MICR2000 - Bacteriology
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
Conjugation
Conjugative R plasmid
DNA transfer from one cell to another
Attach two cells via pilus
transfer one strand of plasmid to recipient cell
synthesis of complementary strands in both cells
cells separate
Can carry AMR genes
Transposable elements
DNA sequences that can move positions within or between DNA strands
Two types:
Insertion sequences (IS)
smallest, encode transposase (endonuclease and integrase activities)
this cuts it out and puts it in somewhere else
IS does not encode any other genes
flanked by inverted repeat (IR) that is recognised by transposase
Transposons
contain transposase and IR as well as non transposition related genes - often AMR
often contain integrons
contain promotor and attachment site for gene cassettes (free floating DNA)
cassettes encoding gene followed by integrase specific recombination site - 59 base element recognised by integrase
integrase integrates cassettes to attachment site and promoter tests what they do
cassettes excised and transferred btwn integrons
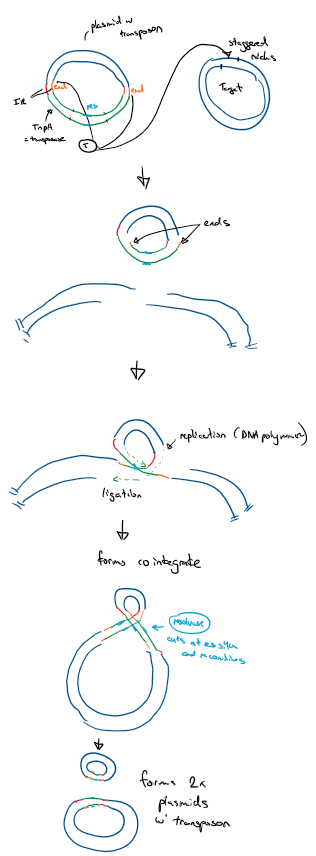
Transposition methods
Conservative - cut and paste
transposase cuts target DNA (staggered nick)
transposon integrates, DNA synthesis to fill in gaps = direct repeats
Replicative - copy and paste
TnpA gene transcribed to make transposase
transposase binds to IR and initiates transposition
cuts donor plasmid at ends and makes staggered nicks on target plasmid
ligation of transposon to target ends
3’ ends replicate through DNA polymerase, replicating the carried genes of the transposon
this forms a cointegrate - a single molecule of DNA between the donor and target plasmid (looks like 8)
contains 2x copies of transferred DNA
Resolvase binds to ‘res’ regions on transposon
cuts and recombines forming 2x plasmids with the transposon
How to stop spread of AMR
stop inappropriate use of antibiotics to reduce selective pressure
remove ineffective antibiotics from use
monitoring, isolation and treatment programs to prevent establishment and spread of multiple resistant pathogens
Normal flora
microbes that live on and in our body without causing infection under healthy conditions
balance btwn enough for microbes to survive and not enough to cause infection
pathogens can be transient members of normal flora
Virulence factors
Bacterial product or strategy contributing to virulence or pathogenicity
colonisation of host
evade immune system
damage host
6 categories
motility
motile bacteria can target host cells in dynamic environments like mucosal environments
ability to contact host cells
adhere to host cells and resist physical removal
pili/fimbriae and adhesins
invade host cells
invasins allow penetration
inside the cell access to nuritents, hide from immune system, divide and multiply
resist phagocytosis and complement
capsules hard for macrophages to attach and engulf, biofilms
evade immune defenses
phase variation of surface
vary surface structures to evade detection
capsules can resemble human tissue
don’t evade - KILL
endotoxin
lipid A in gram -ve outer membrane
released when bacteria attacked (membrane breached) and can be secreted
exotoxin
soluble excreted toxins
toxin genes spread on plasmids
cytotoxins kill or inhibit cells
neurotoxins interfere with nerves
enterotoxins affect epithelial cells of GI tract
requires production of antitoxins
ability to compete for nutrients
compete with host tissue and normal flora for limited nutrients
Staphylococcus aureus virulence factors
g+ve cocci
adhesins - adhere to host cells
secrete exotoxins that kill host cells
secrete enzymes that deteriorate red blood cells and immune system enzymes
neutralise hydrogen peroxide from macrophages - resist phagocytosis
protein a - evade immune system
