in breakdown of the gingival tissue and breakdown of the bone
how do we know all this information?
there were a lot of studies of the bacterial population in health and disease states
these studies were done for:
saliva
buccal cells
tongue
gum biopsy
dental plaque
etc-ease of sampling
many of the early researchers were dentists
*all these sites are easy to obtain samples from in the oral cavity
How can studying disease tell us information?
allows one to understand the changes that occur in the bacterial population as the host transitions to disease
Caries-S. mutans
periodontitis - 70-100 species
how do collect samples so we can identify what organisms are present in the oral cavity?
scraping a surface for a sample
supragingival plaque
subgingival plaque-paper points
Buccal cells
tongue
Inserts
stents
retrievable enamel chips
Biopsy
punch biopsies
Fluid collection
GCF (paper point, capillary)
saliva
what surfaces can be scrapped to get a sample?
supragingival plaque
subgingival plaque-paper points
Buccal cells
tongue
*the signatures of each of these is going to be very different from one another
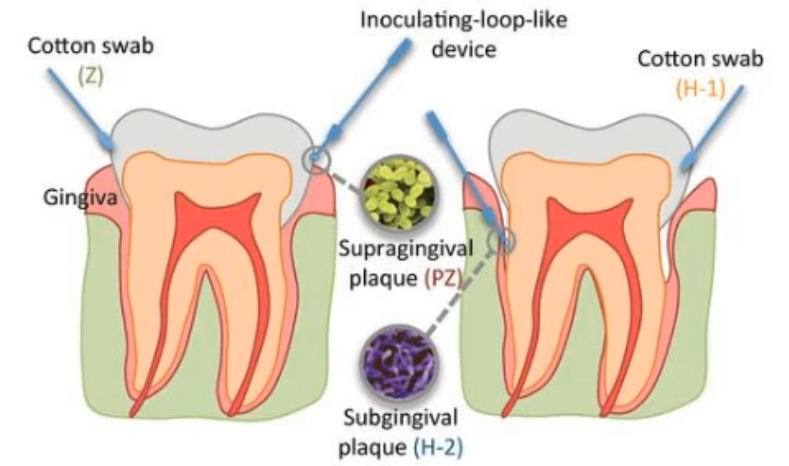
what are inserts?
stents
retrievable enamel chips
