L1 - Cell Cycle Control
5.0(2)
Studied by 10 peopleCard Sorting
1/55
Last updated 8:48 PM on 12/7/22
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
56 Terms
1
New cards
Function of Cell Division
Embryonic Development
Tissue regeneration - to replace naturally dying cells
Homeostasis - to repair
Tissue regeneration - to replace naturally dying cells
Homeostasis - to repair
2
New cards
The cell cycle
The basis of replication, development and growth
Highly conserved across eukaryotes
Requires intricate coordination
Interphase + M phase
Occurs every 24h (ish)
Not every cell goes through it
Highly conserved across eukaryotes
Requires intricate coordination
Interphase + M phase
Occurs every 24h (ish)
Not every cell goes through it
3
New cards
When the cell grows it needs to make more:
Proteins
RNA
DNA
Lipids
Organelles
RNA
DNA
Lipids
Organelles
4
New cards
Interphase
The growth phase
G1 (Gap 1) + S (Synthesis) + G2 (Gap 2)
Over 90% of time spent in this phase
The chromosomes are relaxed and long
G1 (Gap 1) + S (Synthesis) + G2 (Gap 2)
Over 90% of time spent in this phase
The chromosomes are relaxed and long
5
New cards
M phase
The division phase
Mitosis + Cytokinesis
Mitosis + Cytokinesis
6
New cards
Division of cellular components
Most cellular content is divided roughly in two, except for DNA which divided exactly in two
7
New cards
Quiescence
G0
Reversible cell cycle exit
Temporarily leaving the cell cycle
Reversible cell cycle exit
Temporarily leaving the cell cycle
8
New cards
Senescence
Permanent cell cycle exit
In specialised or damaged cells
To prevent cancer by stopping growing entirely
In specialised or damaged cells
To prevent cancer by stopping growing entirely
9
New cards
G1 Phase
Gap 1
Growth phase
Synthesis of RNA and proteins
Where the cycle starts
Growth phase
Synthesis of RNA and proteins
Where the cycle starts
10
New cards
S phase
Synthesis
When DNA replication occurs
Cannot finish until all DNA is replicated
When DNA replication occurs
Cannot finish until all DNA is replicated
11
New cards
G2 phase
Skipped by some cells
DNA repair
Cell prepares for mitosis
At the end of this stage the cell has 2 copies of each 46 chromosomes
DNA repair
Cell prepares for mitosis
At the end of this stage the cell has 2 copies of each 46 chromosomes
12
New cards
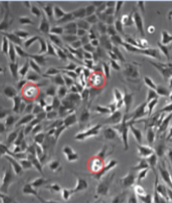
