BIO130 first half
1/169
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
170 Terms
Week 1: cellular diversity: Cell Theory
The cell is the basic unit of life
All organisms 1 or more cells
Cells arise from pre-exsisting cells
Prokarotic cell
No membrane-bound organelles
Smaller than eukaryotes
Less DNA
DNA is compartmentalized but no membrane —> nucleoid
Eukaryotic cell
Nucleus
Membrane-bound organelles
Larger and more complex
Origins of mitochondria
Entangle-engulf-endogenize (E3 ) model
Ancient anaerobic archeal cell and ancient aerobic bacterium
started as ectosymbiote (outside)
then engulfed as endosymbiote
then bacteria membrane broken
(other model show predatory mechanism)
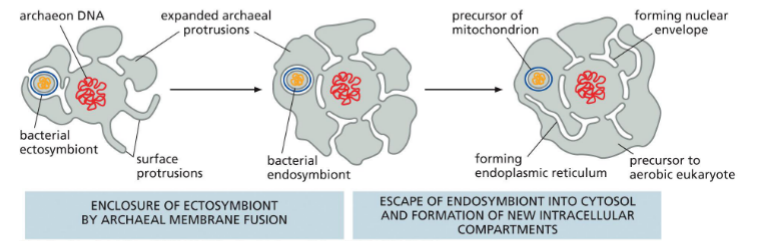
Ancient cell folding protusions
Give way for nuclear envelope and endoplasmic reticulum
Origins of eukaryotes graph
mitochondria first —→ eukaryotes
chloroplast second —→ plants
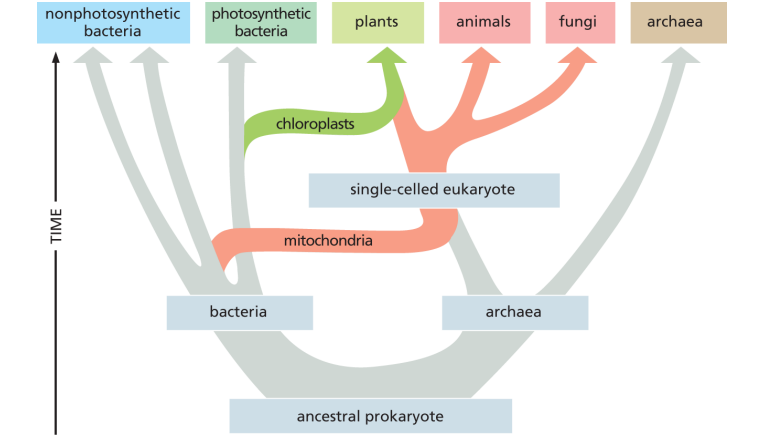
Endosymbiont hypothesis for mitochondria and chloroplasts — evidence
Both have remnants of own genome which resemble modern prokaryotes
Both have kept some of own protein and DNA synthesis components, also resemble prokaryotes
Membranes are similar to prokaryotes and derived from bacteria ancestor
Model organisms and Humans — general attributes of model organisms
Fast development and short life cycles
Small reproductive (adult) size
Readily available
Tractability (manipulation or modification)
Understandable genetics
Examples of model organisms
Ecoli
Brewer’s yeast — simple eukaryote
Arabidopsis — plant
Nematode, drosophila, zebrafish, mice — animals
Lec 2: The Central Dogma of Molecular Biology
Information flow is always in one direction
DNA (transcription) —→ RNA (translation) —→ Protein
.
Refined:
messengerRNA: translation for protein
transferRNA: transport amino acids
rRNA: heart of ribosome, breaking and forming of bonds
Antiparallel and genetic code
DNA, RNA, and proteins are synthesized as linear chains of info with intrinsic direction
RNA translating to amino acid is universal through the genetic code
What are nucleic acids?
Genetic material in a cell
DNA & RNA
Three parts of a nucleotide
Pentose sugar
Phosphate group (1, 2, or 3)
Nitrogenous base
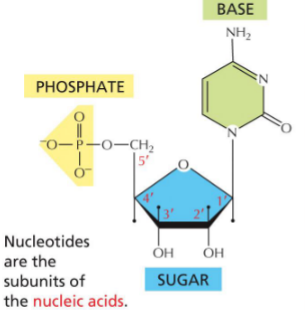
Nucleotide bases
Pyrimidine
Cytosine
Thymine
Uracil
Purines
Adenine
Guanine
Differences between DNA and RNA
RNA:
ribose
OH on 2’ carbon
uracil
DNA:
deoxyribose
H on 2’ carbon
thymine (extra methyl)
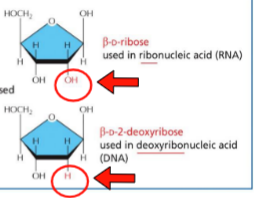
Nucleic acid nomenclature
Nucleoside: sugar + base
Nucleotide: sugar + base + 1, 2, 3 phosphate
