14. Lac Operon
1/74
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
75 Terms
beginning of the RNA chain is
+1 (first synthesized nucleotide - beginning of the 5’ UTR in mRNA)
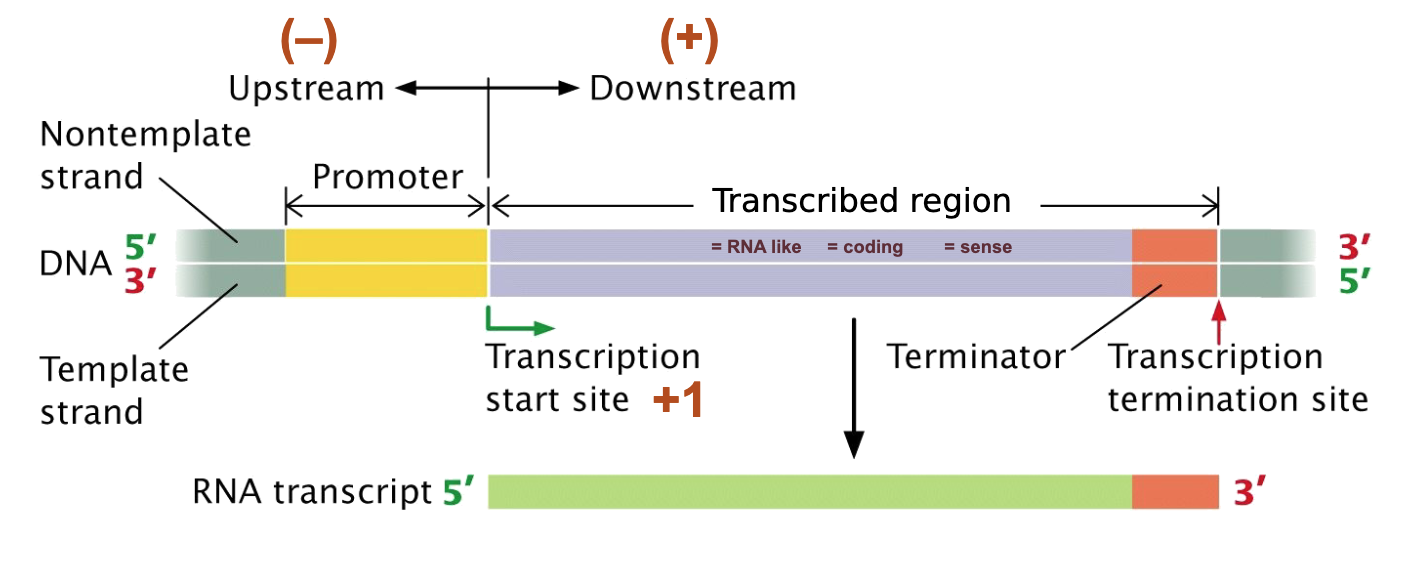
upstream (-)
from the transcription start site to the left (promoter - RNAP binding site etc.)
downstram (+)
from the transcription start site to the right
Initiation: binding of RNAP to the promoter
conformational changes of both promoter and RNAP
formation of open complex (bubble)
RNAP synthesizes approx. 10 bases in this phase
Elongation: enhanced grip on template - polymerization
continues polymerization (faster than first 10 bases)
unwinds DNA in front
does little bit of proofreading (lower fidelity)
anneals DNA behind
dissociates growing chain from the template
Termination: end of transcription
different ways of finishing transcription
regulatory proteins (trans-factors) that bind to
certain regulatory sequences (cis-elements)
regulation of gene expression through two types of trans-factors
activators (“help” RNAP to bind to the promoter) or
repressors (prevent RNAP from binding to the promoter)
bacteria have a
single RNA polymerase
processive enzyme; transcribes about 50 nt/sec
composed of 6 subunits with distinct functions
E. coli RNA polymerase 6 subuntis
2 alpha subunits
2 large Beta subunits
1 omega subunit
one sigma subunit
two alpha subunits
interaction with regulatory proteins (and DNA - control of initiation frequency)
two large beta subunits
Beta and Beta’ are catalytic subunits - inhibited by rifampicin
one omega subunit
required to restore denatured RNA polymerase in vitro to its fully funcitonal form
