Module 11 - RNA and Transcription
the flow of genetic information in cells
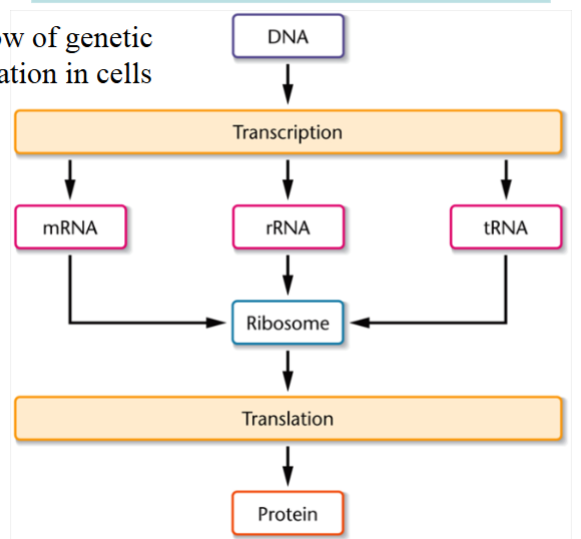
DNA → transcription → mRNA, rRNA, or tRNA → ribosome (all RNAs) → translation → protein
primary structure of a nucleic acid

the sequence of nucleotides
reminders: RNA has U instead of T and an OH group on the 2- C of its sugar instead of an H (more reactive than DNA)
forms secondary structures when folded up
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
41 Terms
the flow of genetic information in cells
DNA → transcription → mRNA, rRNA, or tRNA → ribosome (all RNAs) → translation → protein

primary structure of a nucleic acid
the sequence of nucleotides
reminders: RNA has U instead of T and an OH group on the 2- C of its sugar instead of an H (more reactive than DNA)
forms secondary structures when folded up

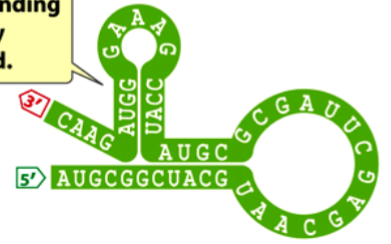
secondary structure
a structure that results when an RNA primary structure folds up due to hydrogen bonding between complementary bases on the same strand
e.g. a stem-loop (aka hairpin) or an alpha helix
types of RNA in all cells
mRNA
rRNA
tRNA
mRNA
messenger RNA
one of the types of RNA found in all cells
rRNA
ribosomal RNA
ribosomal RNA
one of the types of RNA found in all cells
tRNA
transfer RNA
one of the types of RNA found in all cells
types of RNA found only in eukaryotes
pre-mRNA
snRNA
snoRNA
miRNA
siRNA
piRNA
pre-mRNA
pre-messenger RNA
one of the types of RNA found only in eukaryotes
snRNA
small nuclear RNA
one of the types of RNA found only in eukaryotes
snoRNA
small nucleolar RNA
one of the types of RNA found only in eukaryotes
miRNA
microRNA
one of the types of RNA found only in eukaryotes
siRNA
small interfering RNA
one of the types of RNA found only in eukaryotes
piRNA
piwi-interacting RNA
one of the types of RNA found only in eukaryotes
