BIO130 Final Exam
1/175
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
176 Terms
What are the two things animal cells have but plant cells don’t?
Lysosomes → degradation
Extracellular matrix
What are the three things plant cells have but animal cells don’t? And what are their purposes?
Cell wall → cell shape, protection against mechanical stress
Vacuoles (2 types) → degradation (like animal lysosome), storage
Chloroplast → photosynthesis
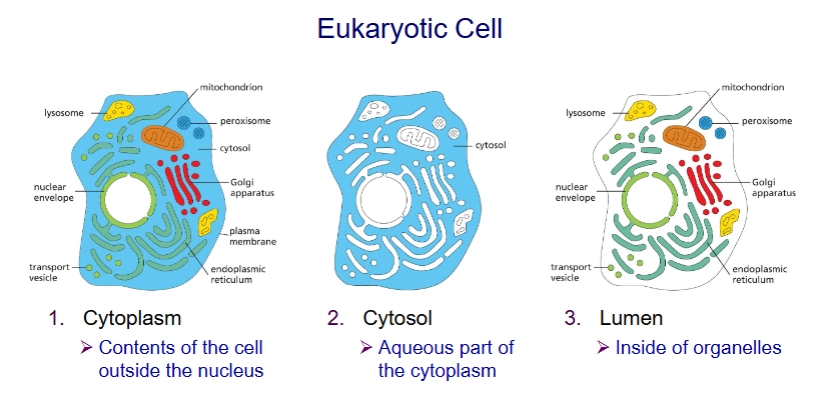
Cytoplasm vs Cytosol vs Lumen
Cytoplasm: everything but nucleus (includes organelles)
Cytosol: everything but membrane bound organelles (aqueous part of cytoplasm)
Lumen: inside of organelles
3 types of lipids that compose membranes
phospholipids, sterols, glycolipids
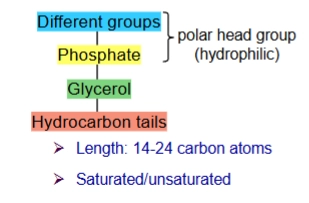
phosphoglyceride and structure
type of phospholipid with glycerol group
glycerol group: 3 C with O connected. 2 Os connects to the hydrocarbon tails, the other connects to the phosphate group.
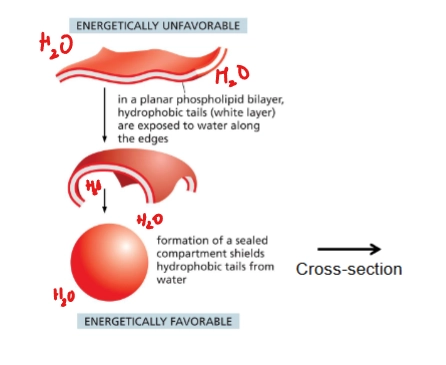
liposome
artificial lipid bilayer in sphere (because that’s energetically favorable)
Uses:
drug delivery into cells
study lipid and membrane protein properties
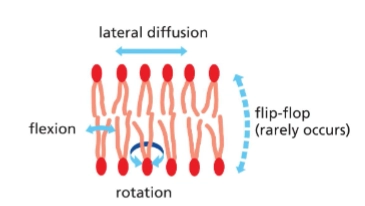
Describe the ways phospholipids can move (fluid)
Phospholipids rapidly move within each leaflet
lateral diffusion (side to side, deeper into plane)
rotate
flex
and rarely “flip flop” - move from one leaflet to another on their own
effect of temperature on membrane fluidity
lower temperature → more viscous, less fluid
factors affecting membrane fluidity (membrane composition)
phospholipid saturation
cis-double bonds increase fluidity (reduce tight packing)
phospholipid tail length (shorter tails increase fluidity because lipid tails interact less)
lipid composition (addition of cholesterol stiffens membrane)
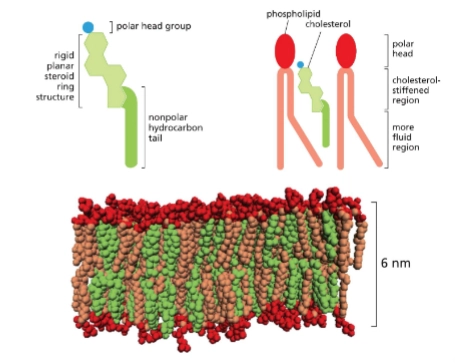
cholesterol
most common sterol in animal membranes.
Decreases mobility of phospholipid tails, makes plasma membrane less permeable to polar molecules.
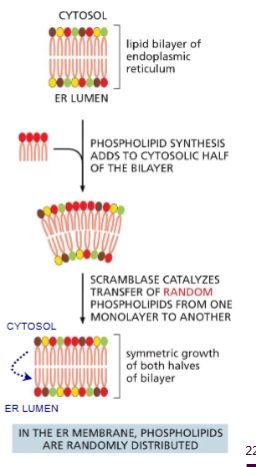
scramblase
aka “Phospholipid translocator”
Enzyme in the ER that catalyzes the flip-flopping of random phospholipids from one leaflet to the other.
Why is scramblase needed?
Phospholipids are synthesized in cytosolic leaflet of ER which means the membrane needs to be evened out.
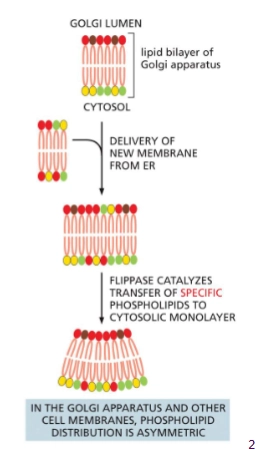
Flippase
The two faces of the plasma membrane have different lipids. This asymmetry is maintain by the enzyme Flippase in the Golgi membrane.
flip flops specific phospholipids to the cytosolic leaflet
4 steps to new phospholipids being made and added to the cell membrane
in ER membrane you make a ton of lipids
there’s some scramblases that flip them to other side to even things out
transport vesicles takes it to Golgi
Flippases sorts things out to get specificity —which from then on stays consistent
note: from then on, “stays consistent”
glycolipids/glycoproteins
Formed when sugar groups are added to lipids/proteins on luminal face. Protects the membrane from harsh environments.
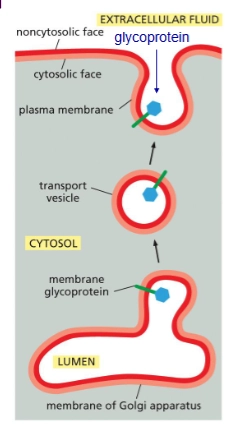
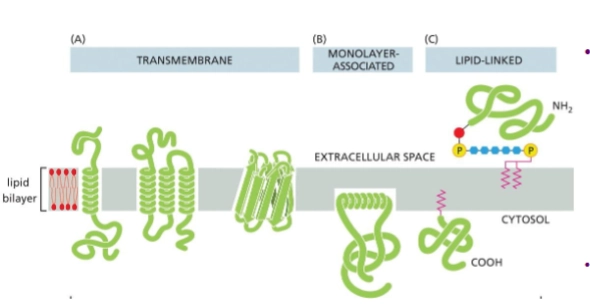
integral membrane proteins
membrane proteins that insert in some way into the lipid bilayer
transmembrane - pass through entire membrane
monolayer associated
lipid-linked
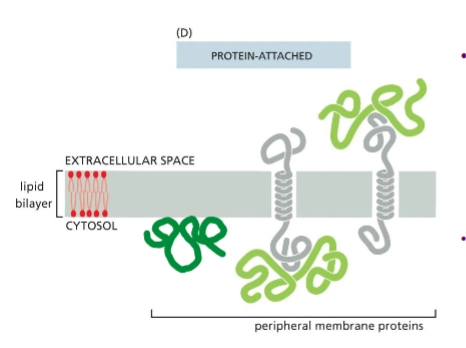
peripheral membrane proteins
no not insert into the lipid bilayer. They are associated with the membrane noncovalently, bound to either lipids or other proteins.
