AP Biology Unit 7: Gene Expression & Regulation
1/83
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
84 Terms
Gene
A physical and functional unit of heredity, a sequence of DNA that codes for a protein
The Central Dogma
Genetic information flows from DNA → RNA → Protein through transcription and translation.
Nucleus
Holds the cell’s genetic information in DNA. mRNA leaves the nucleus through its nuclear pores.
Nucleolus
Builds ribosome subunits from rRNA & proteins which then exit through nuclear pores and into the cytoplasm and combine to form functional ribosomes.
Ribosomes

There are two types of ribosome (free & bound) and they make proteins.
Free Ribosomes
Are suspended in the cytosol and synthesizes proteins that function within the cytosol.
Bound Ribosomes
Are attached to the endoplasmic reticulum and synthesizes proteins for export to membranes.
RNA
Made up of RNA nucleotides. The sequence of the RNA bases and the structure of the RNA molecule determines its function.
mRNA (messenger RNA)
Carries information from DNA to the ribosome.
tRNA (transfer RNA)
Carries amino acids to the ribosome.
rRNA (ribosomal RNA)
Building blocks of ribosomes
microRNA
Small RNA molecules that bind to other RNA molecules to degrade them.
Transcription Overview
Then nucleotide in the DNA is used to make a complementary sequence in mRNA using RNA nucleotides through Initiation → Elongation → Termination.
RNA Polymerase
Uses a single template strand of DNA to make mRNA using free RNA nucleotides after it unwinds the helix. Works in the 5’ to 3’ direction.
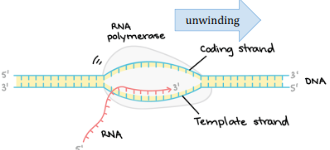
Coding Strand
The side of DNA that is not used in synthesizing the mRNA.
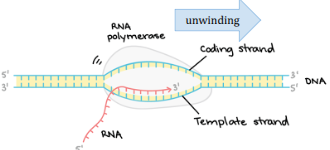
Template Strand
The DNA strand that is used to transcribe the mRNA.
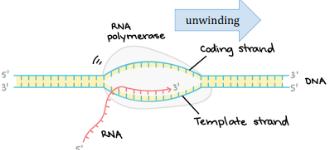
Promoter
In prokaryotes, the RNA polymerase directly binds to this. It is located on the template strand and provides a starting point for reading the beginning of a gene. It also ensures that the DNA is read 3’ → 5’ and mRNA is built 5’ → 3’. Also contains the TATA box.
DNA is read from:
3’ → 5’
mRNA is built from:
5’ → 3’
Transcription Factors
In eukaryotes, they bind directly to the TATA box of the promoter region first, allowing the RNA polymerase to bind on top. They are a suite of proteins that can turn on or off transcription.
TATA Box
A recognition site for transcription factors, located on the promoter region
Termination Sequence
Once the RNA polymerase reaches this, it detaches and various proteins help free the newly transcribed mRNA.
5’ Cap
It’s (a modified guanine nucleotide) is added to the first nucleotide during transcription. It protects the transcript from breaking down and helps the ribosome attach to the mRNA and start reading it.

Poly-A Tail
It is made up of many repeating adenine nucleotides and helps making the transcript more stable while also helping it get exported to the cytosol.

Introns
Stays in the nucleus, and does not code for proteins. Splicesomes “cut” this out.
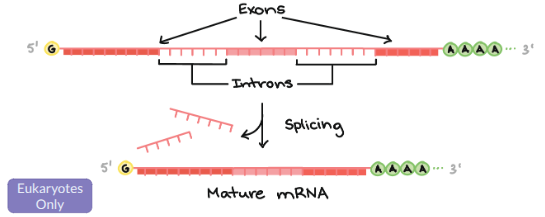
Exons
Exits in the nucleus to go to the ribosomes and do code for proteins.
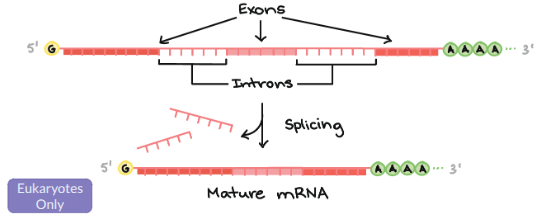
Alternative Splicing
Different mRNA versions result from combining different exons.