types of RNA found only in prokaryotes
crRNA
crRNA
CRISPR RNA
the type of RNA found only in prokaryotes
transcription
the transfer of genetic information from DNA by the synthesis of a complementary RNA molecule copied from the DNA template
requires a gene in DNA (the template), RNA polymerase (the enzyme), and free NTPs
primers are not needed
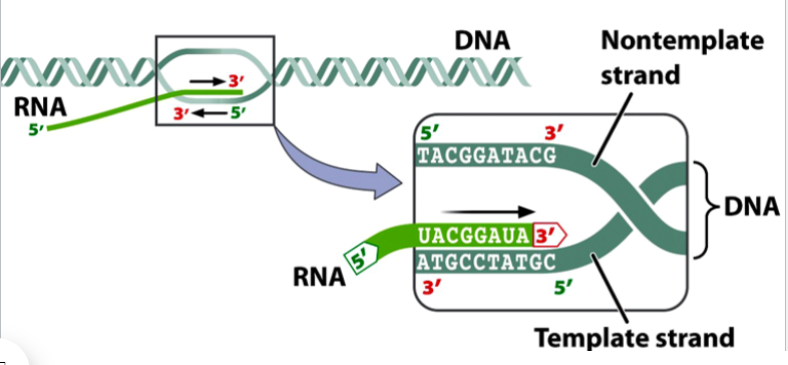
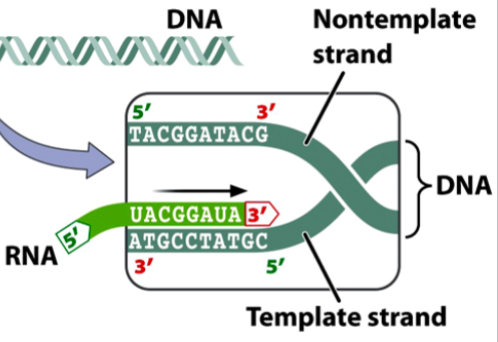
the antisense strand
the template strand of DNA
gets transcribed
bases are complementary to the RNA formed
the bottom dark green strand in the picture
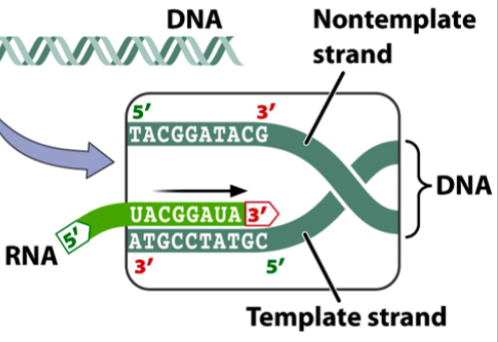
the sense strand
the non-template strand of DNA
does not get transcribed
bases match the RNA formed, except Ts gets swapped for Us
the top dark green strand in the picture
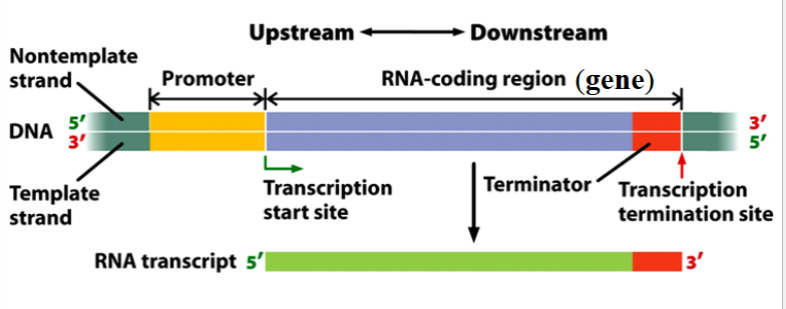
transcription unit
a stretch of DNA that encodes an RNA molecule and the sequences necessary for its transcription
components: promoter, RNA-coding region, terminator
promoter
the binding site for RNA polymerase and the transcription initiation apparatus
one of the components of a transcription unit
depicted as yellow in the picture
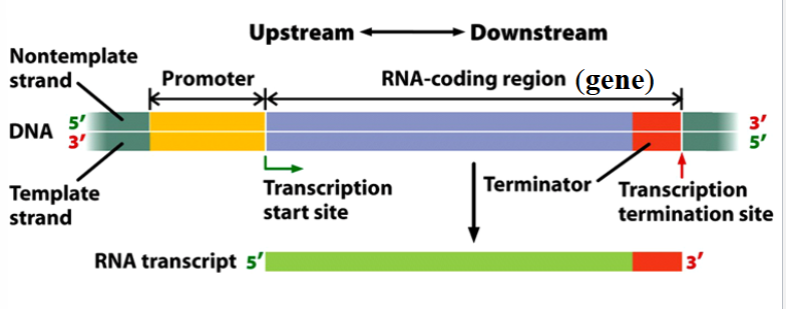
RNA-coding region
a sequence of DNA nucleotides that is copied into an RNA molecule
the gene itself; only the transcribed portions of the transcription unit
one of the components of a transcription unit
depicted as blue in the picture
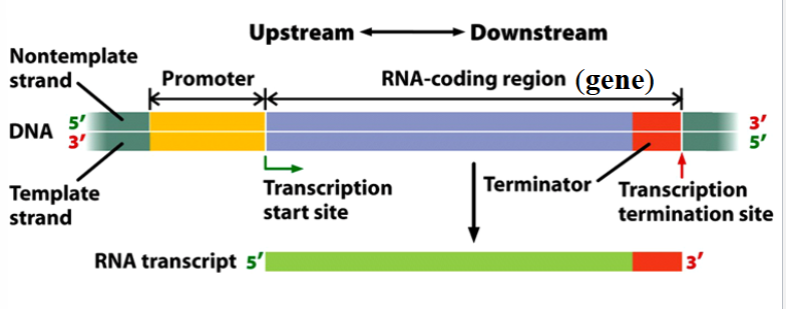
terminator
a sequence of nucleotides that signals where transcription is to end
part of the gene
depicted as red in the picture
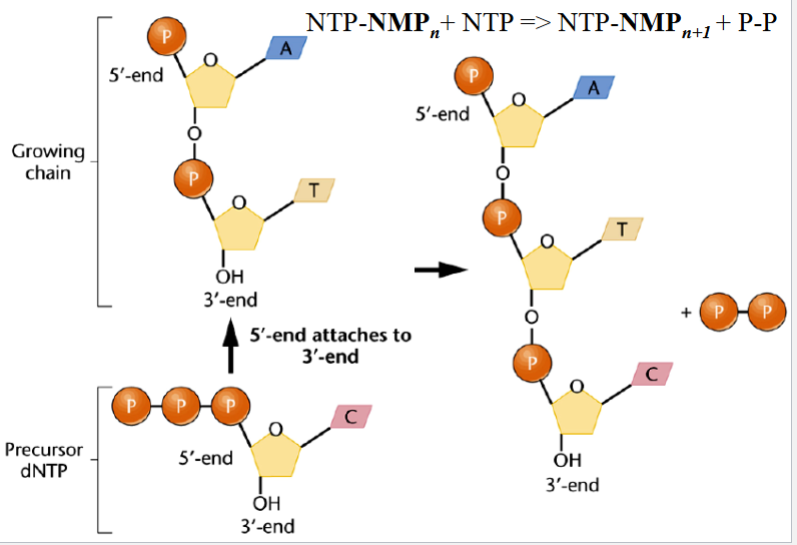
phosphodiester bond formation
one phosphate of an NTP (at the 5’ end of a free molecule) attaches to the 3’ OH of another molecule (in a chain)
the H pops off of the OH, two phosphates pop off
during initiation: NTP + NTP → NTP-NMP + P-P