one sigma subunit
interaction with promoter - initation
6 subunits = holoenzyme and without omega subunit
core enzyme
e.coli has various sigma subuints
sigma subunit directs the holoenzyme to specific promoters
each different omega recognizes a specific promotor sequence
specific gene expression - fast initiation under specific conditions
consensus sequence (s) in general are the
most commonly found nucleotide sequence of DNA or RNA, of related function, found in different locations or organisms
promoters:
sites that are critical for binding of RNAP
promoters are asymmetrical:
direct positioning of RNAP
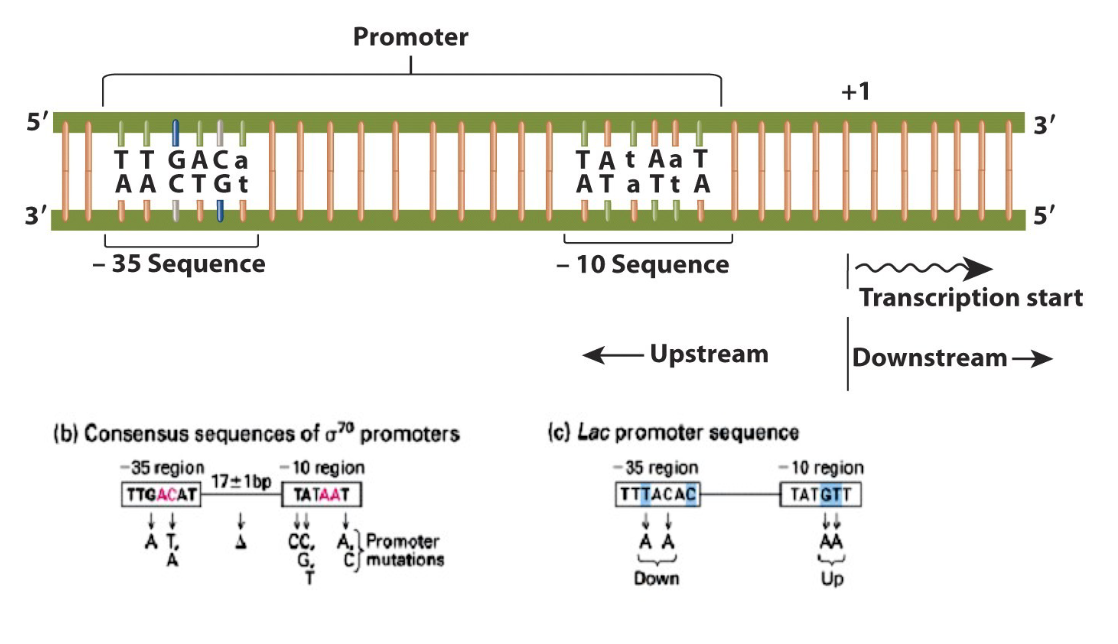
two consensus sequences in RNAP promoters
-10 region (TATA box)
-35 region
RNAP binds to dsDNA however,
nucleotide sequence that is on the coding strand (in 5’-3’ orientation) and precedes teh coding sequence for the mRNA is usually given/written
promoter determines
which strand will serve as a template
starting point for transcription
strength of polymerase binding
promoter strength
measure of how strongly RNAP is bound
strong promoter
high frequency of initiation of transcription
weak promoter
low frequency of initiation of transcription
Francois Jacob, Jacques Monod in 1950s found
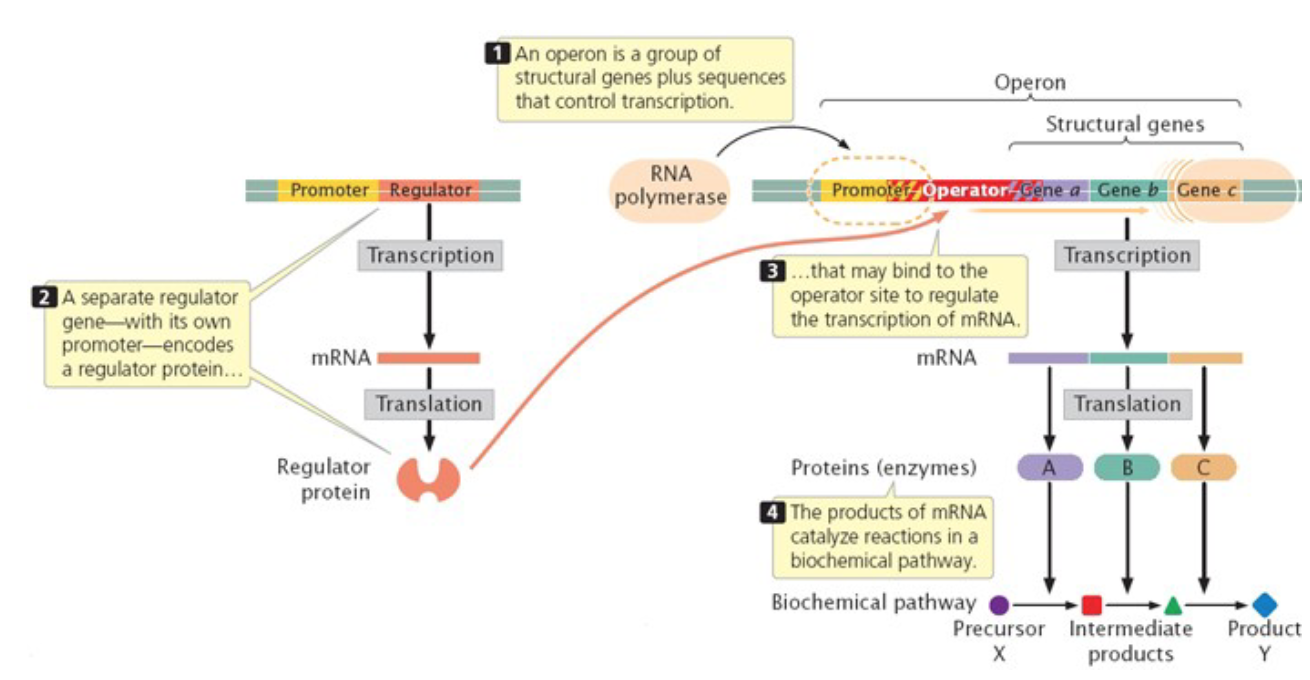
in bacterial cell, genes are organized in operons
Operon - segment of the genomic DNA which consists of structural genes and adjacent regulatory sequences
all structural genes in the operon code for the proteins necessary in the same metabolic pathway
regulatory gene (s) which are not part of the
operon, are involved as well - code for regulatory proteins