Mitotic cells in culture
Round up
In order to go through division cells detach and look round rather than flat
In order to go through division cells detach and look round rather than flat
13
New cards
Mitosis
DNA is partitioned equally during mitosis with the help of centromeres
Results in two diploid daughter cells, identical to the parent
Prophase
Prometaphase
Metaphase
Anaphase
Telophase
Results in two diploid daughter cells, identical to the parent
Prophase
Prometaphase
Metaphase
Anaphase
Telophase
14
New cards
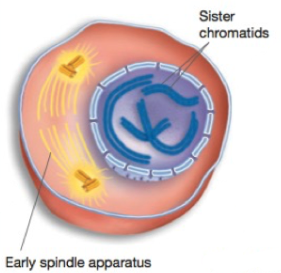
Prophase
Chromosomes condense
Spindle aparatus begins to form
The two sister chromatids lie togehter, attached at the centromere
Spindle aparatus begins to form
The two sister chromatids lie togehter, attached at the centromere
15
New cards
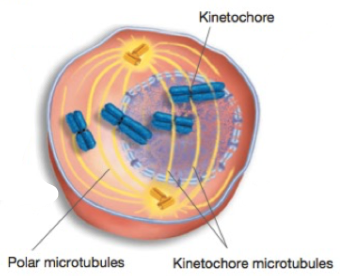
Prometaphase
Nuclear envelope breaks down
Microtubules contact chromosomes at kinetochores
Microtubules contact chromosomes at kinetochores
16
New cards
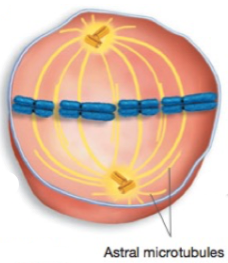
Metaphase
Chromosomes complete migration to the middle of the cell (equatorial plane)
Most highly condensed chromosomes
Most highly condensed chromosomes
17
New cards
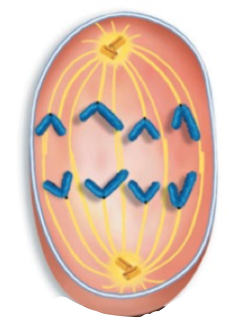
Anaphase
Sister chromatids separate into daughter chromosomes and are pulled to opposite poles of the spindle apparatus, centromere first
End with 92 seperate chromosomes, half near each
End with 92 seperate chromosomes, half near each
18
New cards
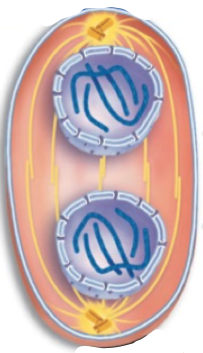
Telophase
The nuclear envelope re-forms and chromosomes condense
Spindle fibres disappear
Spindle fibres disappear
19
New cards
Centrosome
The principal microtubule organising centre (MTOC)
Duplicated and divided exactly once per cell cycle
After duplication the centrosomes migrate to either ends of the cell
Most cells only have one centrosome, but some (e.g. cilia) have multiple
Duplicated and divided exactly once per cell cycle
After duplication the centrosomes migrate to either ends of the cell
Most cells only have one centrosome, but some (e.g. cilia) have multiple
20
New cards
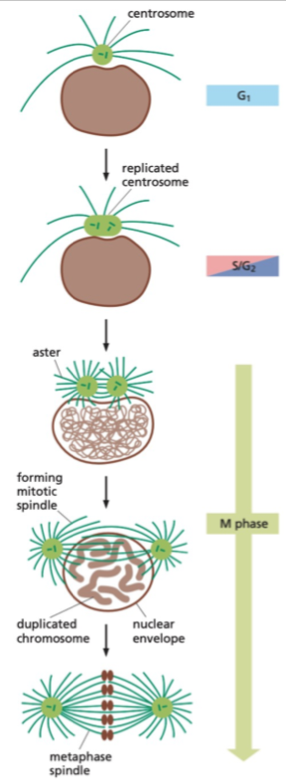
The Centrosome Cycle
Centrosome cycle runs at the same time as cell cycle - shares many features
21
New cards
Cell cycle regulation
Cells cannot return to the previous state after a restriction point