Nucleoside monophosphate
Nucleoside diphosphate
Nucleoside triphosphate
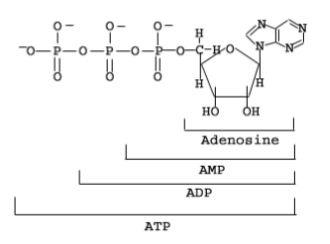
Nucleic acid chains (phosphates provide energy..)
DNA is synthesized from deoxyribonucleoside triphosphates (dNTPs)
RNA is synthesized from ribonucleoside triphosphates (NTPs)
Nucleotides linked by phosphodiester bonds
Phosphates provide energy for bonds
Molecular interactions
Electrostatic attrations (charges attract)
Hydrogen bonds
Van der waals attractions
Hydrophobic force
(Individually weak, sum to be strong)
Three forces that keep DNA strands together
H-bonds (base pairing, G-C stronger cause 3 bonds)
Hydrophobic interactions (phosphate backbone hydrophylic, bases hydrophobic
Van der waals attractions
DNA structure - energetically favourable
DNA will naturally come together, energetically favourable
Proteins can recognize and make contact with specfic sequences in major & minor grooves
Separating DNA strands
DNA can unravel and return to double helix
use energy or enzymes to denature
useful for replication, transcription, or PCR
Lec 3: intro to protein structure (primary, secondary…)
Quaternary: more than 1 polypeptide chain, subunits
Multiprotein complexes: many chains and subunits, machine
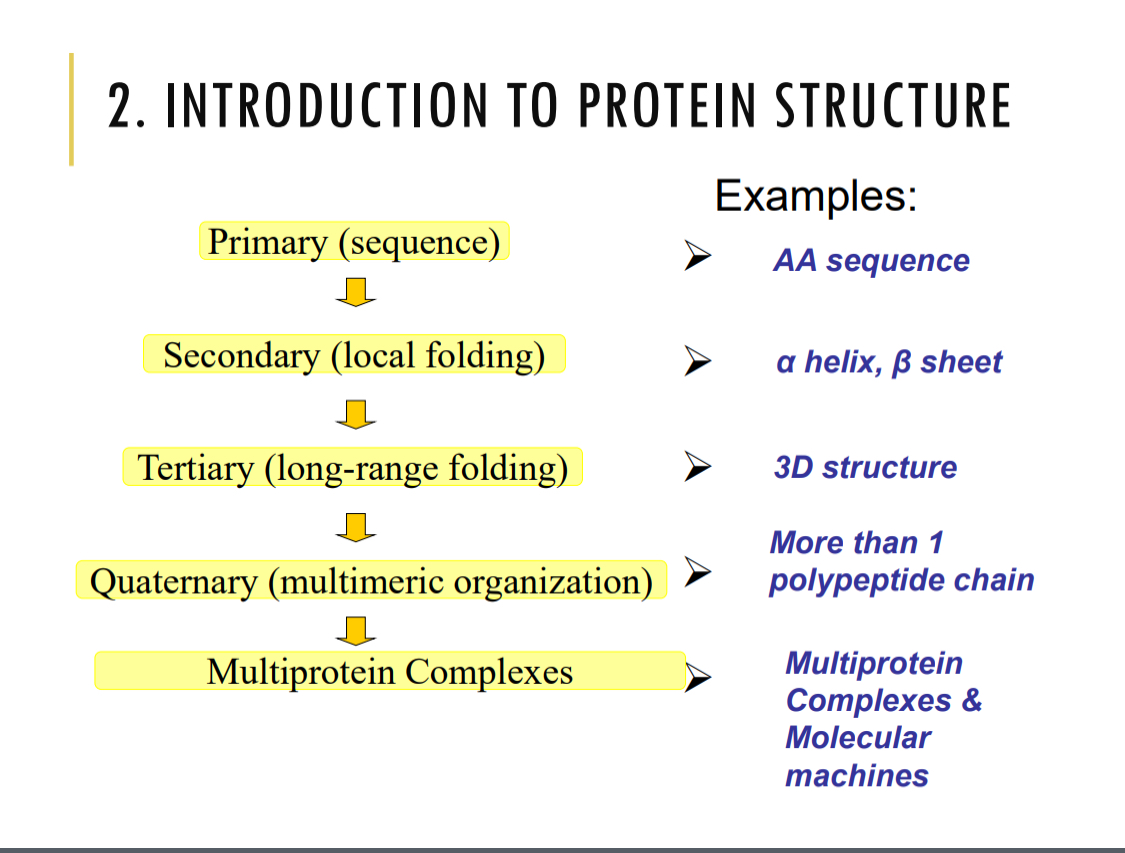

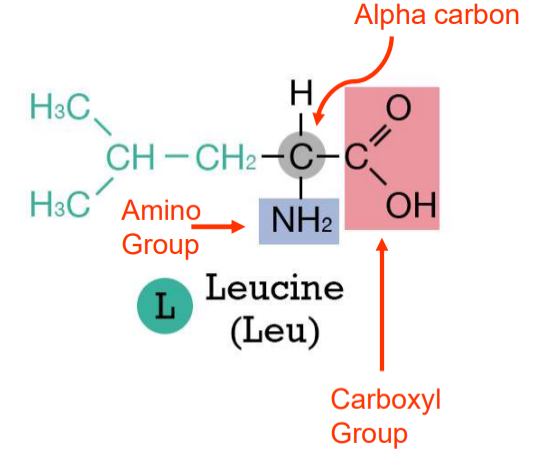
Amino acid structure
proteins composed of amino acids
side-chain/R group is variable and determines the type of amino acid
three major categories
acidic ( - charge)
basic ( + charge)
uncharged polar
nonpolar
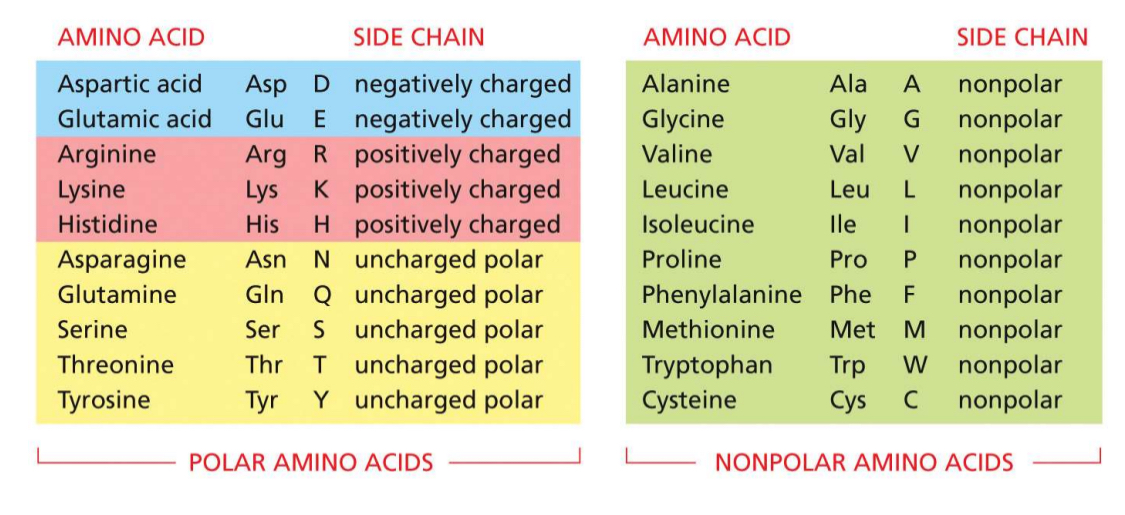
classification system for amino acids
half are polar
5 charged polar
5 uncharged polar
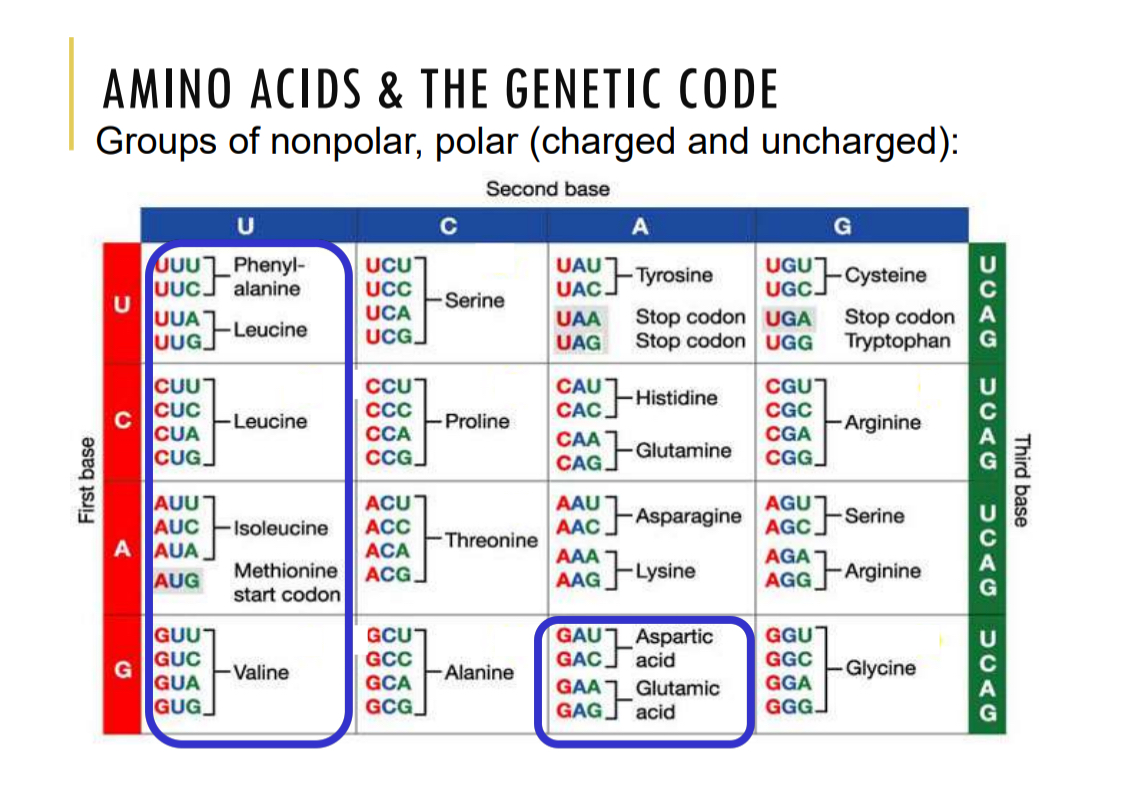
Amino acids and the genetic code
AUG — start codon methione
UAA, UAG, UGA — stop codon
Degenerative code: more codons than AA, more than one can code for the same AA
Flexibility: similar codons for similar AA, can tolerate mutations better
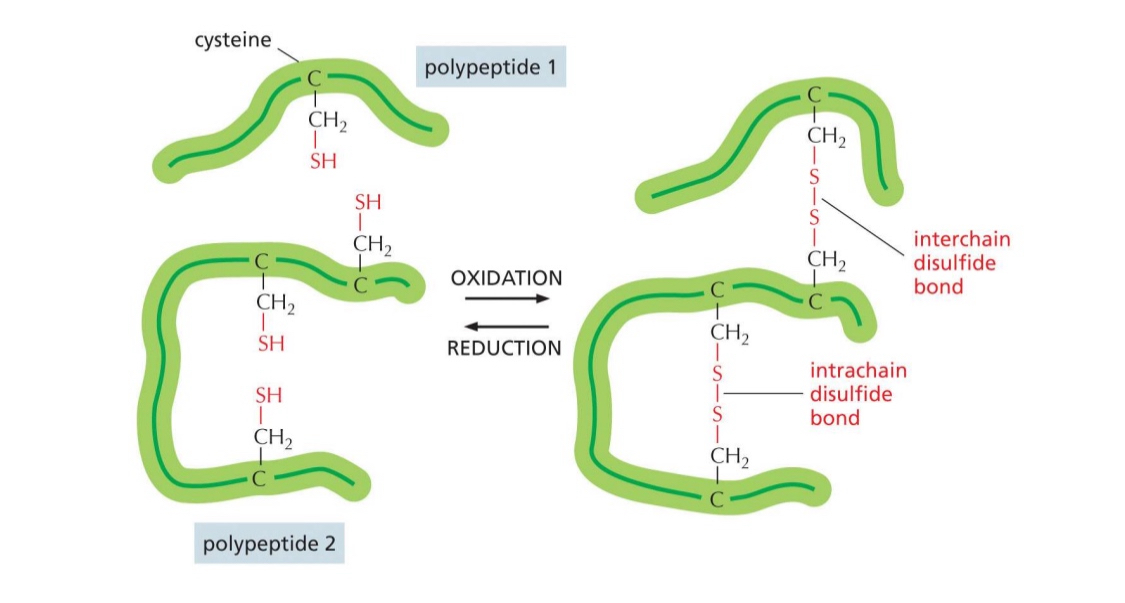
Unique amino acid: cysteine
can form disulfide bonds — (oxidation form, reduction break)
both interchain and intrachain
covalent bond creates stability
“staple”
often used in structural proteins
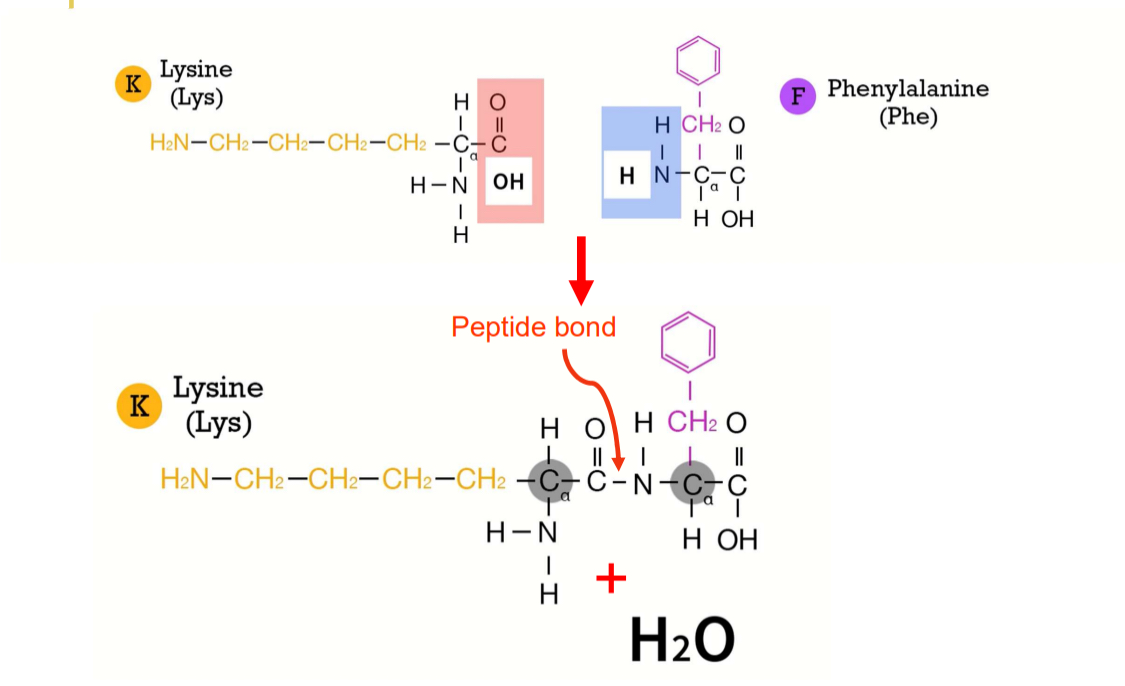
Primary structure: Peptide bonds
catalyzed by ribosome
peptide backbone of C-C-N-C-C-N
Polarity: always grow in the same direction, starting at N-terminus
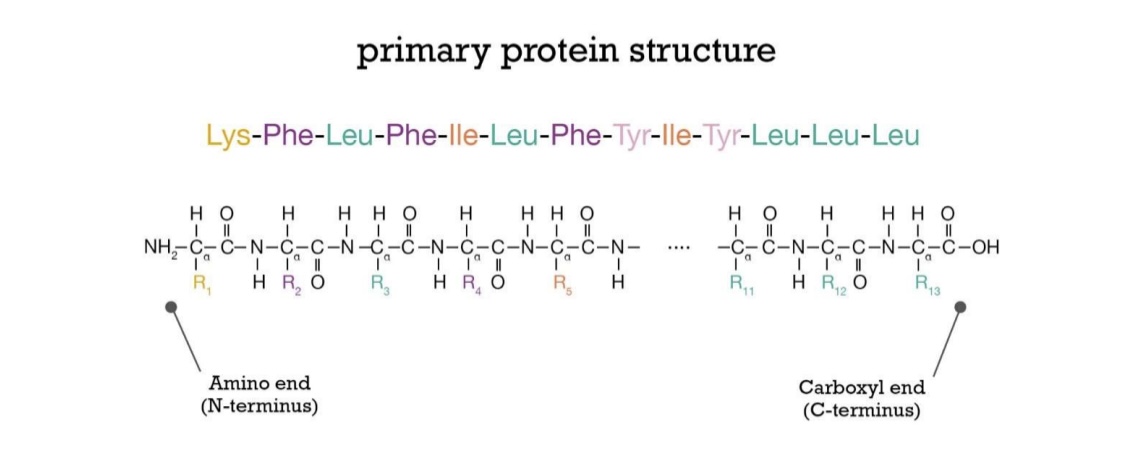
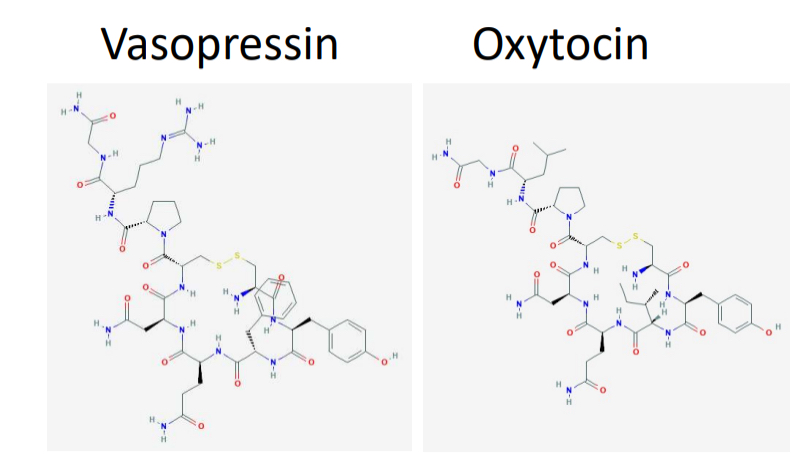
Differences in primary AA sequence matter - vasopressin example
both vasopressin and oxytocin are 9 AA long
both are identical except at two locations
vasopressin controls urine production
