Molecular & Cell Bio. - Exam 3 - Epigenetics
1/31
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
32 Terms
what is epigenetics?
the study of how cells control gene activity without changing the DNA sequence
- reversible changes
- turn genes on or off
- related to chromatin modification and regulation of gene expression
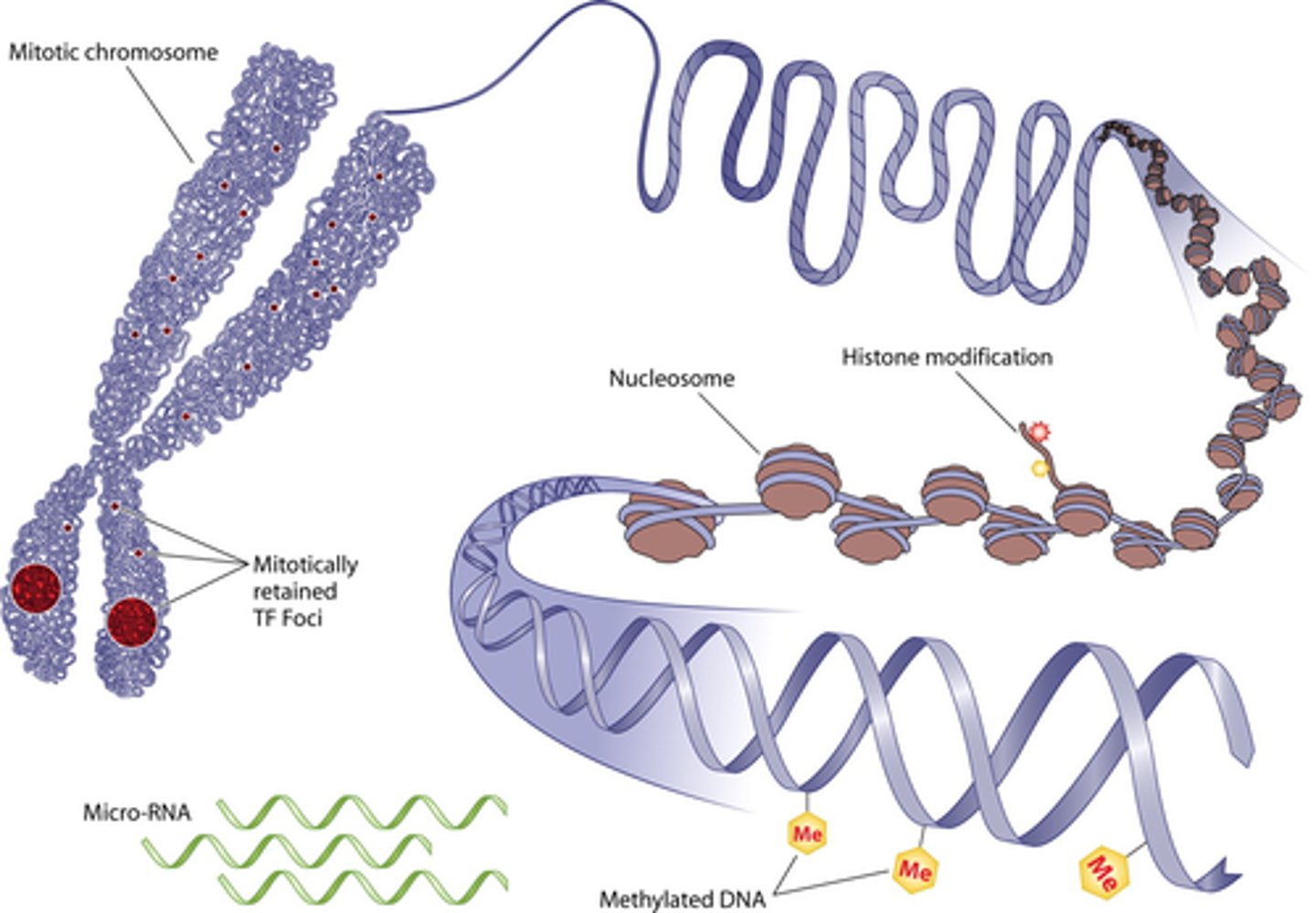
what are 2 examples of epigenetic gene expression mechanisms?