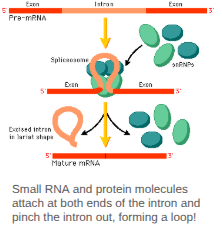
Spliceosome (snRNPs)
An enzyme complex made up of protein and small RNAs. It does the cutting and gluing back together of the pre-mRNA to make it mature.
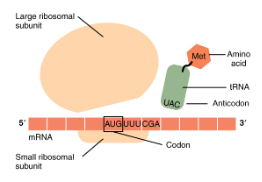
Translation Overview
The mRNA sequence is organized in codons, which are decoded by pairing with complementary anticodons on tRNA molecules to assemble amino acids into a polypeptide chain, forming a protein.
Initiation → Elongation → Termination
Translation - Initiation
Small ribosomal subunit binds to mRNA at the start codon (AUG)
An initiator tRNA is added
The large ribosomal subunit attaches
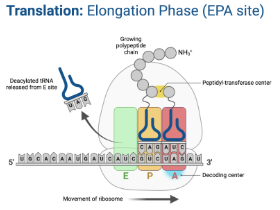
Translation - Elongation
Ribosome moves down the mRNA in the 5’ → 3’ direction
For each codon, a tRNA with a corresponding anticodon brings an amino acid to the ribosome
Amino acids are added to the preceding one by a peptide bond using peptidly transferase through dehydration synthesis
Goes through the APE sites until the ribosome reaches a stop codon in the mRNA
A Site (Aminoacyl-tRNA site)
Holds tRNA carrying next amino acid to be added to the chain
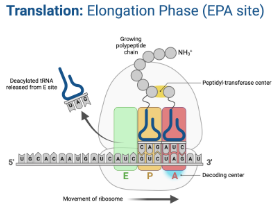
P Site (Peptidyl-tRNA site)
Location at which the amino acid is transferred from its trNA to the growing polypeptide chain.
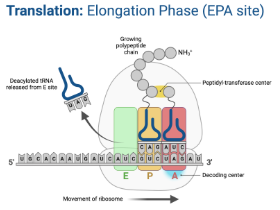
E Site (Exit site)
Empty tRNA leaves the ribosome
Start Codon
AUG
Stop Codons
UGA, UAA, UAG
Translation - Termination
When the ribosome reaches the stop codon, a protein called the release factor, causes the polypeptide chain to separate from the ribosome.
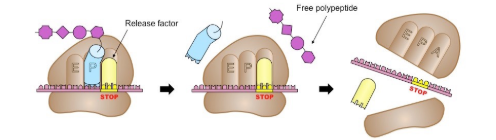
Post Translation
The polypeptide chain folds into a specific, 3D shape based on the amino acid sequence, forming its secondary, tertiary, or quaternary structure. Special helper proteins called chaperoning help this process.
Cells use “targeting signals” to route proteins to the right place.
Post Translation - Proteins going OUTSIDE of the cell (or an organelle)
Made on bound ribosomes on the rough ER.
They are sent to the rough ER where they are folded (and sometimes modified).
Then, they are sent to the golgi and undergo further modification. Vesicles transport these proteins.
Post Translation - Proteins STAYING inside the cell (cytoplasm or organelles)
Made on free ribosomes in the cytoplasm.
They are folded into their functional shape in the cytoplasm
Remain in the cytoplasm or are sent to organelles using specific targeting signals
*Do NOT go through the rough ER or golgi
Protein Synthesis - Prokaryotes
Transcription occurs in the cytoplasm, no mRNA editing, & transcription + translation occur simultaneously.
Protein Synthesis - Eukaryotes
Transcription occurs in the nucleus, mRNA is edited prior to translation, translation occurs after transcription is completed.
Mutations
A change in the DNA sequence that may be caused by factors such as mutagens, errors in DNA replication, or errors in mitosis or meiosis.
Mutagens
External factors such as radiation and reactive chemcials.
Point Mutations
A base is changed, but the number of bases stays the same. There are three types of point mutations: silent, missense, and nonsense.