during elongation: NTP-NMPn + NTP → NTP-NMPn+1 + P-P
note: in equation above, - represents a bond between terms, not a minus sign
note: the initial nucleotide of the chain (5’) remains as a nucleoside triphosphate (NTP)
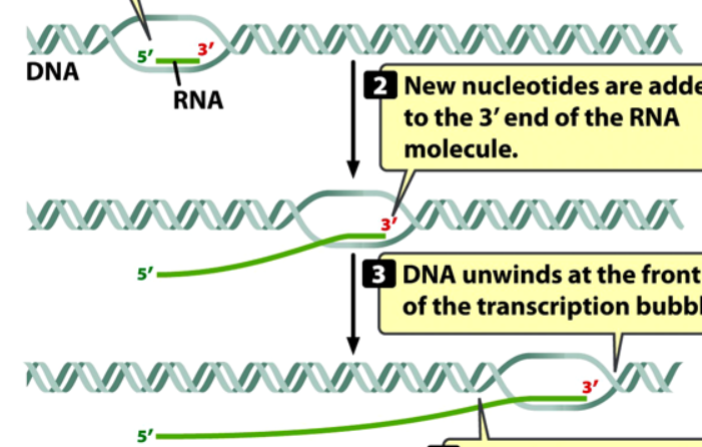
RNA synthesis in transcription
initiation does not require a primer
new nucleotides are added to the 3’ end of the RNA molecule
DNA unwinds at the front of the transcription bubble and then rewinds
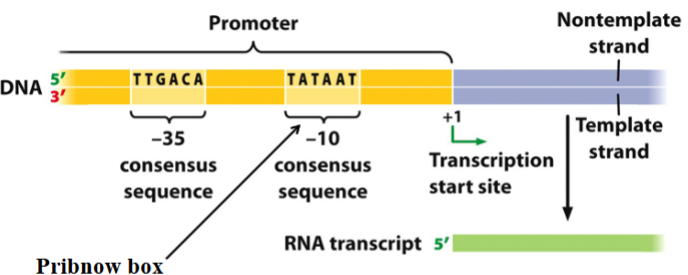
consensus sequence
comprises the most commonly found nucleotides at a specific DNA site
a conserved sequence
e.g. TTGACA, TATAAT
e.g Pribnow box in bacteria (TATAAT), which has similar function as TATA box in eukaryotes
found in the promoter
depicted by light yellow in the picture
process of transcription in bacteria
catalyzed by RNA polymerase
sigma factor recognizes promoter, directs RNA polymerase to it → forms holoenzyme
DNA double helix is unwound and denatured locally
initiation of transcription begins at the transcription initiation site
sigma factor dissociates after initation
transcription continues at ~50 nucleotides per second at 37* until the enzyme encounters the terminator sequence
promoter
the binding site for RNA polymerase on the DNA molecule
right before +1
not transcribed
gets recognized by sigma promoter; the site where the sigma factor binds with RNA polymerase
transcription initiation site
where initiation of transcription begins
+1
the first base to be transcribed
sigma factor dissociates here
termination of transcription in bacteria
depends on the formation of the hairpin loop secondary structure at the terminator site
hairpin loop forms after inverted repeats in the gene are transcribed into the RNA molecule and form intramolecular base pairs
formation of the hairpin loop destabilizes the DNA-RNA pairing; transcription ends