- DNA methylation
- histone modification
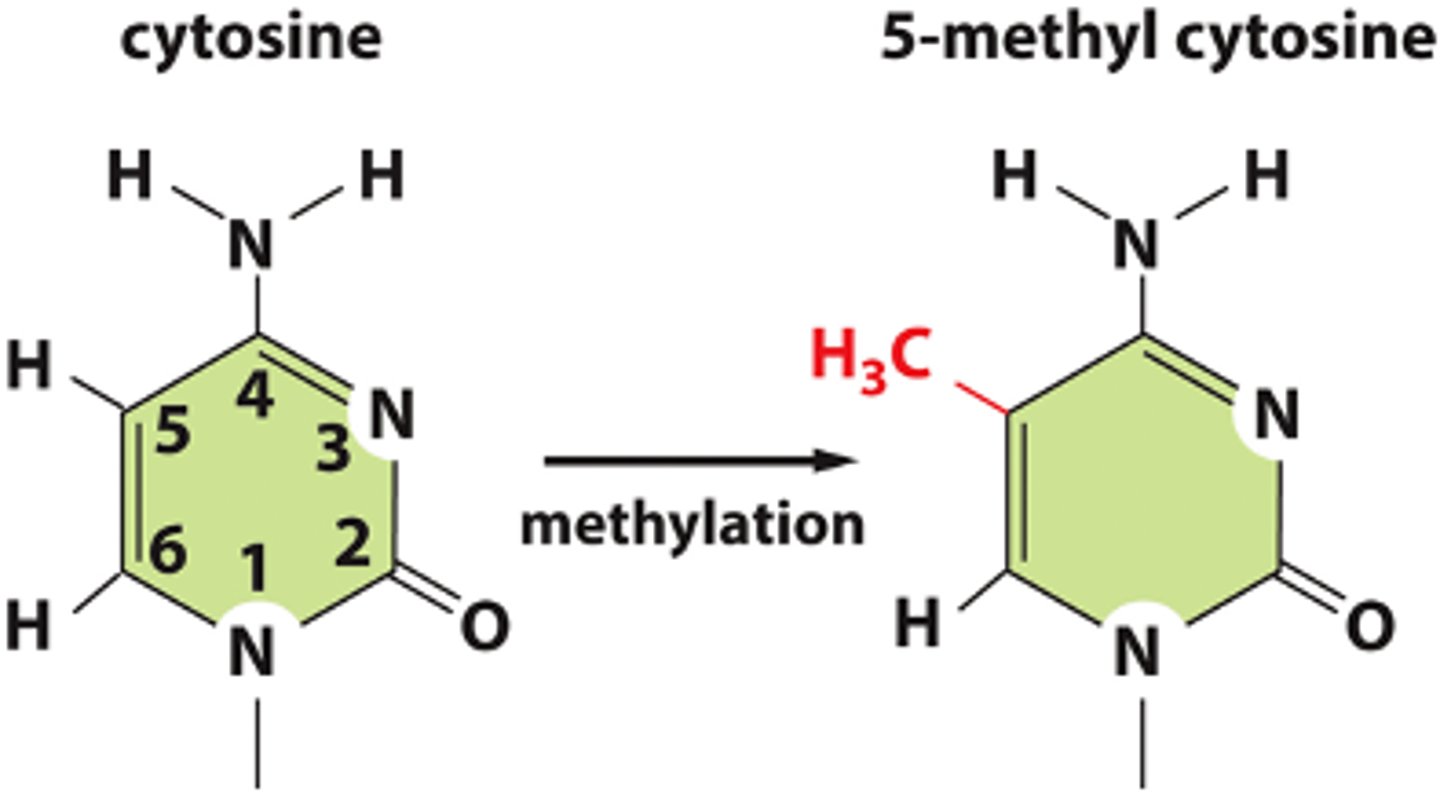
what bases are affected by DNA methylation? what enzyme catalyzes this?
chemical modification to an individual cytosine
- cytosines are located in the CG-rich regions (called CG or CpG islands)
- catalyzed by DNA methyl transferase (DNMT)
- DNA methylation patterns are maintained and propagated during cell division
T/F: DNA methylation is often associated with long-term gene inactivation
TRUE
- signals transcriptional repression
- CA cells often display altered DNA methylation patterns!
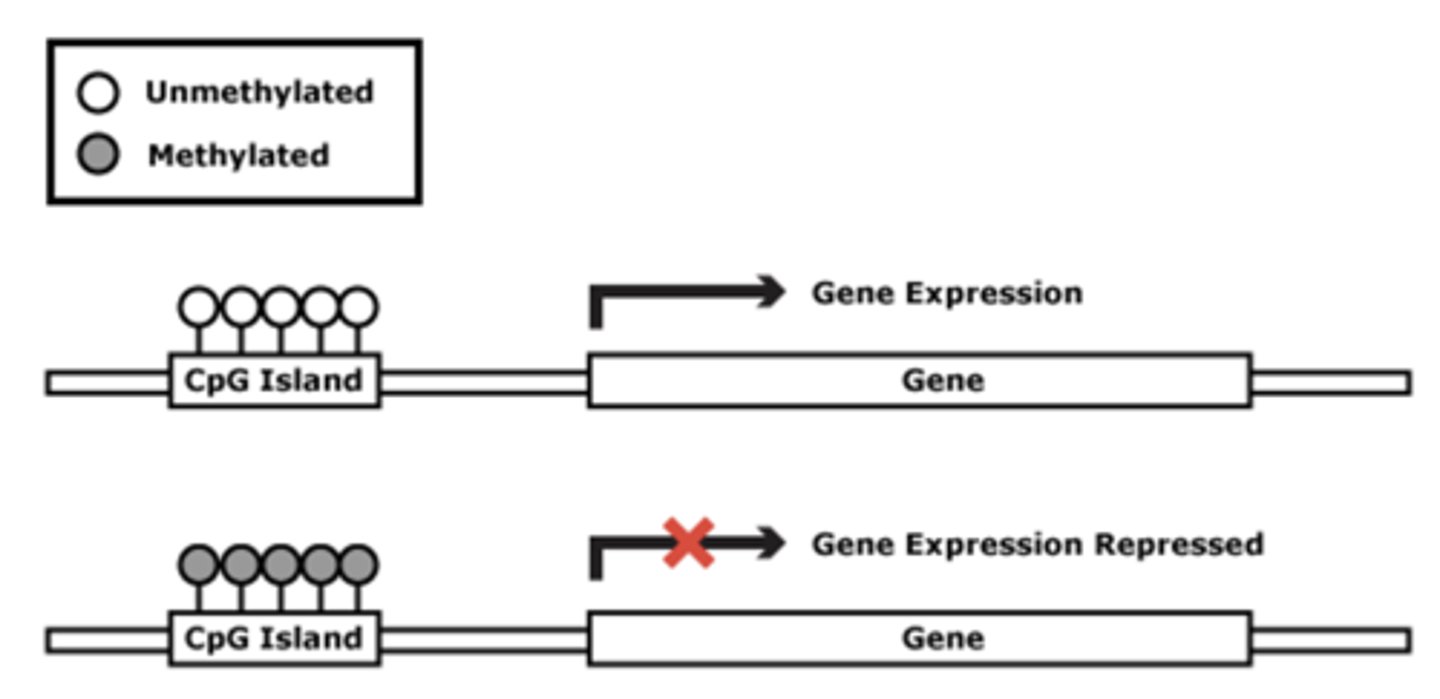
how does DNA methylation exert its repressive effects?
