CELL 201 - UNIT 3: DNA & the Nucleus
1/64
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
65 Terms
what did Mendel discover?
patterns of inheritance
what did Miescher discover?
- first person to isolate phosphorus-rich chemicals from white blood cells using pus from a bandaid.
> named these chemicals "nuclein" because they were isolated from the nuclei of cells. the name was later changed to "nucleic acid" because of their acidic nature.
what did Thomas Hunt Morgan discover?
chromosome theory
> the link between chromosomes and inheritance
what did Barbara McClintock develop and discover?
- developed a chromosomal staining techniques to visualize the different chromosomes.
- discovered transposons (jumping genes)
what are transposons?
jumping genes
> segments of chromosomes that can move from 1 spot on a chromosome to another spot on the same chromosome
(cannot switch between different chromosomes)
what did Erwin Chargaff discover?
that DNA composition varies between species
> in any species the number of A and T bases will be equal, and the number of G and C bases will be equal
(A=T) (G=C)
what did Rosalind Franklin + Watson and Crick discover?
- the structural model of DNA (double helix)
> they did not discover DNA or that it was genetic material. they developed a hypothesis in which they explained how DNA is structures and how it could transmit genetic information.
how did Rosalind Franklin and Maurice Wilkins discover the double helix shape of DNA?
they used a technique called X-ray crystallography
> Franklin's images let Wilkins determine that DNA was helical in structure. the pattern in the image suggested that the molecule was made up of 2 strands, forming a double helix.
what did Franklin conclude about the structure of DNA
there were 2 outer sugar-phosphate backbones, with bases paired in the interiorr
what did Nirenberg discover
the triplet system (reading RNA in groups of 3)
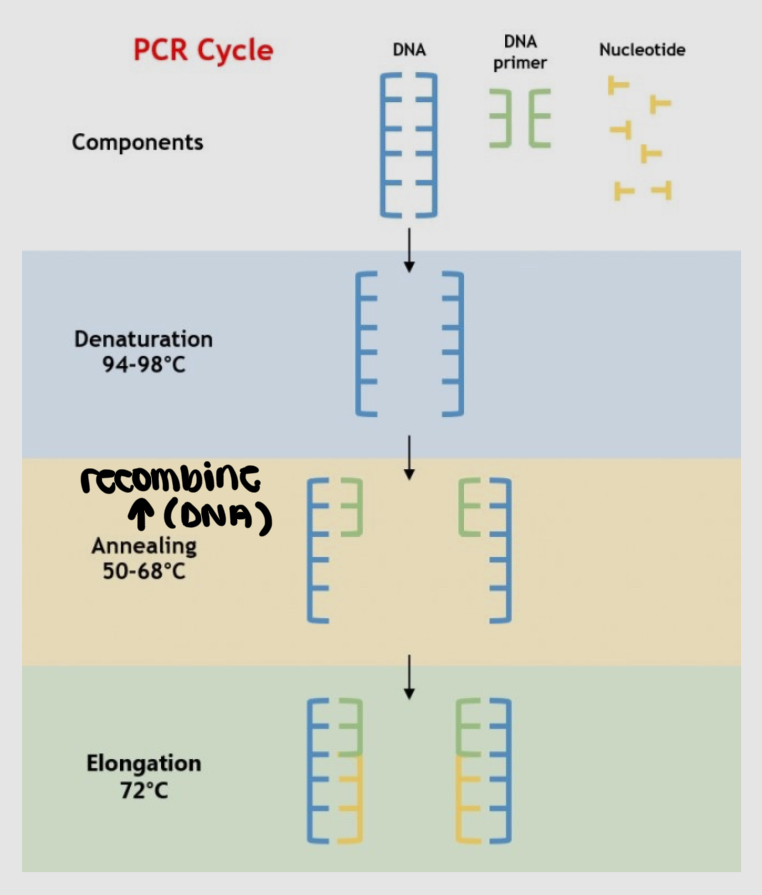
what is the Polymerase chain reaction (PCR) and how does it work
it's a method to rapidly make many copies of a specific DNA sample
what is the human genome project?
a project that had a goal to determine the base pairs that make up human DNA. identifying and mapping all of the genes in a human genome.
