Control of gene expression
1/97
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
98 Terms
What is meant by ESEs/ISEs/ESSs/ISSs?
Exonic or intronic
Splicing
Silencers or enhancers
What is the splicing code?
Regulate excision of introns and joining of exons during splicing
Facilitated by spliceosome
Influenced by the combination of splicing and repressors, where sequence-specific RNA binding proteins can bind to them to regulate recruitment of the spliceosome
What is a constitutive exon?
Always included in the mature mRNA
What are SR proteins?
Activate splice sites
RNA-binding domain and protein-interaction domain
RNA-binding domain binds to splice site
Protein interaction domain recruits U2AF to the polypyrimidine sequence close to the 3’ splice site
U2AF can recruit U2 for spliceosome assembly sequence
What are hnRNPs?
Splicing repressors
RNA binding domain and protein-interaction domain
Bind to the 3’ polypyrimidine tract to prevent U2AF and U2 recruitment, preventing spliceosome assembly
What are the three key factors that determine which splice site is used?
How similar to the consensus sequence the splice sites are (‘strength’)
Whether or not there are nearby enhancer/repressor elements, and any SR proteins / hnRNPs bound to them
Any RNA secondary structures that are masking splice sites
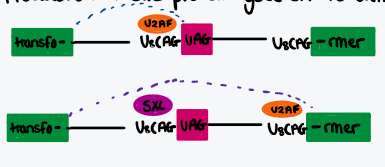
Describe influence of SXL protein in drosophila
Controls expression of transformer gene that controls somatic sex determination and dosage compensation
In males, no SXL present, premature termination means no transformer gene, no dosage compensation (good)
In females, SXL prevents U2AF binding to proximal pyrimidine tract, spliceosome assembles at distal pp tract, full transformer protein translated, dosage compensation (good)
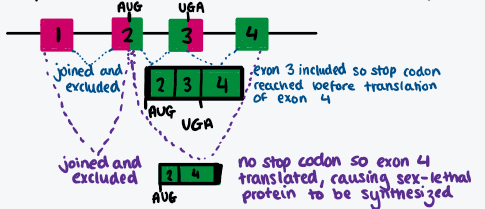
Draw diagram showing how splicing can change whether or not the SXL protein is translated
Draw a diagram showing what happens in splicing the transformer gene when the SXL protein isn’t and is present (good example of the impact of splicing repressor proteins)
What is the GU/AG rule?
Most introns start with GU and end with AG
What are the three key cis elements of a 3’ splice site?
Branch point consensus containing conserved adenine
Polypyrimidine tract
3’ splice site
Which bases are the pyrimidines?
Cytosine
Thymine
Uracil
What do trans factors do?
Bind to cis-elements to cause transcription by recruitment of ribosome or recognition of polypyrimidine tract by U2AF
What are snRNPs?
Small RNA-protein complexes that directly base pair with the mRNA
Involved with spliceosome assembly
Includes the U1-6 proteins (exl. U3?)
What type of ‘factor’ is U2AF and what does it do?
Trans factor
Large subunit binds to polypyrimidine tract
Small subunit associates with 3’ splice stie
How do we know that other proteins assemble to the U1 snRNP?
Antibodies against U1 snRNPs block splicing
Other things must be recruited where those antibodies are binding
What is the chemistry process of splicing?
Two successive transesterifications
-OH of branch point attacks phosphate at 5’ of intron
5’ exon released
Forms lariat intermediate
-OH of 5’ exon attacks phosphate at 3’ of intron
Ligates two exons together
Lariat released, rapidly degraded
No ATP consumed, number of phosphodiester bonds is conserved
What are the four stage of spliceosome assembly?
Early complex
A complex
B complex
RNA rearrangements into catalytic spliceosome
What is the full process of spliceosome assembly?
U1 to 5’ splice site consensus
U2AF to polypyrimidine tract and 3’ splice site
U2 bps to branch point
ATP hydrolysed
U4-U6 and U5 recognise A complex and bind
ATP hydrolysed
RNA rearrangements into the catalytic spliceosome
U4-U6 pairing is broken
U1 unbinds, replaced by U6
U6 also binds to U2
Align pre-mRNA for first catalytic step
Generates catalytic site
How are trans-acting factors identified?
Isolate specific factors that bind to cis-acting sequences
UV cross-linking
How are cis-acting elements identified?