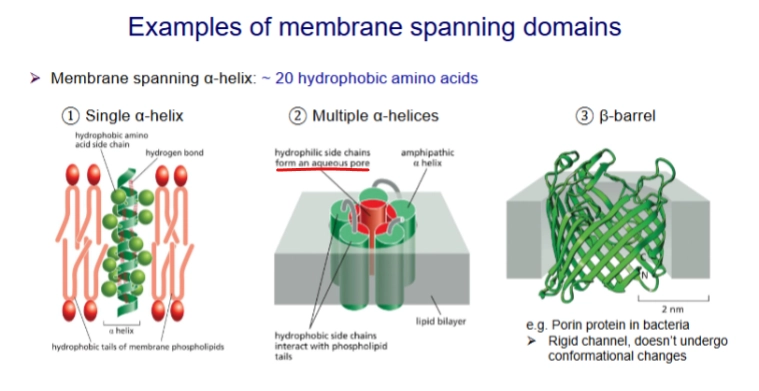
properties of transmembrane proteins
amphipathic - AA side chains are either polar or not polar
the nonpolar region is typically 20-30 hydrophobic amino acids.
Have specific orientations important to function
X-ray crystallography
technique used to determine the 3D structure of proteins
Turns proteins into crystals, light is shown through them, and the diffraction pattern is measured
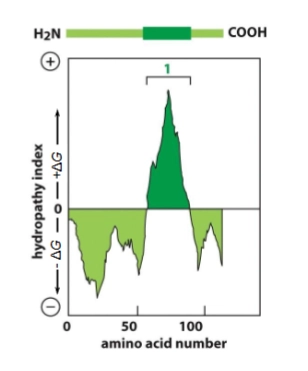
Hydrophobicity plots
x-axis: AA number. Starts on N-terminus, goes to C-terminus.
high y value = very hydrophobic, low y value = very hydrophillic
each stretch of 20-30 hydrophobic AAs is a transmembrane domain
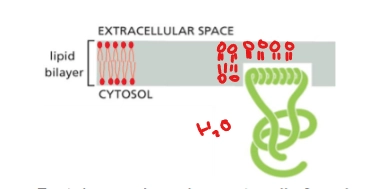
Monolayer Associated Membrane Proteins
A type of integral membrane protein
protein is anchored on cytosolic face by amphipathic alpha helix.
Ex. Sar1 - involved in membrane bending, vesicle formation
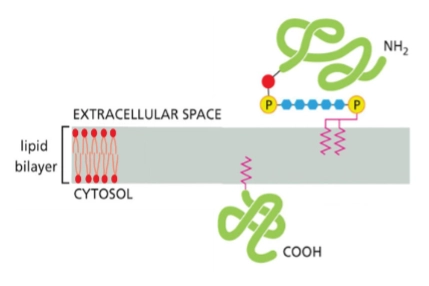
lipid-linked membrane proteins
type of integral membrane protein
2 types:
Protein with GPI anchor on noncytosolic face
Protein with lipid anchor on cytosolic face
Use of detergent
extraction of membrane proteins
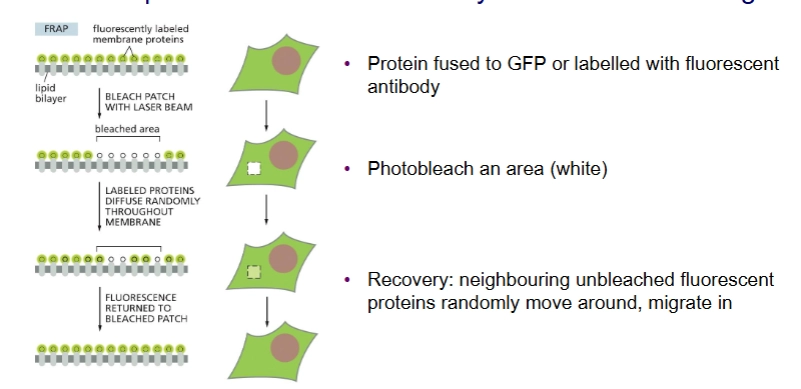
FRAP (Fluorescence Recovery After Photobleaching
Experiment to observe lateral diffusion of membrane proteins.