Checkpoints maintain directionality and as quality control
Mostly involving PTMs, small and reversible, covalently attached, needing an enzyme and being veru strong
Checkpoints maintain directionality and as quality control
Mostly involving PTMs, small and reversible, covalently attached, needing an enzyme and being veru strong
22
New cards
Kinase
Phosphorylate the target
23
New cards
Phosphatase
Dephosphorylate the target
24
New cards
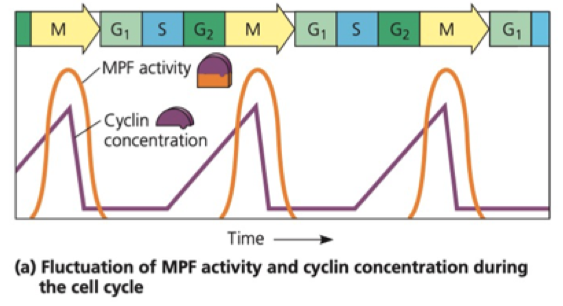
Cyclin
Non enzymatic protein
Levels of cyclin increase and decrease throughout the cell cycle
- Through protein synthesis and cleavage
Levels of cyclin increase and decrease throughout the cell cycle
- Through protein synthesis and cleavage
25
New cards
Cdk
Cyclin dependent kinase
A kinase that can phosphorylate targets
Requires cyclin to function (only active when bound to cyclin)
Can only phosphorylate specific consensus motifs on targets
Levels maintained the same throughout the cycle
A kinase that can phosphorylate targets
Requires cyclin to function (only active when bound to cyclin)
Can only phosphorylate specific consensus motifs on targets
Levels maintained the same throughout the cycle
26
New cards
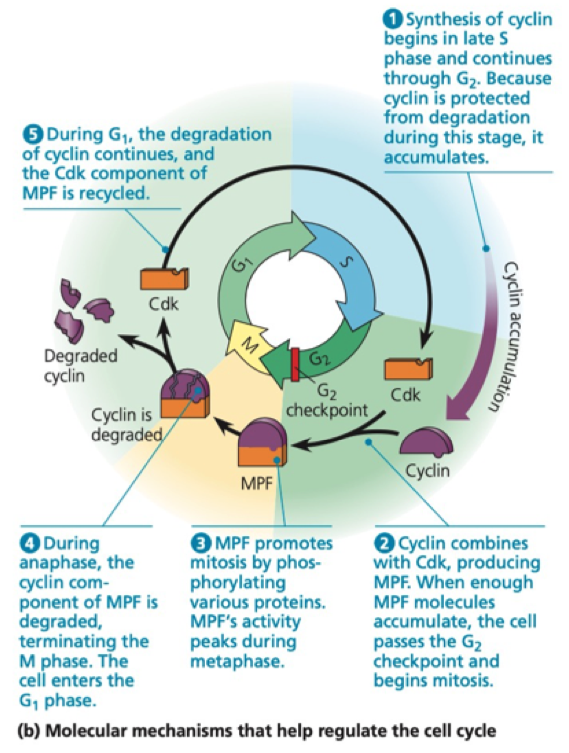
Maturation promoting factor
First discovered in frogs
Fusion of mitotic cells with a cell in any other cycle stage causes premature chromosome condensation - these cells go into mitosis
MPF consists of Cyclin and Cdk
Fusion of mitotic cells with a cell in any other cycle stage causes premature chromosome condensation - these cells go into mitosis
MPF consists of Cyclin and Cdk
27
New cards
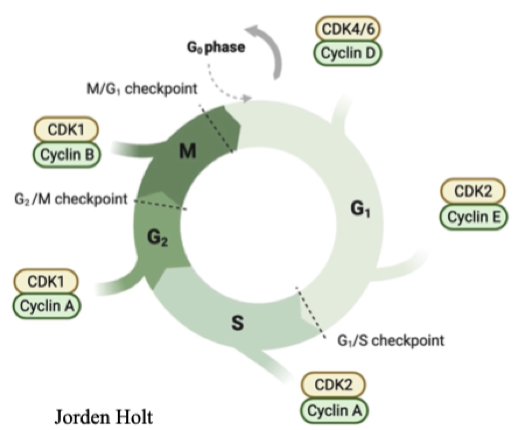
Cyclin expression
Different types expressed at different points in the cell cycle
Confer substrate specificity for CDK
Downstream targets of Cyclin-CDK move the cell cycle forward
Levels decrease sharply after the end of the phase - sharp line between phases
Confer substrate specificity for CDK
Downstream targets of Cyclin-CDK move the cell cycle forward
Levels decrease sharply after the end of the phase - sharp line between phases
28
New cards
Internal influences of cell cycle
Growth
DNA replication
DNA damage
- Little damage must be repaired
- Lots of damage causes senescence or apoptosis
DNA replication
DNA damage
- Little damage must be repaired