*basically a material (piece of plastic, etc) that u insert into the periodontal pocket which allows for colonization to from on it and retrieved to measure the samples that attached to that site; can be be put at different intervals to see how the biofilm grows over time
how do we determine what organisms are present in the oral activity?
culture dependent methods:
take sample and plate in various media conditions
subject to different atmospheric conditions (aerobic, anerobic)
subject to biochemical assays
understanding their metabolism can help with determining their species
early DNA-based culture-independent assays (ushered in by the age of genomics and DNA detection technology)
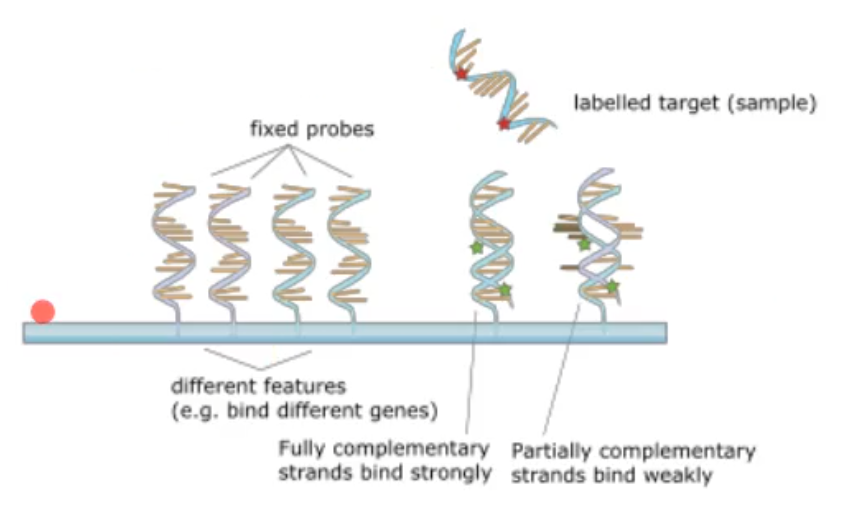
based on the idea that DNA will bind to complementary strand of DNA
this can be done with 16S probes or whole genome DNA probes
based on DNA of an unidentified bacterium matching up and binding to a known DNA sequence
Early DNA-based culture-independent assays
having fixed DNA probes(from either the 16S or whole genome DNA) and attaching them to a slide and taking the sample and label it. If it binds to the fixes probes and lights up it is complementary
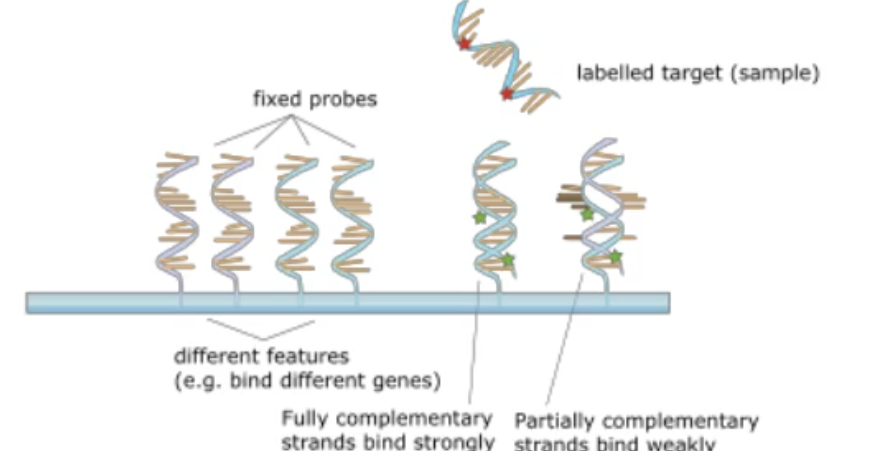
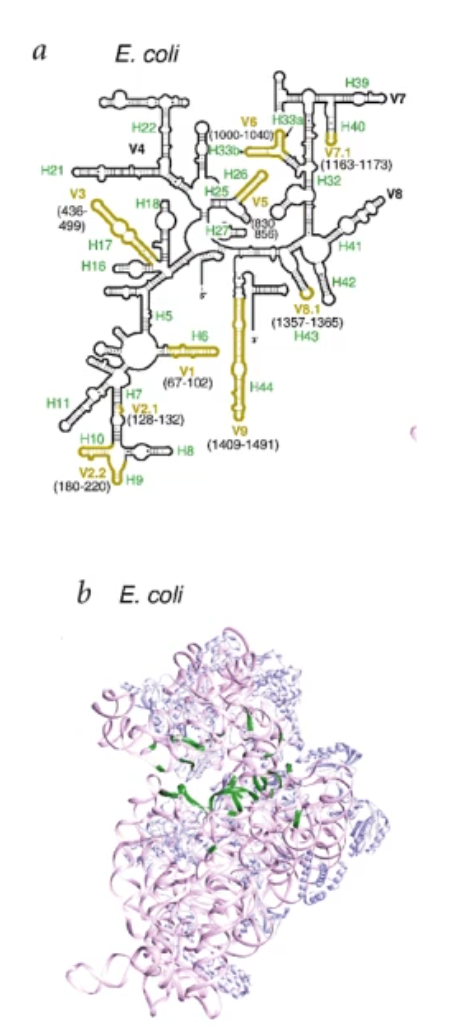
