BIOL 3000 Gene Complexity and Gene Structure
1/45
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
46 Terms
Complexity
The state of having many parts and being difficult to understand or find an answer to
Biological complexity
The result of hierarchical organization of cells
1960’s
The discovery of repetitive DNA sequences
“Junk DNA”
DNA that we do not know what function it has
What defines complexity?
The number and type of cells
The degree of cellular organization
Gene value
The amount of DNA that you have in an organism
C value Paradox
Comes from junk DNA
Excess DNA that is present in the genome that does not seem to be essential for the development or evolutionary divergence of an organism
T or F
Genome size correlates with organismal complexity
True
The Onion Test (T. Ryan Gregory 2007)
Ask the audience who has more complex DNA and see what they say
The onion has more DNA than us
G-value Paradox
The number of genes does not correlate with organismal complexity.
The number of genes because most of the organism has the same number of genes.
Percent of coding DNA sequences in the human genome
2-5% of the human genome (~20,000 protein coding genes)
Eukaryotic advancements
-Alternative splicing
-Transcription factors
Are you more advanced than an onion?
We don’t really know!a
Three classes of nucleotide sequence
Highly repetitive DNA sequence (HR)
Moderately repetitive DNA sequence (MR)
Single copy DNA sequence (Unique)
Highly repetitive DNA sequence (HR)
Comprises of about 10% of the human genome
Most is located in the heterochromatin regions around the centromere/telomere (“non-coding DNA regions”)
Occurs as variable length motifs (5-100 bp), in long tracks of up to 100Mb; there are repeats of certain bases and have long tracks
Present at >106 copies per genome
Postulated functions include structural and organization role to nothing more than junk
e.g. - alpha satellite DNA
Alpha satellite DNA
Two to more then 30 repeats of 171 bp tandem repeats
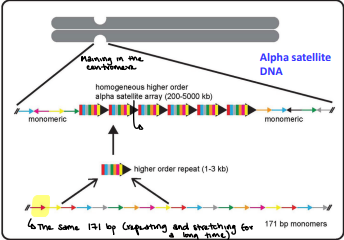
Moderately repetitive DNA sequence (MR)
Comprises of about 30% of the human genome
Found mostly throughout the euchromatin
Average 300 bp in size
Present between 10-105 copies per genome
Also includes ‘redundant’ genes for histones, and ribosomal RNA and proteins, (gene-products present in cell in large numbers)
Functions in length mutability, modulation of transcription, factor binding spanning between promoter elements, and alternates splicing
e.g. Microsatellite DNA, interspersed repetitive DNA
Microsatellite DNA
Repeating sequences of 2-30 bp of DNA
Variable number of tandem repeats typically occurring in non-coding regions of the genomes
Ranging in size of the repeating
Occurs through a mutation process known as “slippage recognition”
Useful genetic markers that tend to be highly polymorphic
Slippage recognition
Happens in DNA replication
The repeats causes the DNA to slip forwards or backwards
Interspersed repetitive DNA
Transposable elements
Single copy DNA sequence (Unique)
Comprises about 1-5% of the human genome
Found throughout the euchromatin
~20,000 protein coding genes
“Coding DNA regions = GENES”
Present at single or low copy number per genome
Percentage of HR, MR, and unique DNA
10% (HR) + 30% (MR) + 5% (Unique) = 45%
Giemsa Stain
Highly repetitive: Heterochromatin (tightly coiled)
Moderately repetitive: scattered throughout Euchromatin
Unique: Euchromatin - genes
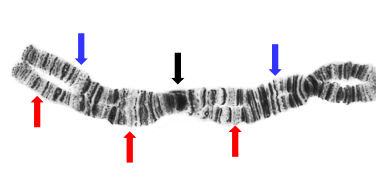
Gene
The basic physical and functional unit of heredity
How one person passes along traits
A sequence of unique nucleotides (genotype) that carry the genetic information which is to be expressed (phenotype)
What are the instruction manuals for our bodies?