- Lots of damage causes senescence or apoptosis
29
New cards
External influences of the cell cycle
Food
Space
Communication with organs - tissue damage and developmental stage
- Hormones
Space
Communication with organs - tissue damage and developmental stage
- Hormones
30
New cards
Cyclin destruction
Through ubiquitination
Causes abrupt decrease in Cdk function after end of phase
Causes abrupt decrease in Cdk function after end of phase
31
New cards
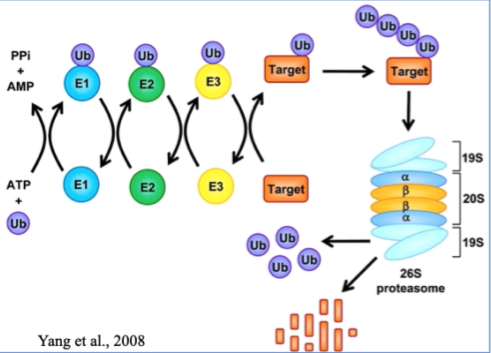
Ubiquitin-Proteasome System
Ubiquitin - small PTM, covalently linked
Polyubiquitation formed on lysines
An entire protein enters the proteasome and amino acids leave
These can then be used to make more proteins
Ubiquitins open the "lid" of the proteasome so it opens and degrades the protein
Polyubiquitation formed on lysines
An entire protein enters the proteasome and amino acids leave
These can then be used to make more proteins
Ubiquitins open the "lid" of the proteasome so it opens and degrades the protein
32
New cards
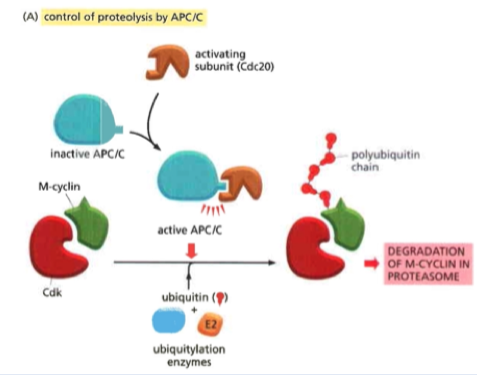
APC/C
Anaphase Promoting Complex or Cyclosome
In the ubiquitin ligase family (joins ubiquin to something)
Key regulator of metaphase to anaphase transition
Cyclin is the most important target
Coactivators:
- Cdc20 (mitosis to metaphase)
-- polybiquinates M cyclin (bound to Cdk) and causes degradation
- Cdh1 (anaphase to the start of S phase)
In the ubiquitin ligase family (joins ubiquin to something)
Key regulator of metaphase to anaphase transition
Cyclin is the most important target
Coactivators:
- Cdc20 (mitosis to metaphase)
-- polybiquinates M cyclin (bound to Cdk) and causes degradation
- Cdh1 (anaphase to the start of S phase)
33
New cards
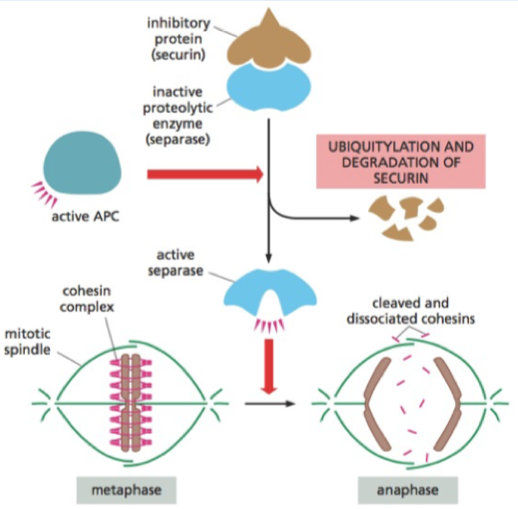
Cohesin
Holds chromosomes together
Cohesin cut by seperase - to go from metaphase to anaphase
Seperase held by securin (inhibitory)
APC ubiquitinates scurin and so releases separase when all chromosomes are lined up in middle of cell
Cohesin cut by seperase - to go from metaphase to anaphase
Seperase held by securin (inhibitory)
APC ubiquitinates scurin and so releases separase when all chromosomes are lined up in middle of cell
34
New cards
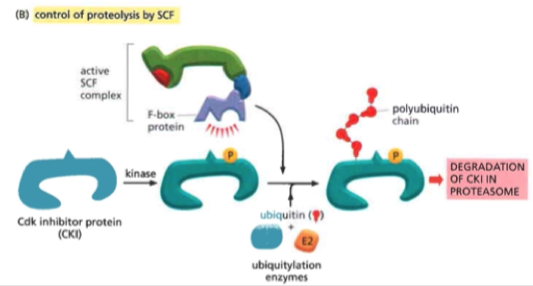
CKI (CDK inhibitor proteins)
Block Cdk function by covering the protein surface (including active site) or covalently attaching chemical groups