what is 16S?
16S ribosomal RNA (or 16S rRNA) is a component of the 30S small subunit of prokaryotic ribosomes (ribosomes make proteins)
all bacteria have it
there are conserved and variable regions (conserved can be all bacteria have that sequence; variable can be specific to a specific species)
this allows for speciation
who is the father of modern periodontal microbiology?
Sig Socransky, wrote “microbial complexes in subgingival plaque”(1998) and understood that their were changes in the oral microbiome that were occurring from healthy to diseased, used a lot of early plating techniques but also had a good understanding go DNA technology and used them to identify the microorganisms, etc
what was the point of “microbial complexes in subgingival plaque”, what question did it ask and how did it find the answer?
they wanted to define the community by asking what clusters together at different sites? The method was to use 40 whole genome DNA probes, DNA-DNA hybridization checker board analysis
took 25 healthy and 160 chronic periodontitis patients samples and extracted the DNA and preformed DNA-DNA hybridization analysis
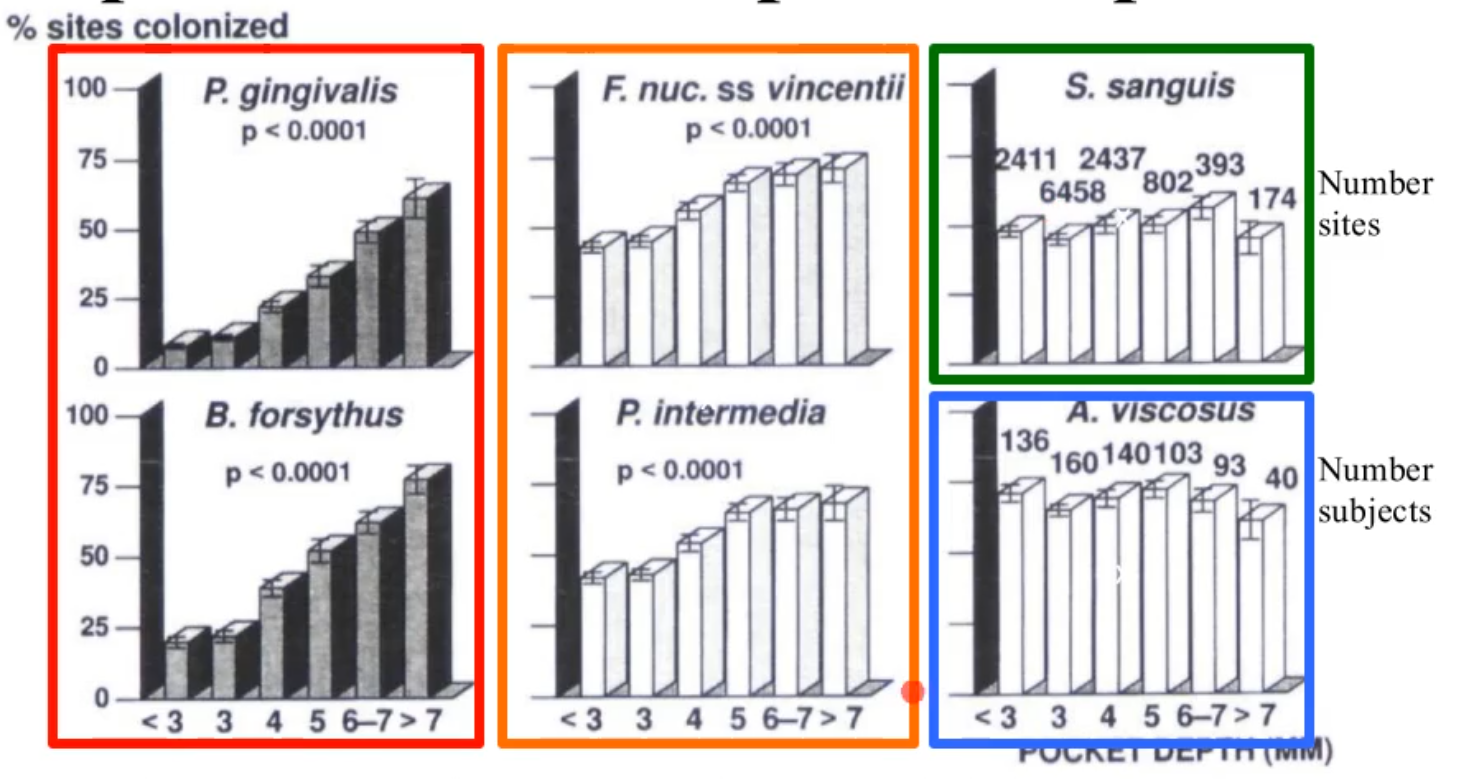
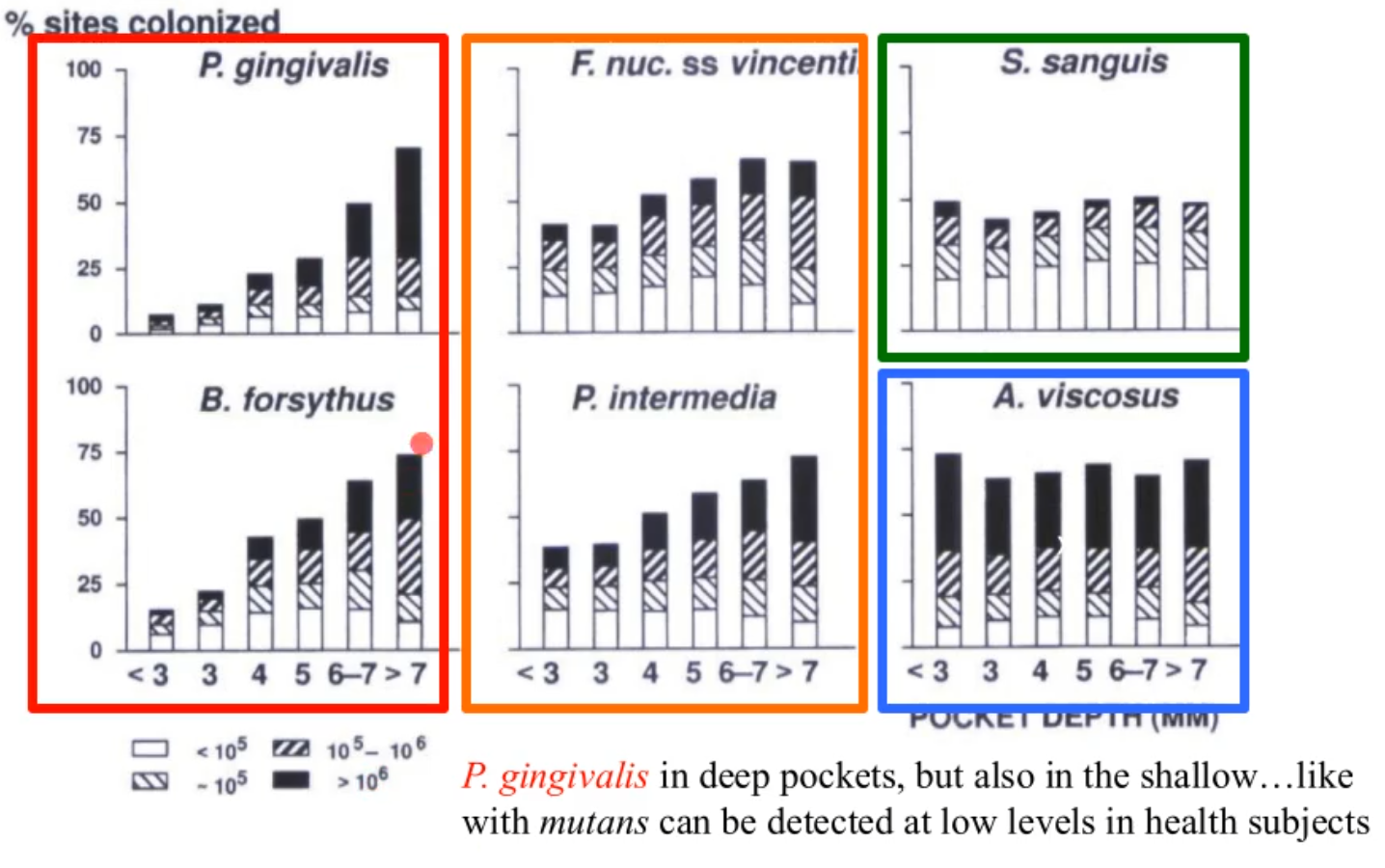
what does this picture show?