catabolic operons
expression of enzymes involved in catabolism of energy source
biosynthetic operons
expression of enzymes involved in synthesis of energy source
all structural genes are transcribed from the
same promoter sequence
structural genes under influence of
same transcription factors (=regulatory proteins) that recognize the same regulatory elements
all structural genes in an operon are
transcribed as part of the same (single) mRNA - polycistronic mRNA
regulatory genes have
their own promoters
examples of catabolic and biosynthetic operons
lactose operon & arabinose operon (catabolic)
tryptophan operon (biosynthetic)
in prokaryotes: gene regulation allows a
single cell to adjust its environment
genes encoding catabolic enzymes are “off” when
not needed and only turned “on” when the substrate is present
prokaryotic cells lacks compartments;
therefore transcription and translation are coupled! → rapid response to changes in environment by rapid change in gene expression
example: after sensing lactose in their environment, bacteria make
> 50,000 copies of needed enzymes in 90 seconds
constitutive genes
continuously expressed - required for “ongoing” survival of the cell - house keeping genes (such as those required for protein synthesis, glucose metabolism, utilization etc.)
regulated or inducible genes
expression is regulated (induced or suppressed) as needed, depending on the changes in environment
bacterial genes - organized into operons
structural genes most commonly
regulatory sequences - binding sites for regulatory proteins
regulatory genes (and the proteins they code for) are separate!!