> the mapping of human DNA.
what is DNA
the structural basis of genetic information
> an antiparallel double helix
what is the sugar in DNA and RNA respectively
in DNA > deoxyribose sugar (H instead of OH on C-2)
in RNA > ribose sugar (OH on C-2)
how is a nucleotide imported onto a DNA strand?
via dehydration
what is a nucleoid?
a region in the cell where the DNA is contained but is not bound by a membrane
how and where is DNA found in prokaryotes?
supercoiled in the nucleoid, and as round plasmids
what is the difference between bacterial DNA as supercoiled and as plasmids
plasmids are optional, they are smaller and easily transferable between bacterias (of the same and different species)
what is the basic structure of a eukaryotic chromosome?
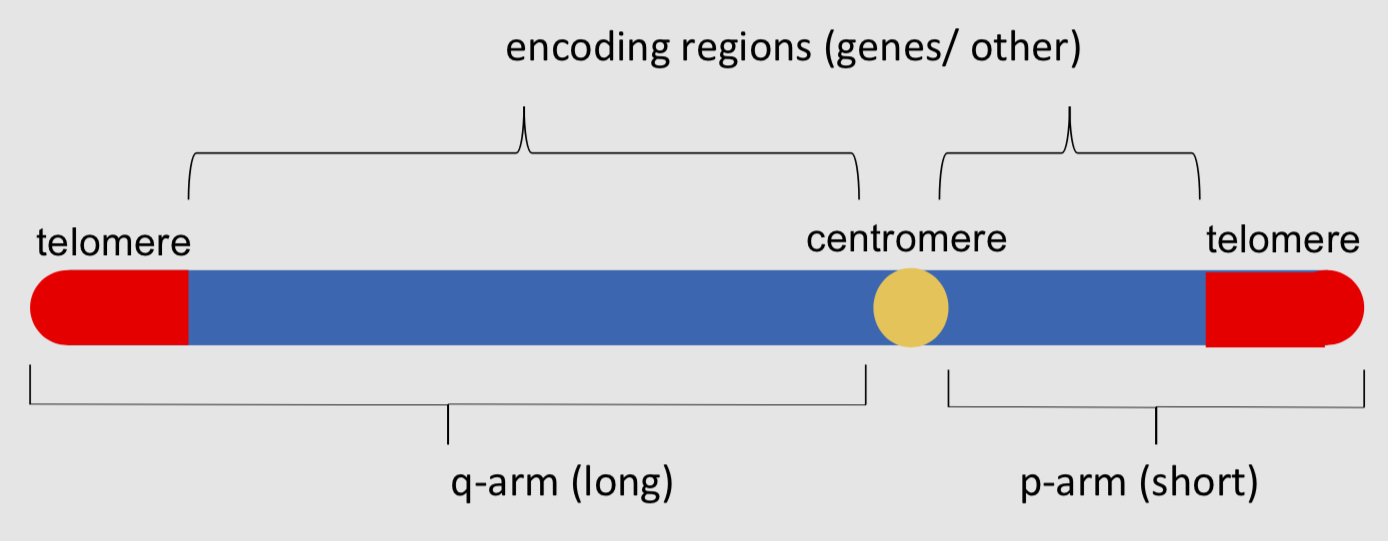
what are the unique and repeated sequences in a eukaryotic chromosome?
unique:
- protein-coding regions ~20%
- genes (introns, exons, promoters) ~25%
repeated:
- simple sequence repeats (telomeres, centromeres)
- segmental duplications (large repeated blocks in 2 or more locations)
- transposons or "jumping genes" (mobile genetic elements, often present in multiple copies)
breakdown of repeated sequences in eukaryotic chromosomes
tandem repeats:
- back to back
- 10-15% of the genome
ex. telomeres, centromeres
interspersed repeats:
- scattered around
- copies are similar but not identical (not all are necessarily active)
- hundreds of thousands of copies
ex. transposons
which ones are cleaved out during RNA splicing, introns or exons?
introns
how can LINEs copy themselves?
through mRNA, the gene is then duplicated and the LINE goes back into DNA
define LINEs
long interspersed nuclear elements
> most common
> can copy themselves
> retrotransposons
define SINEs
short interspersed nuclear elements
> cannot move on their own
what are retroviral-like elements?
possible remnants of viral infection
what is the function of transposons?
regulation of genes by interrupting coding or regulator sequences.