the % of sites colonized by 6 subgingival species at selected pocket depths
S. sanguis and A.vsicosus
regardless of pocket depth they are there and not really changing
F. nuc. ss vincentii and P. intermedia (orange)
still present in shallow pocket depths but increase as u get to deeper depths
B. forsythus and P. gingivalis (red)
still exist in shallow but even more prevalent in deeper depths
*orange and red are what cause disease and are found primarily in the deep pockets and are the g- anerobic bacteria that are mentioned
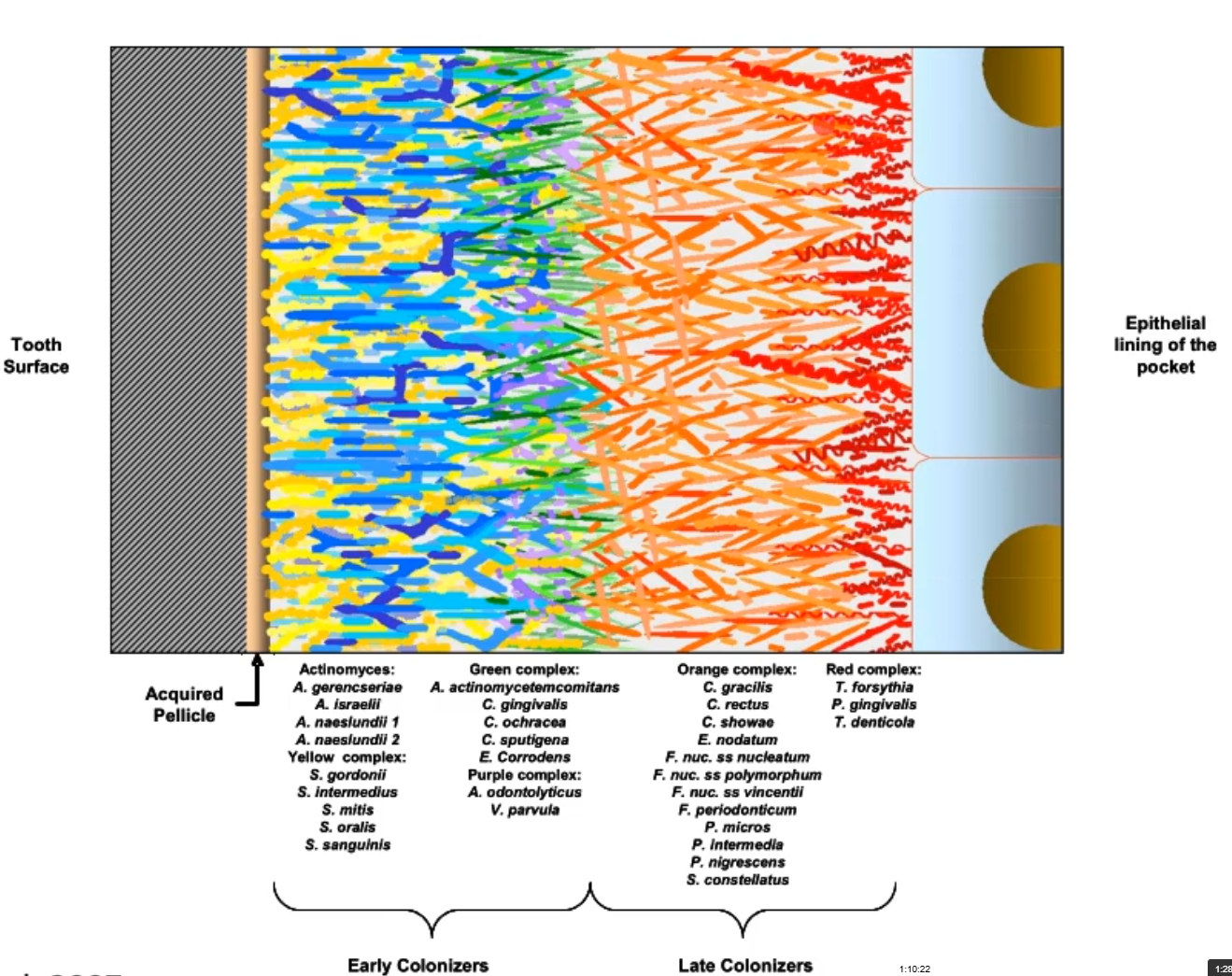
what does this picture show u?
the microbial succession in the oral biofilm
Through the early work, we can state that in many dental diseases (chronic periodontitis):
pathogenic multi-species plaque biofilm that is a dysiotic system that results in out-growth of the pathobionts
