1.5 Nucleic acids
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
29 Terms
What is each nucleotide made of
A pentose sugar, a phosphate group and an organic nitrogenous base
what bond is formed between nuclotides
phosphodiester bonds
draw and label a typical nucleotide
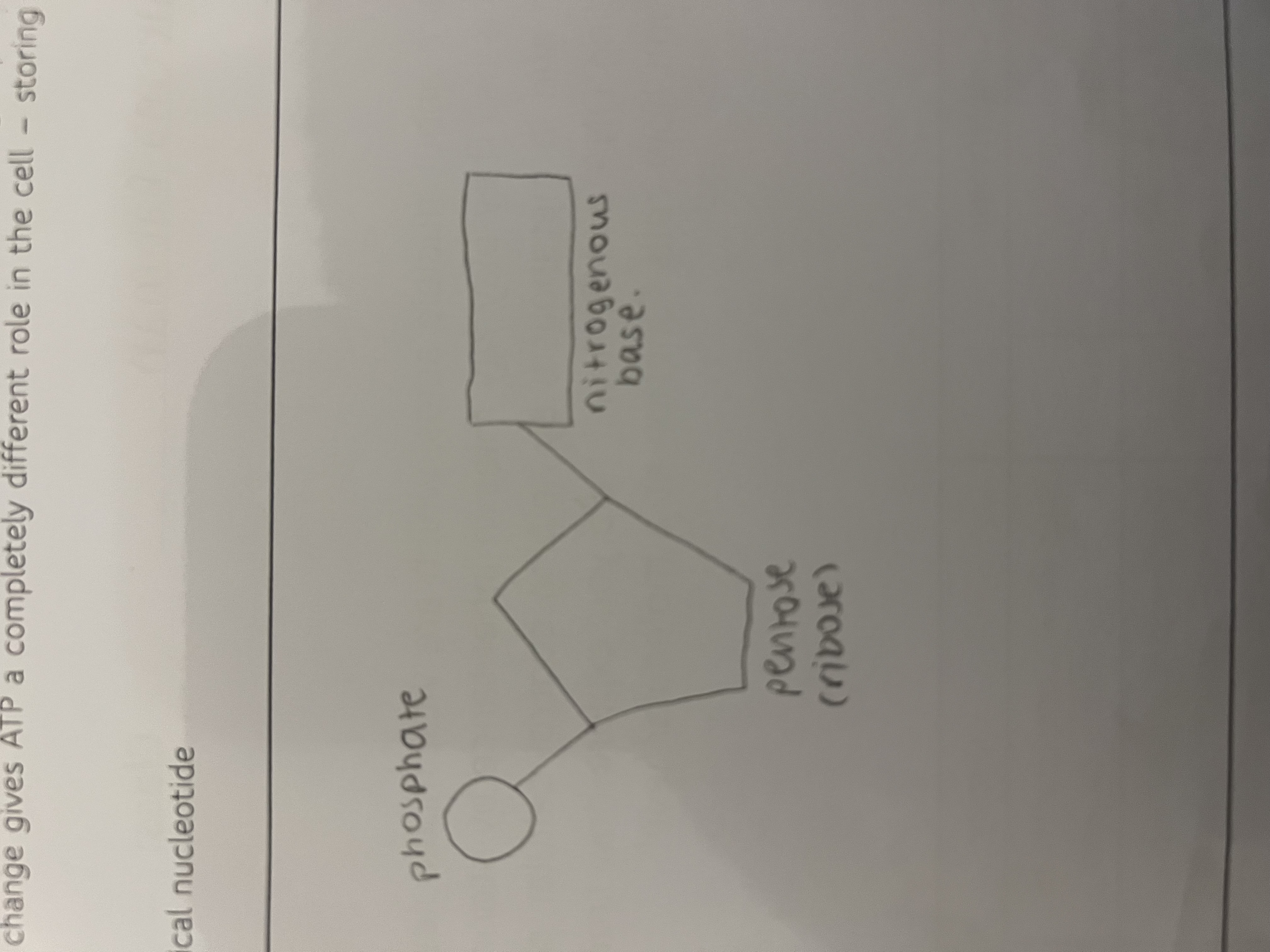
what are the two main categories of nirtogenous base in nucleic acids and give examples of both and defintion
purines: guanine and adenine, they have a double ring in their structure
pyrimidines: thymine, cystosine and uracil, they have a single ring
how many bonds between adenine and thymine, and also cystosine and guanine
A-T two hydrogen bonds
C-G three hydrogen bonds
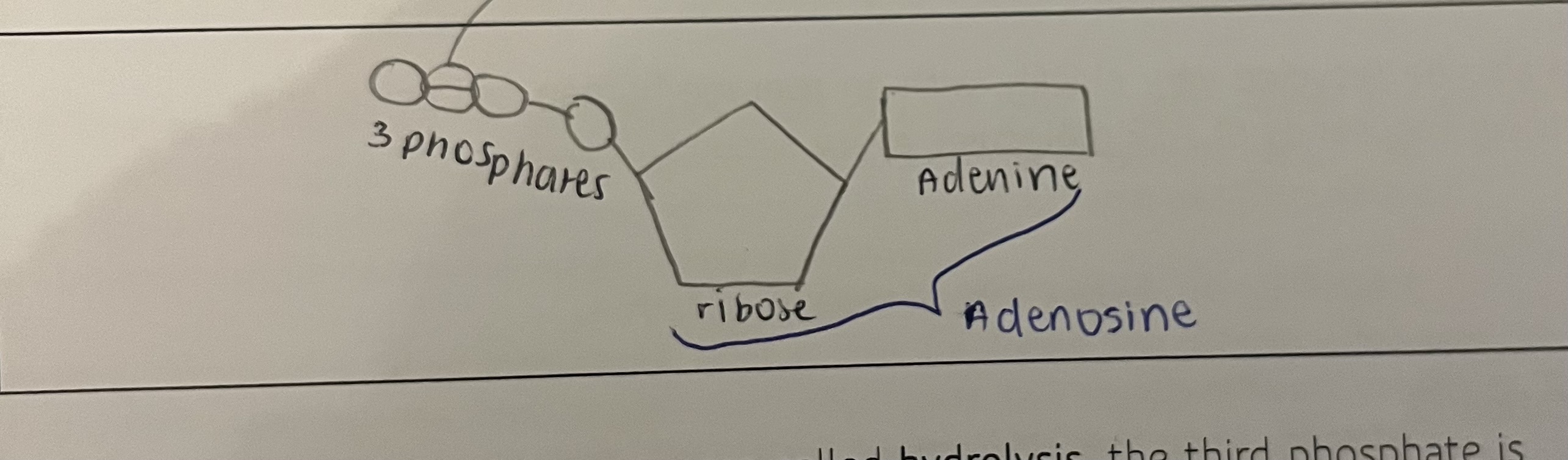
what is ATP made of and draw a pic
adenosine triphosphate
-adenine (base), ribose (pentose sugar) and three phosphate groups
what happens when ATP is broken down
broken down using hydrolysis
-third phosphate is released, forming ADP (adenosine diphosphate) and inorganic phosphate (Pi)
-releases 30.6 kj mole-1 of energy
draw the reversible reaction of ATP being broken down and reformed
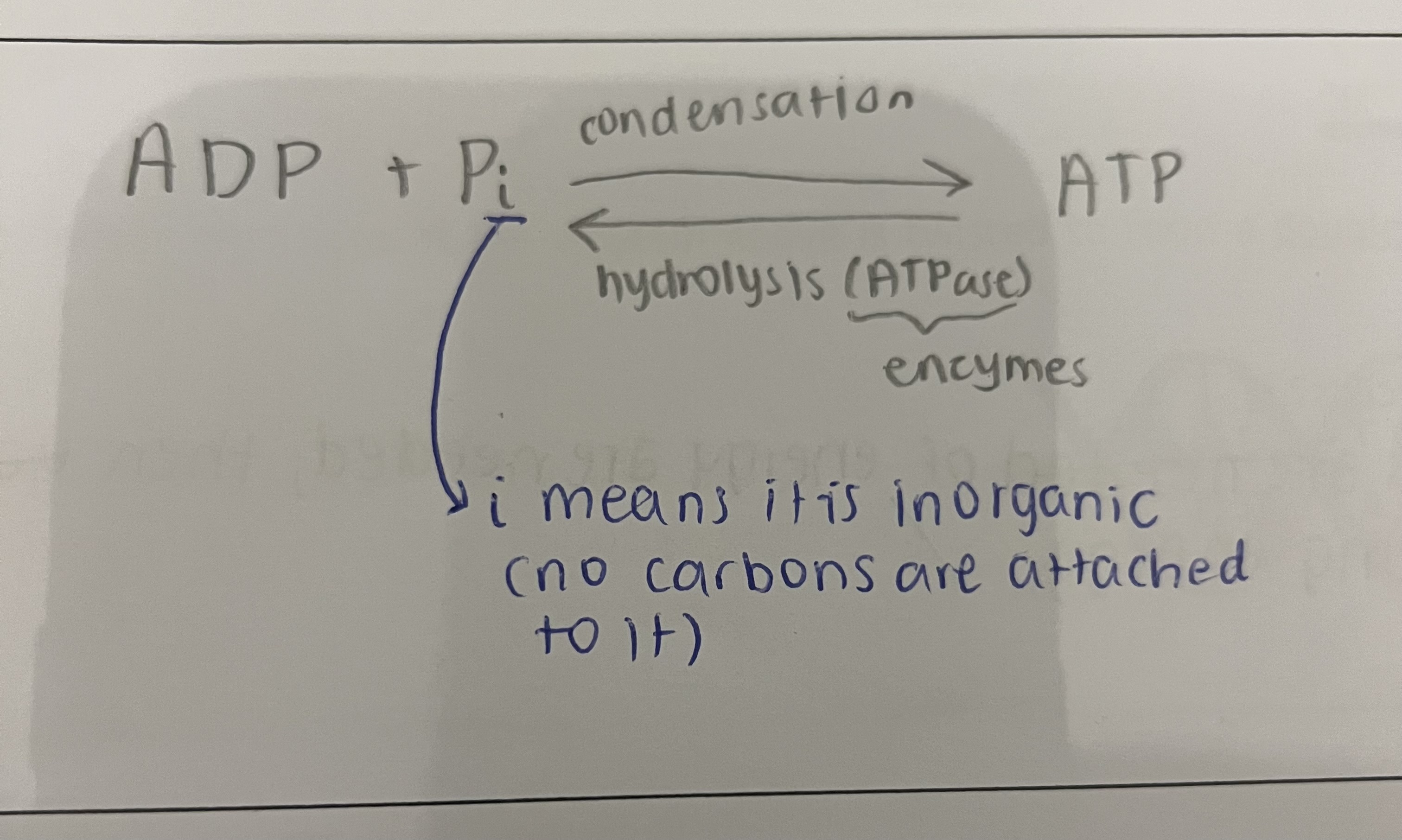
why is ATP called the universal energy currency
used by all living organisms, from bacteria to humans
why is only a small amount of energy released from ATP
-less wasteful of energy
(if it was large amounts and small amounts were needed then a lot would be wasted)
give features of DNA and what it stands for
deoxyribonucleic acid
-double helix
-two long polynucleotide strands that run in opposite direction (antiparallel)
→one strand runs from 5’ to 3’ and the other runs 3’ to 5’
-sugar is called deoxyribose meaning there is less oxygen on the ribose on carbon two
-hydrogen bonds between bases
give features of RNA and what it stands for