by blocking DNA-binding proteins (like transcription factors) from accessing DNA
- also recruits proteins that contain methyl-CpG binding domains (MBD)
DNA is wrapped around histones in a nucleosome core particle and sealed by _____________
histone H1
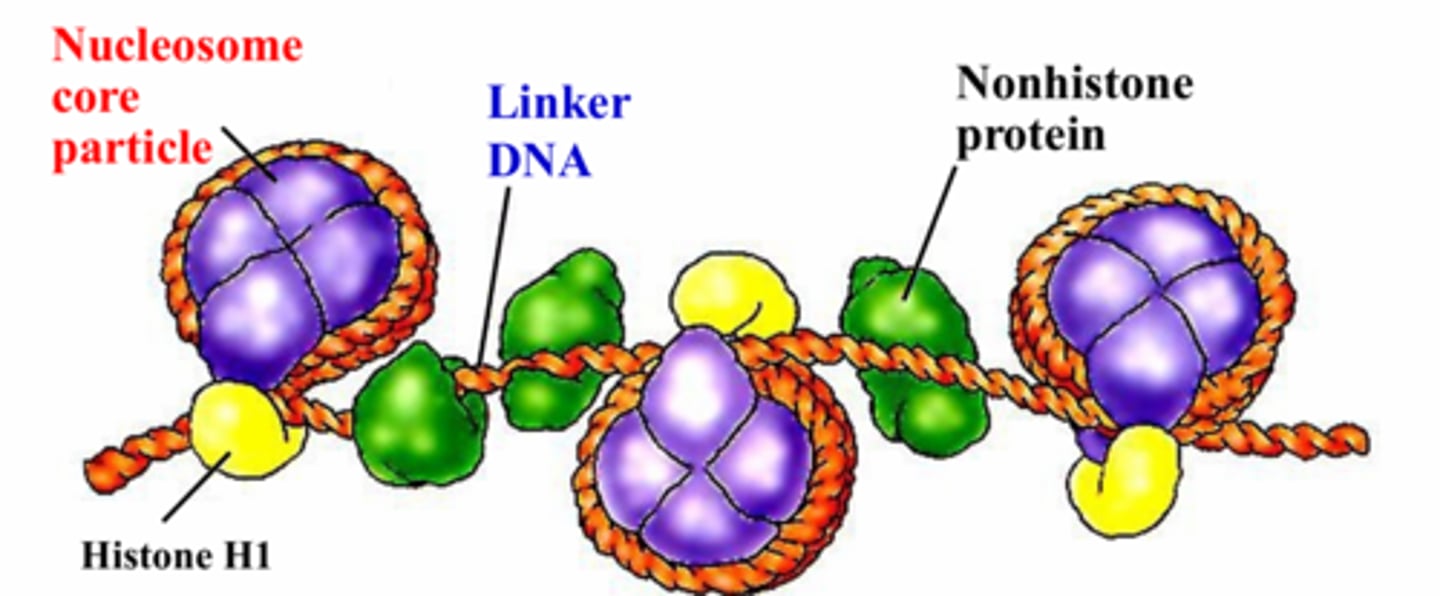
what do nonhistone proteins do in chromatin?