Identify consensus sequences that can be experimented on using mutational analysis
Describe the process of RNA purification / biotin tagging
Use biotinylated UTP to make oligonucleotide identical to sequence querying that proteins interact with
Incubate with protein
Recover using streptavidin beads that bind to biotin
Western blot to identify presence of bound proteins, or use NMR if not a clue
Describe the process of RNA immunoprecipitation (RIP)
Antibody-based technique to map in-vivo RNA-protein interactions
Fix antibody against protein of interest against side of well
Mix protein with various RNA, add to well
Repeat washings so only protein of interest of left
RT to extract cDNA from RNA bound in protein
PCR or seq
Describe the process of cross-linking RNA and proteins
Irradiate with UV
Forms stable covalent bonds between RNA and proteins
What are the components of mature, post-spliced mRNA?
Coding sequence
5’ and 3’ untranslated region
5’ cap
3’ poly(A) tail
What is the purpose of the 5’ cap?
Increases splicing efficiency of introns close to 5’ end
Required for export to cytoplasm
Binding site for eIF4G in efficient translation initiation
Protection from 5’ exonucleases
What is the 5’ cap made of?
N-methyl-guanosine (m7Gppp-)
How is the 5’ cap added co-transcriptionally?
5’ gamma phosphate removed by RTPase
Pol II CTD activates RGTase
RGTase adds GMP to 5’ end
Methylated at position 7 (hence notation m7Gppp)
Final step in yeast, but in mammals some nucleotides are then individually modified
How is 5’ capping made specific?
All and only Pol II transcripts are capped
If Pol II promoter is changed to pol I or III there will be no capping, despite the transcript sequence being the same
Only di- or tri- phosphate ends will be capped, so mRNAs that have already been digested with endonucleases will not be capped
What is the tandem repeat of the CTD of RNA Pol II and how many repeats are there in yeast and mammals?
YSPTSPS (
26 in yeast
52 in mammals (double)
Simply, what does the CTD of RNA Pol II do?
Activates RGTase to add GMP in the second step of capping
What evidence is there that the CTD of RNA Pol II is required for capping?
Amanitin is an inhibitor of Pol II (allows selection for those that have taken up desired pol II)
Cells infected with one of two version of amanitin-resistant pol II: one has standard 52 CTD repeats and other has 5
Endogenous pol II inhibited by amanitin
Incubated
Capped/uncapped mRNA extracted and quantified
In 5 repeat group, equal amounts capped and uncapped (both very low)
In WT group, uncapped mRNA volume the same as in 5 repeat group suggesting uncapped mRNA formed by a different enzyme e.g. pol III
In WT group, much higher capped mRNA amount
What evidence is there that capping enzymes associate with the phosphorylated version of the RNA Pol II CTD?
Pass nuclear extract through one of three affinity columns: WT CTD, mutant CTD, phosphorylated WT CTD
measure capping activity of each sample using northern blotting and radiolabelled probes to the cap
Capping activity only retained in column with phosphorylated WT CTD
What does run-on transcription mean?
Pol II transcription does not terminate at precise regions
Carries on for hundreds bps downstream of what will become 3’ end
What evidence is there for run on transcription?
Incubate nuclei with NTPs and radiolabeled UTPs in vitro
Initiation is inhibited but RNAs that are already being transcribed are completed using the radiolabeled UTPs
Hybridized to probes once fully synthesized
There is radioactive signal much further downstream of the site corresponding to the future mature 3’ end
Signal is not constant and gradually decreases 5’ → 3’
What is the purpose of the poly(A) tail and how long is it in yeast and mammals?
Protects from 3’ exonucleases to control rate of mRNA degradation
Binding site for PABP (poly-a binding protein) which then binds to eI4FG
Evidence that cleavage and polyadenylation can happen independently
Radiolabel RNA substrate
ATP present: substrate is cleaved and polyadenylated
ddATP present prevents elongation when incorporated, substrate is cleaved but not polyadenylated
So polya happens after transcription and requires downstream elements and cleavage is upstream of polya
RNA molecule that mimics pre-cleaved substrate is polya but not cleaved further so cleavage must need some downstream elements
What evidence is there that polyadenylation has two steps
Normal substrate with AAUAAA consensus sequence has polyadenylation of 10A
Mutated consensus sequence has no polyadenylation at all
Mutated consensus sequence, but small poly(A) already present has polyadenylation
So, cleaved substrate with correct consensus is initially pola with a small poly(A) and then adenylated further in a second step
What is the consensus sequence involved in polyadenylation
AAUAAA
Why do proliferating cells have shorter 3’ UTRs
Fewer binding sites for regulators of gene expression
Why do only 105 of ribosomes exist in the dissociated form?