ribonucleic acid
-pentose sugar is ribose →one more oxygen
-uracil instead of thymine
-shorter than DNA
-single-stranded polynucleotide
what are the three types of RNA
-ribosomal RNA (rRNA)
-messenger RNA (mRNA)
-transfer RNA (tRNA)
what is rRNA function
ribosomal RNA
-forms ribosomes with the addition of protein
-made in thhe nucleolus
-highly folded to make a globular structure
what is mRNA and its function
messenger RNA
-single stranded and made in nucleus
-mRNA carries the code from DNA in the nucleus out of the nuclear into the cytoplasm and then attaches to the ribosome
what is tRNA and function
transfer RNA
-single stranded polynucleotide twisted into clover-leaf shape
-transfers correct amino acid to the growing polypeptide during translation in ribosome
-has an anticodon on one end and an amino acid on the other
what do every three bases code for and whats it called
codon
-codes for amino acid
what are the similarities between ATP and DNA+RNA
-both have adenine
-both have pentose sugar
-both have phosphate
-protein synthesis
what are the differences between ATP and DNA+RNA
-different structures
-DNA and RNA have more phosphates
what is a gene
stretches of DNA that code for polypeptides or functional RNA
what is an exon and intron
exon: coding sequences
intron: non-coding sequences
in eukaryotes and prokaryotic cells, is the genes continuous or discontinuous for both
eukaryotes: usually discontinuous (interrupted by introns)
prokaryotic cells: usually continuous (lack introns)
what is genetic code and what are they key features of it and explain
genetic code is the set of rules by which information encoded in the DNA is translated into proteins
-linear: DNA is arranged in straight lines
-triplet: bases are found in triplets called codons, 1 codon is the code for one amino acid