Genes
Molecular level “gene”
DNA sequence that gives rise to an RNA molecule
“Transcriptional unit”
The template for the RNA
Transcribed region
DNA to RNA
Part of this region contains the information that specifies an amino acid sequence
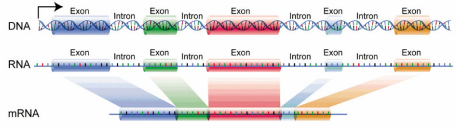
Exon
“Coding sequences” = phenotype
Intron
“Intervening sequences” = areas of genes that generally don’t code for phenotype
5’ Untranslated Region
mRNA that is directly upstream from the initiation codon, before the start codon
3’ Untranslated Region
Section of the messenger mRNA that immediately follows the translation termination codon
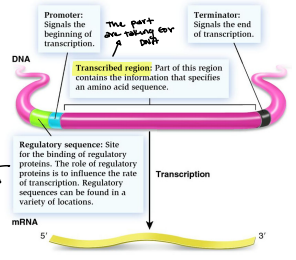
Promoter
DNA sequence onto which the transcription machinery binds and initiates transcription
Signals the beginning of transcription
TATA Box
Highly conserved sequence in DNA serving as the binding site for transcription factor binding
Allows transcription to be turned off or on (on a basal level)
Core DNA sequence is 5’-TATAAA-3'
“ON/OFF” for transcription
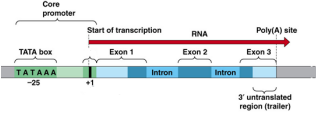
Regulatory Sequences- Enhancers
They enhance or inhibit and affect the rate of transcription
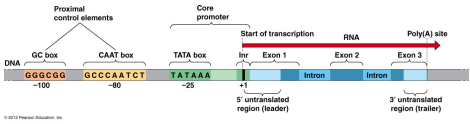
CAAT Box
5’-GGCCAATCT-3’ consensus sequence that occurs upstream by 60-100 bases to the initial transcription site
Typically required for inducible genes to be produced in sufficient amounts
GC Box
Region of DNA that can be bound with proteins (activators) to activate transcription of a gene or genes
Regulator proteins
Proteins bind to them to modify their speeds
“Ramped up” level of transcription
Termination
“The end of the gene”
Tells the DNA polymerase to leave
No one else can turn off transcription, only terminator
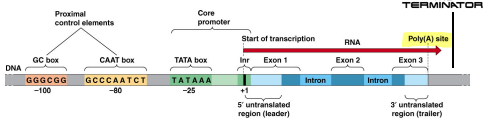
Terminator
A section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription
Regulatory sequence
Site for the binding of regulatory proteins. The role of regulatory proteins is to influence the rate of transcription. Regulatory sequences can be found in a variety of locations. Controls the mechanism
Types of Genes
Solitary genes
Duplicated genes
Multigene families
Pseudogenes
Repeated genes
Solitary genes (unique)
A single copy of a gene (haploid situation); two copies in diploid
Comprises the bulk of euchromatin
Duplicated genes
Process by which a portion of a chromosome is duplicated resulting in an additional copy of a gene
Results in a copy of the original gene called a paralog gene
Either of the two genes may mutate and change the original function of the gene
Usually occurs due to an error during meiosis
Many copies of a single gene (management)
Multigene families
Set of several similar genes, formed by duplication of a single original gene, and generally with similar biochemical functions
Most often located in similar regions of the chromosome
Most often used or synthesized at different times
One class/family genes but they have different functions
Pseudogenes
Dysfunctional relatives of genes that have lost their protein-coding ability
Often the result of multiple mutations within a gene
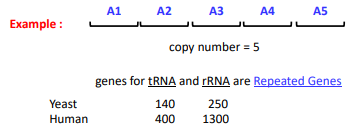
Repeated genes
Multiple copies of small genes clustered throughout the genome at specific sites
Present in high copy number
Many times present in a “head-to-tail” configuration