bind to the linker DNA between nucleosome core particles
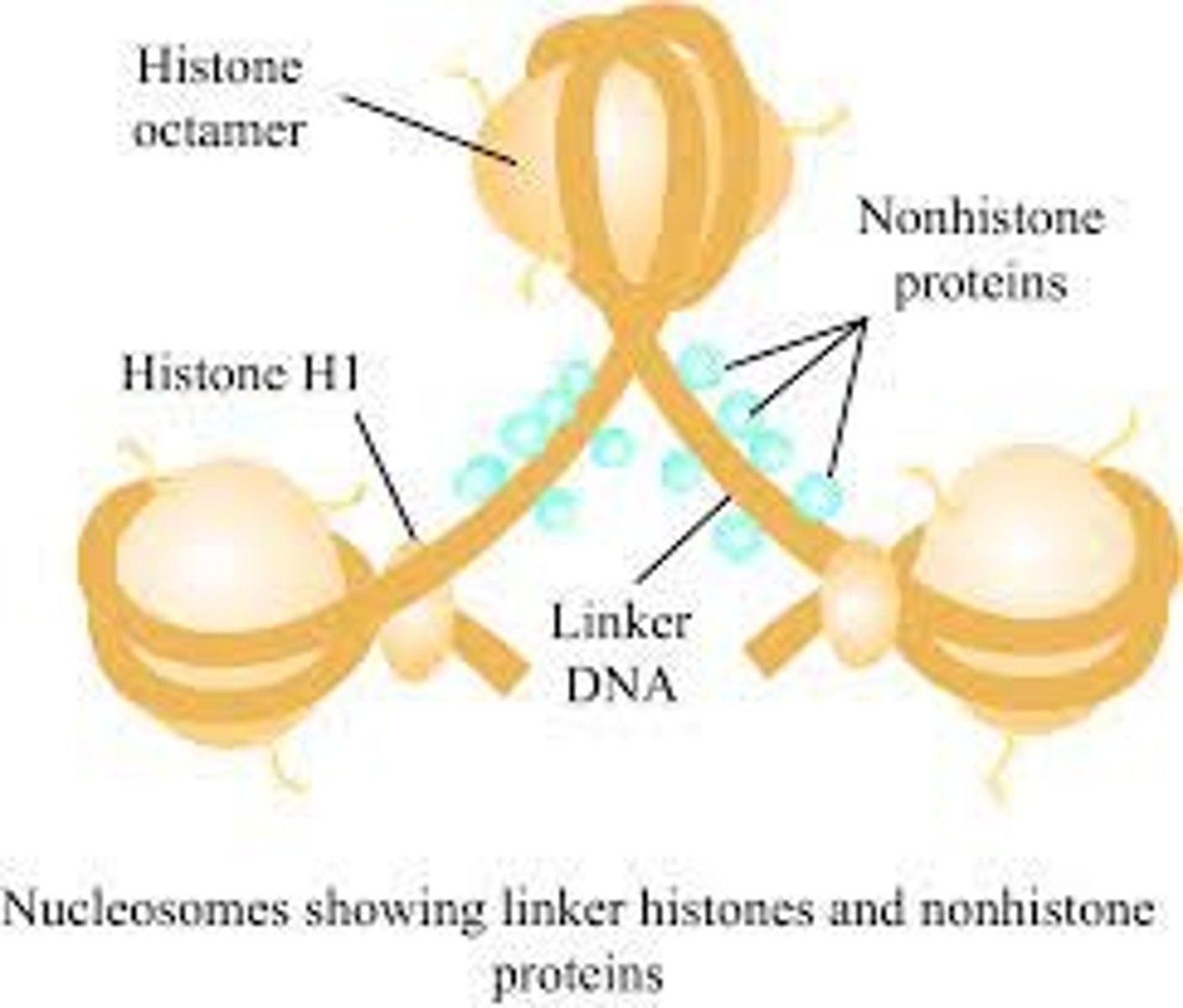
a nucleosome core particle consists of _______ bps of DNA wrapped around a histone
146
describe how chromatin fibers are packaged into nucleosomes to form chromosomes
the packaging of DNA into nucleosomes yields a chromatin fiber ~10 nm in diameter
- chromatin is further condensed by coiling into a 30 nm fiber, containing about 6 nucleosomes per turn
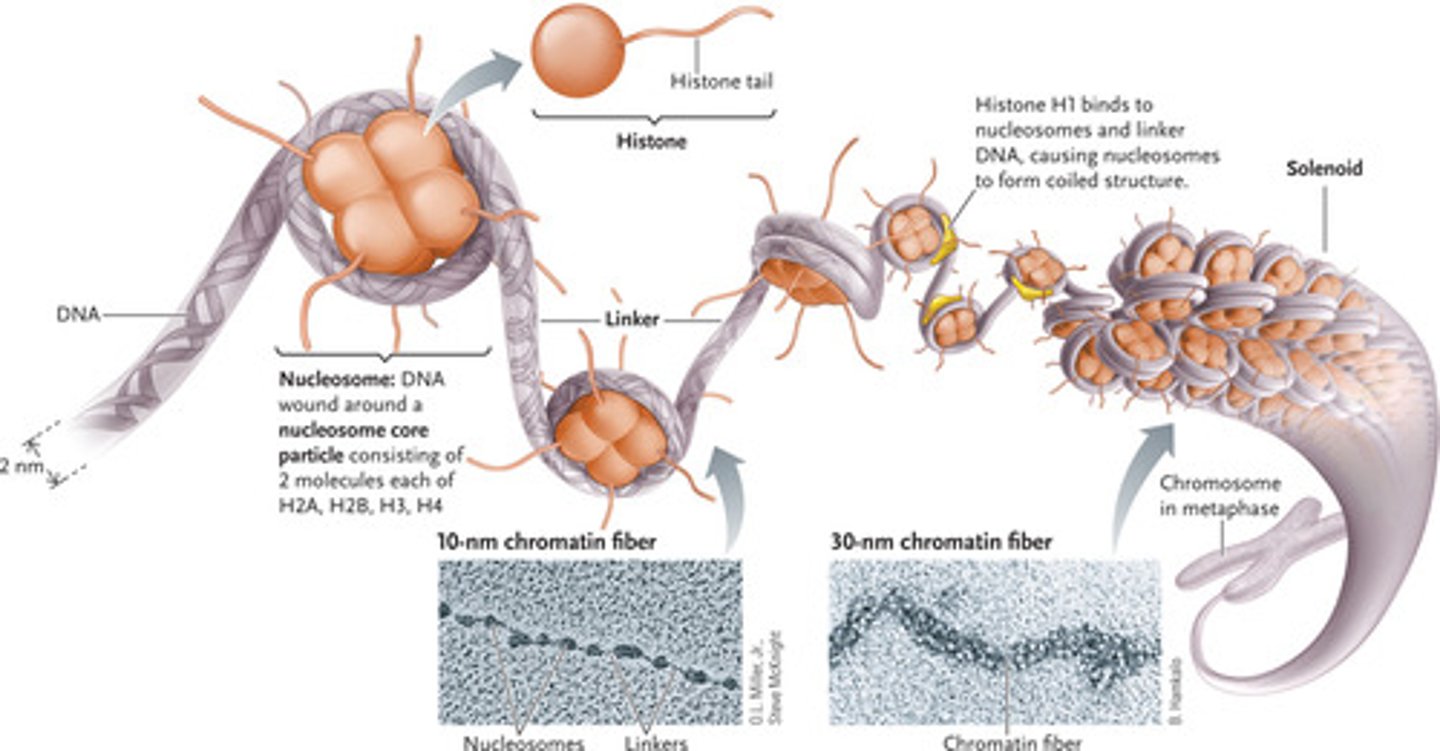
what are the 3 main types of histone modifications?
