BSCI 222 EXAM 2 FLASHCARDS
1/341
Earn XP
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
342 Terms
What is regression analysis used for?
to understand and predict the relationship between one dependent variable and one or more independent variables
In genetics, it is used to predict how the inclusion of a certain trait will then lead to a slew of diseases/phenotypic effects
What is a regression line and why is it important for genetics?
Essentially, Statistical Methods Are Required for analyzing quantitative characteristics. A regression line defines the relationship between two variables
Galton was the one who “invented” the first regression line and thus correlation
The regression line is the line that best fits the points on the graph (an “average” of sorts)
What is an example of how a regression line can be used?
regression graph between a father’s weight and his son's weight
Son was on y axis (dependent variable)
Dad was on x axis (independent variable)
Through studying the regression graph and line, it was shown there was a positive correlation (slope was positive) between father’s weight and sons.
The more dad weighed, the more likely the son was to weigh
What is the regression coefficient and how is it used?
The Regression coefficient represents the change in y per unit of change in x
Essentially the slope of the regression line.
The regression coefficient for many parent vs. Child relationships is about 0.35
We can have various regression coefficients depending on the slope of the regression line
A regression coefficient of .35 means for every 1 unit increase in x, there is a .35 unit increase in y
What is "regression to the mean".
Regression to the mean is a statistical phenomenon that describes how extreme results tend to move closer to average over time
How dramatic/ significant the regression coefficient indicates how likely it is that genetic variation is to be eliminated over time
More dramatic regression coefficient = more likely for traits to be merged/ eliminated
What is a possible explanation for the regression to the mean?
A partial explanation for regression to the mean is that recessive alleles from either parent can be passed through parents (where they are not expressed) to children
Albert Eistein and his wife were both very smart, but their children were not their level of smart, indicating that their traits were not influenced by their parent’s phenotype and thus their coefficient of regression was higher.
What does linkage mean?
The inheritance of the same parental features for more than two generations is known as linkage.
What is an example of linkage?
Flower color and pollen shape in sweet peas
We expect to have a 9;3;3;1 ratio of phenotypes, but this is not the case
Our highest amounts observed are from our parental phenotypes
The “common types” contain the parental configurations of alleles for the trait (i.e. on the chromosome it got from its parents)
The rare types of gametes contain Recombinant configurations of alleles, that is, ones that can only form if a recombination event (crossover) occurs in between the genes.
What are the “common types” in a linkage situation for progeny?
The “common types” contain the parental configurations of alleles for the trait (i.e. on the chromosome it got from its parents)
What are the recombinant types in a linkage situation for progeny?
The rare types of gametes contain Recombinant configurations of alleles, that is, ones that can only form if a recombination event (crossover) occurs in between the genes.
What happens in linkage if the genes are on the same chromosome?
The changes in gametes that can be produced (recombinants) are:
ppll individuals can only produce pl gametes
PpLl individuals can only produce PL and pl gametes (if no crossing over)
What does recombination mean?
Recombination is the variation of parental qualities in the next generation to exhibit non-parental traits. It does not support the law of independent assortment.
This is going to be our “rare”, recombinant configurations that are not a part of our parental phenotypes
Why can’t recombination frequencies exceed 50%?
A recombination frequency of 50% represents the theoretical maximum where genes assort independently
They are either on separate chromosomes or so far apart on the same chromosome that crossing over occurs randomly between them in every meiotic event, resulting in a 50% chance of getting a recombinant allele combination in each offspring;
any value higher than 50% would imply that more recombinant offspring are produced than parental offspring, which is not possible based on the mechanics of meiosis (if genes are linked, then we automatically assume that we will have more of the parental phenotype/genotype)
What are the expected ratios when examining a cross of two unlinked genes for heterozygotes?
1:2:1 genotypic and 3:1 phenotypic for unlinked (monohybrid)
9;3;3;1 phenotypic for unlinked (dihybrid)
What are the expected ratios when examining a cross of two linked genes for heterozygotes?