Covers the active site of the protein
- Temporary non-covalent interaction
- When DNA damage will stop Cdk from acting and holding in the current cell cycle phase
Is broken down by ubiquintation by the SCF complex
Two families in mammals:
- CIP/KIP
- INK4
Covers the active site of the protein
- Temporary non-covalent interaction
- When DNA damage will stop Cdk from acting and holding in the current cell cycle phase
Is broken down by ubiquintation by the SCF complex
Two families in mammals:
- CIP/KIP
- INK4
35
New cards
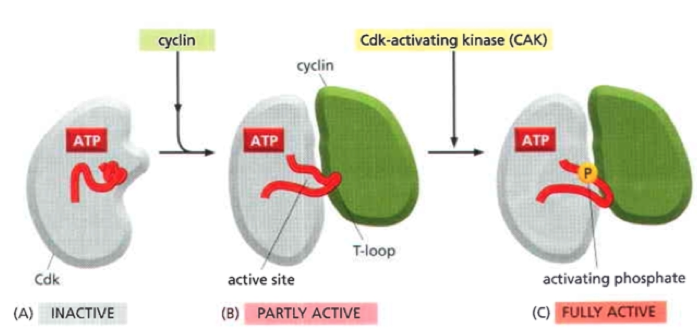
Cdk activation
Cyclin binding is necessary but not sufficient
PTMs act to fine regulate
Active site of Cdk only fully active when phosphorylated by cyclin activating kinase (CAK)
PTMs act to fine regulate
Active site of Cdk only fully active when phosphorylated by cyclin activating kinase (CAK)
36
New cards
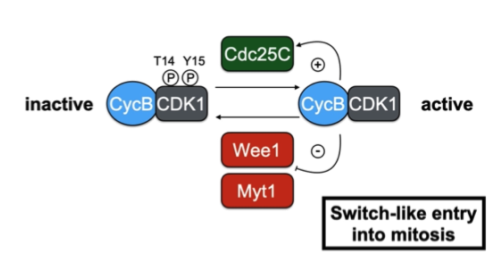
Wee1 and Cdc25
Wee1 - inhibitory kinase
Phosphorylates a neighouring site and blocks the active site
Cdc25 - activatory phosphatase
Removes the inhibitory phosphate
Phosphorylates a neighouring site and blocks the active site
Cdc25 - activatory phosphatase
Removes the inhibitory phosphate
37
New cards
p21
A Cdk inhibitor
Covers the cyclin CDK
Regulated by p53
- p53 is often gone in cancer cells
Covers the cyclin CDK
Regulated by p53
- p53 is often gone in cancer cells
38
New cards
p53
TF to activate p21 transcription
In response to DNA damage, p53 is activated by phosphorylation
Pauses the cell cycle to repair the damage
In response to DNA damage, p53 is activated by phosphorylation
Pauses the cell cycle to repair the damage
39
New cards
Oncogenes
Have a normal function in the cell, when upregulated causes cancer
e.g. E2f or Cyclin E
e.g. E2f or Cyclin E
40
New cards
Tumour supressor genes
When missing on not being created causes cancer
e.g. Rb
e.g. Rb
41
New cards
Positive feedback loop (Mitosis)
Ensure a process keeps going
Activating the activator and suppressing the suppressor
Activating the activator and suppressing the suppressor
42
New cards
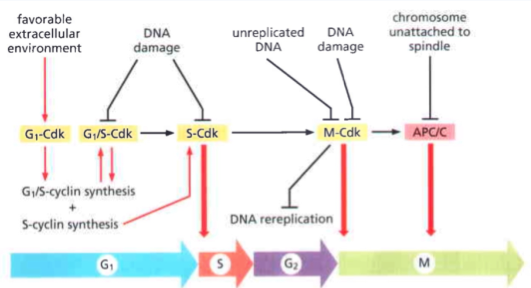
Cell cycle checkpoints
To ensure there is suitable cell state and environment before proceeding to the next stage
Each checkpoint requires a different stimulant and acts through a different Cyclin-Cdk couple