the pathobionts are always there, albeit at low levels in health
the thinner the plaque the less likely have a high number of pathobionts
Reach about 20% of the disease biofilm-highly pathogenic (little virulence bombs)
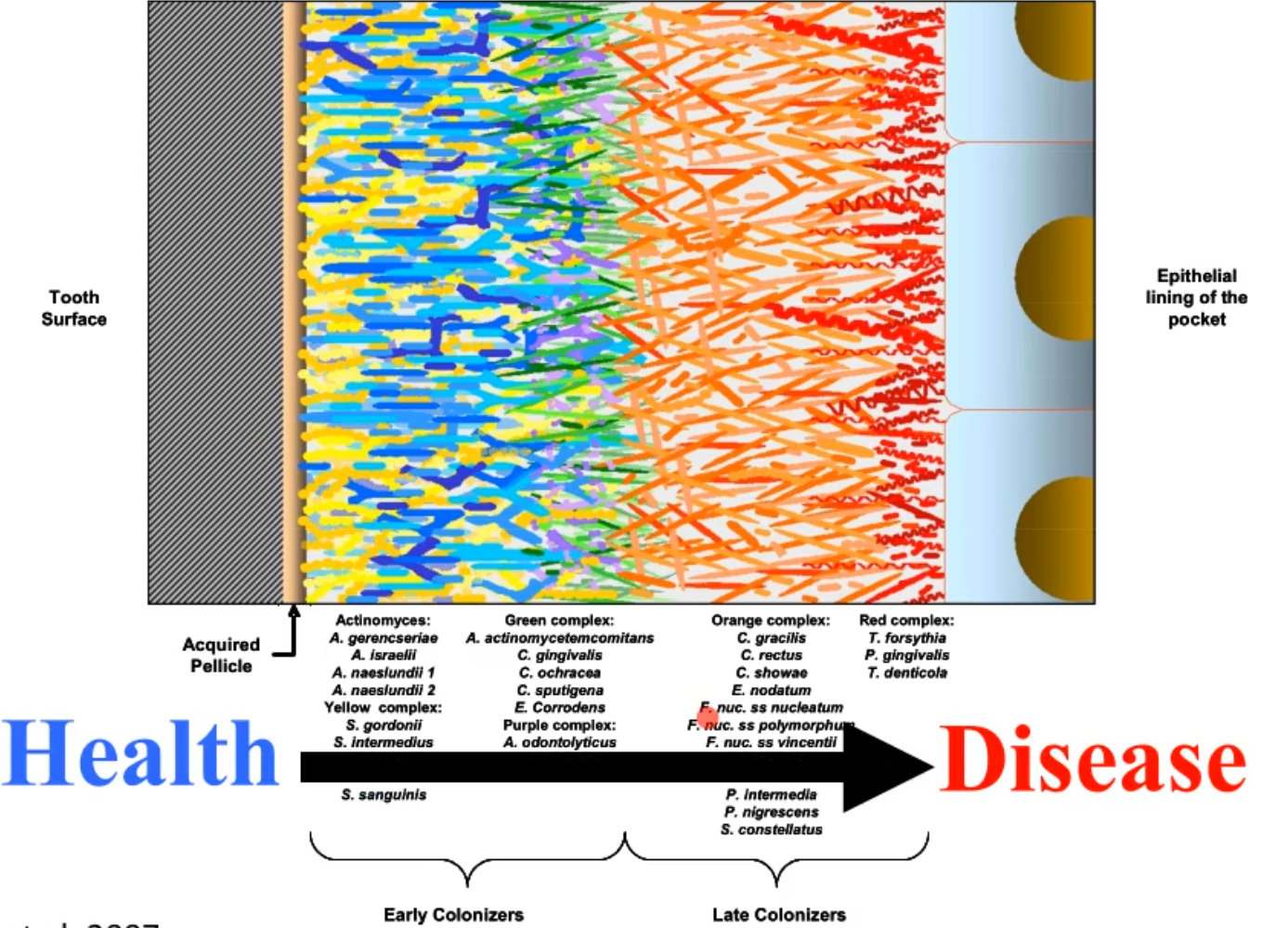
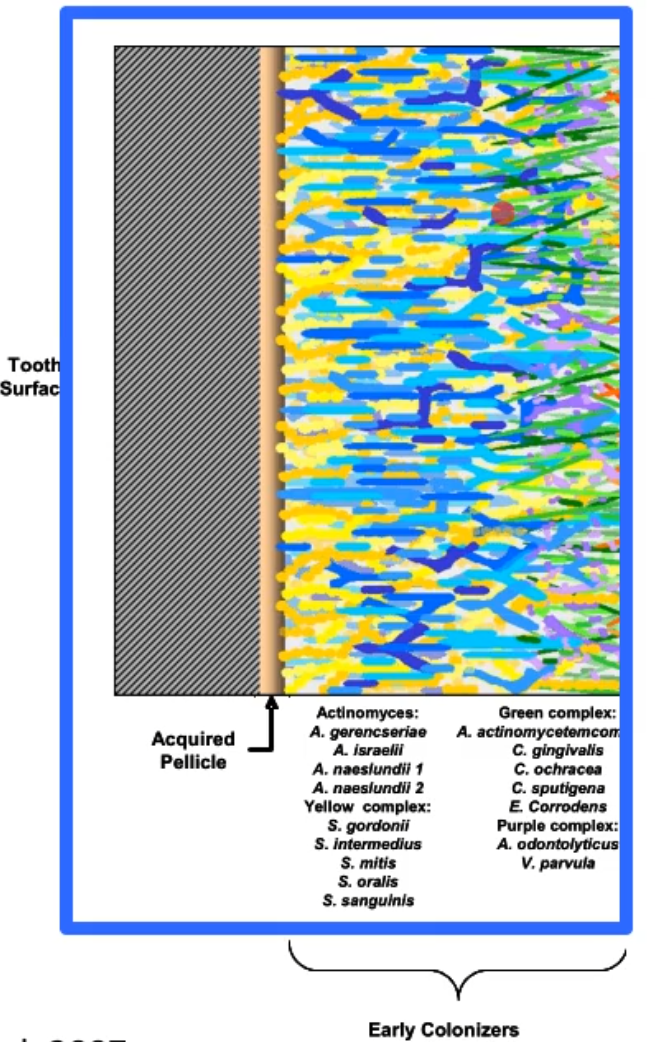
what does this picture represent?
it shows how the goal in treatment is to arrest the plaque to about that point in the picture, basically the goal of dentists
details on how Dental plaque = biofilm
chronic periodontitis is a biofilm mediated pathology
oral biofilms are unique because - in the oral cavity if you can’t attach and forma a biofilm tou can’t hand around
have to make attachment factors that stick to surfaces or other bacteria
GCF is increased in chronic periodontitis, easy to wash away the bacterial aggregates
No second changes!
how can we grow biofilms?