oxytocin involved in birth, lactation, and pair bonding
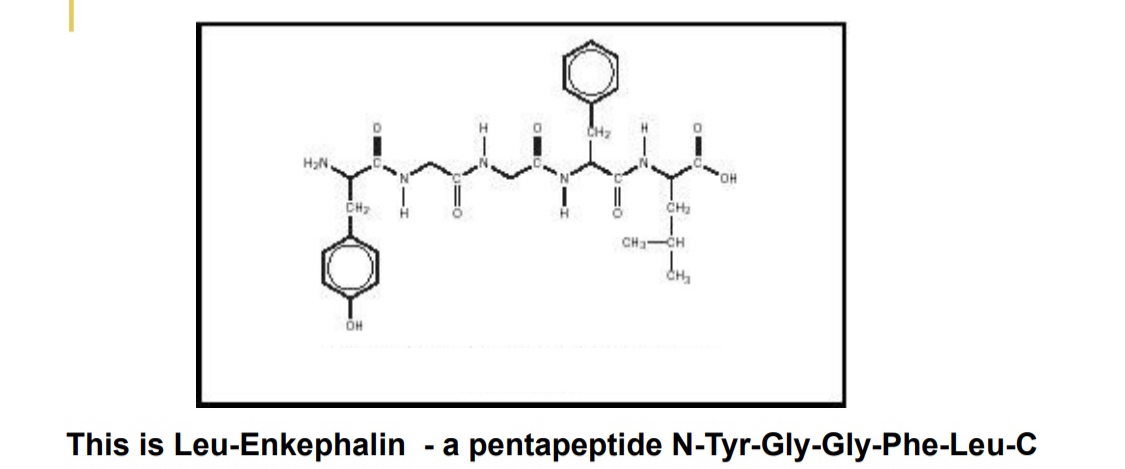
Order of AA is important too - Leu-enkephalin
natural opioid
the opposite order of AA has no pharmalogical effects
the amine-carboxyl orientation essential to function
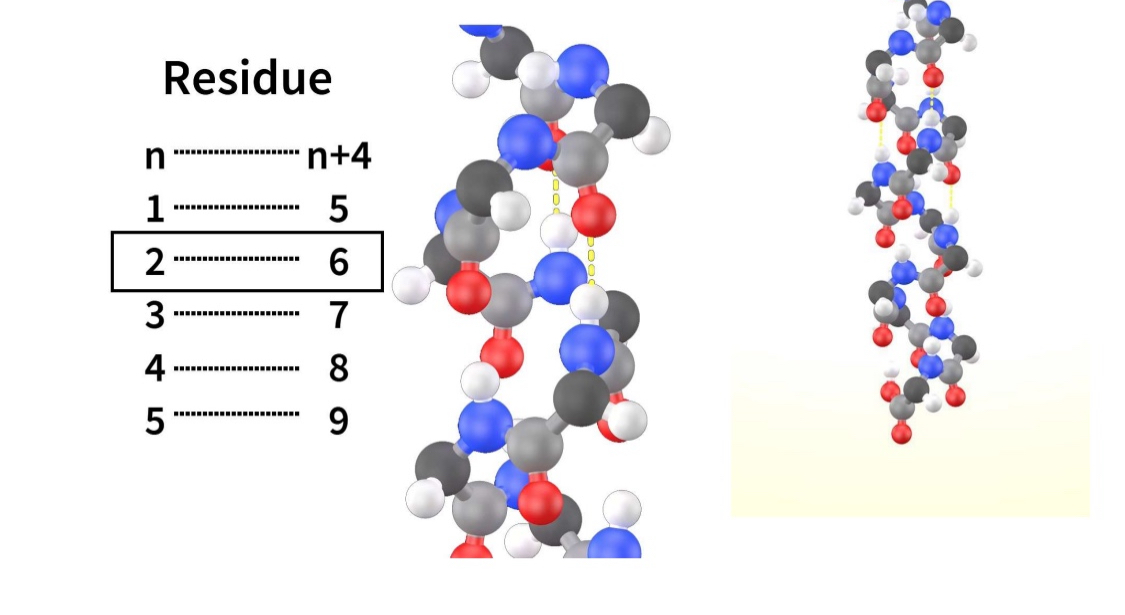
Secondary structure: Alpha-helix
Forms independantly of side chains
carboxyl — h-bonds — amino of AA 4 after
n — n+4
(helical structures are common in biology because they are stable
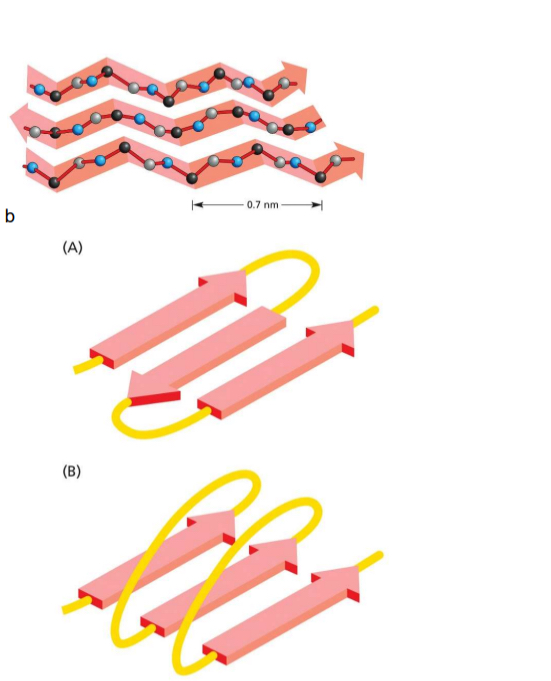
Secondary structure: Beta sheet
Forms independtly to R groups (but they alternatively point up and down, interactions)
H-bond of carbonyl (C=O) with amide hydrogen (N-H) of neighbouring strand
typically contain 4-5 strands but can have more
can be antiparallel or parallel
anti only needs small sequence between
parallel needs more
Counting polypeptide chains
Always count from N-terminus (amino end)
Coiled coil
Alpha helices twisting together
Only form with amphipathic — protein
repeating hydrophobic molecule every 4 peptide bonds — hydrophobic stripe
2 helices will wrap together, push hydrophobic parts into middle
very stable and strong
keratin in hair
myosin motor proteins

Amyloid structure
Beta sheets stacked together
Misfolded proteins can form amyloid structure — neurodegenerative diseases
prions: converts properly folded molecules
Lec 4: Tertiary structure (3D)
Overall 3D structure of a protein
Held together by:
hydrophobic forces
non-covalent bonds
covalent disulfide bonds
Hydrophobic force
non-polar AA in interior of folds
polar AA on exterior
Tertiary structure — continued (energetically favourable + chaperone proteins)
Proteins fold into conformation that is most energetically favourable — spontaneous
H-bonding in:
backbone/backbone
backbone/side chain
side chain/side chain
Chaperone proteins help make process more efficient and reliable in living cells
misfolded proteins cant function
Tertiary structure can have large variety of shapes
globular, filament, etc
But few of the possible chains will be useful
majority 50—2000 AA long
well-behaved, stable
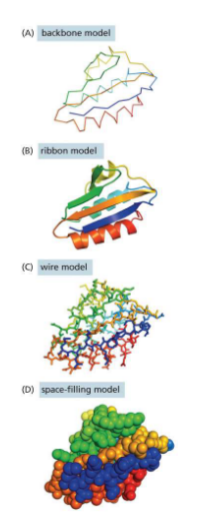
Models for proteins
Backbone model — only backbones
Ribbon model — shows folding
Wire model — shows positions of bonds
Space-filling model — contour map
Protein domains (still single polypeptide) (eukaryotic proteins often have 2 or more…)
Regions of proteins that have specialized functions
single polypeptide
each domain has own tertiary structure and function semi-independently
connected by intrinsically disordered sequences (flexible regions)
.