post-translational modifications to the N-terminal tail of histones
- methylation
- acetylation
- phosphorylation
in what 2 forms can DNA be found within the nucleus? which histone modifications are associated with each?
euchromatin
- open
- induced by acetylation and phosphorylation
heterochromatin
- condensed
- induced by methylation
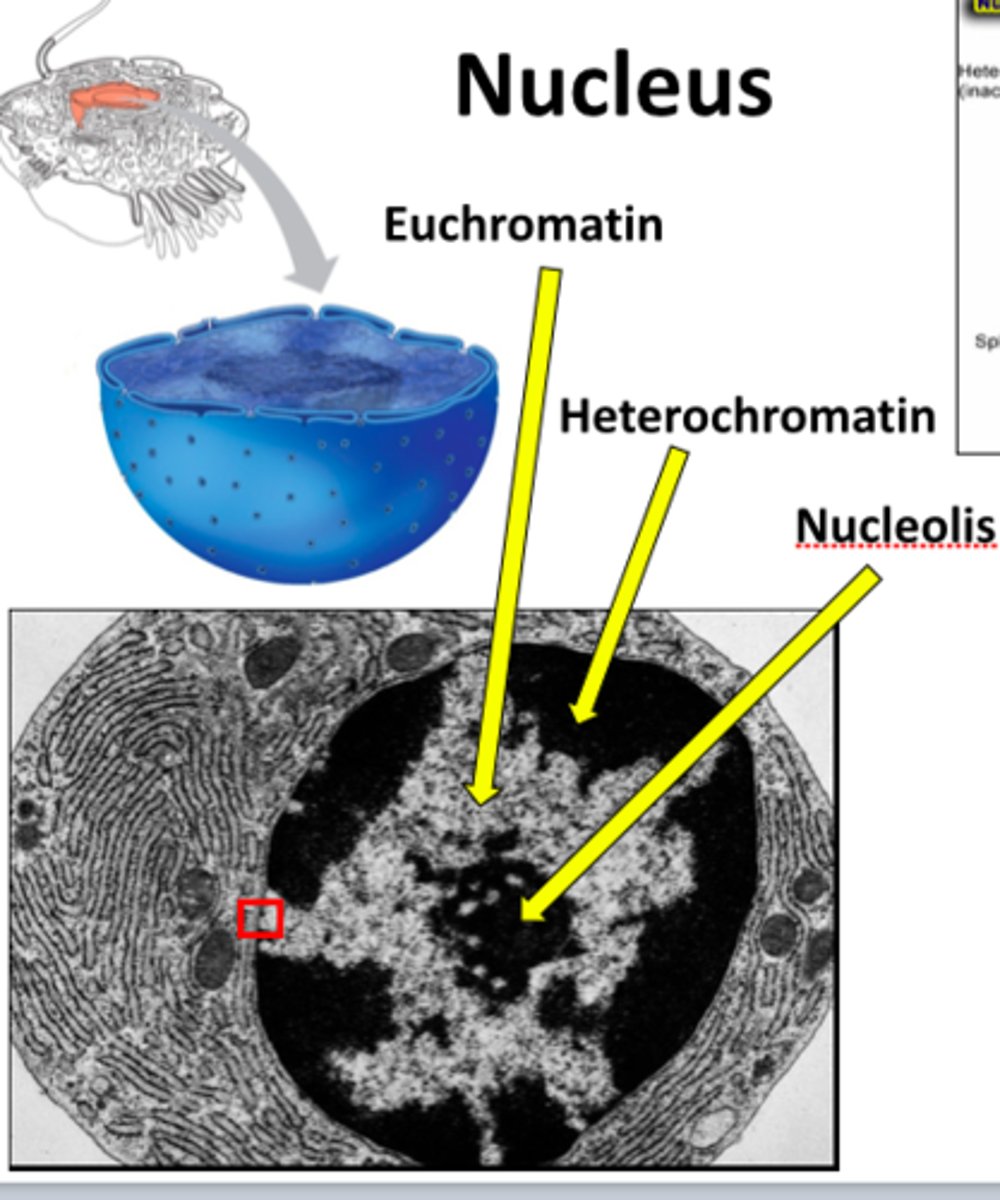
what are epigenetic marks?