Ribosomes are 10x more abundant than dissociation factors
When small and large subunit are associated under physiological conditions it prevents them translating
What are the prokaryotic and eukaryotic initiation factors? What do they do?
Prok: IF3
Euk: eIF3
Bind to small subunit to prevent its reassociation with the large subunit so it can begin the process of translation initiation
What is the order of the sites on a ribosome (5’ → 3’)>
EPA
What is the full process of prokaryotic translation initiation?
30S subunit associates with IF1 and IF3
This complex interacts with mRNA at RBS
IF2 forms complex with GTP and initiator tRNA
These two complexes associate into the 30S initiation complex
Initiator tRNA is at the P site
50S subunit joins
Activates intrinsic GTPase activity of IF2
GTP hydrolysed, all initiation factors are released
70S initiation complex formed
IF2 exchanges GDP with GTP to recycle it
Some facts about the prokaryotic initiator tRNA
All bacterial proteins are synthesised with fMet as their first amino acid as that is the aa on the charged initiator tRNA
Formyl group rapidly lost
50% proteins lose the met entirely by slower enzymatic removal
What is the Shine Delgano sequence (bases)?
AGGAGG
What evidence is there that the Shine-Delgano sequence exists?
Bind ribosome subunits to mRNA’s during initiation to prevent elongation
Digest unprotected mRNA with RNase
mRNA inside ribosome is protected
Isolate fragments (RT) and sequence and map onto genome
16s rRNA binds to AGGAGG roughly 10nt upstream of the the initiator AUG
What is the advantage of having multiple initiation sites for prokaryotes?
Polycistronic translation
Ribosome can be bought to each ribosome binding site individually and does not stop translating when the next SD/RBS is reached
Summarise the three steps of prokaryotic elongation in translation
Binding of aminoacyl-tRNA to A site
Formation of peptide bond
Translocation of ribosome
Fully describe the process of prokaryotic elongation in translation
Elongation factor-Tu (EF-Tu) brings the aa-tRNA to A site, has GTPase activity
GTP-EF-Tu is masking aminoacyl group of aa-tRNA so it cannot react with existing peptidyl-tRNA
If codon-anticodon match is correct, conformational change in ribosome triggers GTPase activity
GDP-EF-Tu is released, tRNA in A site moves towards P site to enable peptide bond formation
If codon-anticodon match is incorrect, GTP-EF-Tu is released with GTP still intact
deacylated tRNA leaves P site, exits through E site as ribosome translocates
Describe how prokaryotic ribosomes translocate using elongation factors
GTP-EF-G only binds when EF-Tu isn’t present
GTPase activity stimulated on binding, causes translocation three nucleotides along, 5’ → 3’
Peptidyl-tRNA moves to P site, deacylated tRNA leaves via E site
Describe the importance of release factors for termination of prokaryotic translation
Recognise termination codons
Induce hydrolysis of the peptide chain from peptidyl-tRNA to release it
Need action of class I (RF1 and RF2) and class II
Describe how class I and class II release factors cause termination of prokaryotic translation
Class I (RF1 and 2) recognise termination codon directly, induce hydrolysis of peptidyl-tRNA
Class II (RF3) binds ribosome when bound to GDP
Exchanges GDP for GTP causing conformational change that releases a class I factor
Subsequent GTP hydrolysis releases RF3
What are the 6 key prokaryotic translation regulation mechanisms
Incorporation of the SD sequence in to a secondary structure
Translational coupling
Repressor proteins
Small non-coding RNAs (sRNAs)
Thermosensors
Attenuation
What is useful about the MS2 bacteriophage mRNA?
Contains three genes (maturase, coat, replicase)
Each gene has its own Shine-Delgano box
Region between each gene has potential to form secondary structure that disguises the SD box
What evidence is there that incorporating the SD box into a hairpin loop inhibits translation?
Incorporate SD box of MS2 bacteriophage into hairpin
Many different point mutations made in coat protein that destabilised or stabilised the hairpin, but did not alter SD sequence or protein sequence
Translation efficiency measured in vivo
Mutations that destabilised the hairpin had more efficient translation and vice versa
How does bacteriophage MS2 show regulation by repressor proteins?