you can measure the speed of lateral diffusion by looking at the slope of recovery
membrane transport proteins
2 classes: channel proteins, transporter proteins
Transport polar and charged molecules (small, nonpolar molecules can just simply diffuse on their own)
selective
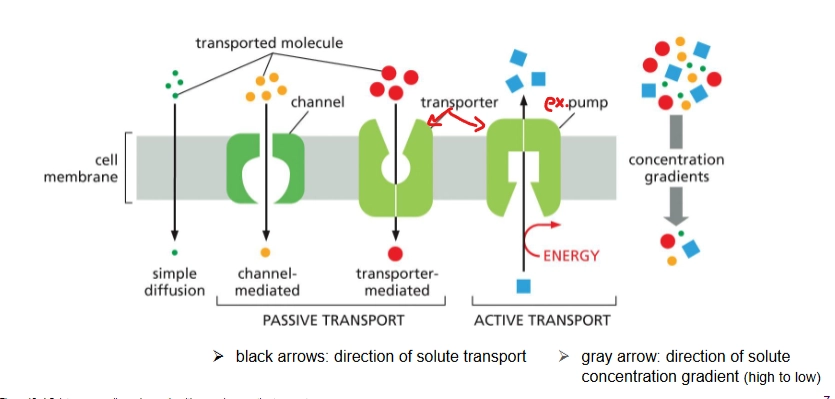
channel proteins
Type of membrane transport protein for passive transport
Binds weakly to transported molecule; does not change conformation a lot
Selective for size and charge
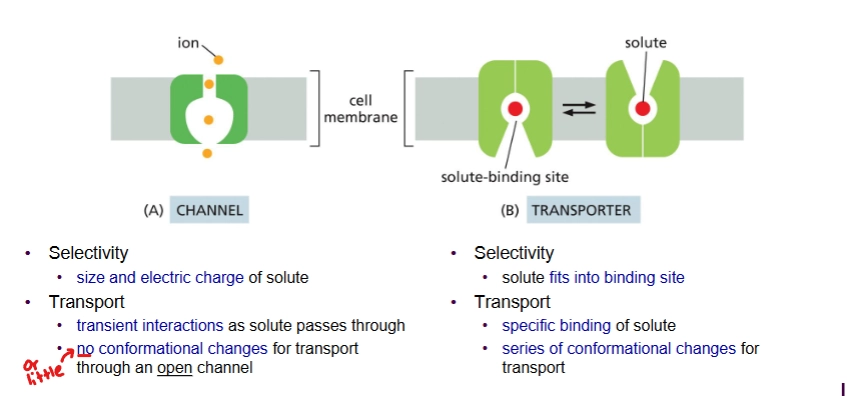
Transporter proteins
type of membrane transport protein - can do passive and active transport
Solute binds strongly; conformation changes a lot during transport
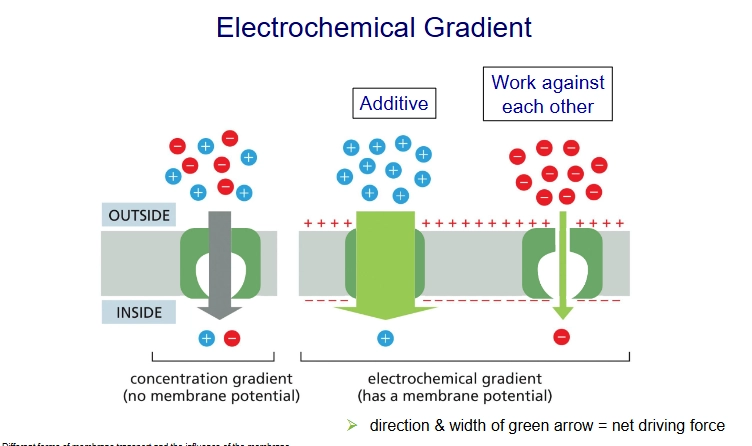
membrane potential
the difference in charge across the membrane
electrochemical gradient
concentration gradient + membrane potential (but we take concentration gradient to be more significant)
describe channel proteins in more detail
passive transport
Hydrophilic pore across membrane
Selective for size and charge
Example = ion channel
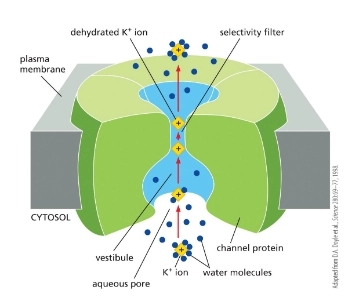
ion channels and their 2 kinds
passive transport of ions - selective for ion size and charge.
2 kinds:
Non-gated Ion Channels - always open
ex. K+ leak channel
Gated Ion Channels - signal required to open channel
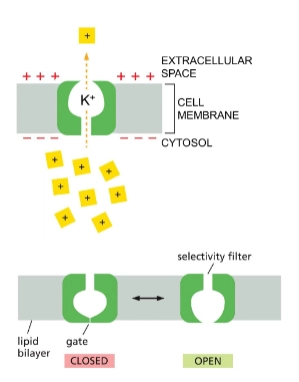
4 types of gated ion channels
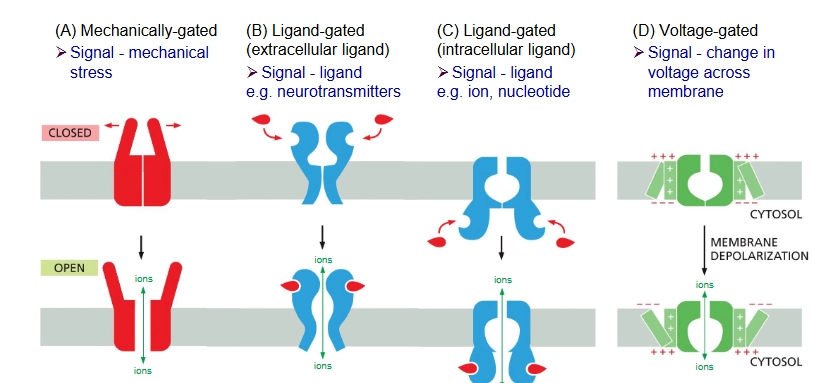
comparison of transporter-mediated diffusion and simple diffusion/channel-mediated transport (rates of transport)
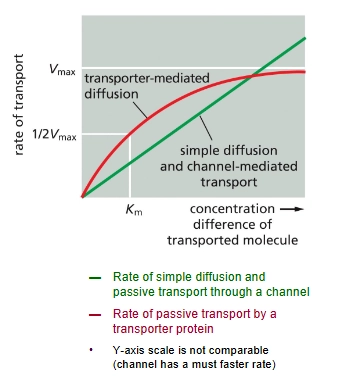
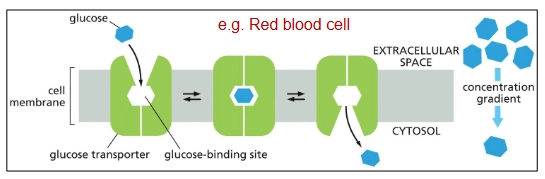
Uniport
Passive transport transporter protein (down electrochemical gradient)
Uni → “one” solute
ex. glucose transporter (GLUT Uniporter)
Gradient driven pump
Type of active transport transporter protein
1 solute down its gradient, 2nd solute against its gradient
ATP-driven pumps (ATPases)
type of active transport transporter protein
Requires ATP hydrolysis to move solute against its gradient
light driven pump
type of active transport transporter protein in bacteria
Describe the 2 types of gradient driven pumps
symport
2 solutes moved in same direction
antiport
2 solutes moved in opposite direction
For both, free energy from 1st solute moving down its electrochemical gradient is used to transport the 2nd solute against its electrochemical gradient.
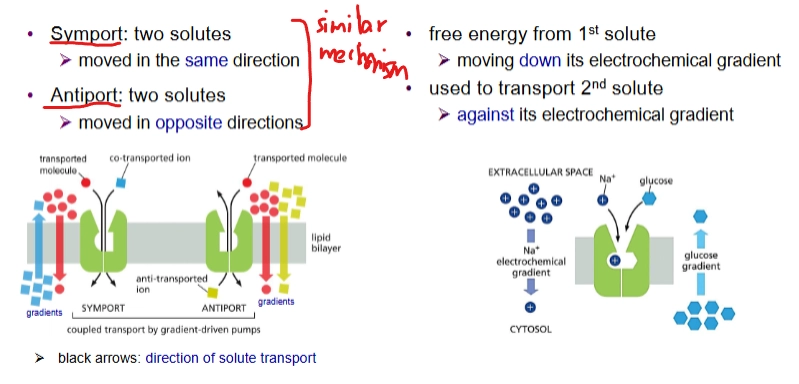
How is the Na+ electrochemical gradient maintained?
Symports and Antiports are present
Na+ - K+ pump (plasma membrane ATP-driven pump) in animal cells.
