nucleic acids
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
41 Terms
what is the molecule of inheritance?
Must be able to replicate accurately
Must contain in a stable form, the information about an organisms structure and function
Must be able to change in order to generate variation
inheritance
Gregor Mendle - Pea guy - hereditary
Proposed the existence of particulate unit factors for each hereditary trait but the molecule was still unknown
chromosomal theory of inheritance
Walther Flemming discovered thread-like structures called chromatin within the nucleus
Theodor boveri studied chromosomes replicating and dividing (in sea urchins)
Each cell has to have the correct number of chromosomes
Chromosomes hold the key to the inheritance of characteristics
chromatin
Chromatin=DNA+protein
DNA wraps around assosiated proteins called histones
Holecule wraps around itself to form dense chromosomes
what is molecule of inheritance?
1928 - Frederick Griffiths - showed presence of a 'transforming factor' that could pass on new characteristics
Took two strains of bacteria - R & S
R-benign
S-lethal
If he heat treated S strain - mouse lived - heating killed disease + characteristic
Mixed heated S and R - lethal phenotype is restored
Characteristics can be transferred from one cell to another
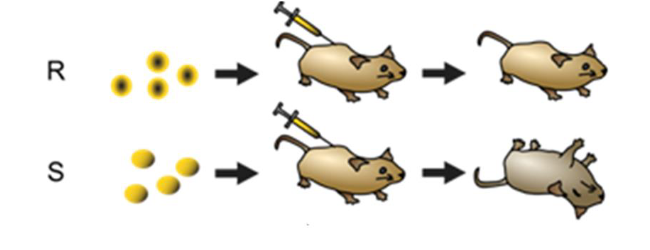
the lethal phenotype could be destroyed by heat
but the lethal phenotype can be transferred to a living presiously non-lethal bacterium
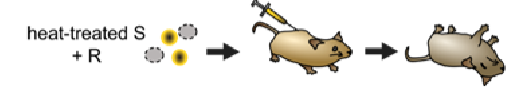
1944 - Avery, McCarty and McCleod - showed that the transforming factor was DNA
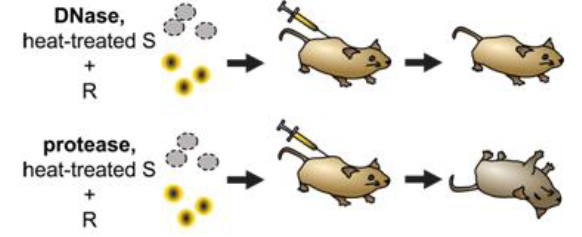
importance of nucleic acids
Key molecules in central dogma of biology
DNA is molecule of inheritance
Individual variation is encoded within our DNA - expressed through the conversion of RNA to protein
Individual genetic variation is key to forensic DNA profiling
Individual genetic variation is key to health - many diseases have underlying genetic cause
structure of DNA - Watson and Crick and Rosalin Franklin
Watson & Crick - proposed DNA structure and double helix
Pentose sugar
Nitrogenous bases
Phosphates
Maurice Wilkins and Rosalin Franklin studied DNA by x-Ray diffraction and concluded that it was helical
Chargaff’s Rule
1950's Chargaff carried out base composition studies on DNA
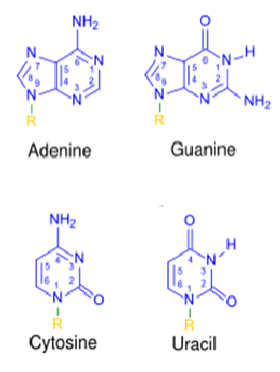
4 nitrogenous bases
50% of bases were purines and 50% pyrimidines
Amount of adenine was equal to thymine and cytosine equal to guanine
DNA structure
Pentose sugar = deoxyribose
Four types of nitrogenous bases, join to the primary carbon in deoxyribose
Purines - adenine and guanine - 9 carbon, double ringed structures
Pyrimidines - thymine and cytosine - 6 carbon, single ringed structures
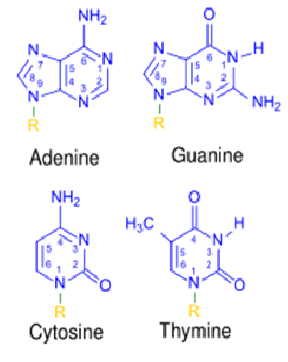
bases attach via covalent (glycosidic) bond to sugar molecule - at C1 position and nitrogen of base
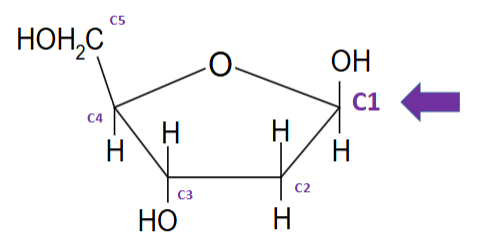
nucleoside (base and sugar)
Deoxyribose and nitrogenous base - covalently bonded
Deoxyadenosine
Deoxyguanosine
Deoxythymidine
Deoxycytidine
nucleotide (base, sugar and phosphate)
Phosphates joined to hydroxyl on C5 of sugar group
Nucleotides have either one, two or three phosphate attached