If you have linked genes, the ratios will not follow what they are supposed to be
You should be given the number of observed offspring with a particular trait
Most common phenotypes/genotypes are parental, common types
Least common phenotypes/genotypes are recombinant types
Use the recombination frequency (total recombinant for that allele over total offspring) to calculate map distance
What are the expected ratios for a test cross?
For a single gene test cross, the expected ratio is 1:1 if the dominant individual is heterozygous.
For a dihybrid test cross, the expected ratio is 1:1:1:1 if the individual is heterozygous for both genes.
Why you would perform a test cross when gene mapping?
When gene mapping, you would perform a test cross in order to determine the phenotypic ratios of certain traits:
You can then convert these ratios into precents, where if a percent is greater than 50% you have two independently assorting genes on different chromosomes/ far apart from each other (they are genetically unlinked)
If a percent is less than 50%, you can add percentages to determine how far apart alleles are if they are linked
What is an example of using a test cross for gene mapping?
two point testcross of flower color and pollen shape (cross PPLl and ppll – parental, or common types)
Has a 7;1;1;7 ratio, which is
44% PPLl (common)
6% PpLl (recombinant)
6% ppLl (recombinant)
44% ppll (common)
Percent recombinance = map units
12% of progeny exhibit new combinations of P and L, so p and l are 12 map units apart
How will linkage alter the results for the expected ratios for a dihybrid cross and test cross?
Linkage will alter the results for the expected ratios for a dihybrid cross and test cross by deviating from the anticipated ratio
The ratios will be more spread out than usual, with the greatest distribution of phenotypes being that of the parents, not necessarily dominant or recessive
This means it is impossible to know which trait is dominant and recessive, and more difficult to test cross to find an unknown parental genotype, for the ratios are not going to always be what you expect them to be
Essentially, the linked alleles will not separate as often during meiosis, leading to a skewed distribution of phenotypes in the offspring.
What is the purpose of using a tester in a test cross?
The purpose of using a tester (homozygous recessive for all the genes of interest) is to ensure that the alleles provided by the non-tester parent fully determine the phenotype, or appearance, of the offspring.
When are genes in the cis configuration for heterozygotes?
Cis (coupling) occurs when the wild alleles on the same chromosome and the recessive are on the other chromosome
W m
W+ m+
Would be represented as PL/pl - dominant and recessive alleles are linked together
When are genes in the trans configuration for heterozygotes?
Trans (repulsion) occurs when the wild allele shares a chromosome with the recessive allele
W m+
W+ m
Would be represented as Pl/pL
What is important to note about the arrangement of alleles (cis or trans) when it comes to test crosses?
The arrangement of alleles determines which phenotypes will be most common in a test cross
How are the phenotypes of offspring are affected by cis (coupling) and trans (repulsion) heterozygotes?
a cis (coupling) heterozygote produces fewer recombinant offspring compared to a trans (repulsion) heterozygote,
the offspring are more likely to inherit the parental allele combinations in a cis arrangement, while a trans arrangement allows for a higher chance of generating recombinant phenotypes due to crossing over during meiosis.
If wild type alleles are mixed in with recessive, non-wild type alleles, there is greater chance for variability if the alleles are linked
What is an example of how configuration (cis or trans) can influence recombinant offspring?
PL/pl (cis) vs. Pl/pL (trans)
PL/pl x PL/pl
PPLL, PpLl,ppll – MOST COMMONLY PARENT PHENOTYPES/GENOTYPES
Pl/pL x Pl/pL
PPll, PpLl, and ppLL – NOT PARENTAL GENOTYPES/PHENOTYPES
What are the things to remember with a 3-point cross? (Sturtevant's fearless ideas)
Mechanism of recombination is crossing over
Chiasmata are the visible signs of crossing over
Result in reciprocal exchange of equal and corresponding segments of homologous chromosomes
A very short distance between two genes is effectively a very small target for crossover events (low RF value)
Recombination frequencies (RF) between two genes depends on physical distance between them
How often the crossing over events occur can be measured by counting the recombinants
What are 3-point crosses?