eukaryotic proteins often have 2 or more
domains are important for the evolution of proteins
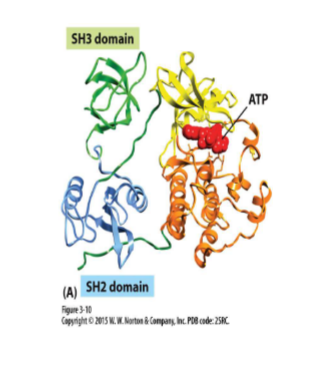
Protein domains — extra example: Src protein kinase
Kinase — phosphorylate proteins (changes activity)
Src protein kinase has 3 domains
SH2 and SH3 regulates kinase
Protein families (similar AA sequences & tertiary structures)
Way to organize proteins — a protein can belong in more than one
Similar AA sequences and tertiary structures
Members have evolved different functions
Most proteins belong to families with similar structural domains
Quaternary structure (subunit)
More than1 polypeptide chain
not all proteins
subunit = separate polypeptide
can get really big
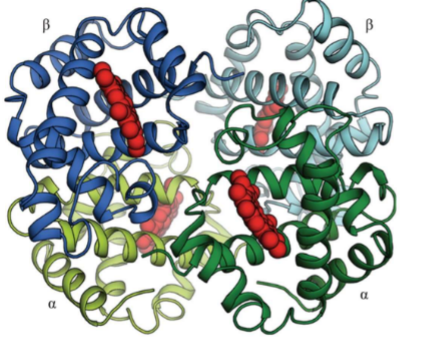
Quaternary structure example: Hemoglobin
Each hemoglobin has 4 subunits (2 alpha, 2 beta)
Sickle cell anemia is caused by mutation in beta subunit
Multiprotein complexes and molecular machines
Can be:
many identical subunits (actin filaments)
mixtures of proteins and DNA/RNA (ribosomes)
dynamic assemblies of proteins to form machines (DNA replication)
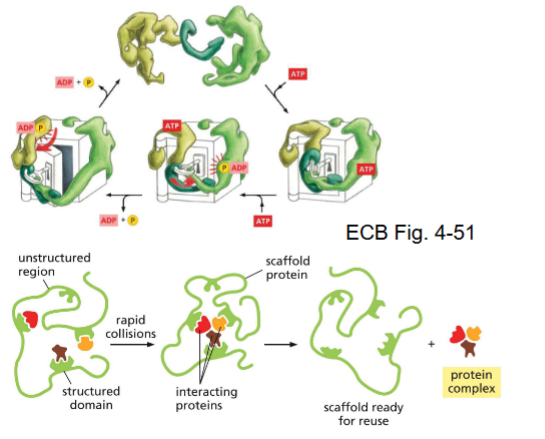
Multiprotein complexes and molecular machines (scaffold proteins)
conformational changes
perform job
often need ATP
.
scaffold proteins
binds proteins together
How are proteins studied?
Past
purify proteins
electrophoresis & affinity chromatography
.
Now
Mass spectrometry
sequenced many genomes
find mass and match to predictions
discover precise 3D structure with other techniques
can also use AI to predict structure using only polypeptide
Protein separation
separate using size, shape, charge, hydrophobicity
Proteomics
Large scale study of proteins
structure
interactions
abundance and turnover
location
Lec 5: sequencing genomes
We are able to sequence genomes
Genomes can come in all sizes
Bacteriophage — 16.9 kb (kilobases)
E.coli — 4.6 mb (megabases: million)
Generating genetic variation
Tinkering not inventing
Mutation of gene
Mutation of regulatory DNA
Gene duplication and divergence
Exon shuffling
Transposition (TE)
Horizontal transfer of DNA (common in bacteria)
Purifying selection and genes
Purifying selection has maintained few hundred fundamentally important genes
one gene in all species: gene for rRNA
Circular DNA
Prokayote DNA is circular
Organellar genomes are also circular
Because of shared ancestry
small size due to relying on host for protein production
mitochondria — 16.5kb
chloroplast — 121kb
Human genome (base pairs + diploid + karyotype)
3 billion base pairs per genome
One genome from each parent = diploid (46 chromosomes)
20,000 protein-coding genes across 23 pairs of chromosomes
Karyotype: organized profile of full set of chromosomes
Comparing genome sizes
Size of genome is loosely related to complexity of the organism — but MANY discrepancies
Bac and archea — smaller
Mammals — bigger
But:
newt, wheat, and amoebas have bigger genomes (amoeba 10X)
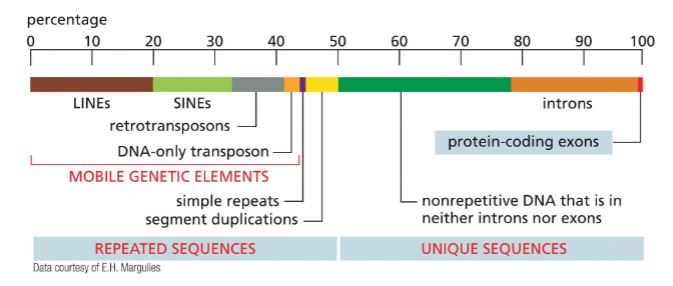
Human genome: repeated sequences and unique sequences
Repeated sequences: ~50% of genome is repetitive DNA
Mobile genetic elements: transposons (TEs)
can move places
copies itself and gets stuck
can also create novel genes
Unique sequences: Less an 1% encodes protein — exons
introns
other non repetitive DNA used for regulation — intergenic DNA
Packaging of DNA — prokaryotes
DNA is condensed through folding and twisting about 1000 folds using proteins
Forms the prokaryotic Nucleoid
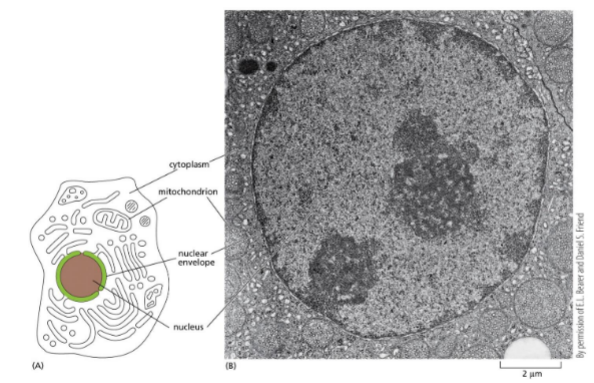
Packaging of DNA — eukaryotes: nucleus picture
Dark circle: nucleoli/nucleolus — parts of chromosomes that encode rRNA
Dark stains around the edge: heterochromatin (transcriptionally silent, most compact)
Light stains: euchromatin (gene rich, transcriptionally active)
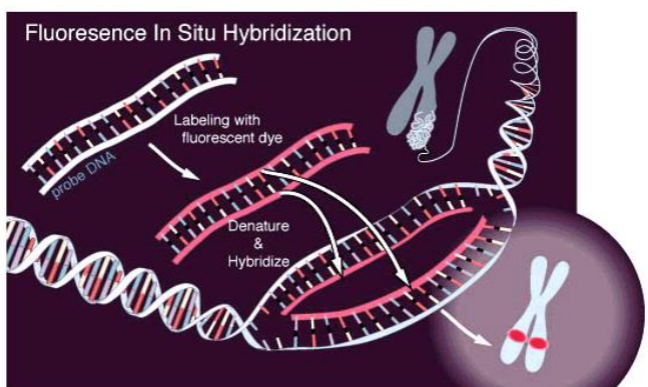
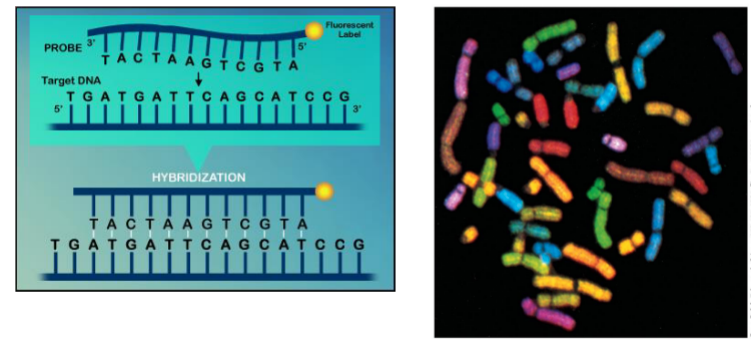
Fluorescence In-situ Hybridization (FISH)
Uses single strand probes labeled with fluorescence or radioactive isotopes
first denature
then hybridize with probe that pairs
can paint chromosome
highlights shape, size and number
Packaging of DNA: Solution — Chromosomes!!