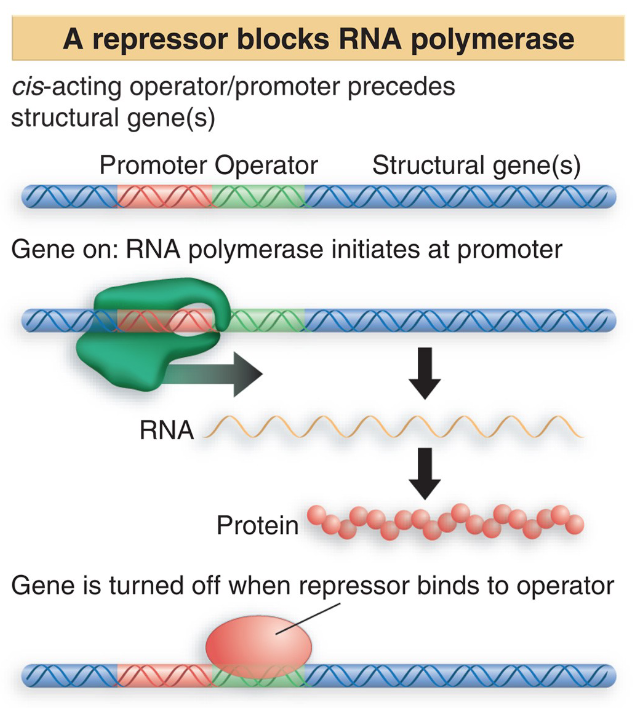
inhibit transcription - negative regulation
regulatory proteins in active state turn “off” the expression of the gene
binding site - operator
regulatory proteins could be repressors or aporepressors
sensitive to changes in cell environment
recognized by effector molecules
two types: their binding changes protein conformation
two types of regulatory molecules
inducers - inactivate repressor
induce transcription (lac operon)
co- repressors - activate aporepressors
stop transcription (Trp operon)
stimulate transcription - positive regulation
regulatory proteins in active state turn “on” the expression of a gene
regulatory proteins called activators
bind to DNA near promoter sequence
influence affinity of RNAP to stay bound to the promoter
binding creates conformational changes in DNA near the promotor = allostery
(from closed to open complex)
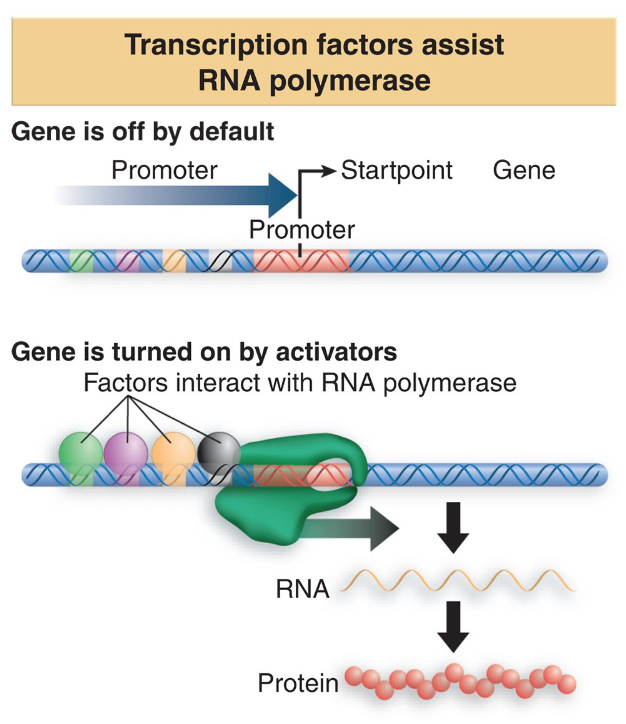
activators sometimes need
effector molecules
three different e.coli operons → three different types of regulatory proteins and methods of regulating gene expression
lactose operon
tryptophan operon
L-arabinose operon
Lactose operon
catabolic
negative regulation
regulatory protein is repressor; effector = inducer is allolactose
tryptophan operon
anabolic
negative regulation
regulatory protein is aporepressor; effector = co-repressor is tryptophan
L-arabinose operon
catabolic
positive and negative regulation
regulatory protein works as both activator and repressor (regulator) effector = inducer is arabinose
structural genes in lac operon of e.coli
code for enzymes for lactose import and catabolism (lactose-energy source when there is no glucose)
inducible genes in lac operon of e.coli
gene products are made in abundance ONLY when lactose is in the media
lac operon lac Z, lac Y, and lac A
enzyme necessary to cleave lactose and to convert lactose into allolactose effector): B-galactosidase coded by lac Z gene