The frequency of crossovers between two genes depends on the distance between them. We can use the frequency of recombination events between two genes (i.e., their degree of genetic linkage) to estimate their relative distance apart on the chromosome.
What are the steps to 3-point crosses?
Identify parental (most common)
Identify double recombinants (least common)
Everything else is a single recombination
What is the first step to creating a gene map?
Determine gene order by:
Identifying the parental genotypes and double recombinant genotypes
Whichever allele switches between the two is the middle allele
Ex: If our parental genotypes are VBN and vbn, and our double recombinants are vBN and Vbn, then V is our middle allele – gene order of BVN
What is the second step to creating a gene map?
Calculate RF (recombination frequencies)
Add the two single recombinants to the double recombinants and divide by the total number of progeny
We have four total single recombination events – two between the first and middle allele, and two between the middle and last allele
EX: To get RF value for BV, add single recombinants for BV to double recombinants and divide by total progeny.
RF VALUE EQUALS MAP UNITS IF WE MULTIPLY BY 100 (to get a percent)
What is the third step to creating a gene map?
Map out the genes using the map units and gene order
What is the fourth step to creating a gene map?
Calculate COC (coefficient of coincidence (C.O.C.)
COC equals observed double crossovers vs expected double crossovers
Expected double crossovers = RF1 RF2 total progeny (anything less than 1 indicates interference)
What is the final step to creating a gene map?
Calculate interference:
I = 1 – COC
tells us the fraction of double crossovers that were expected but NOT observed
Crossing over in one region influences the crossing over that may occur nearby
What is meant by the term linkage group?
all of the genes on a single chromosome. They are inherited as a group; that is, during cell division they act and move as a unit rather than independently.
What is a genetic map?
A genetic map locates genes or markers based on genetic recombination frequencies.
What is a physical map?
A physical map locates genes or markers based on the physical length of DNA sequence. Because recombination frequencies vary from one region of the chromosome to another, genetic maps are approximate.
Why is DNA stored in the nucleus?
DNA is stored in the nucleus because it needs to be protected
DNA is valuable – if it is just hanging out in the cell it could be varied greatly
It contains the cell's genetic blueprint and needs to be protected from damage
Why can RNA leave the nucleus?
RNA can leave the nucleus because its primary function is to carry the genetic information from DNA to the ribosomes in the cytoplasm where proteins are synthesized;
essentially, RNA acts as a temporary messenger molecule that can safely exit the nucleus to facilitate protein production.
DNA is the blueprint OG recipe book, but RNA is the photocopy
Why can’t DNA leave the nucleus?
DNA is also a lot larger than RNA
RNA only is encoding for specific proteins, while RNA is encoding for specific proteins for specific functions – A LOT SMALLER
What is the central dogma of biology?
DNA -> RNA -> Protein
The flow of genetic information in a cell
The information, or “language,” in DNA is ultimately translated into the language of polypeptides
When DNA is transcribed, the result is an RNA molecule
RNA is then translated into a sequence of amino acids in polypeptide
What kind of information are DNA and RNA (analog or digital)?
DNA and RNA are digital – the information flows clearly in one direction (cannot go from DNA to protein directly or from protein back to DNA)
What is the general structure of an amino acid?
Amino acids are joined together by peptide bonds (protein bonds)
Amino acids contain 3 groups - amino group (NH2), Carboxyl group (COOH) and R group (different for each amino acid)
The Amino group of one amino acid is joined together to the carboxyl group of another amino acid
Done through dehydration synthesis, where water is extracted from the molecule and the amino acids are joined together.
The common amino acids have similar structures; they are distinguished by the R groups with their different functional groups.
How does form relate to function?