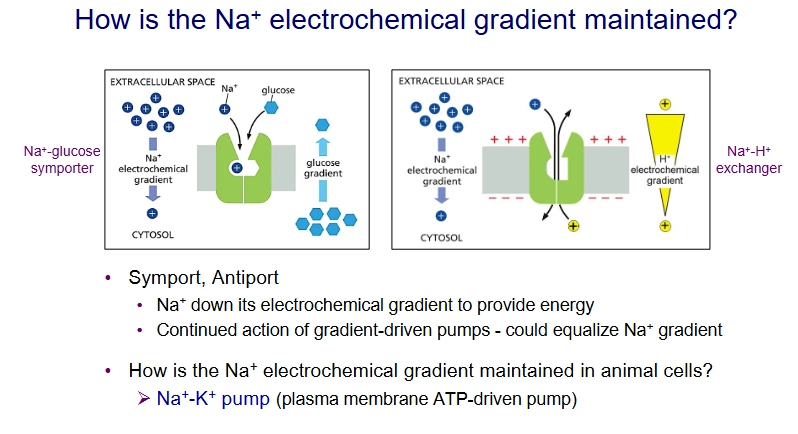
How is cytosolic pH regulated?
Antiport: Na+ - H+ exchanger on plasma membrane
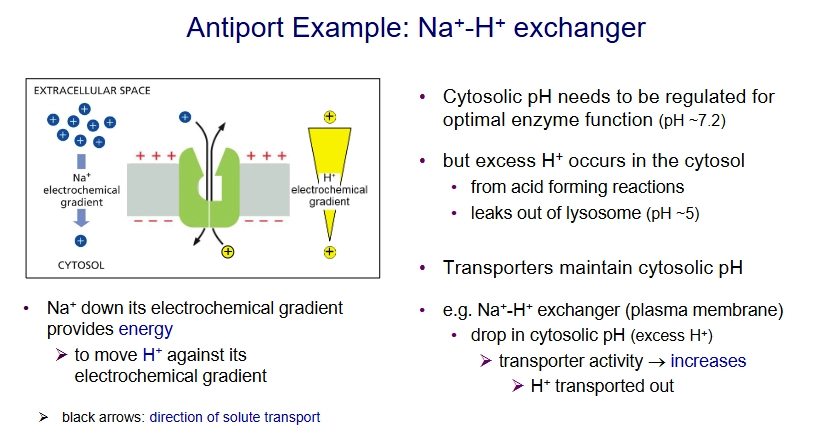
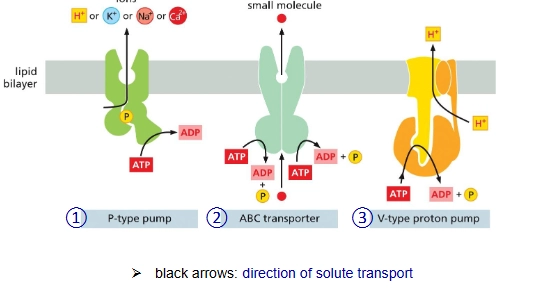
What are the three kinds of ATP-driven pumps"?
P-type pumps
ABC transporter
V-type pump
P-type pumps (ATP driven pump)
Generates and maintains electrochemical gradients
ex. Sodium Potassium pump (3 Na+ out, 2 K+ in)
ex. H+ pump in plant cell plasma membranes
* “pees itself” → phosphorylated during pumping cycle
(note: another example is Flippase)
why is the Na+ gradient important?
Transport nutrients into cells (e.g. glucose)
maintain pH
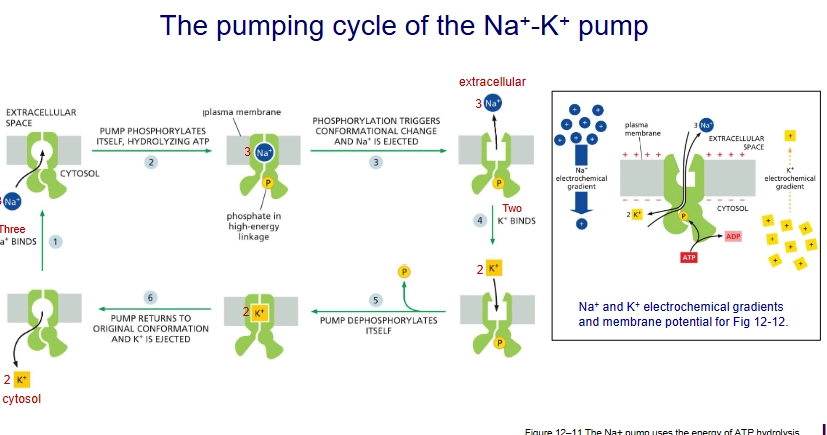
How does the Na+ - K+ pump work?
3 sodiums bind
the pump phosphorylyzes itself triggering conformational change
3 sodiums ions ejected, 2 potassium ions bind
pump dephosphorylates itself, returns to original conformation and the potassium is ejected.
ABC Transporter
type of ATP-driven pump
Uses 2 ATP to pump small molecules across cell membrane
V-type pump
Type of ATP-driven pump
Uses ATP to pump H+ into organelles to acidify the lumen
Found in lysosomes and plant vacuoles
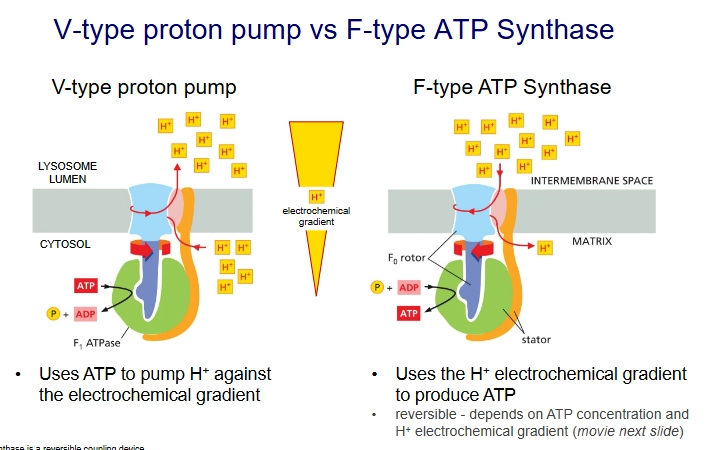
V-type proton pump vs F-type ATP Synthase
They are structurally related but opposite modes of action.
V-type proton pump uses ATP to pump H+ against the electrochemical gradient.
but F-type ATP Synthase uses the H+ electrochemical gradient to produce ATP.
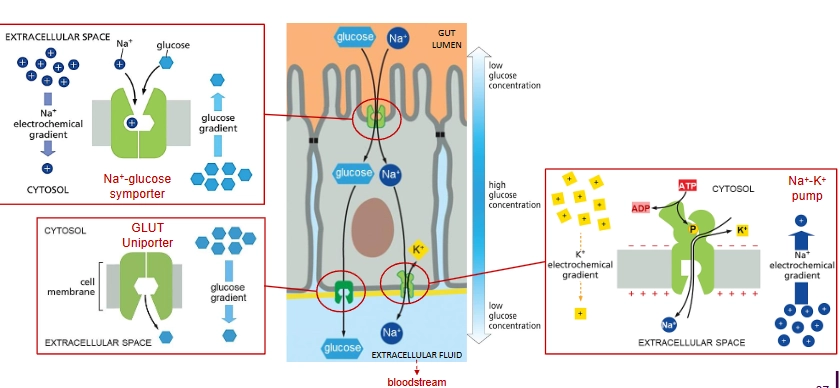
How is glucose transferred from the intestine to the bloodstream?