Deoxynucleotide triphosphates (dNTPs) are the building blocks of the DNA molecule
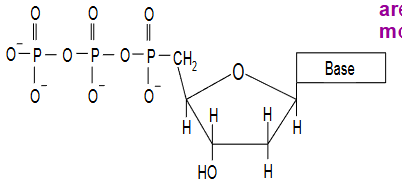
joining nucleotides together
3' hydroxyl and 5' phosphate react to form ester bond releasing 2 phosphate groups
Leads to formation of phosphodiester bond
At end of each DNA chain the 5' sugar has free phosphate group whereas 3' sugar has free hydroxyl group
DNA strands have direction
Sugar phosphate backbone
DNA as a double stranded molecule
Antiparallel strand held together by base pair bonding
Bases lie perpendicular to axis, bound together by hydrogen bonds
Two hydrogen bonds = adenine - thymine
Three hydrogen bonds = guanine - cytosine
Watson and Crick Model
Two polynucleotide chains wound around each other in a right handed double helix
Sugar-phosphate backbone is on outside of helix
Bases point to central axis
Chains run antiparallel
Amount C=G
Amount T=A
Amount purine (A+G) = amount pyrimidine (T+C)
how does DNA replicate accurately?
If one chain is given we can fill in the opposite because of base pairing
DNA Replication - proposed models
Conservative model - original DNA strands separate, copied rejoin respectively
Semi-conservative model - one of each in new
Dispersive model - broken into sections and stitched back together
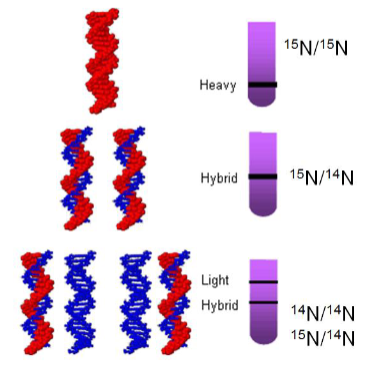
Meselson and Stahls experiment - semi conservative replication
using heavy and light nitrogen
If the conservative hypothesis was correct no hybrid forms would be detected
• If the dispersive hypothesis was correct DNA of intermediate density would be detected
• DNA Replication is semi-conservative
DNA Replication
Occurs in nucleus
Carried out by DNA Polymerase iii
One strand used as template to synthesis a new strand based on Chargaff's base pairing rules
New strand synthesised in 5' to 3' direction
enzymes in DNA replications
Helicase - uncoils separates
DNA polymerase - inserts and forms bonds of nucleotides
Primase - adds RNA primers
Ligase - bonds fragments
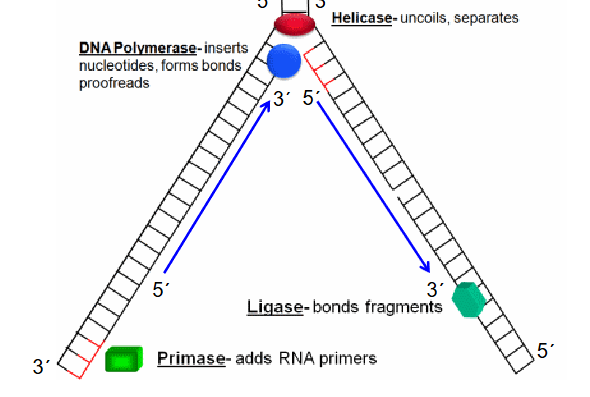
DNA replication
DNA polymerase iii synthesised new DNA strand from multiple start points - forms bubbles in DNA
Replication rate is 500-5000 bp/min in eukaryotic
Error rate = less than 1 in a billion bases
function of DNA
Some sections of DNA contain nucleotide sequences that determines order of amino acids in a protein (genes)
DNA sequence converted by transcription to an intermediate molecule - RNA
structure of RNA
Single stranded
Ribose instead of deoxyribose (2' hydroxyl group)
Uracil instead of thymine
RNA
Sugar phosphate backbone joined 5'-3'
Bases attached at the primary carbon
Synthesised from DNA in transcription
transcription
Occurs in nucleus
H bonds break and DNA unwinds
RNA polymerase read 5'-3' joining ribonucleotides according to their complementary bases
Single stranded RNA molecule released
There are 3 types of RNA
ribosomal RNA (rRNA)
Most abundant type of RNA
Combines with proteins to form ribosomes found in cytoplasm and on the RER
Ribosomes hold mRNA molecules in place for translation
Peptidyl transferase activity of rRNA catalyses formation of peptide bonds
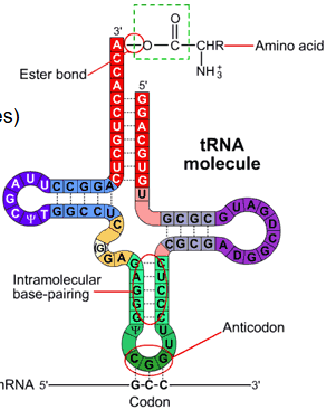
transfer RNA (tRNA)
Smallest form of RNA (73-95 nucleotides)