23 pairs in humans
each chromosome contains a single, long, linear DNA molecule and associated proteins — CHROMATIN
chromatin is dynamic — accessible or transcription, replication, and repair
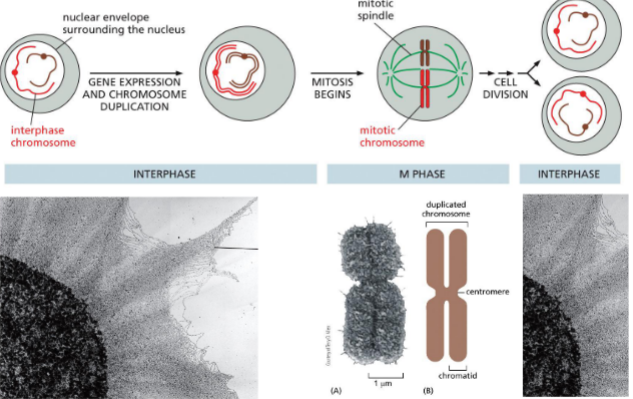
Centromere
Special sequence that allows for attachment of spindles to split sister chromatids during mitosis
tightly condensed
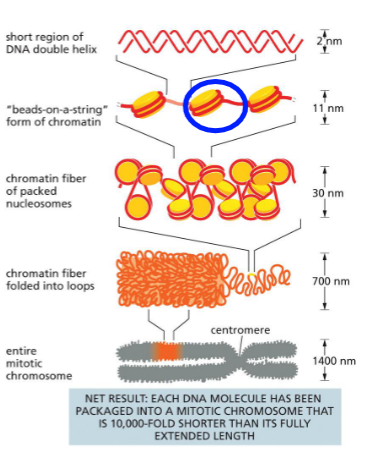
Chromatin organization
DNA double helix
Wrapped 1.7 times around histone protein — nucleosome (beads on a string)
Bent with linker histone H1 — chromatin fiber
Folded into loops with SMC proteins (condensins) — mitotic chromosomes
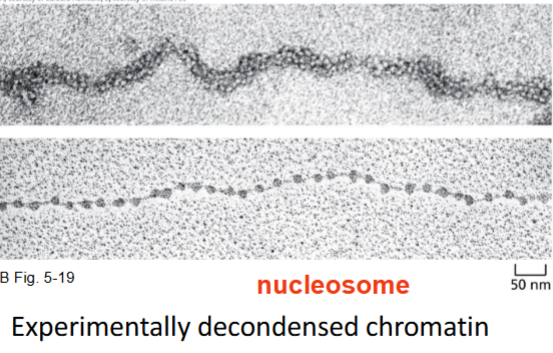
Nucleosome
Nucleosome is the basic structural unit (can be experimentally decondensed from fiber)
each has 147 pairs
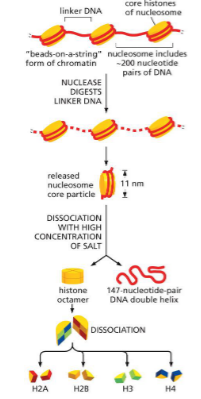
Histones (postive/negative charge)
Small proteins rich in lysine and arginine — positively charged
Matches with negative charge of DNA backbone
Made up of 4 parts
8 polypeptides into dimers
dimers into 4 cores — octamer
One linker histone (H1)
kinks loose DNA into tigher fold for chromatin fiber
Lec 6: Packaging of nucleosomes: cohesins + sequence specific clamp proteins
Chromatin fibers are looped into chromatin loops using sequence-specific clamp proteins and cohesins
sequence-specific clamp proteins recognize specific sites
use ATP to pull proteins together
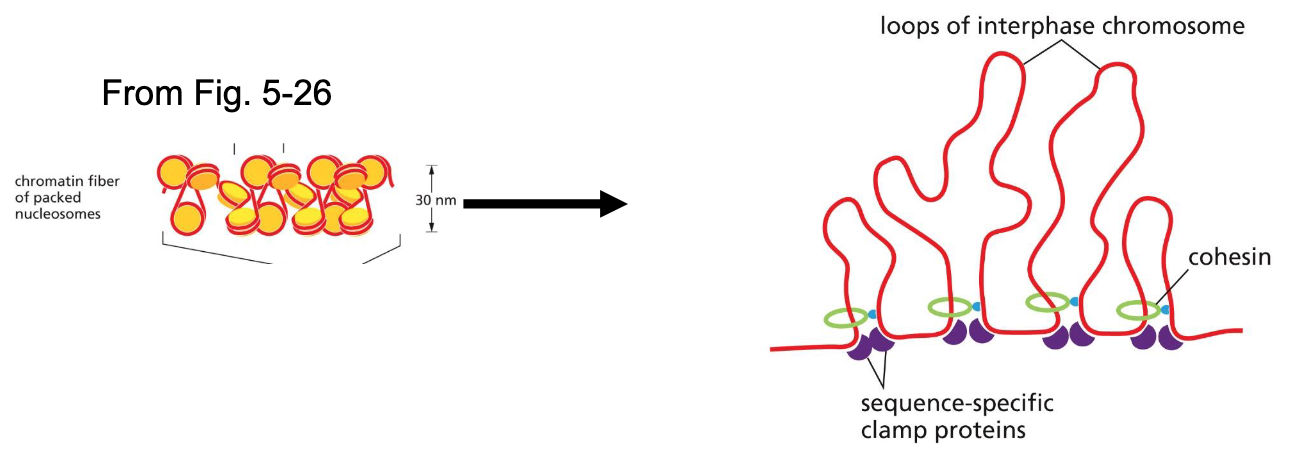
Packaging of nucleosomes: mitotic chromosomes — condensins
Condensins II replace most cohesins to form loops
Condensins I forms loops with loops
More compact
Stacked
Most compact during metaphase
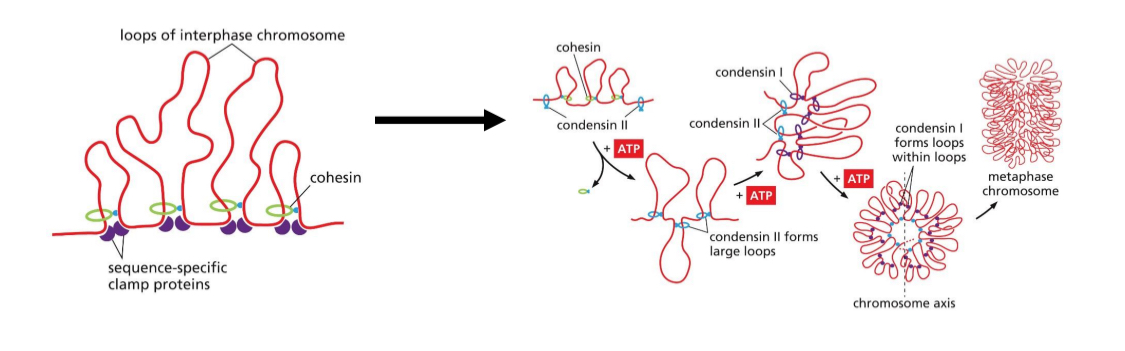
Mitotic chromosomes packing results
10,000 times shorter than extented DNA length
Chromatin modification
Chromatin remodeling complexes + histone modifying enzymes — changes chromatin structure and access to DNA for replication or transcription
Used ATP
Slides DNA on histones
Evict histones
N-terminal tails of histones modified (adding groups, providing docking sites for proteins)
Chromatin remodelling complexes + histone modifying enzymes act as a team (HME recruits CRC)
Heterochromatin (constitutive vs facultative)
Highly condensed chromatin
Meitotic and mitotic chromosomes
Time spent highly condensed varies
Regions of interphase chromosomes that gene expression is suppressed
.
Constitutive heterochromatin — most compact, compact all the time (Telomeres)
Facultative heterochromatin — can be opened, to turn on/off (degree of compaction controls gene expression)
Centromeres are compacted all the time
Euchromatin
Relatively non-condensed chromatin
Degree of condensation varies
Level of activity varies
Areas where genes tend to be expressed
.
Quiescent euchromatin — less active
Active euchromatin — least compact, gene expression
Degree of chromatin condensation is reversible (modifications of histones + chromatin remodeling complexes + RNA polymerase)