capsule - resist phagocytosis
coagulase - slow down immune system through blood clot
Helicobacter pylori and virulence factors
Host adapted pathogen that colonises human stomach and duodenum - inhabits mucosal layer )noninvasive) so not cleared by immune response (persistent infection), can be treated by antibiotics
symptomatic or asymptomatic infection - virulent strains have cag a pathogenicity island
Virulence factors:
Urease
bacteria imports urea from gastric juice (through porin to periplasm, UreI to cytoplasm)
urease in cytoplasm catalyses urea → ammonia reaction
ammonia makes gastric acid more basic allowing H pylori to survive
Flagella
motility
lophotrichous arrangement
move in mucosal lining
Adhesins
BabA and SabA allow adherence to gastric epithelium
Mucinase
degrades gastric mucus locally for easier motility
CagA - cagA pathogenicity island confers high virulence
CagA protein and type IV secretory system transcribed and translated
injected into host cells via type IV secretory system (syringe) to release pro-inflammatory cytokines
increases acid which wears away mucous
Testing for H. pylori
Rapid urease test
pH test for urease catalysing ammonia production
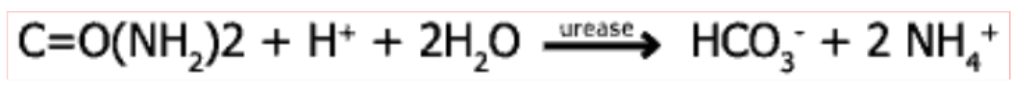
Treatment and prevention of H. pylori infection
Acid lowering drugs
Antibiotics
Group A Streptococcus (Streptococcus pyogenes) diseases, location
Location
skin and throat
Range of diseases
sore throat
localised common infections
cellulitis
impetigo (skin infection)
less common invasive infections
bacteraemia
toxic shock systems
necrotising fascilitis
post streptococcal sequelae - diseases after repeated infection with GAS
kidney failure and acute rheumatic fever (heart failure)
immune sequelae
GAS makes M protein which has anti phagocytic activity
similar to heart myosin - autoimmunity against heart myosin causing rheumatic heart disease
How do GAS infections align/misalign with Koch’s old postulates?
1 - Bacteria present in every case of disease and absent in healthy animals (NO FIT)
GAS present in normal flora
2 - Bacteria must be isolated from host with disease and grown in pure culture (FITS)
GAS can be cultured
3 - Specific disease must be reproduced when pure culture of bacteria is inoculated into a healthy susceptible host (NO FIT)
bacteria absent from post streptococcal sequelae so this doesn’t hold
different strains produce different things
4 - Bacteria must be recoverable from experimentally infected host and found to be same as original (NO FIT)
different strains of GAS
Koch’s molecular postulates
Identifying the gene or gene product responsible for virulence rather than the pathogen
Postulates
shows gene present in strains of bacteria that cause disease and not present in avirulent strains
disrupting the gene reduces virulence and reintroduction restores virulence
introduction of cloned gene into avirulent strain congers virulence
gene is expressed (not methylated)
specific immune response to gene protects against virulence
Is HtrA involved in GAS virulence?
thought HtrA involved in protecting GAS proteins during thermal stress
did test and found virulence disappeared when DNA added but didn’t return when it was returned to normal.
polar effect - DNA downstream affected (frame shift)
double crossover recombination
keeps reading frame the same
deleted mutant had no effect on virulence
Therefore HtrA doesn’t affect virulence
GAS virulence factors
M protein
helps resist phagocytosis
similar to heart myosin - immune sequelae
Fibronectin binding proteins (FBP)
allows GAS to bind to fibronectin in ECM of tissues and colonise that tissue
Phase variation
swap out FBPs
different combinations can contribute to different tissue binding - tissue tropisms
redundancy in FBPs allows infection of more than one tissue