In essence, form creates function
The folding (secondary structure) of amino acids creates the function of the proteins
After ribosomes link all amino acids, the protein folds into shape that determines function.
Amino acid sequence is the primary structure.
This structure folds to create secondary and tertiary structures.
Two or more polypeptide chains associate to form quaternary structure.
How does the form relate to the function of DNA?
The double helix structure of DNA, with its complementary base pairs (A-T, C-G), allows for efficient replication and stable storage of genetic information.
How does the form relate to the function of RNA?
As a single-stranded molecule, RNA is more flexible than DNA, enabling it to participate in various cellular processes like transcription (copying DNA information) and translation (converting genetic code into proteins
How does the form relate to the function of nucleotides?
The building blocks of DNA and RNA, nucleotides (composed of a sugar, phosphate, and nitrogenous base) determine the sequence that encodes genetic information
How does the form relate to the function of amino acids?
These are the building blocks of proteins, and their specific sequence, determined by the DNA code, dictates the protein's structure and function.
How does the form relates to the function of proteins?
The complex 3D structure of a protein, formed by folding of its amino acid chain, is critical for its function, including enzymatic activity, transport, and structural support
In summary, how does form relate to function for various key components of storage and replication of genetic information?
The form of DNA, RNA, amino acids, proteins, and nucleotides directly relates to their function, where the specific arrangement of their building blocks allows for the storage, transfer, and expression of genetic information, ultimately leading to the creation of functional proteins with diverse roles within an organism;
essentially, the sequence of nucleotides in DNA dictates the amino acid sequence in proteins, which then folds into a specific shape to perform its designated function.
What is a polypeptide?
long chain of amino-acids joined by peptide bonds; Are extremally diverse-a typical polypeptide contains 15 amino acids so 20150 possible sequences!! Each sequence will have different properties
What does DeepMind's program AlphaFold do, and why does it matter?
AlphaFold 3 can predict the structure and interactions of all of life’s molecules in seconds
What would take someone an entire lifetime to complete now takes a computer seconds
Allows us to analyze and see the structure of obscure proteins and thus analyze their functions
Its AI program, AlphaFold, can predict how proteins fold into 3D shapes, a fiendishly complex process that is fundamental to understanding the biological machinery of life.
What is an example of how Alphafold is used?
Demis Hassabis and John M. Jumper were part of a Google DeepMind team whose A.I. technology predicts protein shapes. The University of Washington’s David Baker designed “new protein that was unlike any other,” the Nobel committee said
If we can create new proteins, then we could possibly synthesize new proteins to fight disease, cure cancer, ANYTHING is possible
What is paleogenetics?
The scientific study of ancient genetic material, essentially examining preserved DNA from the remains of extinct organisms to understand the history of life on Earth, including the evolution and migration patterns of past populations,
What does paleogenetics allow us to do?
By comparing their genetic sequences to modern organisms, it allows researchers to learn about our ancestors and how they adapted to different environments through time.
The remarkable stability of DNA makes the extraction and analysis of DNA from ancient remains possible, including Neanderthal bones that are more than 90,000 years old and the genome of a 700,000 year horse fossil (preserved in permafrost).
What is an example of paleogenetics?
Genetic analysis of Neanderthal dental plaque from 42,000 to 50,000 years old Neanderthals from Spy Cave consumed woolly rhinoceros and wild sheep, supplemented with mushrooms. Those from El Sidrón Cave showed no evidence for meat consumption, but had a largely vegetarian diet, comprising pine nuts, moss, mushrooms and tree bark - showing quite different lifestyles between the two groups.
Allows us to understand that older groups are not necessarily more primitive, given that the tree bark they were eating was of an aspirin tree.
What is environmental DNA (eDNA)?
Enviromental DNA (eDNA) is genetic material that is shed from plants and animals - for example, from skin cells or droppings - and accumulates in their surroundings.
What is eDNA used for?