can be rho-dependent or rho-independent
rho-dependent termination
rho protein (a helicase) is needed to break the DNA-RNA pairing and terminate transcription
rho binds to an unstructured region of RNA; moves toward its 3’ end
when RNA polymerase encounters a terminator sequence, it pauses, rho catches up
rho unwinds the DNA-RNA hybrid using helicase activity, brings transcription to an end
rho-independent transcription
the weak DNA-RNA pairing at the terminator site is enough to destabilize the interaction and terminate transcription
terminator contains an inverted repeat followed by ~6 A nucleotides
inverted repeats are transcribed into RNA
the string of Us causes RNA polymerase to pause
the inverted repeats in RNA fold into a hairpin loop, which stabilizes the DNA-RNA pairing
the RNA transcript separates from the template, terminating transcription
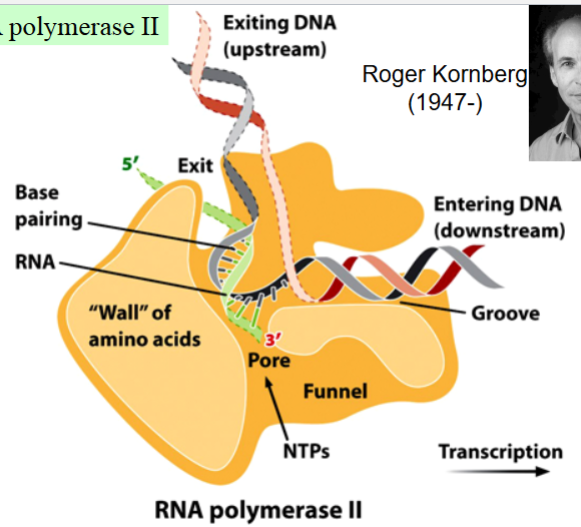
RNA polymerase II and transcription elongation in eukaryotes
RNA polymerase maintains a transcription bubble during elongation
in the bubble, ~8 nucleotides of RNA remain base-paired with the DNA template strand
the DNA double helix enters a cleft in the polymerase, gets gripped by jaw-like extensions of the polymerase
the two DNA strands get unwound
RNA nucleotides enter the polymerase through a pore, get added to the 3’ end of the growing RNA molecule
the DNA-RNA hybrid funnels through the polymerase, hits a wall and bends → keeps bubble open, positions the DNA-RNA hybrid at the polymerase’s active site
the newly synthesized RNA separates from the DNA, runs through another groove before exiting the polymerase
termination of transcription of protein-coding genes in eukaryotes
does not occur by itself
requires the activity of Rat1 exonuclease
after the protein-coding region of the gene is transcribed, an RNA endonuclease cleaves the RNA molecule at a consensus cleavage site