G0/G1 = Mitogen stimulation
G1/S = Restriction point
S/G2 = DNA damage
G2/M = Antephase checkpoint
M/G1 = SAC
Each checkpoint requires a different stimulant and acts through a different Cyclin-Cdk couple
G0/G1 = Mitogen stimulation
G1/S = Restriction point
S/G2 = DNA damage
G2/M = Antephase checkpoint
M/G1 = SAC
43
New cards
Mitogen stimulation
Mitogen = a molecule that pushes the cell into mitosis
Without this it will not enter the cell cycle (will remain in G0)
e.g. growth factors
Without this it will not enter the cell cycle (will remain in G0)
e.g. growth factors
44
New cards
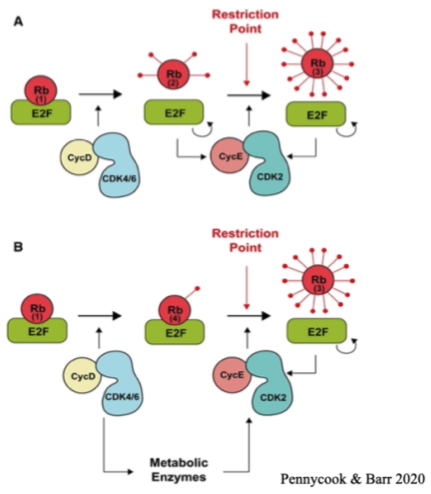
Restriction point
The point at which the cell decides whether to go through with mitosis
Mitogen signal acts through G1 and G1/S Cdks - phosphorylating Rb and releasing the Rb targets
Rb covers the transcription factor target E2F
The uncovering of this causes the transcription of genes needed for cell proliferation
Mitogen signal acts through G1 and G1/S Cdks - phosphorylating Rb and releasing the Rb targets
Rb covers the transcription factor target E2F
The uncovering of this causes the transcription of genes needed for cell proliferation
45
New cards
DNA damage mediated arrest
Can occur at any time in the cell cycle
After a cascade of signals and sequential phosphorylations, Cdk-cyclin complexes are inhibited - through recruitment of CKIs
After a cascade of signals and sequential phosphorylations, Cdk-cyclin complexes are inhibited - through recruitment of CKIs
46
New cards
Spindle Assembly Checkpoint (SAC)
Activated upon nuclear envelope breakdown
To prevent premature segregation of sister chromatids and therefore ensure genome stability
Mitotic checkpoint complex (MCC) is the SAC effector molecule, generated at unattached kinetochores
To prevent premature segregation of sister chromatids and therefore ensure genome stability
Mitotic checkpoint complex (MCC) is the SAC effector molecule, generated at unattached kinetochores
47
New cards
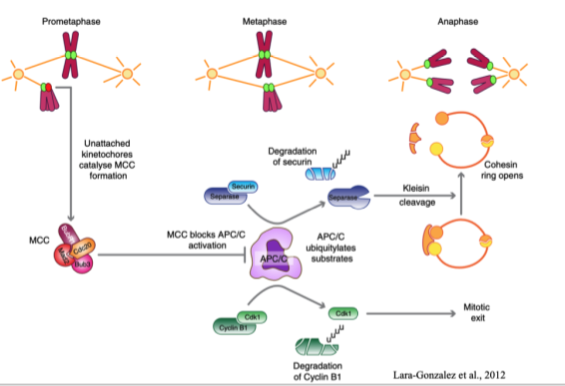
Mitotic checkpoint complex (MCC)
iThe SAC effector molecule, generated at unattached kinetochores
Inhibits APC/C in order to prevent premature anaphase and unequal segregation of DNA
No longer generated with kinetochores are attached, so APC/C can degrade key substrates