flow conditions
static growth
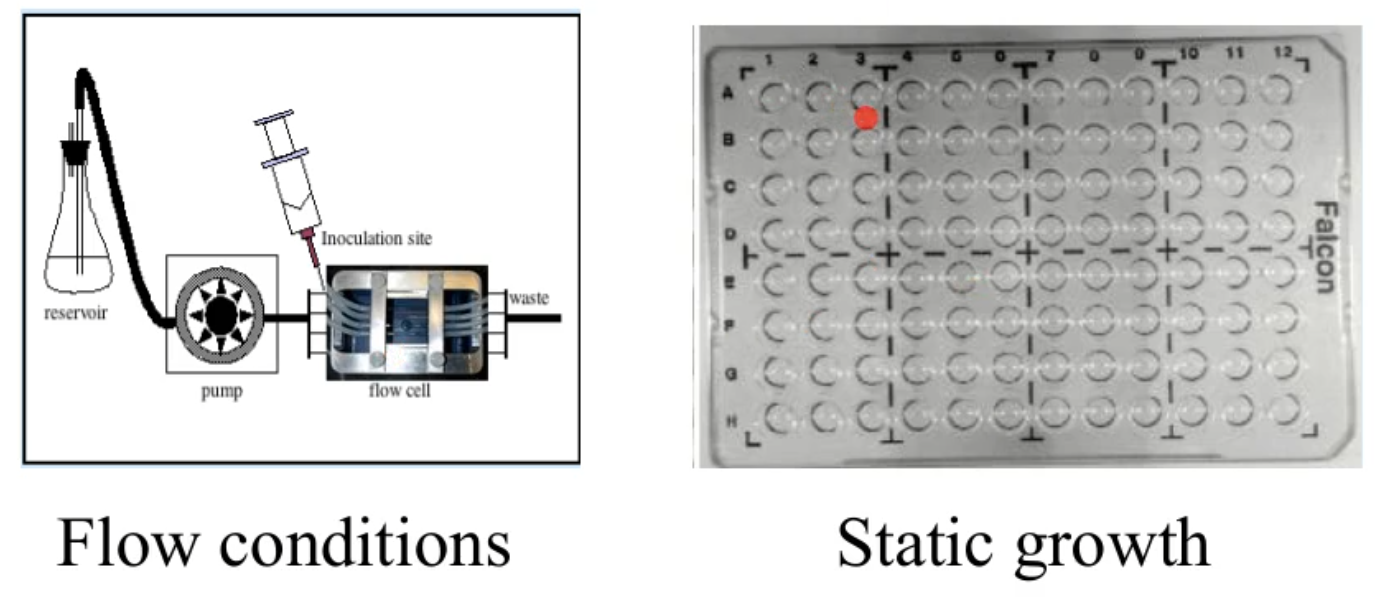
how can we visualize biofilms?
stains
microscopy
light
confocal
SEM
TEM
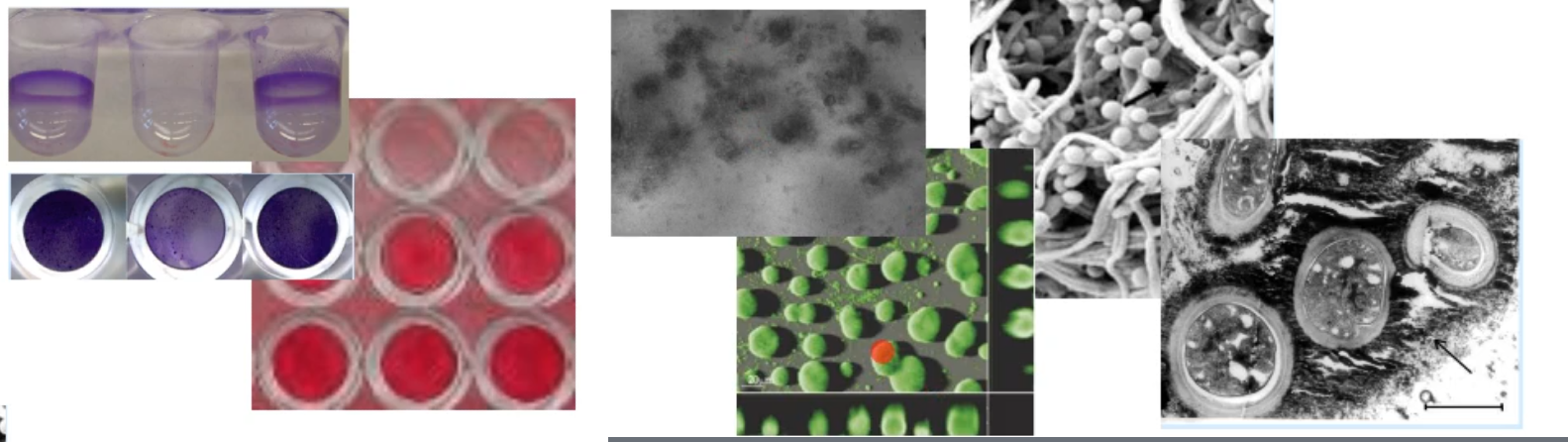

what are single species and co-cultural analysis used for?
lets you break down what is happening in heath vs disease, this is done by:
single bacteria that is chosen by researcher for additional study
co-culture of organisms that are known to interact
culture with mammalian cells to address whether it is pathogenic or protective
use basic plating methods, molecular microbiology, genetics, biochemistry, cell biology and immunology based assays
what are some issues with the older ways of studying bacteria and what new technology is fixing this?
in culture independent research like DNA-DNA hybridization, you have to have a known 16S sequence or genome probe which can give out biased information and you might miss a species
Sequencing can fix this because have and will sequence everything (16s sequencing, metagenomics, transcriptomics) and are able to directly visualize the microbial interactions (no culture biases)
what makes sequencing so effecive?