It is used for measuring biodiversity in showing the plasticity of biological organisms in terms of where they can live and the plants or animals that can live together, is larger than what we thought
What is an example of how eDNA is used?
Two million years ago, North Greenland was about 11-19C hotter than today.
The mix of temperate and Arctic trees and animals suggests a previously unknown type of ecosystem that has no modern equivalent — one that could act as a genetic road map for how different species might adapt to a warmer climate
What is Chargaff’s rule?
Chargaff's Rule A=T, C=G
If you know the composition of one nucleotide, you have the compositions of all of them because of this idea that the level of A equals the levels of T and the level of C equals the levels of G
Ex: If we have 14% frequency of A, we have 14% frequency of T, and 36% frequency of C, and 36% frequency of G.
Which nucleotides pair in DNA and how many hydrogen bonds form between them?
A-T (2 H bonds)
G-C (3 H bonds)
Which nucleotides pair in RNA and how many hydrogen bonds form between them?
A-U (2H bonds)
G-C (3H bonds)
Describe the structure of DNA
A DNA molecule is made up of two linked strands that wind around each other to resemble a twisted ladder in a helix-like shape. Each strand has a backbone made of alternating sugar (deoxyribose) and phosphate groups. Attached to each sugar is one of four bases: adenine (A), cytosine (C), guanine (G) or thymine (T).
What are the different forms of DNA?
A-DNA is a short, wide, right-handed helix.
B-DNA, the structure proposed by Watson and Crick, is the most common conformation in most living cells.
Z-DNA, unlike A- and B-DNA, is a left-handed helix.
What is meant by anti-parallel ?
the term 'antiparallel' means that the strands run in opposite directions, parallel to one another. The antiparallel strands twist in a complete DNA structure, forming a double helix. It runs 3'-5' and 5'-3' linkage.
What are the 3’ and 5’ ends of DNA?
In DNA, the 5' and 3' ends refer to the orientation of the nucleotide strands, which are important for understanding DNA structure and replication.
5' end: This end of the DNA strand has a phosphate group attached to the 5' carbon of the sugar molecule (deoxyribose).
3' end: This end has a hydroxyl group (-OH) attached to the 3' carbon of the sugar molecule.
Describe the structure of RNA
RNA is typically single stranded and is made of ribonucleotides that are linked by phosphodiester bonds. A ribonucleotide in the RNA chain contains ribose (the pentose sugar), one of the four nitrogenous bases (A, U, G, and C), and a phosphate group
What is the historical significance of Pauling, Franklin, Wilkins, Watson and Crick?
These four scientists—Crick, Franklin, Watson, and Wilkins—codiscovered the double-helix structure of DNA, which formed the basis for modern biotechnology
What are the purines?
The purines in DNA are adenine and guanine, the same as in RNA.
Purines are larger than pyrimidines because they have a two-ring structure while pyrimidines only have a single ring.
What are the pyridines?
The pyrimidines in DNA are cytosine and thymine; in RNA, they are cytosine and uracil.
Adenine
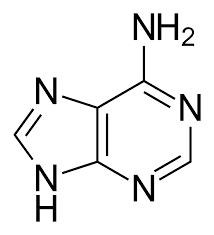
Guanine
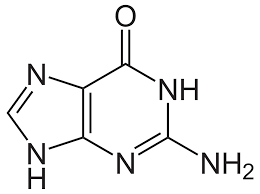
Thymine
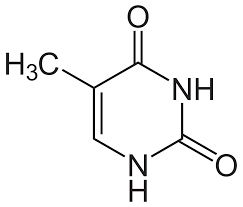
Cytosine
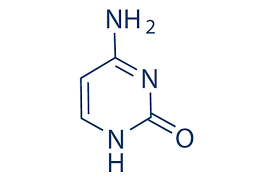
Uracil
What is meant by semi-conservative DNA replication ?
All DNA Replication Takes Place in a Semiconservative Manner
Complementary bases (C-G, A-T) that maintain the clarity and precision of DNA replication
DNA replication does not get fuzzy over time no matter how many times you copy a single strand of it – the first copy and last copy are identical
What is importance of the origin of replication?