Lining the gut lumen are epithelial cells, each separated by tight junctions (so glucose can’t go between them).
Na+ driven glucose symporter on the apical domain brings Na+ and glucose into the cell, then Na+ - K+ pump and passive glucose uniport (GLUT Uniporter) on the basal domain remove Na+ and glucose from the cell.
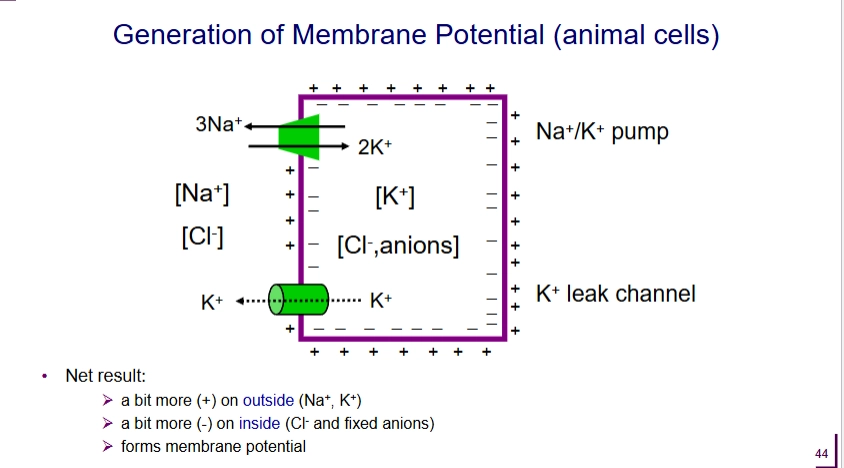
Which transporter proteins are involved with the generation and maintanence of membrane potentials?
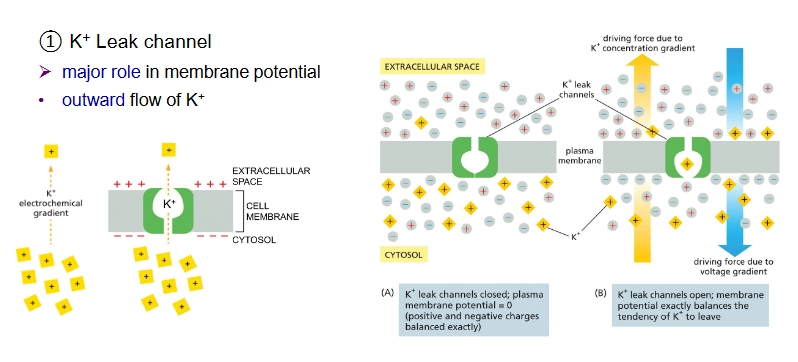
K+ leak channel (passive transport)
outward flow of K+
Na+ - K+ pump (P-type pump)
Net 1 (+) ion pumped out
^ net result = outside is more + than outside
for plants: H+ pump (P-type pump)
K+ leak channel
Important channel for maintaining membrane potential
Na+ - K+ pump
~10% of membrane potential
maintains Na+ gradient w low cytosolic [Na+] and K+ gradient with high cytosolic [K+]
Electrogenic:
3 Na+ ions pumped out
2 K+ ions pumped in
Net 1 (+) ion pumped out
![<p>~10% of membrane potential</p><ul><li><p>maintains Na+ gradient w low cytosolic [Na+] and K+ gradient with high cytosolic [K+]</p></li></ul><p></p><p>Electrogenic:</p><ul><li><p>3 Na+ ions pumped out</p></li><li><p>2 K+ ions pumped in</p></li><li><p>Net 1 (+) ion pumped out</p></li></ul><p></p>](https://assets.knowt.com/user-attachments/d0be8579-d020-4517-80bb-6d1f5c3ae288.png)
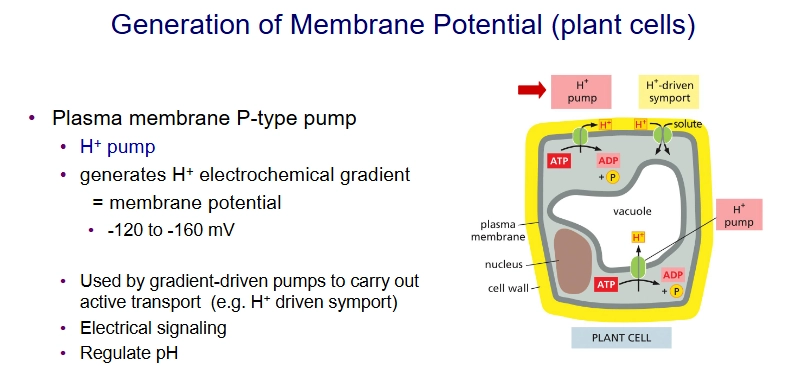
H+ pump and generation of membrane potential in plant cells
Generates H+ electrochemical gradient, which is used by gradient-driven pumps like H+ Driven Symporter.
good for electrical signaling and pH regulation
resting membrane potential
When membrane potential is as equilibrium - Voltage difference is steady
for animals, is from -20 mV to -200 mV
for plants, is from -120 mV to -160 mV
note: is from perspective of the inside of the membrane
What molecules diffuse rapidly?
small, nonpolar, and sometimes if small enough uncharged polar
ex. O2, CO2
briefly what are the functions of the organelles?
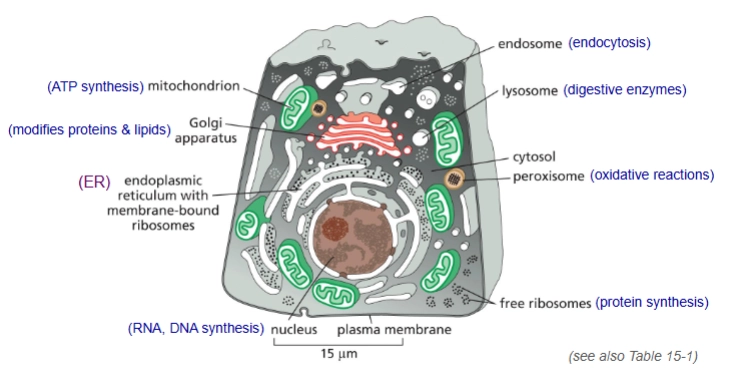
What is cytosol?
50% of the cell volume
Protein synthesis and degradation
many metabolic pathways
contains cytoskeleton
pancreatic exocrine cell
secretes digestive enzymes
rough ER and smooth ER
rough
has membrane-bound ribosomes
involved in synthesis of soluble proteins and transmembrane proteins for the endomembrane
smooth
phospholipid synthesis
detoxification
Which organelles are not membrane bound?
Nucleolus and Centrosome
Simply, how are proteins sorted?
If they have a sorting signal called a signal sequence they are directed to the right place.