enzyme necessary to transport lactose into the cell: permease coded by lac Y
Thiogalactoside transacetylase coded by lac A - unknown function
Gene I
codes for repressor protein, has its own promoter
O - operator
element, binding site for repressor
P - promoter
binding site for RNAP
scenario 1: lac operon; lactose is not present in a medium
RNAP is blocked by repressor
however, expression of lac operon genes is “leaky”
RNAP manages to occasionally bind to promoter (it is dynamic system) & proceed with low level of transcription (weak promoter)
just enough of permease is made to allow entrance of lactose into the cell if it is present in a medium
scenario 2: lac operon; lactose is getting into medium
repressor protein comes off
RNAP binds
B-galactosilase, permease and transacetylase produced
how did jacob and monod figure out regulation of lac operon
model based on 2 types of mutants
constitutive - express all three structural genes from the operon even without lactose
noninducible - do not express structural genes from the operon even when lactose is present
apart from mutation int he promoter (p) - RNAP binding site, they found two other loc involved in expression
Locus O - mutations lead to constitutive expression
Locus I - depending on which part of repressor is affected by mutation the outcome is either constitutive expression or noninducible expression (no expression)
peek at J&M results promoter mutaiton Oc mutation
RNA polymerase cannot bind: structural genes are not transcribed, even when lac is present
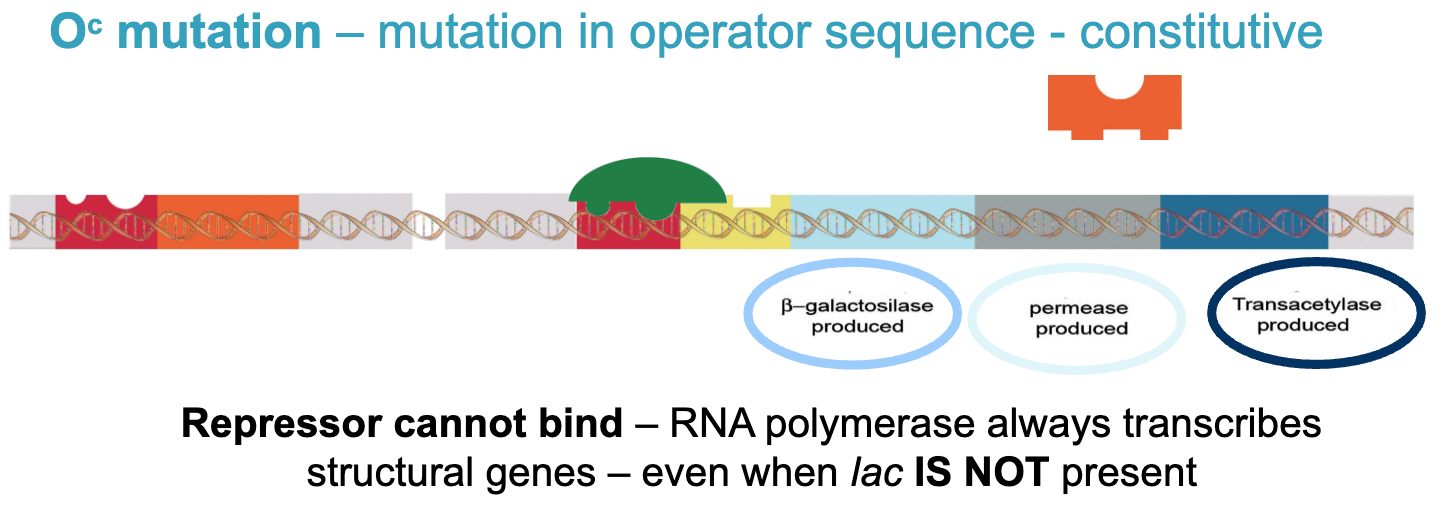
Oc mutation - mutation in operator sequence - constitutive
repressor cannot bind - RNA polymerase always transcribes structural genes - even when lac is NOT present
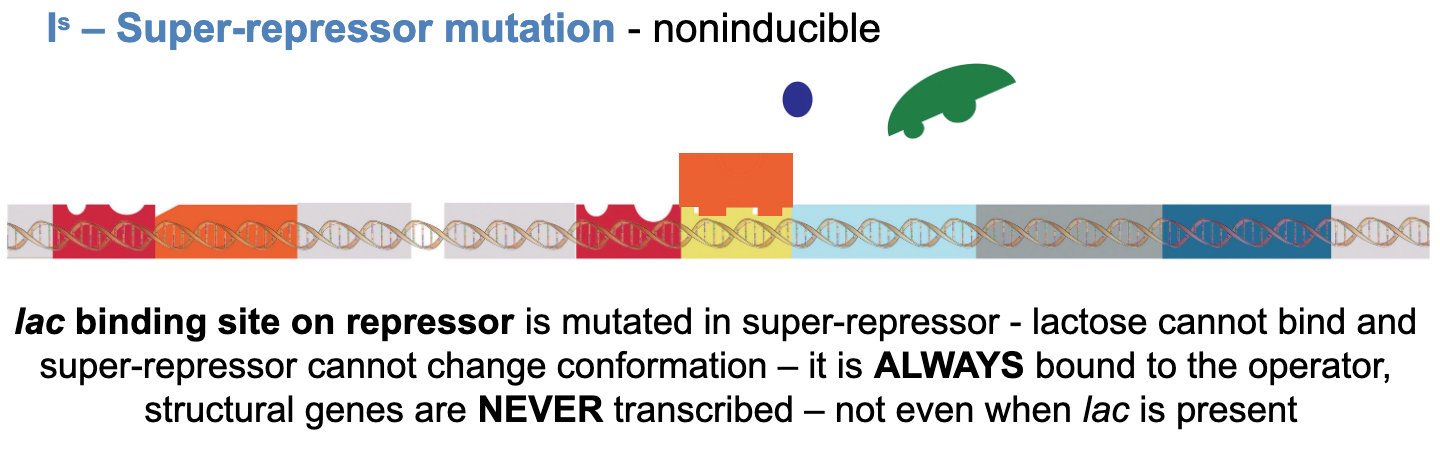
Is - super-repressor mutation - noninducible
lac binding site repressor is mutated in super-repressor - lactose cannot bind and super-repressor cannot change conformation -it is always bound to the operator, structural genes are never transcribed - not even when lac is present