Replicase gene is only required to be translated in early stages of infection, but its translation is coupled to coat protein translation
When coat protein accumulates, it is implied that enough replicase has been made
Coat protein binds to stem loop holding SD complex of replicase gene to stabilise it and repress translation
Prioritizes cellular components for maturase and coat proteins that are required in greater numbers
Explain prokaryotic translational regulation by sRNAs
Bind to mRNA
Activate translation by competing with ds regions in secondary structures, opening the secondary structure to reveal the RBS
Repress translation by directly binding to or masking the RBS
Describe prokaryotic regulation of translation by thermosensors
Secondary structures that block access to SD sequence are sensitive to temperature if proteins (e.g. heat shock proteins) that are only synthesized in response to temperature change, interfere with the stem loop
Observed in synthesis of prfA protein in some Listeria strains
Describe the principle of prokaryotic translation regulation by attenuation
Amino acid synthetic operons are switched off if the corresponding tRNA is abundant
Attenuation represses translation by premature termination of transcription (coupled)
Describe the process of attenuation in the Trp operon
Trp level is sensed by having two Trp in sequence in a short ORF
If trp is low ribosome pauses, 2:3 region hairpin forms
Prevents formation of RNAP-terminating hairpinn
Allows transcription and translation of the rest of the gene
If trp is abundant, no pause
3:4 RNAP-terminating hairpin forms
RNAp cannot bind, transcription terminated
Describe prokaryotic Rho-Independent Transcription termination
GC rich hairpin followed by run of 6 U
Mutations that disrupt hairpin of 6U disrupt temrination
Hairpin causes pausing
Weak interaction between A of template DNA and 6U on RNA causes fall off of template
Describe prokaryotic rho-dependent transcription termination
Rho protein has RNAdepenent ATPase that can act as helicase
Rho recognises Crich regions followed by a hair pin that causes transcriptional pausing (this is a cis-acting region)
Rho translocates along RNA until reaches the RNAP that is being stalled at the hairpin, causes RNAP dissociation
Compare prokaryotic vs eukaryotic initiation
Both: use start codons and a dedicated tRNA
Both: bind the small subunit to RNA first using initiation factors
Difference: recognition of mRNA
Pro: Small subunit binds SD box and initiation codon
Euk: small subunit binds 5’ cap and scans for first initiation codon
Difference: GTP and ATP usage
Pro: Only GTP for formation of 30S initiation complex and release of initiation factors
Euk: Both GTP and ATP
Simply, what is the eukaryotic translation scanning model?
43S initiation complex binds to 5’ cap and scans 5’ → 3’ until first AUG
60S subunit joins, elongation starts
What evidence is there for the eukaryotic scanning model?
90% of mRNAs initiate at first AUG
Insert a new AUG between 5' cap and original AUG, initiation begins at new inserted AUG
Inserting hairpin loop between 5’ cap and AUG inhibits translocation, so must be translocation of 43S between the two
What evidence is there that the 5’ cap is used in eukaryotic translation initiation?
Uncapped mRNA are translated accurately but ineffeciently
Addition of cap analogue reduces translation efficiency, so 43S is interacting with WT cap
Briefly, what are the three major steps of eukaryotic translation initiation?
Formation of 43S initiation complex
Formation of cap-binding complex at 5’ of mRNA
Binding to cap-binding complex and scanningD
Describe the full process of eukaryotic translation initiation:
1: Formation of 43S initiation complex:
40S subunit associates with eIF3 and eIF2
2: Formation of cap-binding complex at 5’ of mRNA
eIF4A, eIF4E and eIF4G all associate together to form the cap-binding complex eIF4F (43S)
Assembly is on mRNA
3: Scanning
43S binds to cap-bound complex
Scans mRNA until first AUG
Helicase activity of eIF4A unwinds initial secondary structure, requiring ATP
GTP in eIF2 is hydrolysed and released, releasing all imitation factors
eIF3
Prevents reassociation of ribosomal subunits
eIF2
GTP-bound
Brings initiator tRNA to the complex
eIF4A
Helicase
eIF4G
Binds to polyA binding protein for transition between initiation to elongation
eIF4E
Binds to 5’ cap
Recruits eIF4G
What evidence is there that eukaryotic mRNAs are monocistronic
Ribosomes scan starting at 5’ cap and eIF4A helicase activity unwinds first AUG, therefore cannot access downstream cistrons
Describe the transition between eukaryotic translational initiation to elongation
PolyA tail is coated in PABP
eIF4G omn 5’ cap binding complex binds to the PABP and mRNA is pseudo-circular
Interaction stimulates elongation
Describe internal initiation within viruses
Picronaviruses have uncapped mRNA and long 5’ UTRs with many AUGs and stable secondary structures
Prevents access to authentic AUG by scanning ribosome
mRNA is not translated by scanning but instead by direct ribosome binding to internal ribosome entry site (IRES)
Allows cap-independent translation
What evidence is there for IRES in viruses?