> move genes around and create genetic variability ("jumping genes")
what is chromatin and where is it found?
DNA + all protein
found in the nucleus of eukaryotic cells
what are histones?
DNA binding proteins that help compact and fold DNA into chromosomes
what are nucleosomes
DNA wound around histones
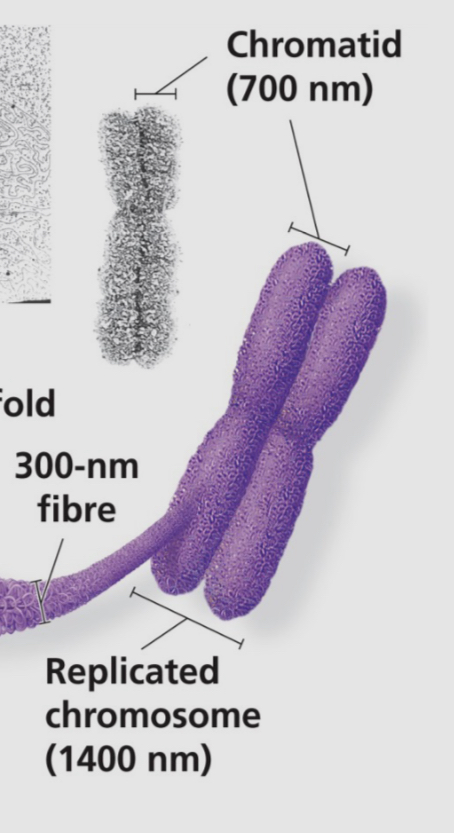
what is the process DNA goes through to compact itself into a chromosome?
DNA > wound around 8 histones into a nucleosome > 30nm fibre > loops into itself > scaffold > 300nm fibre > chromosome
what is a chromatid?
1 part of a chromosome pair (replicated chromosome)
what are the pros and cons about packing the DNA into chromosomes?
pros:
- it helps the DNA with organization and replication (makes it easier)
cons:
- dense packing makes it difficult to express genetic information.
what is heterochromatin and how is it formed?
condensed chromatin, which is looped and folded, bound by cohesin proteins. transcriptionally inactive.
how is chromatin present during interphase
as euchromatin and heterochromatin
what is the difference between euchromatin and heterochromatin?
euchromatin:
- loose structure
- transcriptionally active
heterochromatin:
- tight structure
- transcriptionally inactive (because the DNA is inaccessible)
what is constitutive heterochromatin?
regions of DNA that are permanently condensed and inactive.
what are the structural sections of constitutive heterochromatin?
telomeres and centromeres
centromeres
- constrictions of chromosomes
- where sister chromatids attach
- maintain cohesion between sister chromatids
- assembly site for kinetochores where MT attach during mitosis and meiosis
- defined by a highly repetitive centromere sequence (CEN) > varies between species
telomeres
- highly repetitive
- found at the tips of chromosomes
- protect the ends of chromosomes from degradation during replication
- highly conserved sequence of repeats (TTAGGG)
what are the 2 processes that "loosen" and "tighten" histone binding?
acetylation "loosens" chromatin to allow for transcription
> by histone acetyltransferase
> reversed by histone deacetylase
methylation "tightens" chromatin packing
> by histone methyltransferase
together, the 2 processes create and "histone code" which can modify transcription activity
what are chromatin remodelling proteins and what do they do?
they can alter the position of nucleosomes, and enable or disable transcription.
what are the 4 main mechanisms for epigenetics?
- chromatin remodelling
- histone modification
- DNA methylation
- non-coding RNAs (ncRNAs)
what is DNA methylation, how does it occur, what does it do?
> DNA methylation is adding a methyl group to DNA which inactivates it.
> done by DNA methyltransferases (DNMTs).