16s sequencing, metagenomics, transcriptomics, proteomics:
no culture bias
no probe bias
method to detect the unculturables
can generate entire bacterial genomes with out growing the organism
can ask the questions as to what gene or gene products are present in health vs disease
what questions can u ask with 1 DNA smaple?
what are they doing?
can sequence all genes
compare to database of genes with known functions
tells you pathogenic and metabolic capabilites
whose there?
sequence 16s genes
compare to database of known 16s genes and this lets you either:
identify species
classify as new species
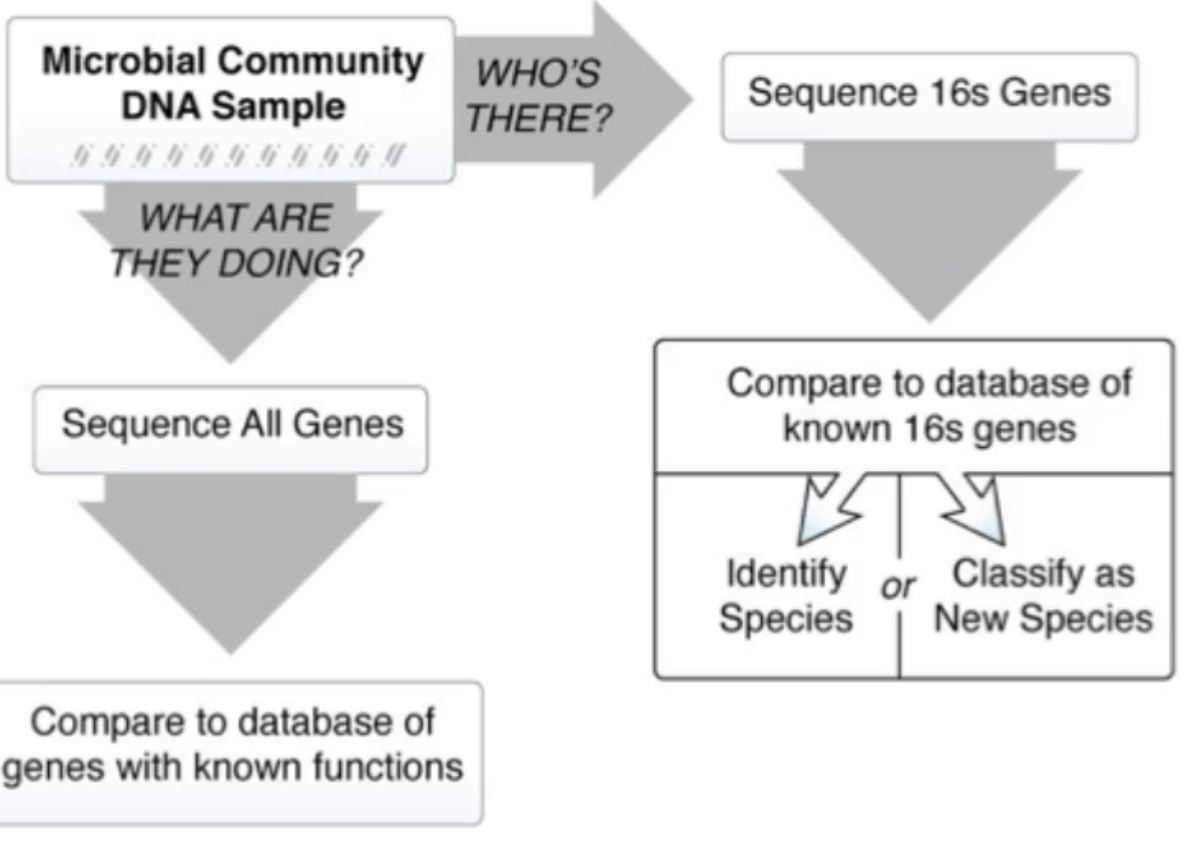
steps for 16s sequencing
environmental samples
DNA extraction
Genomic DNA
PCR and sequencing
16S rRNA sequencing
Sequence comparison
Phylogenetic trees
what confirmations did 16S sequences have on previous studies that complex held true?
streptococci and gram positives predominate in health
anaerobic, gram negatives predominate in periodontitis added a few more species, but the “big 3” still show up
P. gingivalis
T. fosythia
T. denticola
what can you do in combination with 16S sequencing?
