A3.2: Classification and Cladistics | IB Biology HL
1/19
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
Define “taxonomy.”
A3.2.1: Need for classification of organisms
Taxonomy is the study of the classification of living things.
List the levels of classification in the traditional hierarchy of taxa.
A3.2.1: Need for classification of organisms
A3.2.2: Difficulties classifying organisms into the traditional hierarchy of taxa
Domain
Kingdom
Phylum
Class
Order
Family
Genus
Species
Outline the benefits of having a system of classification of organisms.
A3.2.1: Need for classification of organisms
Organisms need to be organized because of the large amount of diversity of species.
Classifying organisms gives a more efficient way to see:
If a species has already been discovered
The similarities between species
Deduce common ancestry
Predict features that undiscovered but related species should have
Discuss the limitations of the traditional classification system.
A3.2.2: Difficulties classifying organisms into the traditional hierarchy of taxa
The traditional classification system is subjective and arbitrary, so it doesn’t always reflect common ancestry and genetic connections.
The classification system also restricts the classification so that the ranked system is respected. If two species are in the same family, then it must be in the same genus.
The hierarchy system also often doesn’t work as more organisms are discovered. For instance, hybrid organisms that can create offspring or undergo introgression don’t fit neatly into one species. Cloned organisms also don’t fit neatly into a species.
Define “synapomorphies.”
A3.2.3: Advantages of classification corresponding to evolutionary relationships
Synapomorphy is a shared, derived characteristic, common between an ancestor and its descendants
Discuss advantages of a classification system that corresponds to evolutionary relationships.
A3.2.3: Advantages of classification corresponding to evolutionary relationships
A classification system that corresponds to evolutionary relationships shows which species are more closely related and how they should be classified together. It also helps scientists determine how a species developed its characteristics and came into its present form. Researchers can use it to theorize why species descended from certain ancestors instead of others. This system is also not based on arbitrary, subjective, or contrived categories because it reflects species’ natural gene sequences.
Define clade.
A3.2.4: Clades as groups of organisms with common ancestry and shared characteristics
A clade is a group of organisms (or viruses) that all have evolved from a common ancestor.
Identify a clade as a branch in a cladogram.
A3.2.4: Clades as groups of organisms with common ancestry and shared characteristics
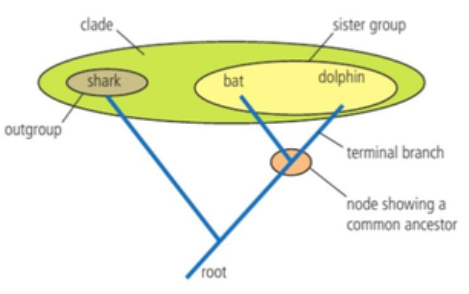
A clade is a branch in a cladogram.
A clade has a single common ancestor, a node, and all of its descendants.
List evidence used for placing organisms in a clade.
A3.2.4: Clades as groups of organisms with common ancestry and shared characteristics
The most objective evidence for placing organisms in the same clade comes from base sequences of genes or amino acid sequences of proteins
Relationships can be estimated by comparing DNA, RNA, or protein sequences
DNA sequences accumulate mutations as organisms evolve & diverge → Scientists compare mutations using sequence alignments to reconstruct evolutionary history
Outline the relationship between time, evolutionary relationships and biological sequences (nitrogenous base or amino acid).
A3.2.5: Gradual accumulation of sequence differences as the basis for estimates of when clades diverged from a common ancestor
More time → more mutations accumulated → biological sequences change → new species diverge → evolutionary relationships formed
The more time has passed, the more biological sequences change, creating new evolutionary relationships.
State the source of differences between biological sequences (nitrogenous base or amino acid).
A3.2.5: Gradual accumulation of sequence differences as the basis for estimates of when clades diverged from a common ancestor
Differences in the base sequence of DNA (and therefore proteins) are the result of mutations, which change biological sequences
Outline the use of a “molecular clock” to determine time since divergence between two clades.
A3.2.5: Gradual accumulation of sequence differences as the basis for estimates of when clades diverged from a common ancestor
Molecular clocks only allow scientists to estimate when species diverged using:
Generation time: Species with shorter generation times → more opportunities for mutations to occur → faster molecular clocks than species with long generation times
Population size: Mutations are more likely to be fixed in small populations → faster molecular clocks
Selective pressure: Strong selective pressure (high intraspecific competition) → mutations are more likely to be fixed in a population
Define “cladogram.”
A3.2.7: Analyzing cladograms
A cladogram is a tree diagram that shows the most probable divergence between species.
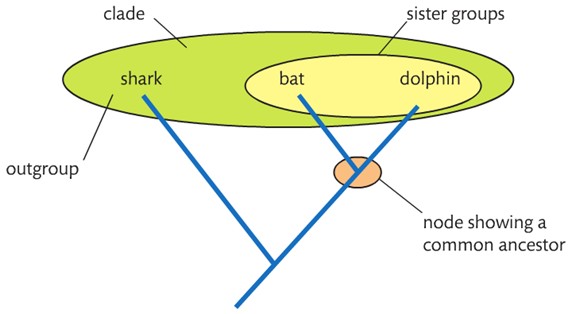
Outline what is represented by a “root”, “node”, “terminal branch”, “sister group”, and “outgroup” on a cladogram.
