mRNA Localization
1/55
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
56 Terms
Why is mRNA used to make proteins instead of DNA?
RNA is more expendable than DNA; we wouldn’t want to damage the DNA, Another reason is that the more steps there are in a process, the more control we have over it
How can we track protein expression?
Clone a gene for a protein next to a fluorescent marker so it has it attached and we can see it
How do we visualize gene expression form mRNA? (YFP/MS2)
Clone YFP into the gene of interest. YFP will bind to MS2 viral RNA that has a hairpin loop structure that will bind to viral MS2 protein that is albled with fluorescence
How can protein localization be controlled?
Either the mRNA translates a protein and it is transported or the mRNA is transported then translated (could also be getting translated as it moves)
Why is localizing the mRNA to the region the protein is needed more economical?
More cost effective, limits protein damage, protein only expressed where needed, harder to transport protein than mRNA
What is an example of RNA localization to create protein gradients?
Drosophila embryo development
How does mRNA localization work in Xenopus?
Vg1 mRNA is repressed until it is localized and anchored to the pole of the cell, important for embryo development
Do yeast produce asexually or sexually?
Both, budding off is asexual, in order for there to be sexual reproduction they must be opposite mating types
What are yeast mating types?
“a” and alpha
How does a mother yeast cell produce cells to mate with? (mRNA localization explanation)
Mother cells will begin to produce ash1 and it will all be localized to the daughter cell, this will prevent the daughter cell from switching mating types (will be the same as the mothers), after a while the mother cell will be able to switch it’s mating type and mate with the daughter cell as all the ash1 preventing mating type switching has been localized to the daughter cell
how does mRNA transport relate to axons and neurons?
Neuron is the center with axons spreading out and away, the neuron requires B-actin at these axon location, it is hard to transport protein and it will be diluted so the cell transports B-actin mRNA to the site instead
How does mRNA localization help fibroblasts?
Fibroblasts move through ECM to help with tissue repair, they move through the ECM by producing pseudopods that contain B-actin, and the actin binds outside and helps with movement, this is orientation and direction specific, this means that B-actin mRNA needs to be localized to correct location to aid with movement
How does mRNA localization help the mitochondria?
Hard to import proteins into the mitochondria so it is easier to move the mRNA in and then translate it in the mitochondria
How does IN-SITU hybridization work for mRNA localization visualization?
Take a cell, mic the cell, make RNA probe with radiolable and hybridize it to RNA/DNA of interest and watch localization
What is true of radiolabels?
they have a high signal to background ratio
What is FISH?
This is in situ hybridization but uses a fluorophore instead of a radiolabel, better signal to background ratio, only one fluorophore per probe so hard to see single mRNAs but easier for clusters
What are mRNA stains?
stain all mRNA, hard to differentiate, can synthesis mRNA then stain and introduce but is very inefficient
What is smFISH?
make multiple probes for the same mRNA at different points each with the fluorophore so the signal for single mRNA strands is much higher
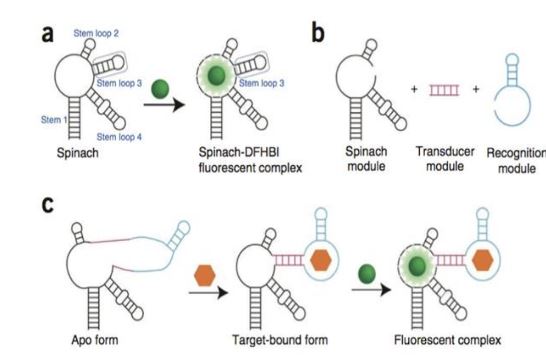
How do aptamers work for studying mRNA localization?
Aptamers are pieces of mRNA and these mRNAs bind to a fluorophore when they bind to their mRNA target, you take the aptamer and bind a transducer module (mRNA) and a recognition molecule (mRNA), the protein of interest will bind to the recognition molecule and cause a conformational change that allows a fluorophore to bind the aptamer and produce a signal
What are fluorescent tagged RNA binding proteins?