Localized covalent modifications of histones + chromatin remodeling complexes + RNA polymerase = modulate the reversible switch along chromosomes
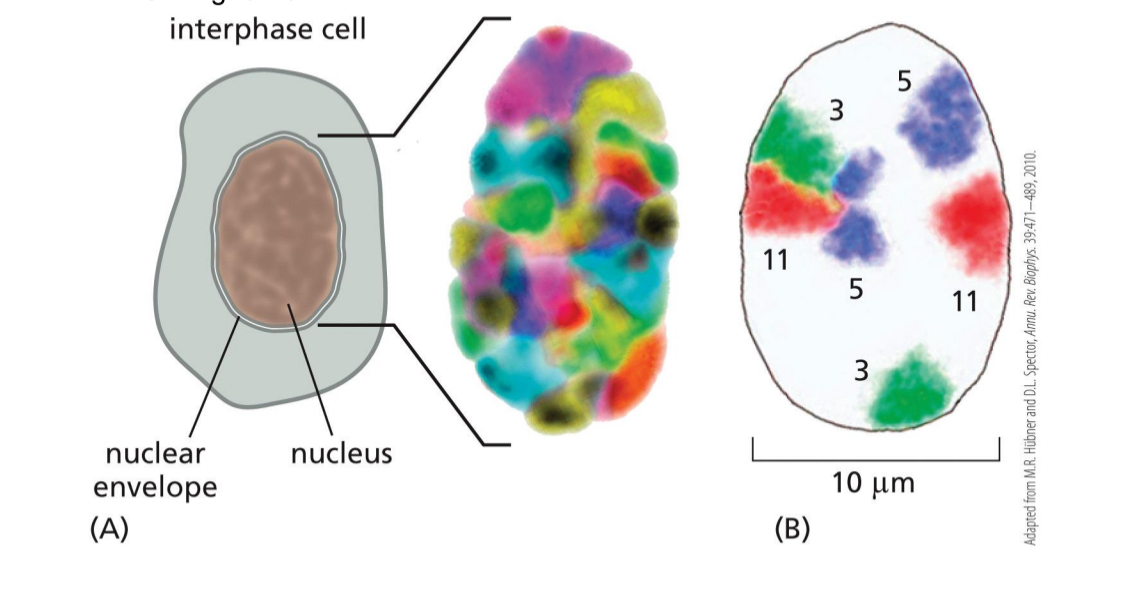
Interphase chromosomes in discrete regions of the nucleus
(shown using FISH)
Chromosomes have territories
Some interact
Some are more on the sides or the middle
Nucleolus — carry genes that encode for rRNA
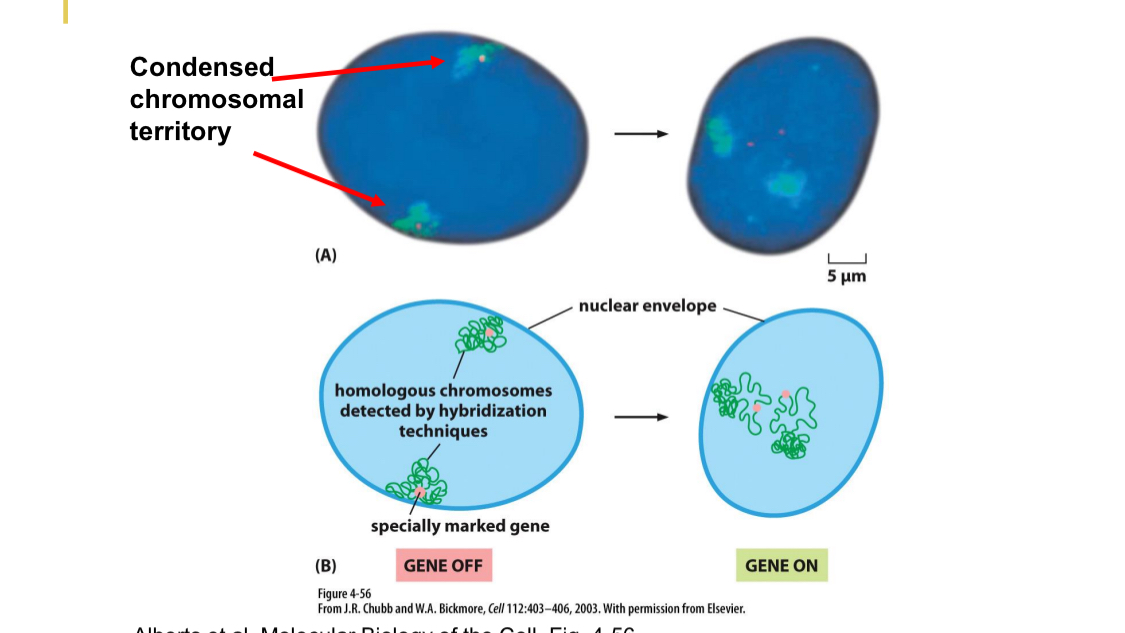
Expressed gene is re-orientated within the chromatin
OFF
Within periphery chromosomes
ON
Genes are now internalized, more in the middle
Chromosomes are more unfolded
Cell nuclei is dynamic
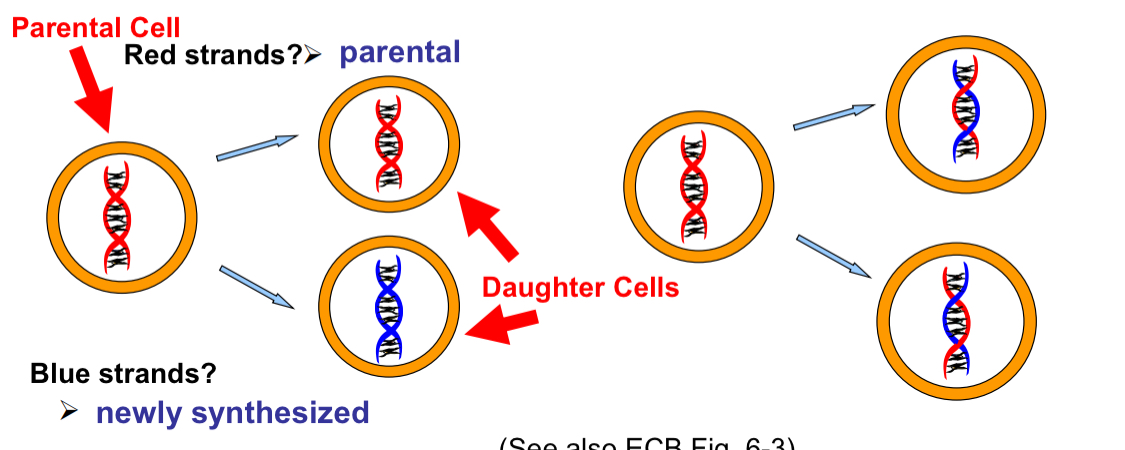
Conservative vs Semi-conservative DNA replication vs Dispersive
DNA replication is actually semi-conservative
Direction of DNA replication - overview
Bidirectional growth from one starting point
DNA is synthesized 5’ — 3’
DNA is split using initiator proteins
2 replication forks
DNA polymerase uses template to add deoxynucleoside triphosphates (dNTPs) to the 3’ OH end
phosphodiester bond
base pairing
energy comes from dNTP — pyrophosphate out
Leading strand
Lagging strand — okazaki fragments
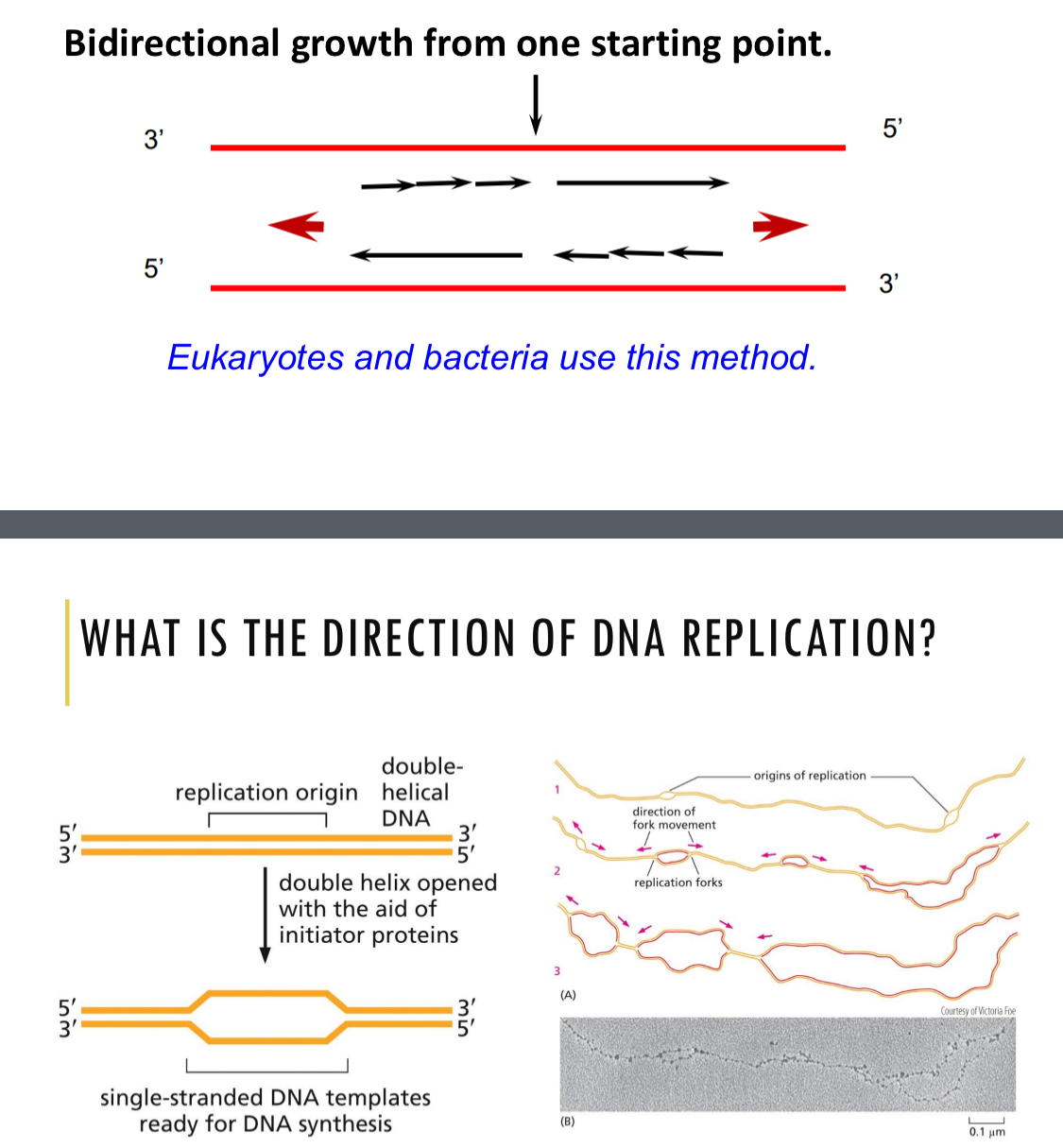
Always start from the same location on DNA
A -T rich sequences, easy to open
Recognized by initiator proteins that bind to the DNA
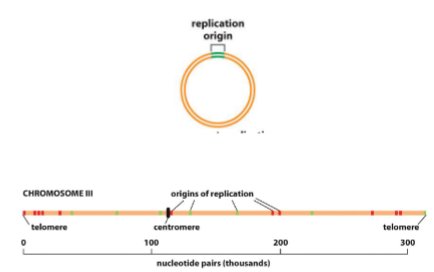
How many origins of replication & speed
Bacteria - 1 (1000 pairs/sec)
Eukaryotes - multiple (100 pairis/sec in humans)
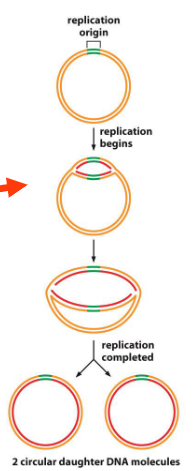
DNA replication in bacteria
Two forks that increase until two new circular genomes are formed
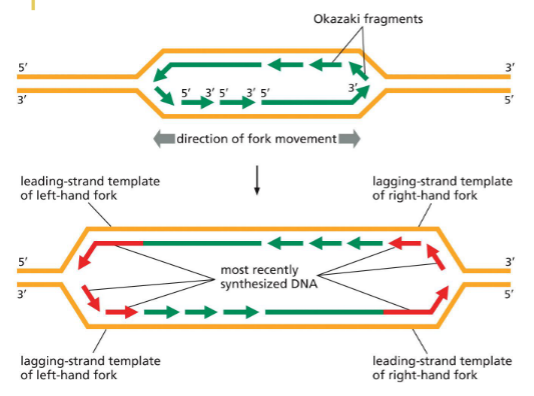
What happens at the DNA replication forks?
Replication fork is asymmetrical — antiparallel with parental strand
Leading strand synthesized continuously
Lagging strand discontinuously — okazki fragments
Fork keeps unzipping as it goes
Procedure and Ingredients for DNA synthesis
Procedure:
separate DNA strands
synthesize DNA
proofread newly synthesized DNA
.
origin of replication — specific
primers — RNA primer
dNTPs
ATP (energy)
DNA polymerase
Accessory proteins
Many mechanisms are similar with eukaryotes and prokaryotes
Many mechanisms are similar with eukaryotes and prokaryotes
origin of replication
initiator proteins
helicase
single-strand (DNA) binding proteins
RNA primers made by primase
DNA polymerase
sliding clamp holds polymerase
nick sealing by DNA ligase
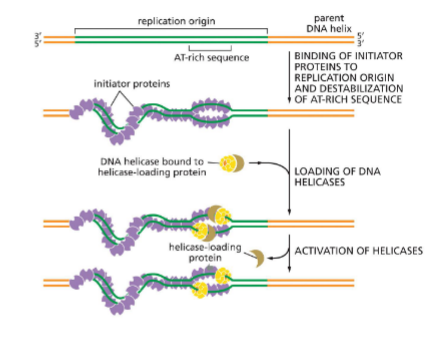
Initiator proteins for replication (E.coli)
Process highly regulated — want start to finish completion
Bind to origin
A-T rich sequences (very specific)
attracts initiator proteins
destablize helix
has ATP but dont use it until synthesize begins (for regulation, no one else can get on strand)
.
Helps helicase bind
attract helicases and bind
and helicase-loading protein
.