> it regulates gene expression by:
- recruiting proteins that repress genes
- preventing transcription factors from binding to DNA
what is the nucleolus?
the site for rRNA synthesis
> can fluctuate in size based on the need for ribosome synthesis (25% of the nucleus)
what are fibrils?
sites for active rRNA synthesis
what are granules
sites for packaging proteins for export from the nucleus
what is the perinuclear space?
the space in-between the inner and outer membrane of a cell.
(the space in the nuclear envelope)
> it is continuous with the lumen of the ER.
what is the lumen of the ER?
the space inside the ER.
> it is continuous with the perinuclear space of the nuclear envelope
what is Fluorescence In Situ Hybridization (FISH)?
a specialized microscopy technique for labelling DNA in cells
> DNA probes are developed based on base pair rulings
> the probes are synthesized with fluorescent tags
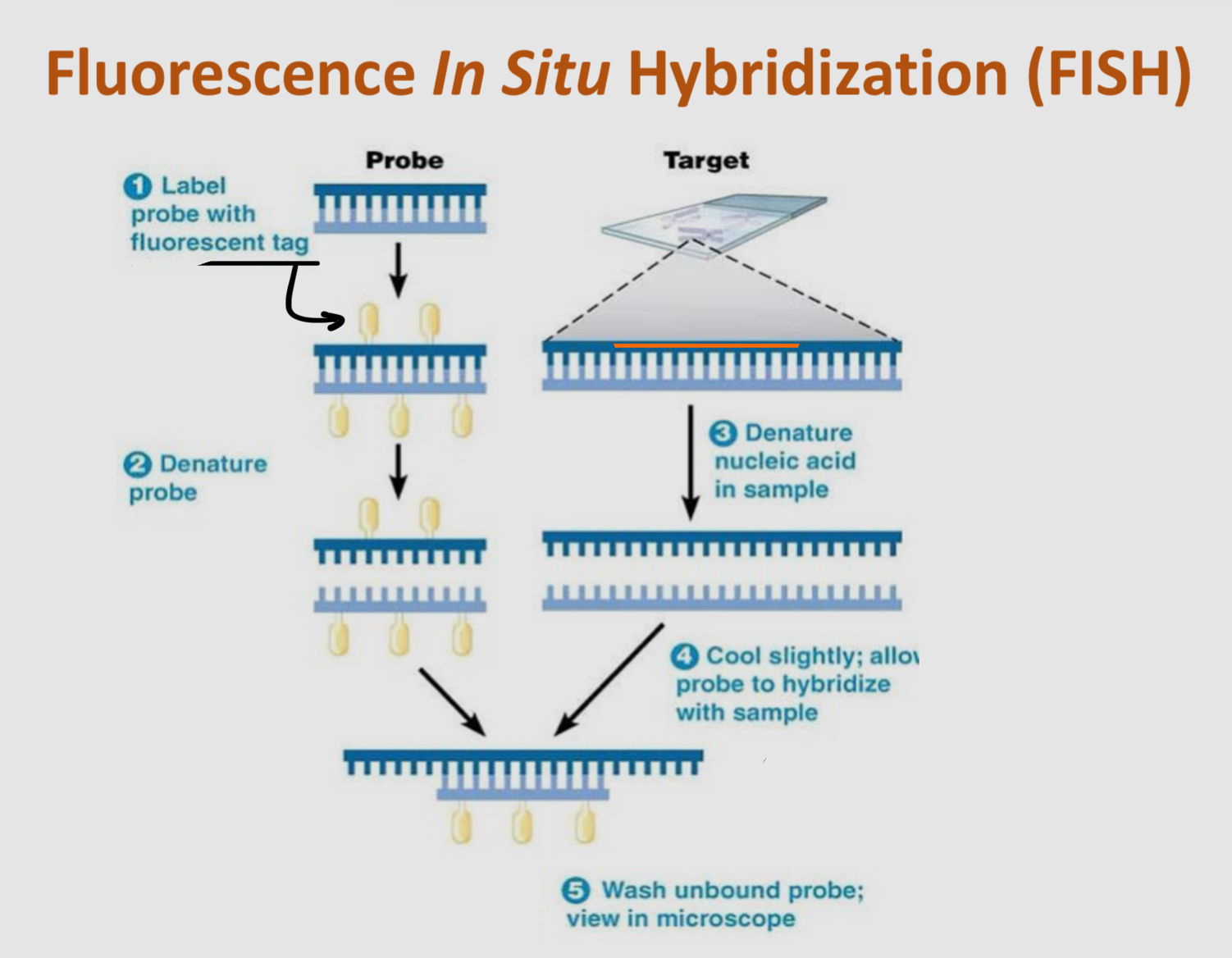
how can FISH be used to paint chromosomes in a cell?