-non-overlapping: the codons are kept seperate
-degenerate: more than one codon can code for the same amino acid (64 combination of bases but only 20 amino acids)
-unambiguous: each codon always codes for the same amino acid
-universal: all living organisms have the same codons and amino acids
what are the two main functions of DNA
-protein synthesis
-replication
what are the three proposed models of DNA replication and what do each mean
1.Conservative replication
-original remain intact and a completely new model is made from scratch
2.Dispersive replication
-DNA strands are a mix of old and new segments so each strand contains scattered pieces of original and new DNA
3.Semi-conservative replication (supported by evidence)
-each new DNA molecule contains one original strand and one newly synthesised strand
Describe the process of semi-conservative replication
1.original DNA (double helix) held together by hydrogen bonds
2.the enzyme DNA helicase breaks apart the hydrogen bonds between the bases which makes DNA split into two separate strands
3.the free nucleotides in the nucleoplasm join to their complementary base via hydrogen bonds
4.the hydrogen bond alone is not strong enough so the enzyme DNA polymerase joins the new nucleotides together with phosphodiester bonds to create a new sugar-phosphate backbone
this creates two identical DNA molecules with one strand from each DNA being from the orginal
describe the experiment to prove semi-conservative replication and explain the results and why it disproved the other methods
1.they grew E. coli bacteria in a medium containing heavy nitrogen (N15) which made DNA denser than normal
2.then, they moved the bacteria to a medium with light nitrogen (N14) and allowed them to replicate