small chemical tags that sit on top of chromatin and help determine if it will be in an open or condensed formation
in heterochromatin, transcription is ______________
repressed
- gene promoter is inaccessible
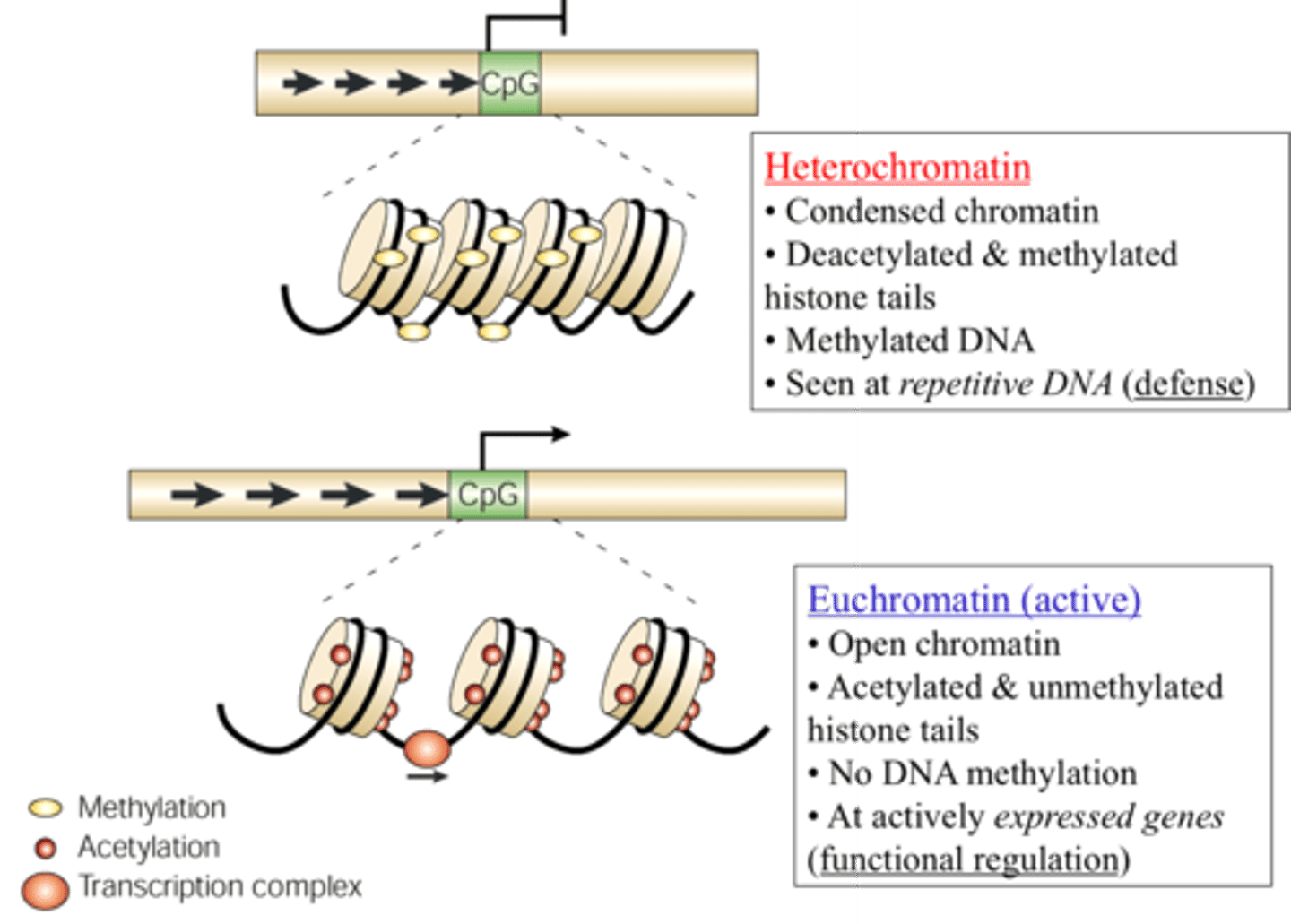
T/F: eukaryotic genes are in an active conformation by default
FALSE
- in a repressed conformation by default
- transcriptional activation requires heterochromatin to DE-COMPACT into euchromatin to allow RNAP II to access the promoter region
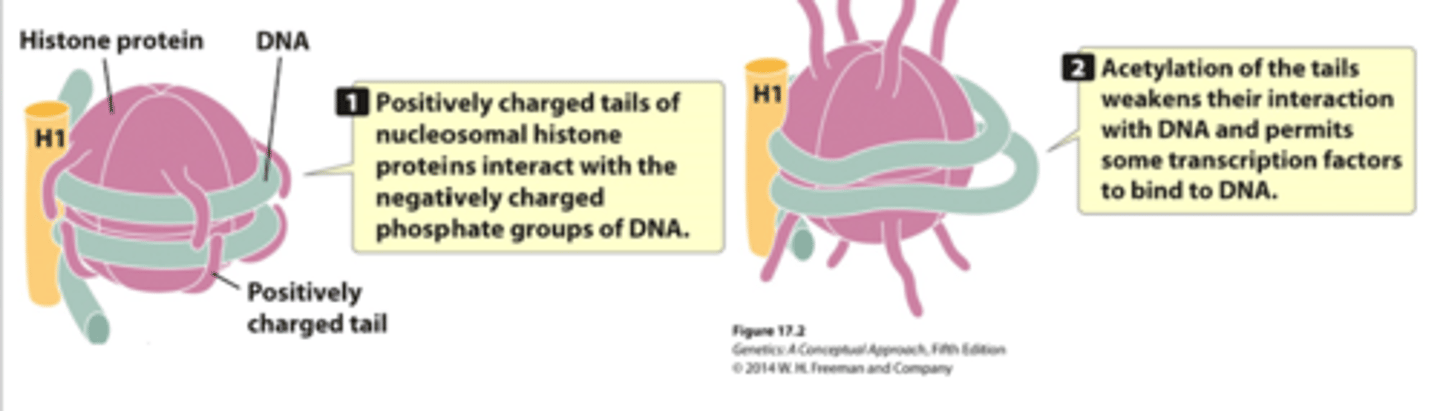
what is the role of histone acetyltransferase (HAT)?
histone code-writers
- acetylates conserved lysine amino acids on histone proteins by transferring an acetyl group from acetyl-CoA to form N-acetyl lysine
- act as coactivators of RNAP II
- induces gene transcription by promoting euchromatin formation (de-condenses heterochromatin)
FYI - what are some examples of HAT enzymes?