by making different probes for each type of chromosome
what is the structure of the nuclear envelope?
- proteins to anchor the nucleus to the cytoskeleton.
- outer membrane is studded with ribosomes.
- nuclear pores on the outside of the envelope which make it continuous with the nucleoplasm.
what is the nucleoplasm?
a gel-like substance that fills the nucleus of a cell
what is the nuclear pore complex (NPC)?
a large protein complex that spans 2 bilayers of the nuclear envelope.
what is the central granule in context of the nuclear pore complex?
it is called the transporter
> plays a role in the active transport of macromolecules
what are FG nucleoporins in context of the nuclear pore complex?
they line the pore and are involved in transport
where does protein synthesis occur?
exclusively in the cytoplasm
where does transcription occur?
exclusively in the nucleus
what do nuclear pores do?
they drive import/export of macromolecules across the nuclear envelope.
what are nuclear localization signals (NLS)?
specific amino acid sequences that allow large proteins to be imported into the nucleus.
> they might bind to nucleoporins for uptake across the nuclear pore.
what is importin and what does it do?
an NLS binding protein that can act as a transporter to shuttle proteins through the NPC
what is RAN in the context of the NPC?
a GTPase exporter that can shuttle out importin once it has unloaded its cargo.
what is the RAN/Importin transport cycle?
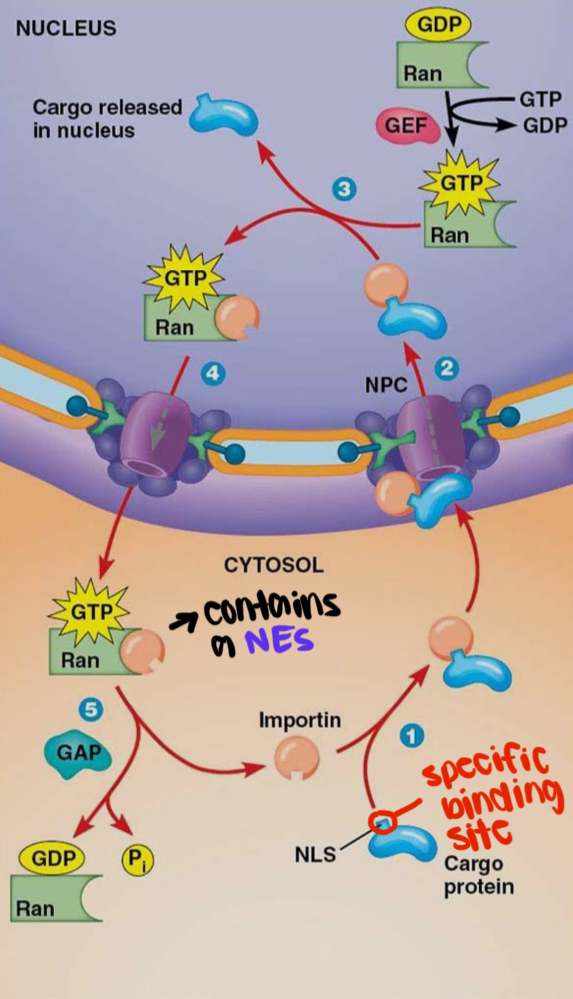
what is the function of the nuclear lamina?
maintains the shape of the nucleus and plays a role in transcription by organizing chromatin
what are laminopathies?
mutations of the lamina proteins
> associated with severe muscle wasting or premature aging (progeria)