Requires ATP
start hydrolyzing when replication is about to begin
no other can start replication until initiator proteins get more ATP
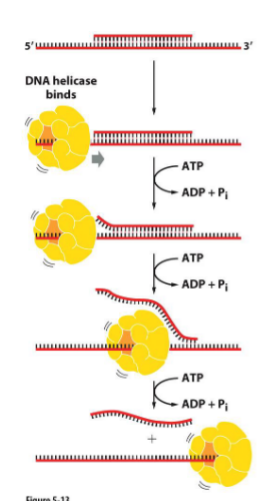
Unwinding by helicase
Two types of helicases exist — predominant moving along the lagging strand 5’ —> 3’
Requires ATP — breaking hydrogen bonds
(helicase has quaternary structure)
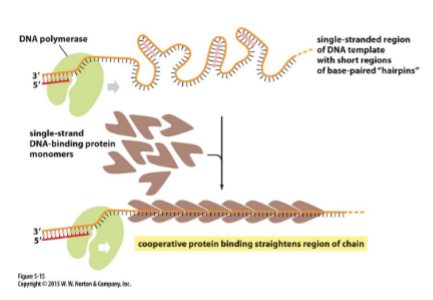
Binding of single-strand binding proteins
Keeps DNA from reannealing or getting tangled — a single strand can H-bond (form hairpins) (important for PCR too)
separates strand by binding ssDNA, coats/cups strand, particularly the lagging strand
proteins start attracting more proteins (cooperative, multiprotein assembly)
prevents strands from H-bond with eachother or by itself
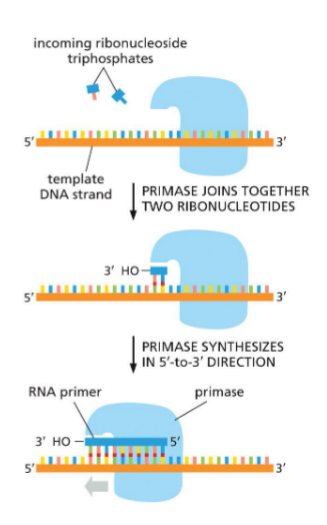
RNA primers made by primase
DNA polymerase needs RNA primer to start synthesizing
primer only need nucleotide + template
polymerase needs 3’ OH + nucleotide + template
.
To begin, DNA polymerase needs bound primer
Primase synthesize an RNA primer (joins ribonucleotides)
Primase synthesizes in 5’ —→ 3’ (antiparallel to template, moving 3’ — 5’ on template strand)
primer is only temporary, cell doesnt like
Primase is on helicase
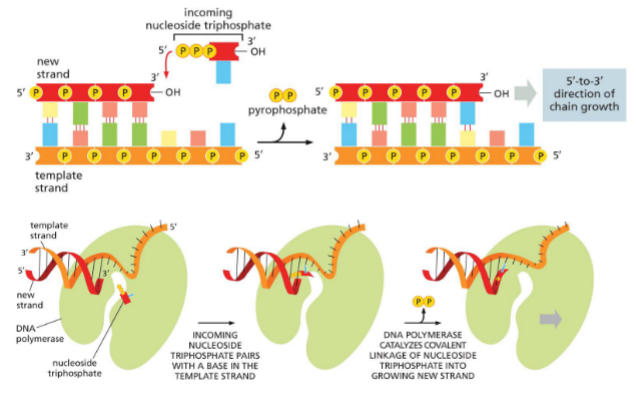
DNA polymerase
Adds nucleotides to 3’ OH end
correct base pairing
clips off 2 phosphates — pyrophosphate
catalyzes phosphodiester bond
move onto next
DNA is synthesized complementary and antiparallel
Grows 5’ —→ 3’
Sliding clamp holds polymerase onto DNA (loaded by clamp loader)
Why? If DNA polymerase holds too tightly, hard to release quickly
Evolved DNA polymerase to not hold on tightly, sliding clamp will let it fall off
Circular protein — doesnt impede polymerase progress
loaded by clamp loader (also holds polymerases together, moves then together)
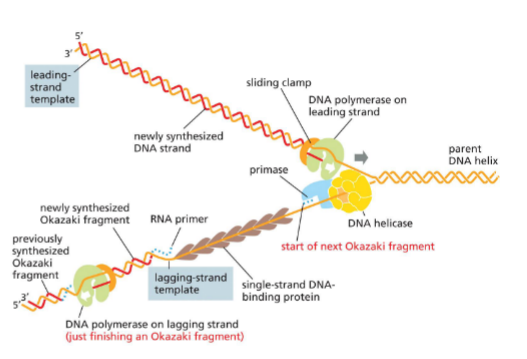
Replication fork picture
lagging strand keeps making primers for okazaki fragments
sliding clamp releases polymerase when reach other okazaki fragment 5’ end
overall moving towards fork
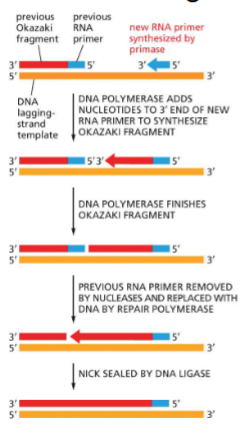
Nick sealing by DNA ligase (ATP & AMP)
A special DNA repair system is responsible for removal of RNA primer and replacing with DNA sequence
Nucleases remove primer
Repair polymerase replaces DNA onto 3’ end of okazaki
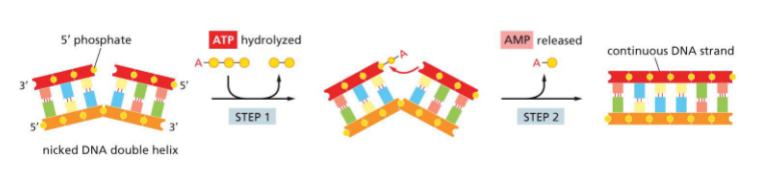
Nicks sealed by Ligase — seal phosphodiester bond (doesnt need more nucleotides)
uses ATP — hydrolyzes
attaches amp to nick (adenosin monophosphate)
release amp
phosphodiester bond
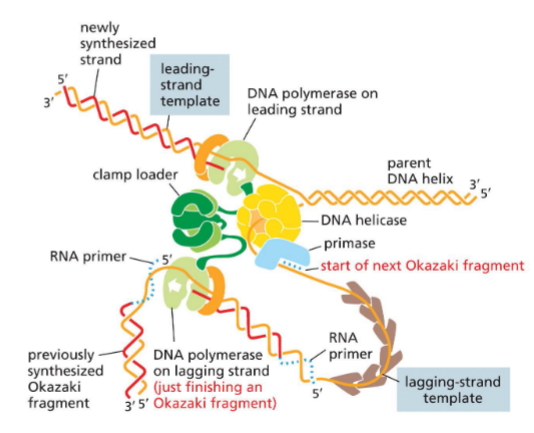
Summary picture (bac replisome + primosome)
whole thing: bacterical replisome — molecular machine