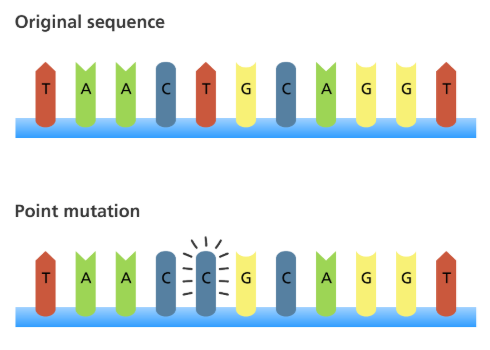
Silent Point Mutations
Still codes for the same amino acid.
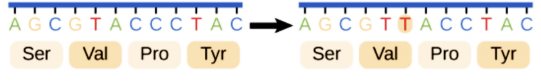
Missense
Codes for a different amino acid, which changes its property, and ultimately, the protein’s shape and function.
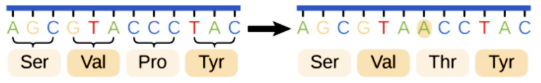
Nonsense
Codes for a stop codon early, so the remainder of the codons will not be read.
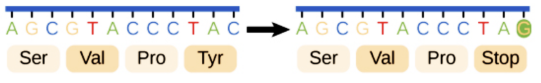
Frameshift Mutations
The number of bases changes due to the addition or removal of one or more nucleotides in the DNA sequence. This causes the reading frame to shift, thereby changing all the codons after the mutation.
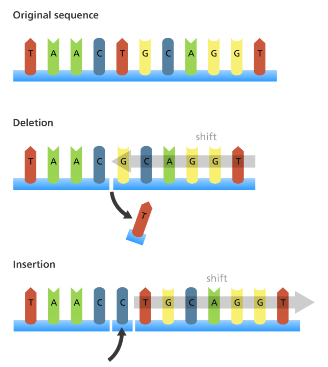
Operon
A group of genes of related function, used to regulate gene expression by controlling related genes at the same time. The transcribed mRNA contains the sequence for all the genes in the operon.
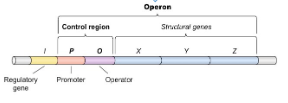
Lac Operon
Controls genes needed to break down lactose. It is inducible, meaning it is turned on when lactose is present.
Ara Operon
Regulates genes needed to metabolize arabinose sugar. It can be both activated and repressed depending on environmental conditions.
Regulatory Gene
Produces transcription factors (proteins) that regulates if the gene is transcribed or not.
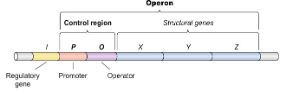
Operator
On/off switch where regulatory proteins bind.
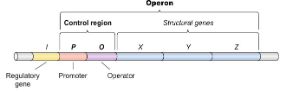
Promoter
Where RNA polymerase attaches.
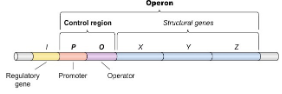
Negative Control
Repressor blocks transcription and RNA polymerase (transcription OFF) → gene is off unless repressor is removed.
→ Inducible Operons & Repressible Operons
Positive Control
Activator helps RNA polymerase bind (transcription ON)→ gene is on or transcribed more.
→ Activators
Inducible Operon
Off → On
Negative Control
Produces enzymes only when nutrients are available. They avoid making proteins that have nothing to do and are usually used in catabolic pathways.
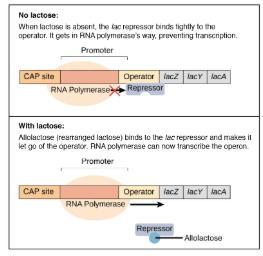
Repressible Operon
On → Off
Negative Control
When the end product is present, transcription is repressed (turned off) to allocate resources to other uses. Usually functions in anabolic pathways.
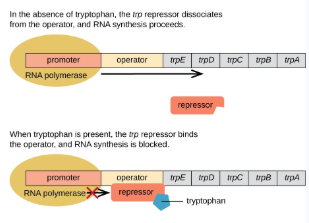
Activators
Positive Control
Activators are proteins that bind near the promoter. They increase transcription by helping RNA polymerase bind to the enhancer (near the promoter)
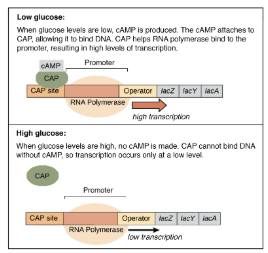
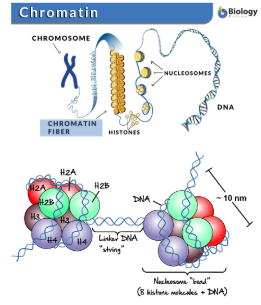
Chromatin
The packaged form of DNA inside the nucleus
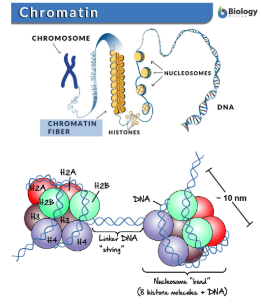
Chromatin Structure
Made up of coiled and folded nucleosomes.