The origin of replication is the specific location (site) on the DNA where replication always starts
Recognized by their sequence
is a crucial DNA sequence where DNA replication begins, ensuring accurate duplication and transmission of genetic material during cell division.
What is an example of the importance of the origin of replication?
E. Coli has a single origin of replication about 245 bps long of mostly A-T base pairs (weak hydrogen bonding)
Special initiator proteins bind to this site and open the DNA (DNAa in E. coli do this)
The origin of replication contains the DNA sequences that are essential for the bacteria to make copies of the plasmid. There are many different origins, and slight DNA sequence differences result in a profound difference in how many copies of the plasmid are maintained in each cell.
What are the components required for DNA replication in bacterial cells?
Initiator protein, DNA helicase, SSBPs, DNA gyrase, DNA primase, DNA polymerase I and III, DNA ligase
What is theta replication?
Circular DNA, E. Coli; as DNA opens, two y shaped structures called replication forks are formed, together making up what’s called a replication bubble
The replication forks move in oppositive directions as replication proceeds
Replication takes place at high speeds
Ex: E. Coli replicates at >1000 nucleotides/second
What components are required for linear eukaryotic DNA replication?
All the same as for bacterial (Initiator protein, DNA helicase, SSBPs, DNA gyrase, DNA primase, DNA polymerase I and III, DNA ligase)
Multiple origins of replication, allows us to “catch up” to the rate of theta replication even with our plethora of genes and proteins
Requirements: New DNA is synthesized from deoxyribonucleoside triphosphates. The newly synthesized strand is complementary and antiparallel and held together by hydrogen bonds
What is the function of an initiator protein?
binds to origin of replication, separates DNA
What is the function of DNA helicase?
unwinds DNA at replication fork
The 1st replication enzyme to load on at the origin of replication
Move the replication forks forward by unwinding the DNA (breaking the hydrogen bonds between the bps)
What is the function of SSBPs (single strand binding proteins)?
attach to single stranded DNA and prevent secondary structures from forming
Coat the separated strands of DNA keeping them from coming back together in a double helix
What is the function of DNA gyrase?
moves ahead of replication fork, making and resealing breaks in DNA
A topoisomerase II, prevents the DNA double helix ahead of the replication fork from getting too tightly wound as DNA is opened up
What is the function of DNA primase?
synthesizes RNA primer
What is the function of DNA polymerase III?
elongates new nucleotide
Responsible for synthesizing DNA: they add nucleotides one by one to the growing DNA chain, incorporating only those that are complementary to the template
Can only add nucleotides to the 3’ end of an existing strand
Cannot start making a new DNA strand from scratch but require a pre-existing chain or short stretch of nucleotides called a primer (made by DNA primase)
What is the function of DNA polymerase I?
removes RNA primer
What is the function of DNA ligase?
joins Okazaki fragments.
What is the direction of DNA synthesis?
The direction of DNA synthesis occurs in the 5’ to 3’ direction, meaning new nucleotides are added to the 3' end of a growing strand.
There is a leading and lagging strand because there is continuous and discontinuous replication in eukaryotes
What is the leading strand?
The new strand that runs 5’ to 3’ towards the replication fork, is made continuously, because the DNA polymerase is moving in the same direction as the replication fork
5- phosphates are much more unstable than 3- nucleotides (hydroxyl), meaning that the DNA is more vulnerable on this strand at this stage and more easily able to be synthesized
What is the lagging strand?
Made in fragments because as the fork moves forward, the DNA polymerase (which is moving away from the fork because it cannot add to the 5’ end) must come off and reattach to DNA
Small fragments are called Orakzai fragments, and they need a new primer
We don’t want DNA that contains RNA or gaps, so RNA primers are removed and replaced by DNA through the activity of DNA polymerase I that has 3’ to 5’ and 5’ to 3’ exonuclease (removes nucleotides) actively as well as being a 5’ to 3’ polymerase
The nicks in the sugar-phosphate backbone that remain after the primers are replaced get sealed by DNA ligase.