Inhibits APC/C in order to prevent premature anaphase and unequal segregation of DNA
No longer generated with kinetochores are attached, so APC/C can degrade key substrates
48
New cards
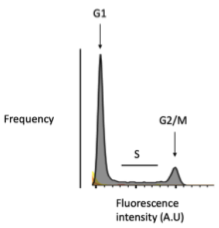
Flow cytometry
Label DNA in samples with dye then fix and sort
49
New cards
Immunohistochemistry (IHC)/Immunofluorescence IF)
Antibodies raised against known mitotic markers
Used to observe cell cycle stage
Used to observe cell cycle stage
50
New cards
Cytokinesis
Cytoplasmic division
Roughly in half
Roughly in half
51
New cards
Sister Chromatids
Identical chromosome pairs
Often exchange material during interphase
Often exchange material during interphase
52
New cards
Meiosis
Produce 4 haploid cells from one diploid cell
Two cell divisions
Two cell divisions
53
New cards
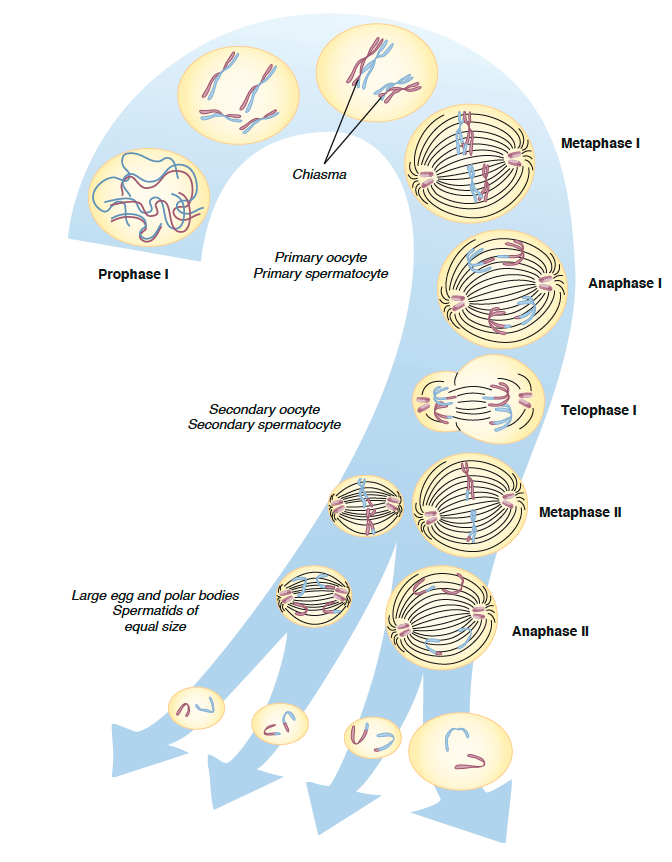
Meiosis I
Reduction division stage
2 haploid from 1 diploid
Interphase I - replication of DNA
Prophase I
- Chromatin coils
- Homologous chromosomes pair up and chromatins intertwine
- Formation of chiasmata - attachments between homologous chromosomes
- Chromosomes begin to move to centre, spindle apparatus begins to form
- Nuclear membrane disappears
Metaphase I
- Spindle formation
- Chromosomes align - two centromeres on opposite side of equatorial plane
Anaphase I
- Chiasmata disappear
- Homologous chromosomes are pulled by spindle fibres to opposite parts of the cell (one of each pair of autosomes and one of sex chromosomes)
Telophase I
- Chromosomes reach opposite ends of cell + slightly uncoil
- Nuclear membrane begins to form
- Cytokinesis (males = equal division, females = unequal - one is polar body)
2 haploid from 1 diploid
Interphase I - replication of DNA
Prophase I
- Chromatin coils
- Homologous chromosomes pair up and chromatins intertwine
- Formation of chiasmata - attachments between homologous chromosomes
- Chromosomes begin to move to centre, spindle apparatus begins to form
- Nuclear membrane disappears
Metaphase I
- Spindle formation
- Chromosomes align - two centromeres on opposite side of equatorial plane
Anaphase I
- Chiasmata disappear
- Homologous chromosomes are pulled by spindle fibres to opposite parts of the cell (one of each pair of autosomes and one of sex chromosomes)
Telophase I
- Chromosomes reach opposite ends of cell + slightly uncoil
- Nuclear membrane begins to form