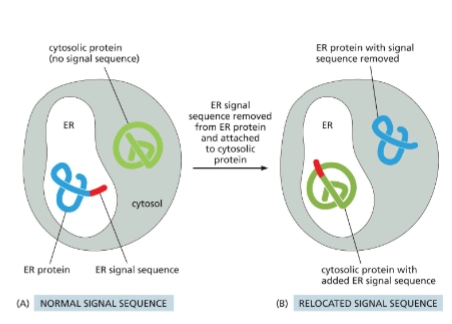
Signal sequence
AA sequence in a protein that directs protein to the correct compartment
recognized by sorting receptors
encoded in the genome
sorting receptors
recognize signal sequences and take proteins to their destination
2 ways protein sorting is done:
post translational sorting
proteins are already fully synthesized in the cytosol before sorting
co-translational sorting - for proteins going to ER
have an ER Signal Sequence
proteins complete synthesis on the ER membrane
Post-Translational Protein Sorting
Depending on destination, proteins are either folded or unfolded
folded: nucleus, peroxisomes
unfolded: mitochondria, plastids
How does protein sorting into nucleus work?
Post-translational. Protein already folded.
Travel through nuclear pores.
Protein has nuclear localization signal
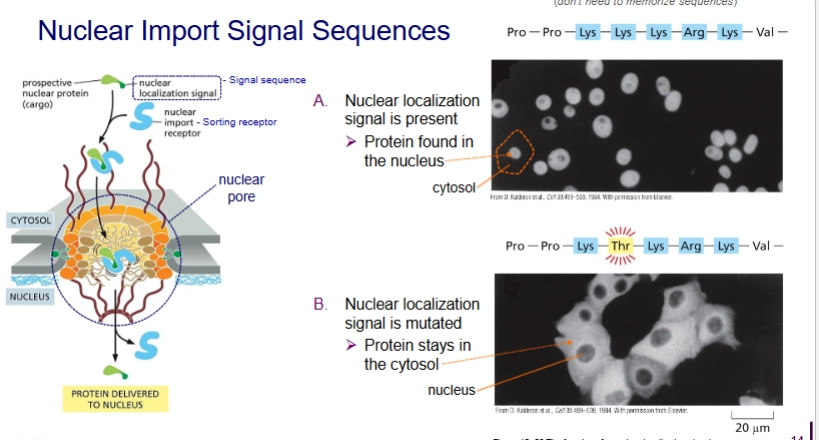
how are proteins sorted into peroxisomes?
Post translational. Folded.
Proteins imported through transmembrane protein complex.
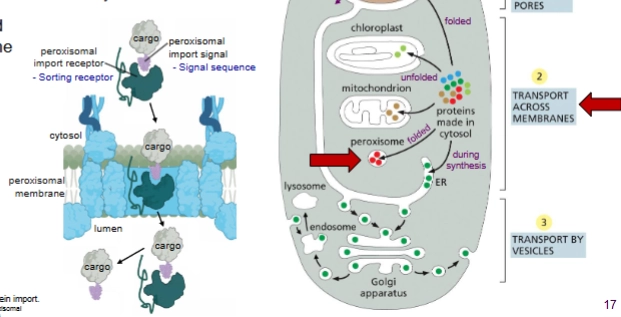
How are proteins sorted into mitochondrion and chloroplasts
Post translational. Unfolded.
Proteins are unfolded for import by hsp70 chaperone proteins
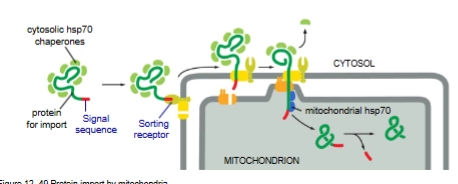
Co-Translational Protein Sorting
For proteins that enter the ER - the entry point to the Endomembrane system
ER Signal sequence is hydrophobic
Endomembrane system
ER, Golgi apparatus, endosomes, lysosomes
The intracellular compartments exchange lipids and proteins.
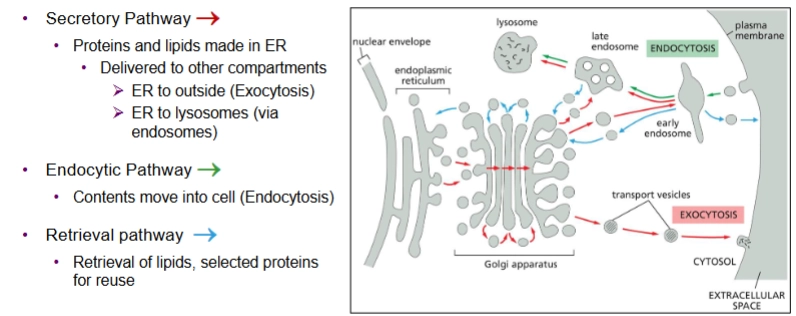
Has 3 pathways:
secretory pathway
endocytic pathway
retrieval pathway
In general what are the steps of co-translational sorting?
Translation starts on ribosomes in the cytosol per usual, but then once ER Signal Sequence is translated the protein is inserted into the ER (“co-translational translocation)
The process looks different depending on if the protein is:
soluble
or transmembrane
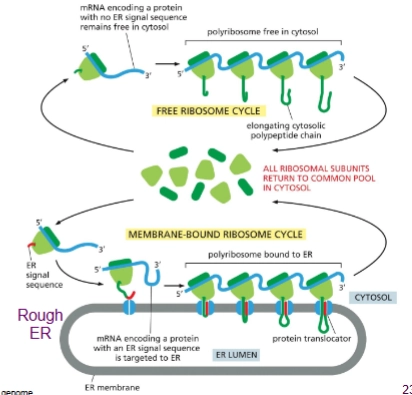
Describe the process of co-translational sorting of a soluble protein
SRP recognizes and binds to ER signal sequence once translated, translation stops. This binds to ribosome too.
This complex docks onto SRP receptor on membrane.
SRP leaves and protein synthesis continues with protein transfer into ER lumen.
The ER signal sequence is cleaved by signal peptidase.
Finished. Protein released into ER lumen and translocon closes. The ER signal sequence is eventually degraded.
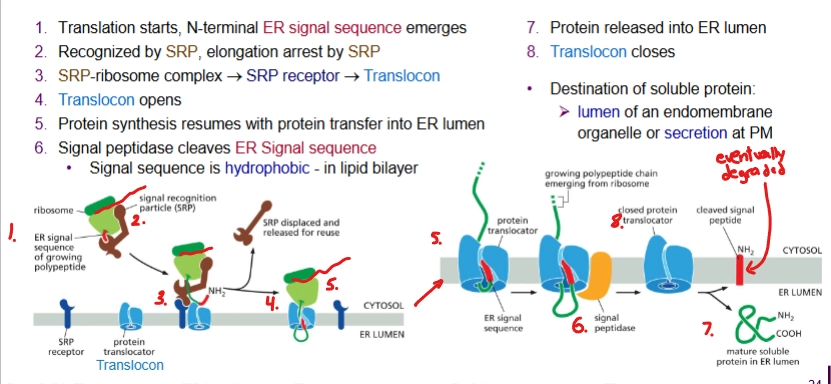
translocon
A gate for the polypeptide chain to be threaded through to get to the lumen. The ribosome docks on it.