- HAT1
- p300/CBP family
- GNAT family (hGCN5, PCAF, ELP3)
- MYST family (TIP60, MYST 1-4)
- TFIIIC90, TAF1
- SRC1, ACTR, p160, CLOCK
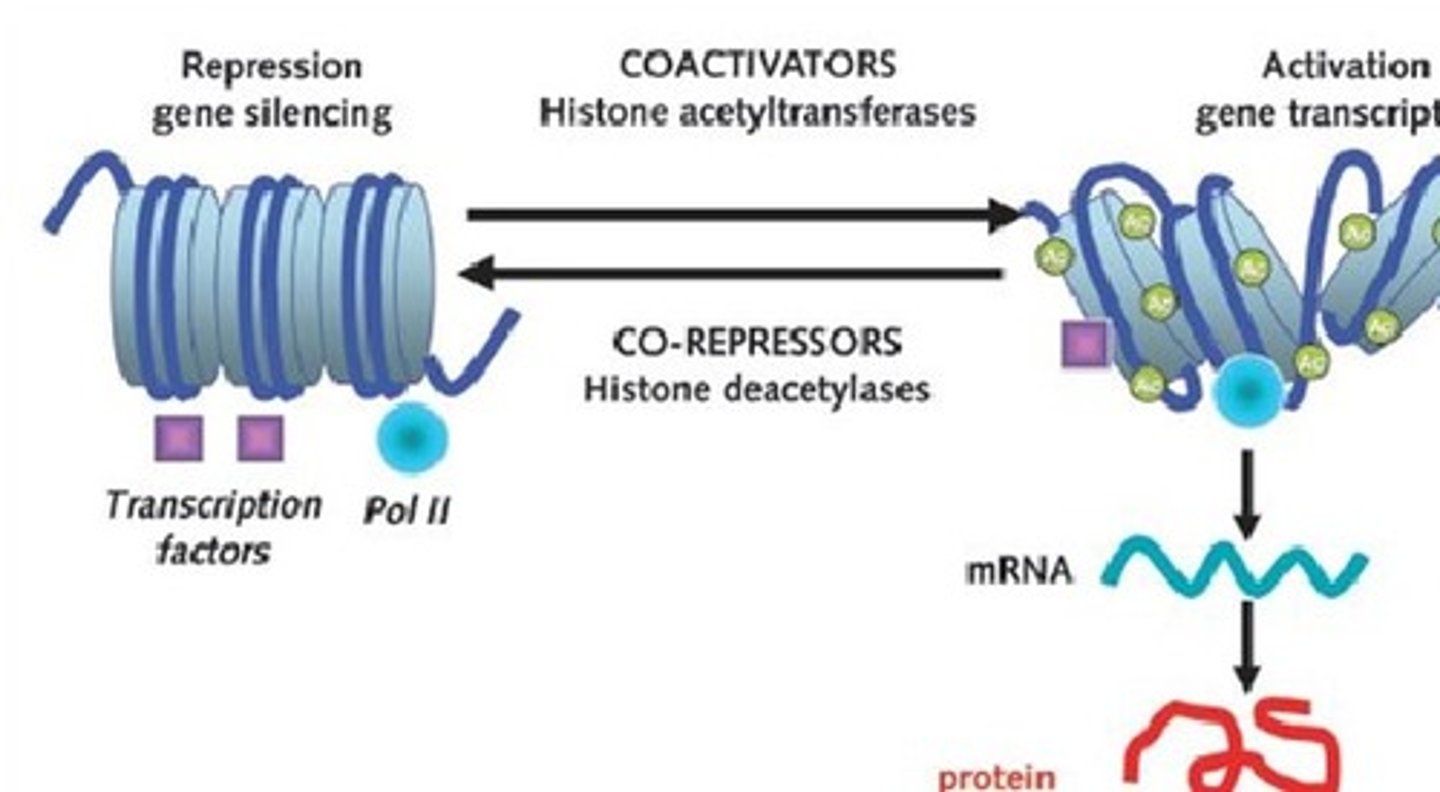
what is the role of histone deacetylase (HDAC)?
histone code-erasers
- remove acetyl functional groups from the lysine residues of both histone and nonhistone proteins
- leads to the formation of heterochromatin from euchromatin (re-condenses euchromatin)
- blocks RNAP II accessibility/activity
FYI - what are some examples of HDAC enzymes?
- class I: HDAC1/2/3/8
- class IIa: HDAC4/5/7/9
- class IIb: HDAC6/10
- class III: SIRT1-7
- class IV: HDAC11
T/F: histone methylation mainly occurs on the side chains of lysine and arginine
TRUE
what is the role of histone lysine methyltransferases (HKMT)?
methylate lysine within the N-terminal tails; lysines may be mono-, di-, or tri-methylated
- methylation at different lysine residues: H3K27, H3K4, H3K9, H4K20
- methylation at arginine residues: H3R17
K-me1's will mono-methylate residues, while K-me2's will di-methylate, and K-me3's will tri-methylate the epsilon-amine group of K
at which lysine residue will methylation ENHANCE gene transcription?