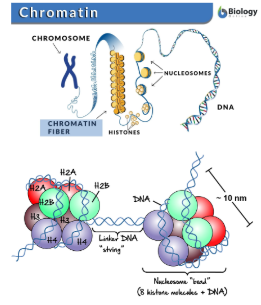
Nucleosomes
Strucutures formed by DNA wrapping around proteins called histones.
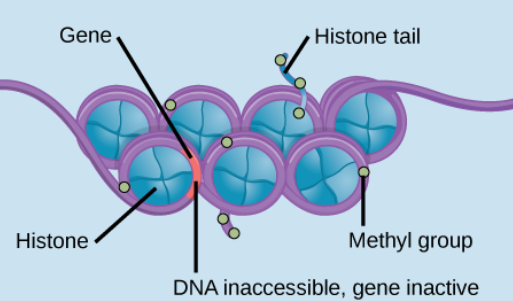
Heterochromatin
Tightly packed chromatin, making genes less likely to be expressed.
This is because DNA is less accessible, and the transcription machinery cannot easily bind to it. The methyl groups are present, and transcription cannot occur.
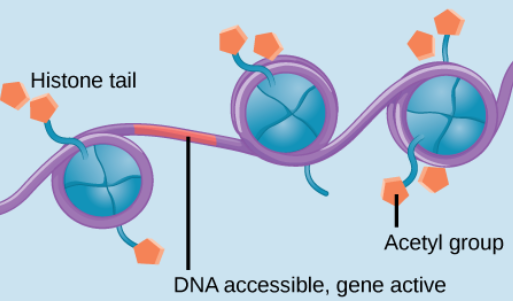
Euchromatin
Loosely packed chromatin, making genes more likely to be expressed.
This is because DNA is more accessible, and the transcription factors + RNA polymerase can bind. The acetyl groups are present, and transcription can occur.
DNA Methylation
Genes OFF
Methyl groups (-CH3) can attach to DNA bases, preventing transcription by blocking transcription factor binding and recruiting silencing proteins.
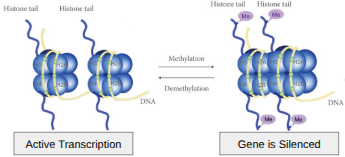
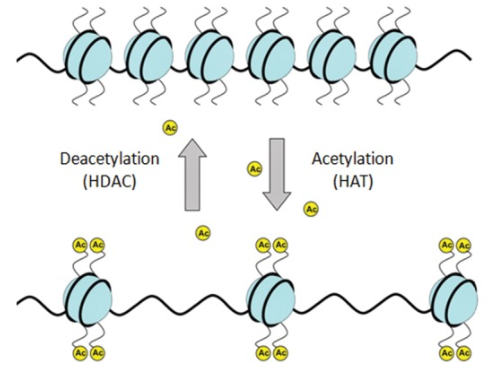
Histone Acetylation
Genes ON
Acetyl groups (-COCH3) are added to histone, preventing them from binding to the DNA as tightly, making room for proteins to bind for transcription.