helicase + primase = primosome
Primosome
Helicase + primase
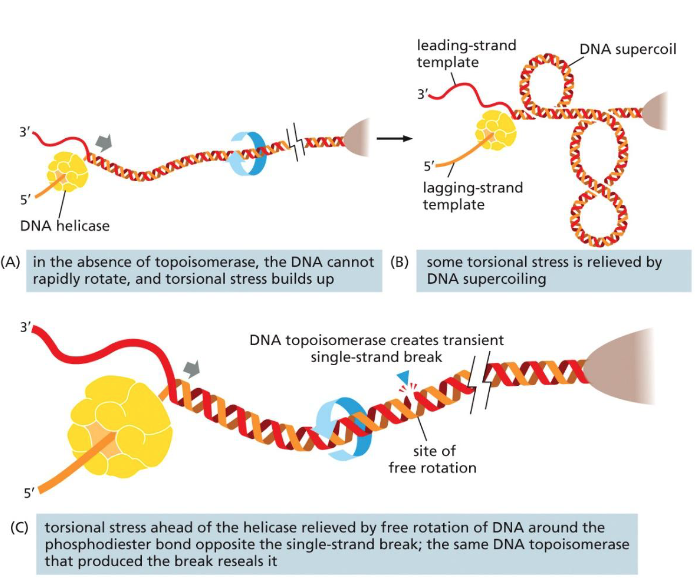
Lec 8: issues in DNA replication: Unwinding problem
As helicase unwinds DNA, supercoiling & torsional strain increases
Problem in circular chromosomes and large eukaryotic chromosomes
Solved by DNA topoisomerase
binds to a location
cut a nick in the backbone of 1 strand (does not cut H-bonds)
allows for rotational freedom
Issues in DNA replication: Ends of linear chromosomes
On the lagging strand, after the removal of the last primer, nothing to replace lost segment — loss of sequence information on the 5’ end of daughter DNA
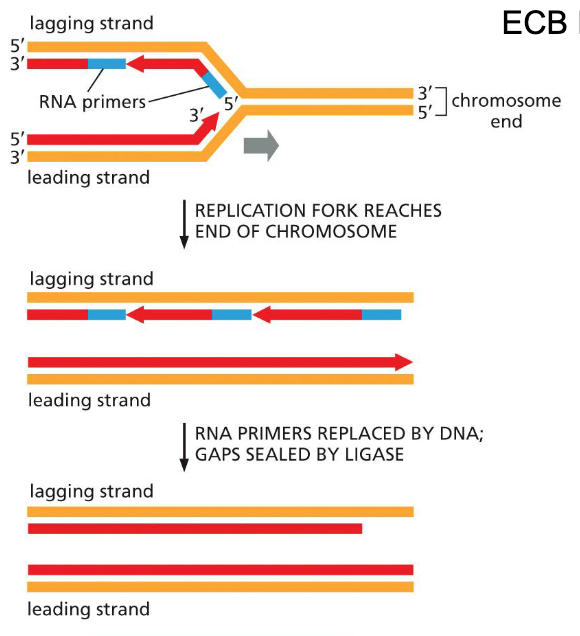
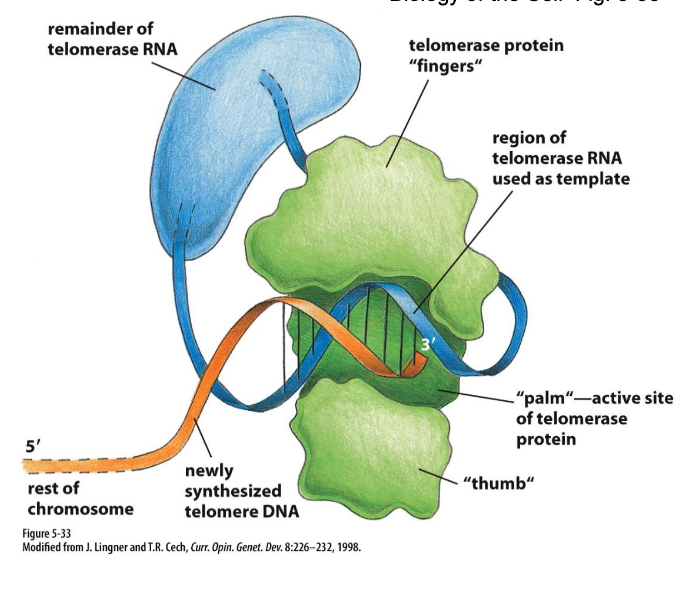
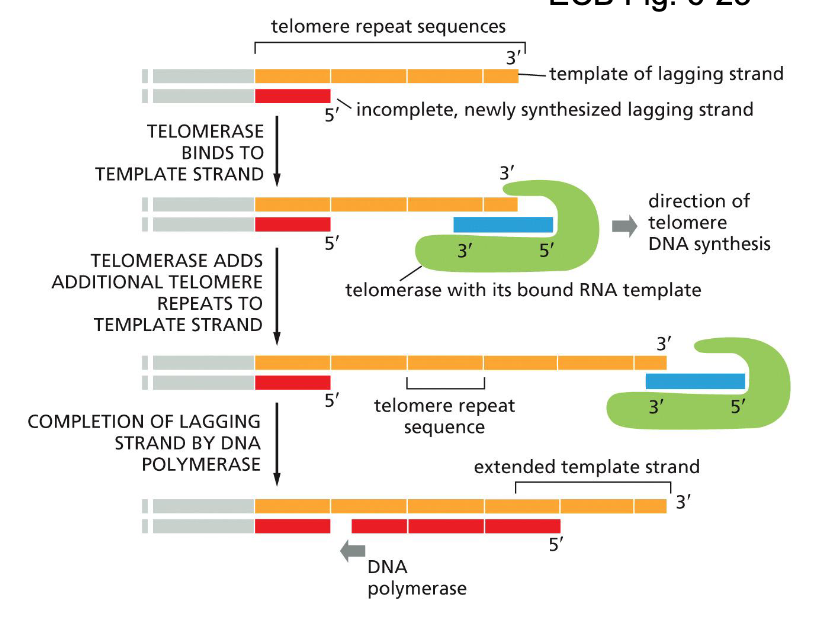
Ends of linear chromosomes solution: Telomerase & telomere replication (G rich ends…)
Repetitve sequence is added to the 3’ end of the parental strand (lagging strand template)
added by telomerase
complementary to an RNA template (telomerase RNA)
resembles reverse transcriptase
G-rich ends (stable)
Daughter strand will still be shorter but no important DNA information will be lost
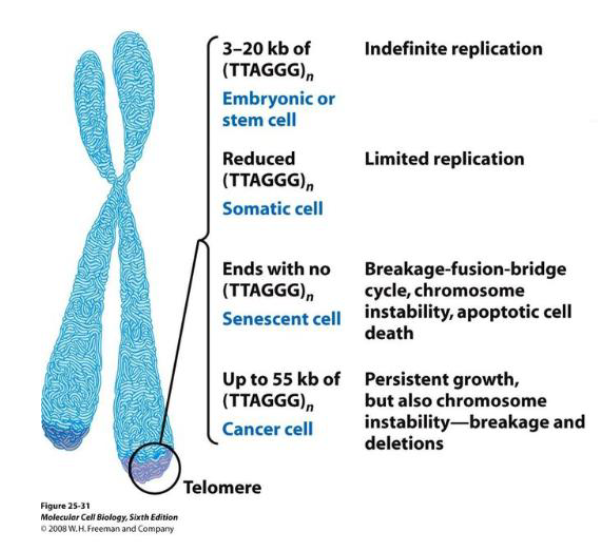
Telomeres and cells
Stem and germ-line cells — indefinite, abundant
Somatic cells — limited, can stop
Senescent cells — not making telomeres, stop dividing when reaches signal
Cancer cells — high levels of telomerase (more than stem/germ), persistent growth
issues in DNA replication: Correcting mistakes
If mistakes are not corrected, it will lead to permanent mutations as the strand divides again
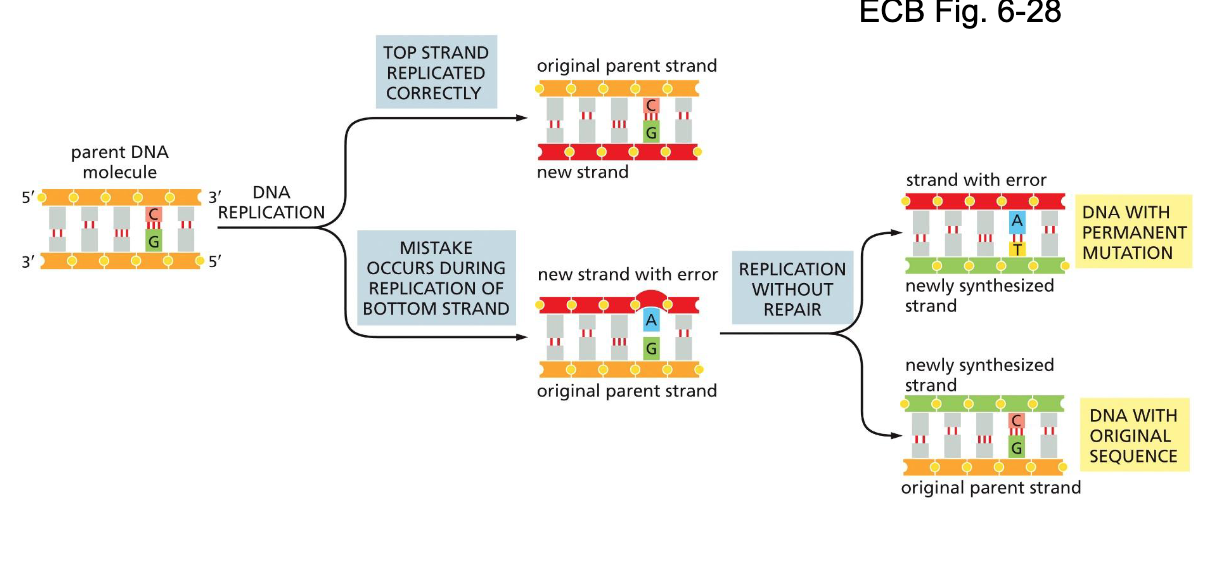
High fidelity of DNA replication
RNA polymerase have error 1 in 1000
DNA polymerase have error 1 in 109
Human genome only changed by about 3 nucleotides per divide
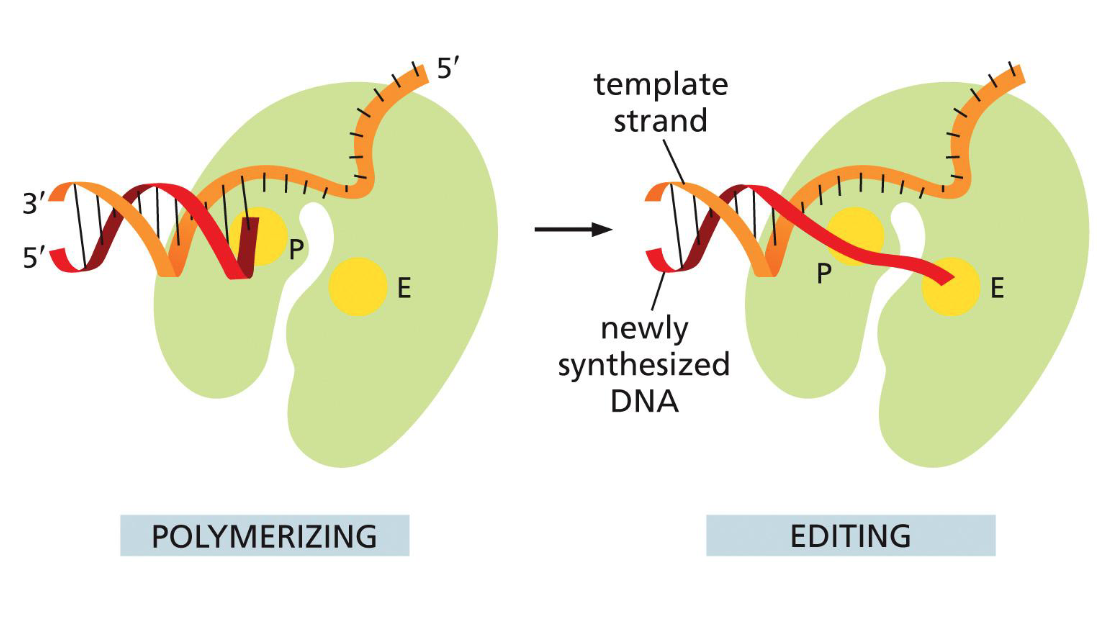
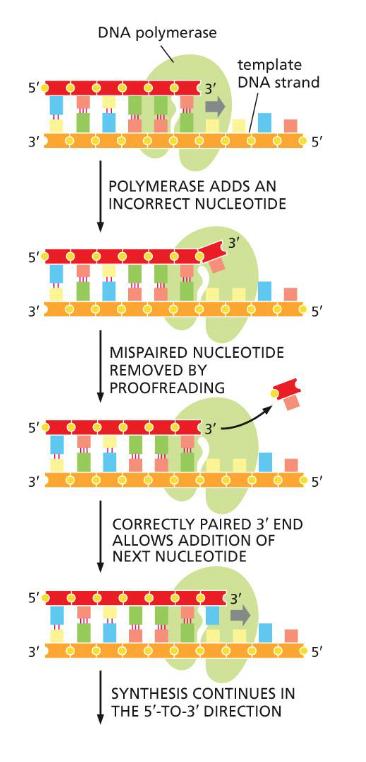
DNA proofreading and repair: 3’ to 5’ exonuclease
DNA polymerase has proofreading exonuclease activity
3’ - 5’ direction
same time as synthesis
checks if previous is correct
if not, clips off mispaired
adds new nucleotide
(this is why DNA replication can only happen 5’ — 3’)
(extrude mistake to editing site, needs to unravel a little bit)
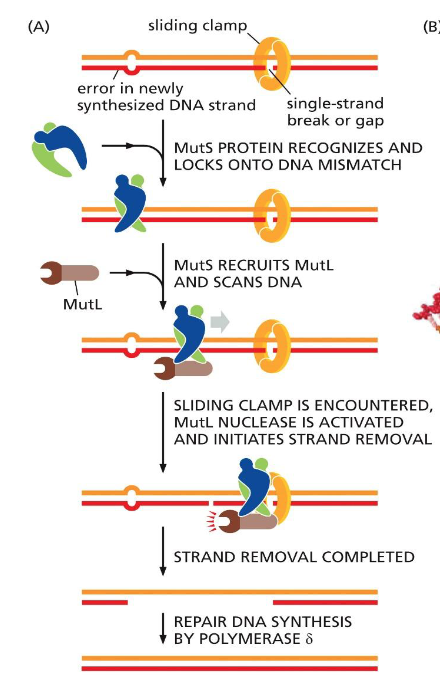
DNA proofreading and repair: Strand-directed mismatch repair (MutS and MutL)
If proofreading fails, error repair
MutS recognize distortion in geometry caused by mismatched pairs
Locks onto DNA
Recruits MutL and scans DNA
Reach sliding clamp, MutL nuclease activate
Clips new strand twice
Filled in with Polymerase δ
Recognizes new strand either by methylation on new strand or by clamp on nick