RESULTS
3.after one round of replication, the DNA was of intermediate density (this ruled out conservative since by now it would have shown one light and one heavy)
4.after two rounds they saw two bands, one light and one intermediate (rules out dispersive which would have shown only intermediate bands even after multiple rounds)
describe the first step of protein synthesis
Transcription: DNA to mRNA
takes place in nucleus and involves copying a gene from DNA into mRNA
1.DNA helicase breaks hydrogen bond between base pairs, unwinding the double helix and exposing the template strand
2.RNA polymerase binds to the template strand and brings free nucleotides (U instead of T) following the rules of complementary base pairings and joins them with phophodiester bonds
RNA polymerase continues to the end of the template strand (til it reaches a stop codon) which leaves the molecule called pre-mRNA
post-transcriptional modification removes the introns, producing functional mRNA that can leave the nucleus
describe the second step of protein synthesis
Translation: mRNA to protein
translations occur in the cytoplasm at the ribosome and involves converting the mRNA code into a sequence of amino acids to form a polypeptide
1.initiation: mRNA leaves nuclear pore, goes to ribosome, ribosome attahes to start codon at one end of the mRNA molecule. The first tRNA with anticodon matches to the codon of the mRNA and join with hydrogen bonds
2.elongation: two tRNA, the two amino acids are close enough for peptide bond. The first tRNA leaves the ribsome leaving the amino acid and the next tRNA binds
3.termination: process continues until a stop codon is reached
-stop codon is a codon which does not have a complementary anti-codon so no amino acid is brought.
4.polypeptide goes to golgi body