- Cytokinesis (males = equal division, females = unequal - one is polar body)
54
New cards
Meiosis II
Equatorial divison
Each haploid cell replicated
Interphase II - V brief, no replication
Prophase II
- Chromosomes thicken and coil
- Nuclear membrane disappears
- Spindle fibers form
Metaphase II
- Spindle fibers pull chromosomes to equatorial plane
Anaphase II
- Centromeres split and carry single chromatid to either side
Telophase II
- Chromosomes begin to uncoil
- Nuclear membranes formed
- Cytokinesis (males = equally, females = unequal - one becomes polar body(
Each haploid cell replicated
Interphase II - V brief, no replication
Prophase II
- Chromosomes thicken and coil
- Nuclear membrane disappears
- Spindle fibers form
Metaphase II
- Spindle fibers pull chromosomes to equatorial plane
Anaphase II
- Centromeres split and carry single chromatid to either side
Telophase II
- Chromosomes begin to uncoil
- Nuclear membranes formed
- Cytokinesis (males = equally, females = unequal - one becomes polar body(
55
New cards
Spermatogenesis
Constantly occurring
Spermatogonia = diploid
Mitosis -> primary spermatocyte (diploid)
Meiosis I -> 2 secondary spermatocytes (haploid - 23ds)
Meiosis II -> 4 spermatids (haploid - 23ss)
Spermatids lose cytoplasm and develop tails
Spermatogonia = diploid
Mitosis -> primary spermatocyte (diploid)
Meiosis I -> 2 secondary spermatocytes (haploid - 23ds)
Meiosis II -> 4 spermatids (haploid - 23ss)
Spermatids lose cytoplasm and develop tails
56
New cards
Oogenesis
Mostly before birth
Oogonia = diploid
Mitosis -> primary oocyte (diploid) - during foetal development
Held in prophase I until birth, continues when ovulated
Meiosis I -> 1 secondary oocyte + 1 polar body (haploid - 23ds)
Emerges from follicle and proceeds down fallopian tube w/ polar attached
Meiosis II -> 1 mature ovum + 1 polar body (haploid - 23ss)
- Only happens if fertilised by a sperm
Polar bodies disintegrate
Oogonia = diploid
Mitosis -> primary oocyte (diploid) - during foetal development
Held in prophase I until birth, continues when ovulated
Meiosis I -> 1 secondary oocyte + 1 polar body (haploid - 23ds)
Emerges from follicle and proceeds down fallopian tube w/ polar attached
Meiosis II -> 1 mature ovum + 1 polar body (haploid - 23ss)
- Only happens if fertilised by a sperm
Polar bodies disintegrate