Discistronic reporter where finishing cistron A translation causes ribosome detachment
Insert IRES between cistrons A and B
Allows translation of cistron B
State four methods of regulating translation in eukaryotes
eIF2-GDP phosphorylation
eIF4E-eIF4G interactions
Gene-specific RNA-Binding proteins
RNA degradation / decay
Describe how eIF2-GDP phosphorylation regulates eukaryotic translation
Regulates initiation (otherwise would be wasteful and risky)
eIF2-GTP hydrolysis in the 43S initiation complex releases all initiation factors, beginning elongation
Recycling back to eIF2-GTP requires guanine exchange factor eIF2B
If eIF2-GDP is phosphorylated quickly, it will bind to eIF2B and prevent it recycling GTP of both that complex and other complexes
Also prevents eIF11 binding to 5’ cap, preventing initiation
Very little eIF2B present, so very rapid prevention of GTP recycling
How is eIF2-GDP phosphorylation part of the antiviral response?
Protein kinase R phosphorylates eIF2-GDP
Activated by presence of dsRNA e.g. with rotaviruses
Prevents translation of viral RNA
Describe how eIF4E-eIF4G interactions regulate eukaryotic translation
Once bound to 5’ cap, eIF4E recruits eIF4G
Can be modulated by 4E-BP which binds to eIF4E before eIF4G can bind
Phosphorylation of 4E-BP releases it
Changes in cellular mRNA levels can be achieved by….
modulating transcription rates, decay rates, or simultaneously both
How can mRNA degradation be measured experimentally?
Block transcription of the gene by inhibiting RNA pol II or cloning gene of interest under a regulatable promotor
Changes in mRNA levels are then reflected only by degradation as no synthesis
Describe non-specific mRNA decay and the two pathways that achieve this
Shortening of polyA tail and then:
Decapping or
3’ → 5’
Describe the non-specific decapping pathway of mRNA degradation
polyA tail is shortened
Lsm protein binds to shortened polyA
Promotes decapping
Can now be degraded 5’ → 3’ by XRN1 exonuclease
Describe the 3’ → 5’ pathway of mRNA degradation
Exosome degradation
Describe transcript specific mRNA decay
Involves both cis elements and trans factors
Trans factors: RNA binding protein recruits exosome to destabilizing cis element
Cis element involved: AU-rich elements (ARE)
Recognized by 3’ → 5’ action of exosome
Describe the biogenesis of miRNAs in animals
Genetically encoded imperfect dsRNA duplexes processed by drosha in nucleus into pre-microRNA
Exported to cytoplasm
Cut by dicer
Mature microRNA is 21nt ssRNA
Describe the biogenesis of siRNAs in animals
Less than 30bp dsRNA will not cause an interferon response
Or can insert small hairpins
Cut by dicer into siRNAs
How to identify binding sites of RNA binding proteins
CLIP-Seq
Proteins and RNA are cross-linked in vivo using UV
Extract
Treat with RNase so only RNA fragments protected by proteins remain
Add antibodies specific to RNABP to capture
Precipitated, RT RNA into DNA, sequence DNA
Map sequences onto genome
Where do miRNAs bind to mRNA and do they function in prokaryotes or eukarytoes?
3’ UTRs near cap (causes decapping)
Eukaryotes only
How are cells able to identify an siRNA as an siRNA?
2nt 3’ overhangs
21nt
5’ phosphate, 3’ hydroxyl
How do siRNA cause mRNA degredation
Loaded on RISC, associated with Ago2
Binds anywhere in the mRNA
Perfect complementarity
Causes formation of A-form helix which allows the duplex to align with the catalytic residues inside Ago2
Endonucleolytic cleavage
Why do microRNAs not cause endonucleolytic cleavage?
Bind with imperfect complementarity
Not form A-form helix
Does not align with catalytic residues inside Ago2
How do microRNAs cause mRNA degredation?
Bind in 3’ UTR
First 2-8 nt ‘seed’ bind perfectly, the rest is imperfect
Need multiple miRNA binding for mRNA degradation (causes recruitment of 3’ → 5’ exonucleases)
What is different about microRNA usage in plants?
Binds with perfect complementarity, which then allows Ago2 function