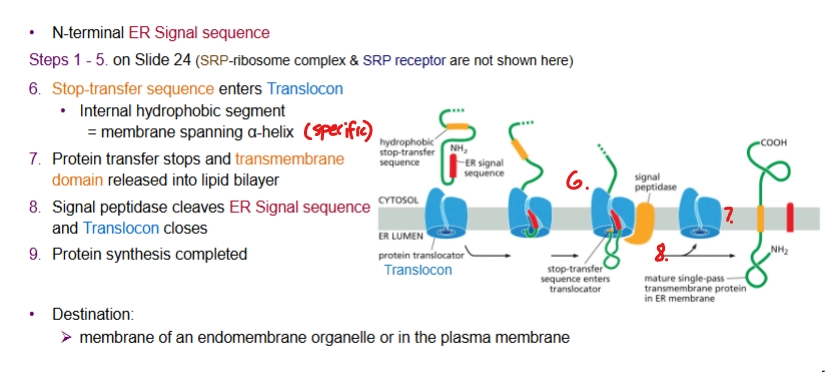
Describe the process of co-translational translocation of a transmembrane protein with N-terminal ER Signal Sequence
Starts the same as for soluble protein, but then a Stop-transfer sequence (internal hydrophobic segment) enters the Translocon.
Protein transfer stops and transmembrane domain is released into the lipid bilayer. Done → signal peptidase cleaves ER signal sequence and Translocon closes.
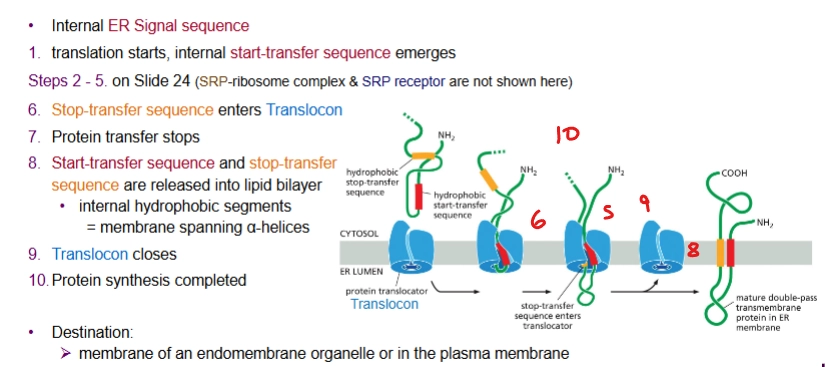
describe the process of co-translational translocation of a transmembrane protein with an internal ER signal sequence
almost the same as for when there’s a N-terminal ER Signal Sequence but the start-transfer sequence emerges first then the stop-transfer sequence. Transmembrane protein makes two passes at least (threaded through)
note: the start-transfer sequence is not removed
secretory pathway of endomembrane system
Proteins and lipids made in the ER then delivered to other compartments
Exocytosis = ER to outside
via vesicles. Vesicle membranes become part of the plasma membrane
or ER to lysosomes
2 kinds:
constitutive exocytosis pathway
regulated exocytosis pathway
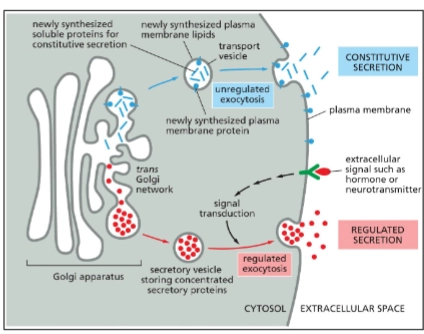
Endocytic Pathway of the endomembrane system
contents move into the cell (endocytosis)
via vesicles. Vesicle luminal contents come from extracellular space. The vesicle membrane is formed from the plasma membrane
Constitutive Exocytosis Pathway (1 type of secretory pathway)
In all eukaryotic cells
Continual delivery of proteins (both transmembrane and soluble) and lipids to the plasma membrane.
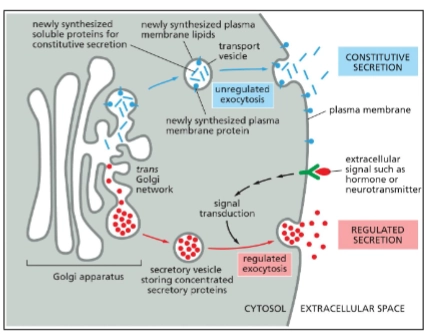
Regulated Exocytosis Pathway (1 type of secretory pathway)
Regulated secretion in specialized cells
Solute sorted in specialized secretory vesicles.
An extracellular signal is needed for vesicle fusion with PM and contents to be released.
ex. pancreatic beta-cells → insulin release w increased blood glucose
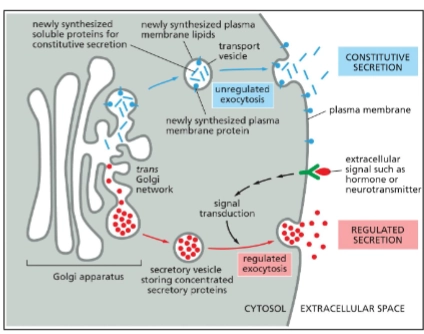
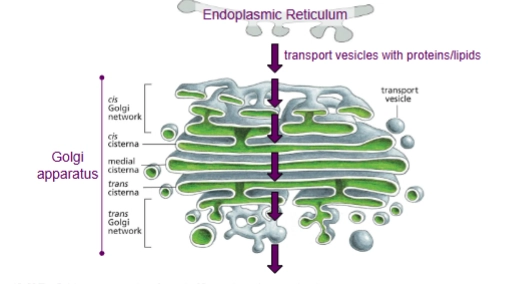
Golgi Apparatus’s role in the endomembrane system
Receives proteins and lipids from the ER, modifies them, and then dispatches them to other destinations in the cell.
Protein Glycosylation
one of the protein modifications that occurs in the Golgi Apparatus.
In the ER, a single type of oligosaccharide is attaches to many proteins.
Then these oligosaccharides are modified (glycosylation) in the various golgi sacs.
there’s different enzymes in each cisterna
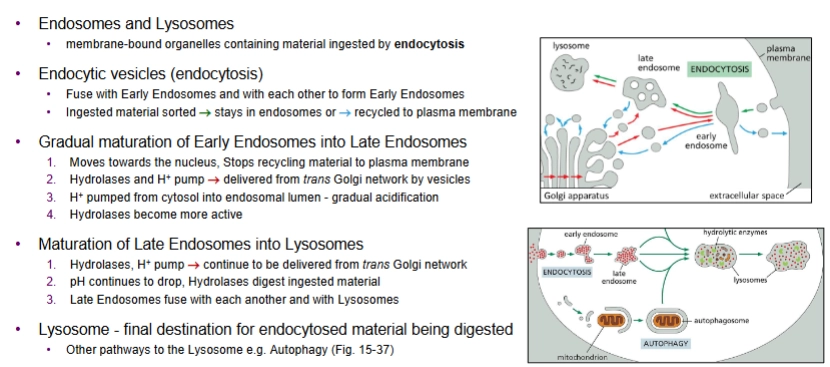
endosomes and lysosomes
membrane-bound organelles containing material ingested by endocytosis
describe the endocytic pathway
Endocytic vesicles fuse w Early Endosomes and with each other. Ingested material is sorted—either stays in endosomes or recycled into plasma membrane.