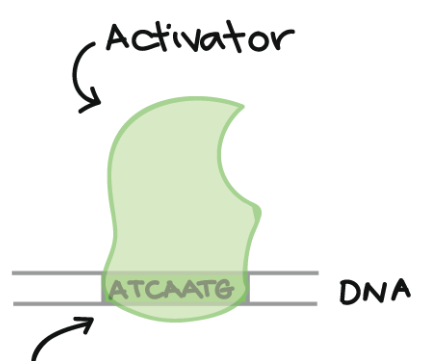
Enhancer
A sequence of DNA nucleotides that is the binding site for an activator
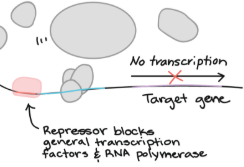
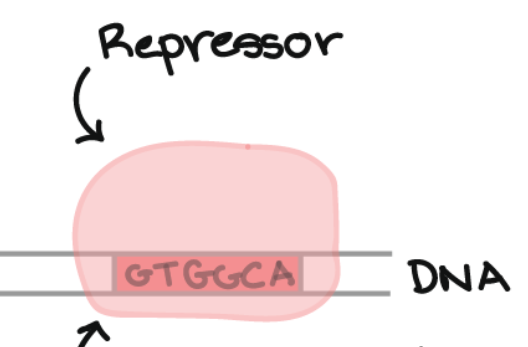
Repressors
Transcription factors that bind to the silencer. This prevents RNA polymerase from binding to the DNA, decreasing or completely stopping the rate of transcription.
Silencer
A sequence of DNA nucleotides that is the bonding site for the repressor.
Small Interfering RNAs (siRNA)
Short segments of RNA about 21-28 bases that bind to mRNA to create sections of double-stranded mRNA. This tags mRNA for degradation and turns off the gene so no protein can be produced.
Controlling how long mRNA lasts regulates how much of a protein is made.
Blocking Translation Initiation
Occurs when a ribosome does not form or mRNA cannot attach (translation cannot occur). Regulatory proteins can attach to the 5’ end of mRNA to prevent attachment of ribosomal subunits and initiator tRNA.
Ubiquitin
A molecule that triggers proteasomes to break down the protein if it is not needed.
*Not common
Gene Regulation in Prokaryotes
Regulates a cluster of genes (operon) and regulation only occurs in the cytoplasm and only at the transcriptional level. There are no histones and introns.
Gene Regulation in Eukaryotes
Regulates individual genes and regulation can occur in the nucleus and cytoplasm and at many levels. DNA is wrapped around histons and introns are involved in alternative splicing.
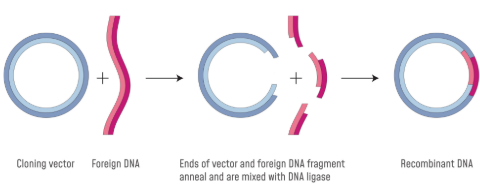
Restriction Enzymes
Cuts DNA into segments at specific sequences.
Can be used to cut a specific gene out of a DNA strand and simultaneously cut a plasmid vector, allowing DNA ligase to join them together, forming recombinant DNA.
Polymerase Chain Reaction (PCR)
The process of making many DNA copies for analysis by amplifying specific DNA sequences
Steps: Denaturation → Annealing → Extension
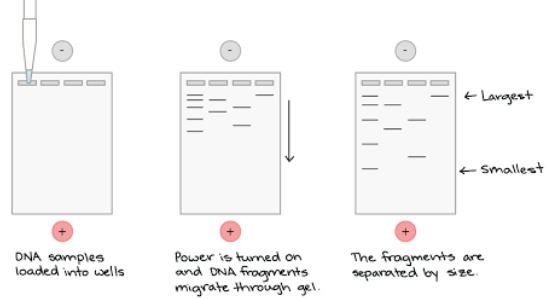
Gel Electrophoresis
When electricity is used to separate DNA fragments of different sizes to be used in DNA fingerprinting, diagnosing genetic diseases, etc.
DNA is negatively charged, so it moves towards the positive end of the electrophoresis chamber.
DNA Sequencing
When fluorescent markers are added to PCR, and the fragments are then separated by size. The order of colors from the markers are recorded, each color representing a different base.
This is used in disease/medical diagnosis, newborn screenings, paternity testing, etc
Virus Replication
Viruses cannot replicate on their own, instead they hijack their host’s cellular machinery to make more DNA & RNA.
Prophage
A viral DNA joined with the host cell DNA.
Lytic Cycle
When new viruses are produced.
Retrovirus
A virus that uses RNA as their genetic material, so they use RNA to make DNA with the help of the enzyme reverse transcriptase. Then the DNA becomes integrated with the host genome to be used to make more viruses.
Can be used as vectors in gene therapy to deliver functional genes into a patient’s DNA.
Gene Therapy
The technique of using genetic material as a drug to treat or prevent disease by replacing, silencing, or correcting faulty genes.
Can be used to treat conditions such as cystic fibrosis, sickle cell anemia, etc.