you can use 16S probes in combination with microscopy and this is called
Fluorescence in situ hybridization (FISH)
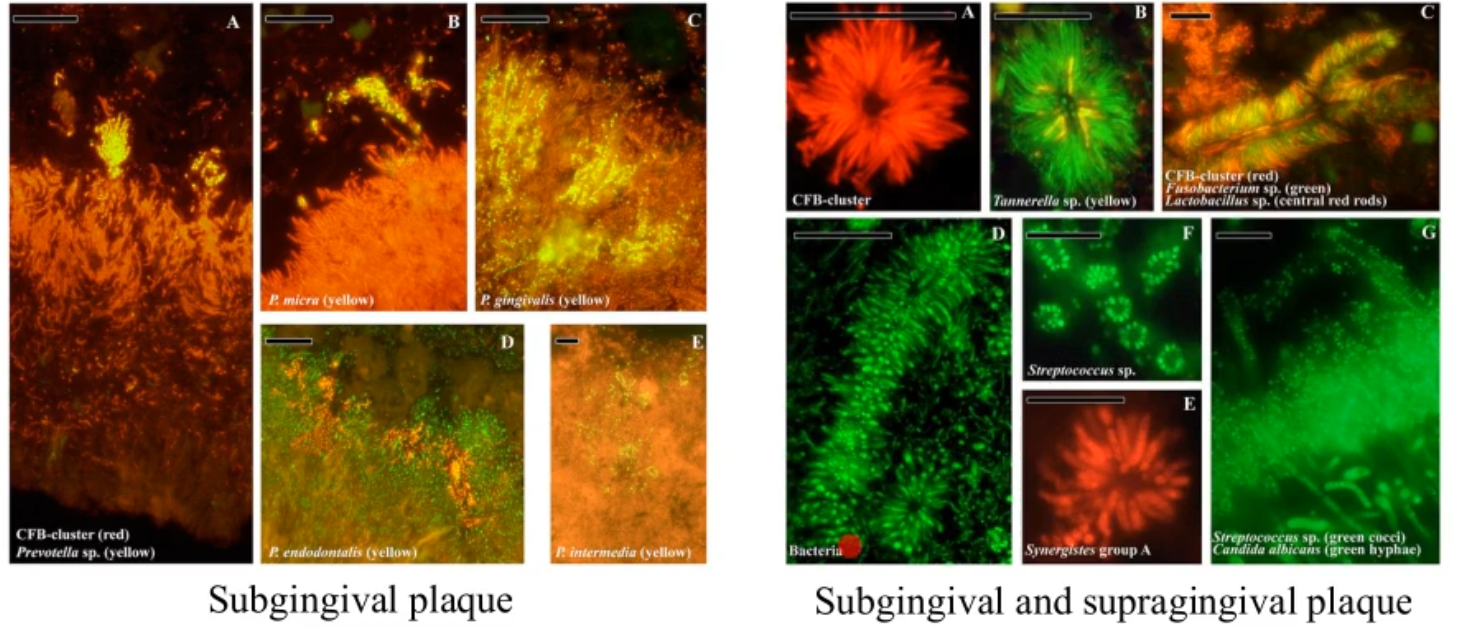
what is FISH benefitable or nice to use?
can show you the complexes that are formed in the plaque
probes can tell you the species (using conserved and variable regions)
microscopy fluorescents can help you visualize the complex
confirms
the predominate species in the plaque
localization and associations
metagenomic analysis using sequencing
take all the DNA in a sample and sequence i, this allows one to determine the genetic metabolic potential of these communities
no culture bias
no probe bias
method t detect the unculturable
can generate entire bacterial genomes with out growing the organism
*have to be able to analyze the data, this is the hardest part, need someone that is good at bioinformatics to be able to analyze for you
*what if everything is dead? - cant tell you if anything is happening/metabolism, can tell you the potential, but the DNA hangs around for a while, but can use transcriptomics instead
Additional “omic” analyses
transcriptomic:
use RNA sequencing to determine what RNA is being made by genes
can ask the question what genes are turned on or off in health vs disease in both the host and bacteria
can do this in many different conditions
proteomics:
protein sequencing to determine what proteins are made
e.g the human salivary proteome wiki
large scale curated analysis of salivary proteome
driving biomarker detection
helps understand the porteins that are present in different conditions
metabolomic
detection methods to determine what metabolites are present
helps to answer the questions: What genes are tuned one or off in health vs disease in both the host and bacteria? what proteins are present or absent in health vs disease? what metabolites are present in health vs disease?
It is crucial to use these types of studies since we know presence alone is not enough to be able to predict disease. Need to know what changes in the bacteria occur prior to disease
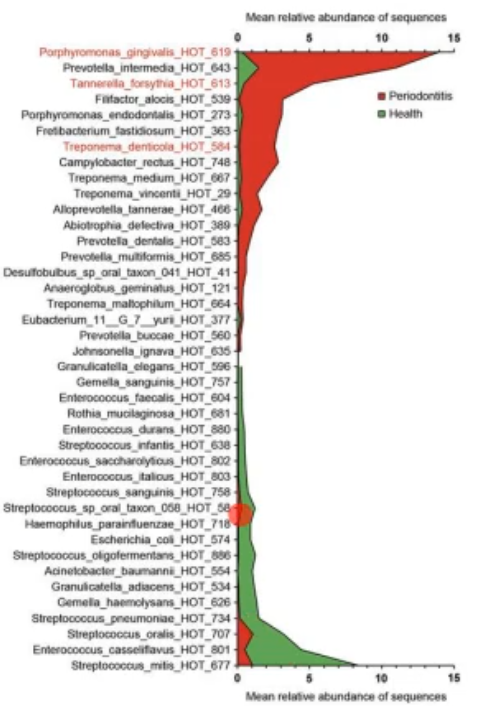
what does this picture show/tell you?
this shows you what you can do with transcriptomics, it shows the amount of genes expressed in disease vs health
Practical approaches for studying disease
differential plating for bacterial growth
good for when the infectious agent is unknown
establish if there is any antibiotic resistance
need the infectious agent alive and need to know culture conditions
16S sequencing
the infectious agent does not need to be alive
can take DNA directly from the site
Metagenomic/transcriptomics/proteomics/metabolomics
Do not need organism to be alive or grown
can take the material directly from the site
good to detect global changes
detect both host and infectious agent changes
Practical approaches for studying disease
what work for the various disease situations?
Bacteremia - blood infection
need to identify live bacteria
jus DNA isolation is not sufficient
Dysbiosis
“omics” analysis works to identify the shift
16S is useful to identify the population
Acute infection
16S is useful to identify
Differential plating to identify antibiotic resistance
metagenomics can be used to identify Ab resistance
Refractory disease (i.e. antibiotic therapy not working)
16S to identify if the correct organism
differential plating to identify organism to verify proper treatment or if infectious agent is resistant