Rat1 binds to the unprotected 5’ end of the trailing, non-coding RNA fragment, begins to degrade it until it catches up with the RNA polymerase (note: RNA polymerase is still transcribing the DNA)
complete degradation of the trailing fragment terminates transcription
Rat1
a 5’→3’ RNA exonuclease involved in the termination of transcription in eukaryotes
binds to the 5’ end of trailing, non-coding RNA and degrades it until it reaches RNA polymerase
CRISPR RNA (crRNA)
clustered regularly interspaced short palindromic repeats
the basis of a form of adaptive immunity found in bacteria and archaea (prokaryotes)
DNA arrays consisting of a number of short palindromic sequences separated by spacer sequences
3 stages: acquisition, expression, interference
spacer sequences
DNA sequences derived from invading DNA molecules, e.g. bacteriophages or plasmids
separate the palindromic sequences in crRNA
a memory of the invading DNA
three stages of the CRISPR-CAS system
acquisition
expression
interference
acquisition
the first stage of the CRISPR-CAS system
foreign DNA enters the cell; gets identified, processed, and inserted into the CRISPR array as a new spacer
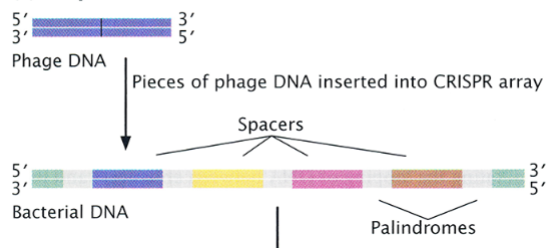
expression
second stage of the CRISPR-CAS system
the entire array gets transcribed into a long CRISPR precursor RNA, then cleaved by a CAS protein into crRNAs
each crRNA contains 1 spacer homologous to a foreign DNA
each crRNA combines with a CAS protein to form an effector complex
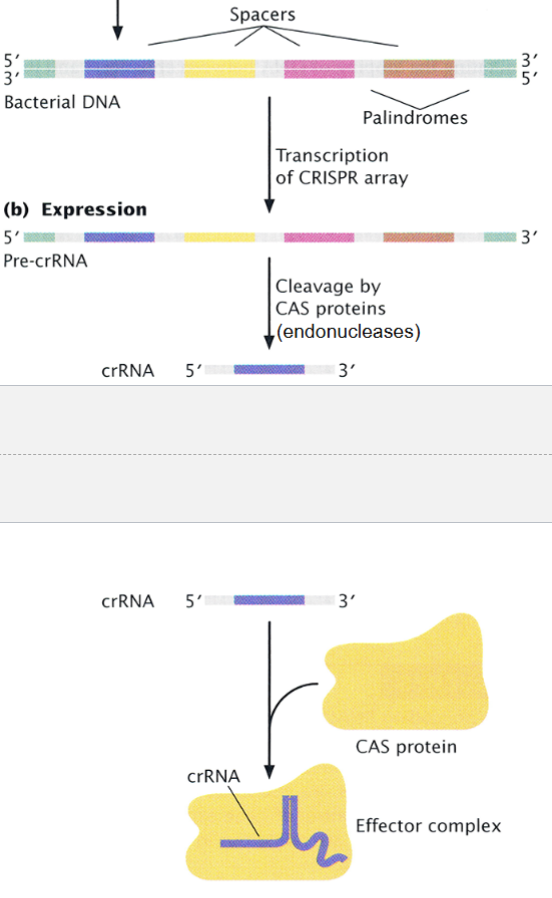
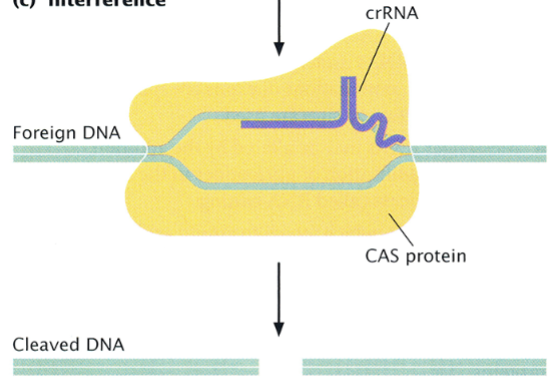
interference
the third stage of the CRISPR-CAS system
if the same foreign DNA enters the cell again, effector complex recognizes it by its base-pair complementarity to the crRNA
CAS protein cleaves it with its endonuclease activity