What is the importance of an 3’-OH group?
The 3'-OH group is crucial in DNA and RNA synthesis because it initiates the formation of phosphodiester bonds and driving the 5' to 3' polymerization process
This phosphodiester bond links the new nucleotide to the growing chain, extending the strand in the 5' to 3' direction – THIS IS THE RNA PRIMER
DNA primase synthesizes a short RNA primer to provide a 3'-OH group for the attachment of DNA nucleotides
What is the energy source for DNA replication?
The energy source for DNA replication comes from the hydrolysis of deoxynucleoside triphosphates (dNTPs), which are the building blocks of DNA.
When a dNTP is incorporated into the new DNA strand, the bond between the two terminal phosphate groups is broken, releasing energy.
The energy released from this hydrolysis drives the polymerization of dNTPs, forming the new DNA strand.
What is a picture of a DNA replication bubble?
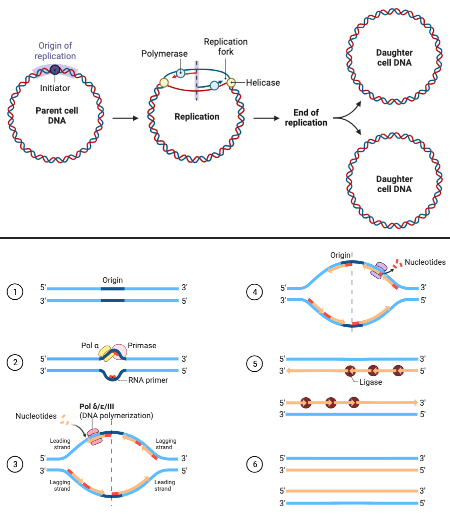
What are the major components of the DNA repliation bubble?
origin of replication, replication forks, DNA molecules produced
What is the a replication bubble?
A replication bubble is the entire region of unwound DNA where replication occurs
Forms when the double-stranded DNA unwinds and separates at specific locations called origins of replication.
Helicase enzymes separate the DNA strands, creating the bubble.
The bubble expands in both directions from the origin, allowing for replication to occur.
What is a replication fork?
A replication fork is the Y-shaped structure at the ends of the bubble where DNA strands separate and new strands are synthesized.
The Y-shaped structure that forms at the ends of the replication bubble.
These are the sites where DNA strands separate and new DNA strands are synthesized.
Replication forks move in opposite directions as replication proceeds.
What is the main differences between prokaryotic and eukaryotic DNA replication?
While DNA replication in both prokaryotes and eukaryotes involves similar mechanisms like strand separation and complementary base pairing, key differences stem from the complexity and size of their genomes, including origin of replication, replication rate, and enzymes involved
Describe DNA replication in prokaryotes
DNA Structure: Circular DNA, typically located in the cytoplasm.
Origins of Replication: Single origin of replication.
Replication Speed: Rapid replication.
Enzymes: DNA polymerase III is the main enzyme for replication, along with other enzymes like DNA ligase and helicase.
Telomeres: Prokaryotes lack telomeres, as their DNA is circular.
Replication Location: Occurs in the cytoplasm.
Describe DNA replication in eukaryotes
DNA Structure: Linear DNA, packaged with proteins (histones) into chromosomes, located within the nucleus.
Origins of Replication: Multiple origins of replication along each linear chromosome.
Replication Speed: Slower replication compared to prokaryotes.
Enzymes: Several DNA polymerases are involved, including polymerase alpha (initiates replication), delta (elongates lagging strand), and epsilon (elongates leading strand).
Telomeres: Telomeres, specialized DNA sequences at the ends of chromosomes, are maintained by the enzyme telomerase.
Replication Location: Occurs in the nucleus.