Early Endosomes gradually mature into Late Endosomes.
moves towards nucleus, stops recycling material to PM
Hydrolases and H+ pump are delivered from the trans Golgi network (compartment of Golgi apparatus) by vesicles.
H+ acidifies endosomal lumen
hydrolases become more active
Late Endosomes mature into Lysosomes
H+ pumps and Hydrolases continue to be delivered from the Golgi, so pH continues to drop and Hydrolases digest ingested material
Late Endosomes fuse with each other and with lysosomes
lysosome = final destination for endocytosed material being digested
How do lysosomes work?
Have ~40 types of hydrolytic enzymes (add water to break down molecules) (ex. proteases, nucleases, lipases)
Lysosomes need to be acidified by H+ pump
It’s membrane bound so protects rest of cell from digestion
and has glycosylated lysosomal membrane proteins on lumenal side to protect itself from its proteases
Transport proteins in lysosomal membrane transfer digested products (AAs, sugars, nucleotides) to cytosol.
When are signal sequences not removed after arrival to their destination?
Nuclear proteins, because nuclear membranes break down during cell division and the proteins need to “save their ticket” to return.
What are the 3 components of the cytoskeleton?
Actin filaments, Microtubules, Intermediate Filaments
which two components of the cytoskeleton are involved with cell division?
Actin filaments and Microtubules
Which type of microscopy should we use to see the cytoskeleton in detail? Describe how it works.
Transmission electron microscopy
uses beams of high energy electrons
resolution limit of ~ 1 nm
Which type of microscopy can be used to determine the location of proteins within a cell? Describe how it works.
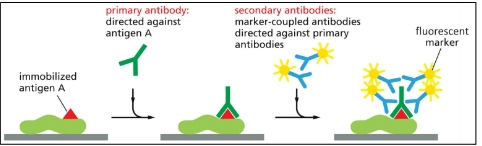
Immunofluorescence microscopy
cells are fixed
uses a primary antibody that binds to the target protein
uses a secondary antibody that binds to the primary antibody that is covalently tagged to a fluorescent marker.
microscope excites the marker.
rank the relative diameters of the three types of cytoskeleton filaments
actin filaments = small
intermediate filaments = intermediate
microtubules = large
What are Intermediate Filaments used for?
structural support
Describe the two types of intermediate filament proteins
Cytoplasmic IFs
in animal cells subjected to mechanical stress
provides mechanical strength
Nuclear IFs
nuclear lamina - 2D meshwork
formed by lamins - in all animal cells
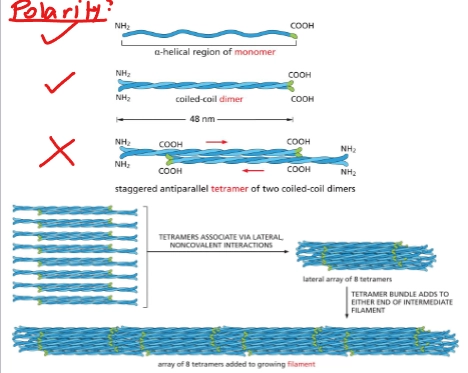
Describe the structure of cytoplasmic intermediate filaments
proteins coil together, ropelike - noncovalently:
monomer = protein with alpha-helical central rod
2 monomers = coiled-coil dimer
2 dimers = nonpolar staggered antiparallel tetramer
8 tetramers = 1 filament (nonpolar)
Keratin Filaments
The Intermediate Filaments in epithelial cells
They are anchored inside each cell at desmosomes (cell-cell junctions) and connect to neighboring cells.
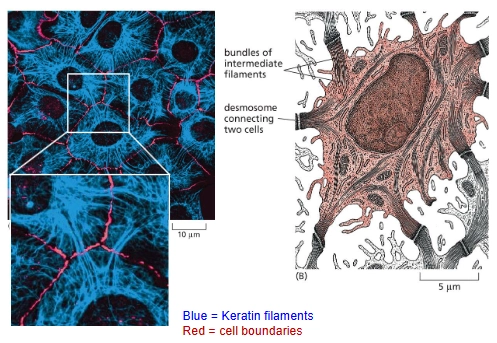
epithelium
sheet of cells covering an external surface or lining an internal body cavity
What are microtubules made of? Describe its structure.
Tubulin - long, stiff hollow tubes
Subunit = two closely related globular proteins
alpha-tubulin and beta-tubulin that both bind to GTP and to each other to form a polar tubulin heterodimer
13 protofilaments per tube. ← all noncovalent bonding
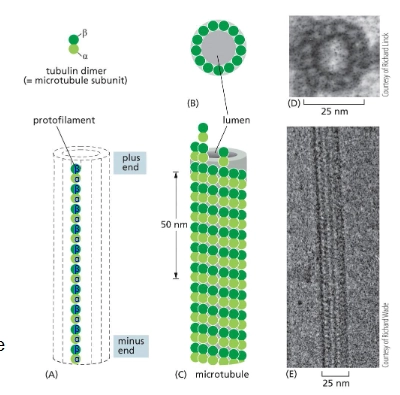
At which end of a microtubule does growth and assembly occur more rapidly?
+ end
Describe the importance of beta-tubulin.
After it’s been in the protofilament for a while, beta-tubulin cuts GTP to GDP (T-form/GTP heterodimer → D-form/GDP heterodimer). This has weaker binding.
So fast growth means there’s a strong GTP cap.
Microtubule Organizing Centers (MTOCs)
Where microtubules grow out of (has nucleating sites).
the (-) end is stabilized at the MTOC
ex. centrosome in animal cells
describe how dynamic instability works for microtubules - growth
growth: αβ-tubulin dimers bound to GTPs are added at the (+) end. Shortly after dimer is added, β-tubulin hydrolyzes GTP to GDP. If growth is fast, this hydrolysis is slower than the addition of new dimers so that there’s a strong GTP cap to be added upon.
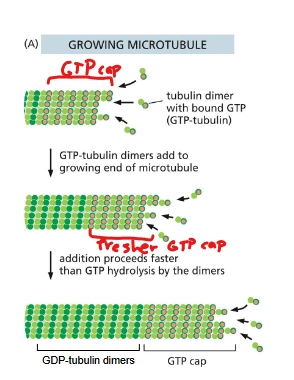
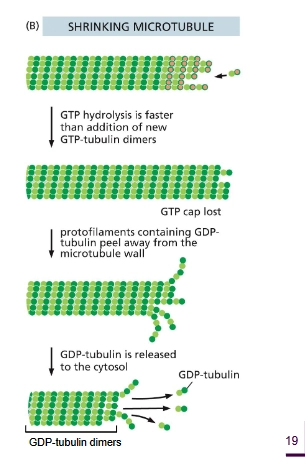
Describe how dynamic instability works for microtubules - shrinking
GTP hydrolysis is faster than the addition of αβ-tubulin dimers, the GTP cap is lost. The microtubule disassembles due to weaker binding.