Target protein that binds the mRNA with GFP and watch the light as it binds
How does the mRNA MSP complex work?
label an mRNA so when it is transcribed it will have MSP hairpin loops in it’s structure that bind to MCP tagged with GFP
What is the difference between active and inactive genes?
RNA pol is more effective at binding the promoters of active genes
What happens before splicing? How is splicing important to mRNA localization?
5’ capping and poly A tail, they may bind and circularize to protect from degradation pre splicing, splicing is important so localization is correct and protein synthesis is correct
How do NUPs check for proper splicing before mRNA diffusion across the pore?
Some NUPs scan mRNA and look for EJC (spliceosomes) at each exon intron junction, if there is not the mRNA is rejected and doesn’t diffuse and is degraded by RNase
What serve as physical contact for transport of mRNA through NPCs?
Spliceosomes (EJCs)
What are the three types of mRNA that can be transported?
Lnc (long non-coding mRNA), intronless mRNA, processed mRNA with EJCs
What must Lnc and intron less mRNA have to be transported?
GC rich region at 5’ end so it can bind to RBM that binds to TREX, TREX binds NXF1
How does processed mRNA get transported?
EJCs have conformational change when splicing is correct, attract TREX, spliceosome falls off and TREX attracts NXF1 for transport across NPC
What happens when the mRNA leaves the nucleus?
it is remolded and export proteins are removed and new proteins for cytoskeletal binding are added, removes TREX and adds RBPs
What is different about RNA in the cytoplasm vs nucleoplasm?
In the nucleus it is free floating and in the cytoplasm it is bound to the cytoskeleton
What are RBPs and RNPs?
RNA binding proteins and RNA nuclear proteins
How can we visualize/purify RNPs?
Can take the mRNA and spin it to separate the RNPs, can purify the mRNA and crosslink the RNPs then remove them and analyze by mass spec
What is RNA capture?
Used to identify RNPs, purify an mRNA with antisense zip code sequence, the peptide will bind to the mRNA and has a SRP sequence that directs it to a given location (binds to SRP), you take a column with anti-SRP and can separate the proteins you want, then analyse by mass spec
What is RNA-affinity purification strategy for RNP analysis?
Clone the gene of interest into a plasmid and you have the MSP sequence fused to the gene in the plasmid MSP has lots of hairpins that bind to MSP binding protein MS2 (you have a MSP tag on your gene of interest), you have it so the MS2 is bound to biotin and then run the whole thing through a column with streptavin that binds biotin to sperate.
What is the role of SMN1 in spinal muscular dystrophy?
SMN1 and SMN2 are responsible for motor neuron genes, when there is a mutation in exon 7 there is binding of a protein that recruits improper splicing proteins that leads to dysfunction
How is spinal muscular dystrophy treated?
hnRNP that binds antisense oligonucleotide and blocks the antisense splicing event (fix that shit)
What is important for the function of zip codes?
Some zip codes have primary sequences (sequence dependent) and other rely on the structure of the motif
What happens if there is a zip code vs no zip code?
No zip code is translated by free ribosomes in the cytoplasm, stress granules, and p-bodies basically they go to RNA depots while those with zip codes are targeted to organelles (SRP sequence for ER)
What is an anchoring site?
Zip code can deliver an mRNA to a pole and have it anchored there for added protection and so it stays, an example is NANOS

Explain the Oskar Nanos system?