Forms clover leaf structure
Carries specific amino acids (bound to 3' end) to the ribosome
tRNA 'reads' genetic code
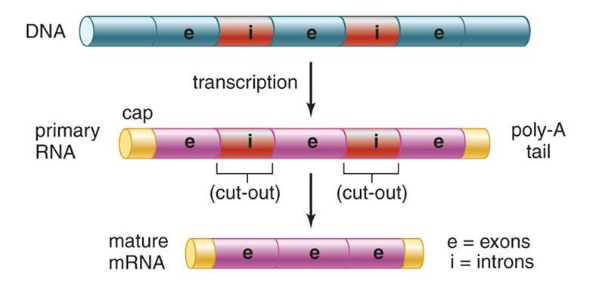
messenger RNA (mRNA)
contains instructions for making proteins, following transcription mRNA molecules are processed, ready for translation
DNA transcribed into RNA
DNA is double stranded:
One strand carries code for RNA - known as coding or sense strand
Opposite (non-coding) strand acts as template for RNA polymerase
Resulting RNA has same sequence as coding strand (but w U instead of T)
how is information encoded?
Crick and Brenner (1961) induced mutations into Bacteriophage DNA = Three nucleotides code for one amino-acids
cracking the code
In 1960's Marshall Nirenberg and others cracked the code
Synthetic RNA using all 64 possible triplet codes identified which triplets coded for which amino acids
The code is degenerate (one amino acid may be coded by more than one triplet of bases

mRNA is translated into protein
Requires coordination of all three types of RNA:
mRNA = the sequence to be translated
tRNA = carrier molecules that bring amino acids together
rRNA = structural and enzymatic component of ribosomes, rRNA has Peptidyl transferase activity (the only non-protein enzyme) which catalyses the formation of peptide bonds between amino acids
The order of bases in mRNA molecule (the genetic code) determines the order of amino acids in a protein (its primary structure)
translating the genetic code
Several signals involved
Initiation codon - usually AUG
Codons - triplet order of bases in mRNA determines order of amino acids in protein
Stop codons - UAA, UGA, UAG
The tRNA anticodon binds to its complimentary triplet (codon) in the mRNA inserting the correct amino acid into the protein
translation
mRNA held on the ribosome allowing tRNAs to bring amino acids together and create peptide bonds between amino acids
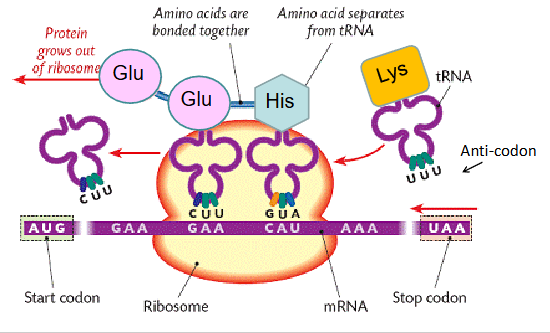
the genetic code
64 possible triplet combinations
Redundancy: >1 triplet codes for the same amino acid
3rd base 'wobble': the first two bases of the codon usually determine the amino acid
the double helix
Replication - strict base pairing gives a simple mechanism for making an exact copy
Storage of information - the order of bases form the triplet genetic code (codon) carrying the instructions to make a specific protein
Stability - strong phosphodiester bonds keep the sequence intact, weak h bonds allow message to be read
Variation - in DNA sequence results in different protein sequences (alleles)
nucleic acids in forensic science
All cells will contains RNA (but its presence may be transient) - limited application in forensic biology
All cells (except red blood cells) contain a nucleus therefore could provide DNA for analysis such as:
Blood
Semen
Saliva
Bone
Teeth
Hair
Tissue
DNA in forensic science
DNA sequence (the order of nucleotides) vary from person to person
There are enough differences to allow us to identify an individual
Link a suspect to bio evidence left at a scene
Identify missing persons through matching to personal items
Variants are inherited from parent to offspring
Paternity testing
Kinship testing - missing persons through relatives