A3.2.7: Analyzing cladograms
A root is the base of the cladogram showing the common ancestor of all the clades in the cladogram.
A node is a split in a cladogram (branches split) that represents a hypothetical common ancestor.
A terminal branch is an end of a branch that represents the most recently evolved organism in the clade.
A sister group is a group of closest relatives.
An outgroup is the most distantly related species in the cladogram.
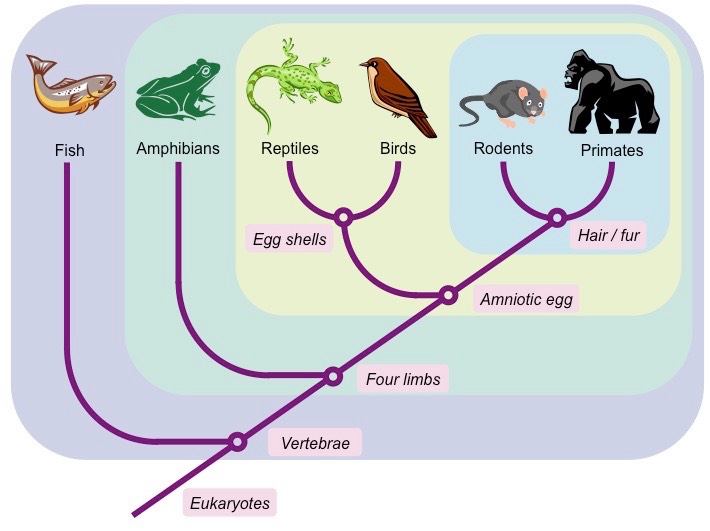
Explain why the development of cladistics lead to the reclassification of some species.
A3.2.8: Using cladistics to investigate whether the classification of groups corresponds to evolutionary relationships
The development of cladistics may lead to the reclassification of some species as the genetic relationships between groups may become clearer. As a result, organisms may not be grouped by morphology anymore, but by evolutionary relationships.
As zones of DNA markers are analyzed, we may realize that an old classification is not monophyletic (taxa didn’t share a most recent common ancestor), and we may have circumscription (moving branches of the tree of life).
Outline the reason and evidence for the reclassification of the figwort family.
A3.2.8: Using cladistics to investigate whether the classification of groups corresponds to evolutionary relationships
Scrophulariaceae of the figwort family were formally classified by morphological features: how flower petals were arranged in the bud before the flower opens (aestivation), whether the petals overlapped or are in a spiral, or the morphology of the nectaries (organs that produce nectar).
However, after the mid 1990s, a DNA analysis of the plants, analyzing zones of DNA markers (such as ribosomal internal transcribed spacer (ITS) region) revealed that the old classification was not monophyletic (taxa didn’t share a most recent common ancestor — the old system grouped plants that belonged in separate branches. Paraphyletic organisms are species that are on separate branches. They used to be in the same family, but are no longer (foxgloves are no longer figworts). Circumscription is the process of placing taxa where they clearly show monophyletic groups (indicating they all share a recent common ancestor, moving branches of the tree of life).
List the three domains of life.
A3.2.9: Classification of all organisms into three domains using evidence from rRNA base sequences.
Archaea
Bacteria
Eukarya
Discuss evidence from rRNA base sequences that led to the reclassification of life from two cell types (prokaryotic and eukaryotic) to three domains.
A3.2.9: Classification of all organisms into three domains using evidence from rRNA base sequences.
Carl Woese, compared the subunits of the ribosome 16S rRNA of typical bacteria and methanogens. Many sequences present in typical bacteria were unexpected in methanogens (some sequences never in bacteria were found in methanogens, or some sequences not found in methanogens were almost always in typical bacteria). Thus, many sequences that Woese found in methanogens (expected to match those found in bacteria) did not match bacteria. This provided evidence that the prokaryotic group is actually made of 2 separate domains — archaea and bacteria.
Outline what separates archaea from bacteria.
A3.2.9: Classification of all organisms into three domains using evidence from rRNA base sequences.
Differences in the subunit of their ribosomes called 16S rRNA → grouping archaea & bacteria difficult to justify
Metabolic reactions carried out by archaea that no bacteria can perform
Some archaea can synthesize methane & some can use hydrogen gas a source of energy
Transcription in archaea (the ways archaea read DNA to produce RNA) & translation (produce proteins from RNA) share more characteristics with eukaryotic transcription & translation than with bacterial transcription & translation
Some physical features of archaea are difficult from bacteria (like the types of molecules used to build their cell membrane & cell wall
Interpret the tree diagram that illustrates the evolutionary relationship between organisms of the three domains.
A3.2.9: Classification of all organisms into three domains using evidence from rRNA base sequences.
The previous classification only had 2 domains — prokaryotes and eukaryotes.
The current classification has 3 domains.