Oskar protects Nanos at the pole, Nanos binds to an anchor after being loclaized to a pole in the oocyte and is stuck there, SMAUG is a protein that degrades nanos and is active everywhere in the cell except this pole, this is beacuse Oskar protects nanos from smaug by degrading it at the pole, so oskar must also be localized to the pole aswell as nanos

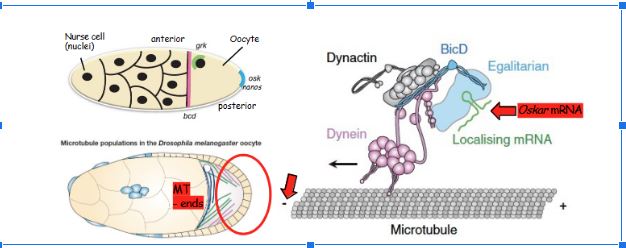
How is Oskar localized to the pole?
Cytoskeletal microtubules, Oscar mRNA is made at one end of the developing cell (old cell to new) and conatins a hairpin structure that contains a zip code, Oskar mRNA zip code binds to eggalitarian and BicD that are responsible for binding to dyeinin and dynactin
What is nanos anchored to?
Actin
How does smaug degrade nanos?
Binds to the 3’ UTR zip code and degrades it

How does dyeinin and dynactin based transport work? What uses this?
This is how mRNAs are transported on microtubules, dynactin is bound to the complex and is required for loading the dyeinin that has two legs, and the loading makes the leg walk further on the filament, one leg is moving while dynactin reloads the ATP to make the other dyeinin leg move, Oskar transport

How does Oskar based transport work in regard to movement along the microtubule filament?
Oskar has a zip code, and this zip code binds 2 eggalitarian molecules (must be a dimer to work), then BicD binds to a dimer of eggalitarian (not single), binding the dimer opens the BicD tail so it can bind to dynactin, then BicD and dynactin bind to dyeinin
How does actin based movement work?
Myosin binds to kinesin and walks along the actin filaments only towards the positive end
What is different between microtubule based transport and actin based transport?
Microtubule uses dyeinin and dynactin and also goes both directions, actin bases is only one direction and uses myosin (towards the positive end)
What do both actin and microtubule transport do the same?
They both move along filaments by binding kinesin, but actin-based does this through myosin and microtubule-based uses dyeinin
How does Ash1 transport work in relation to actin-based movement?
Ash1 prevents breeding type switch, the Ash1 has a zip code that binds myosin and transports it along an actin filament in a single direction to the mother cell, Ash1 is covered in RNPs that inhibit transcription and ZIP binding proteins that help myosin binding
What do RNPs that bind mRNA during localization do?
Help localization, transport accorss nucleus, bind to cytoskeletal elements, ZIP binding proteins for localization, inhibit translation
What are the RNPs of Ash1, what do they do?
RNPs bind to the ZIP code and help bind to mysosin for transport to the daughter cell, some inhibit transcription, the ZIP code is a hairpin structure and forms an RNP complex, Loc1 binds and helps cross the NPC then dissociates, then PUF binds to block transcription and Shep2 and Shep3 bind ZIP to help myosin binding, once in the daughter cell Ash1 mRNA will anchor
How are translation inhibitors removed?
By phosphorylation
What is cargo hitchhiking in mRNA localization?
This is when organelles are moving across microtubules and mRNA hitches a ride, an example is Vg1 mRNA attaching to the ER as they both localize to the pole of the cell in frogs
What is endosomal localization? What is it used for?
endosome is a dynamioc organelle that is transported, mRNA enters the endosome and is transported with it, this is for mRNA that is co-tanslated as it is moved around the cell, this is so that it can deposit protein across the cell as it is localized
How is spetin mRNA transported?
Spetins form cytoskeltal rings important for hyphae development, they are spread out evenly in the cell, this is co-translation as the mRNA binds to Rrm4 that binds Upa already bound to the endosome, a ribosome atrtaches and septins are translated and deposited ad the endosome is transported, however it turns around at the pole (microtuiubles go both ways), this extra time allows for more spetin to be deposited the poles
How do we know that ZIP proteins bind to mRNA?
Take the septin mRNA and attach it to viral MS2 that is fused to a transcriptional activator and it will bind to Rrm4 bound to another activator so when they bind and come close you get transcription of a gene that you can detect and see that it worked