summary: repressor stuck ON the DNA → genes always off
I- mutation - constitutive
operator binding site is mutated in I- repressor - mutated repressor cannot bind to operator - RNAP always transcribes structural genes - even when lac is not present
P-
if RNAP cannot bind to the promoter, structural genes will NEVER be expressed
Oc
dominant over O+ (WT) - bacteria always cleaves lactose; constitutive mutation - structural genes constantly expressed
Is
is dominant over I+ (WT) - bacteria cannot cleave latose; noninducible mutation - structural genes are never expressed
I-
is recessive; I+ (WT) is dominant over I-; constitutive mutation - structural genes constantly expressed
Result: theory that
O is the binding site for regulatory protein I
J & M: proof through complementation testsn
transformation of the bacterial cells with plasmids carrying lac operon (whole or parts)
result is a partially diploid cell = merodiploid
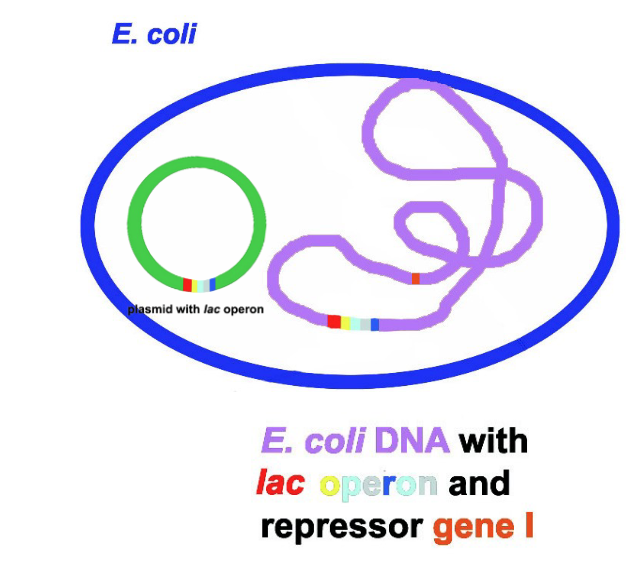
Purpose: artifically create 2 sets of alleles for lac operon genes,
complementation is enabled
plasmid can carry I gene AND operon or only I gene (and not operon)
different combinations of mutations are possible
also in combination with mutations in structural genes (since its the products of structural genes that are being measured)
J & M: discovered and defined
cis elements and trans factors
Repair fo constitutive mutation lac I- complementation in action
genomic DNA: lac I- mutation produced repressor cannot bind to operator = constitutive mutation; normal B gal always synthesized
but plasmid carries normal lac I+ also carries normal Operator (O+) mutated B gal (Z-)
plasmid lac I+ gene codes for a diffusible element that acts in trans by binding to any operator it encounters regardless of chromosomal location
synthesis of normal Beta gal (coded by genomic DNA) will become regulated