K4
which protein arginine methyltransferase (PRTM) produces symmetrical di-methylated arginines? which produces asymmetrical?
- symmetrical: R-me2s
- asymmetrical: R-me2a
list the modification possibilities that could convert euchromatin into heterochromatin (open to condensed)
- DNA methylation
- histone methylation (H3K9me3)
- histone deacetylation
- corepressor complexes
list the modification possibilities that could convert heterochromatin into euchromatin (condensed to open)
- histone methylation (H3K4me3)
- histone acetylation
- loss of H1
- coactivator complexes
what region of a gene must be methylated in order to repress transcription?
promoter
- methylation within the gene itself will not affect expression
list 3 common tumor suppressor genes
- p53
- p16
- E-cadherin
what is the target of some anti-CA drugs in terms of post-translational modifications?
target DNA methyltransferase (enzyme inhibitors)
- ex: 5-Aza 2' deoxy-cytidine (5-Aza-dc) has been approved to treat myelodysplastic syndrome
how do HDAC inhibitors function as anti-CA agents?
- block the deacetylation function of HDACs to allow for the expression of genes that are involved in regulating cell division
- slows down the growth of CA
- ex: Vorinostat, Belinostat (for T cell lymphomas); Panobinostat (for myeloma)
what is miRNA? describe how it is converted into its active form
micro RNA; transcribed from DNA, but does not encode proteins
① once transcribed, the pre-miRNA complexes with RNA-specific endonucleases (Drosha) in the nucleus to create its stem-loop structure (70-80 nucleotides long)
② the pre-miRNA is then exported from the nucleus into the cytoplasm, where it complexes with another RNA endonuclease (Dicer) to produce mature miRNA
③ the mature miRNA is incorporated into the RNA-induced silencing complex (RISC)
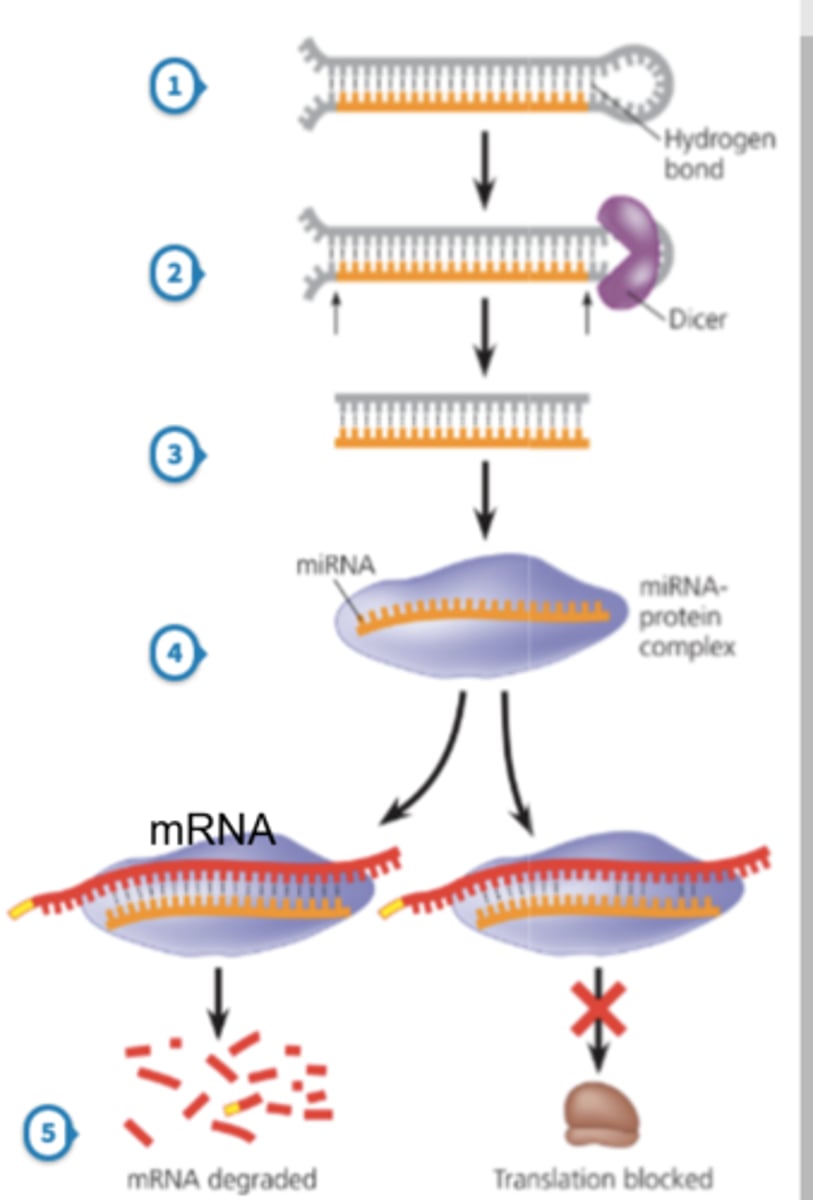
what is the function of miRNA? how does miRNA bind to target genes?
represses translation and induces mRNA cleavage
- binds to the 3' UTR of a target gene via base pairing
T/F: miRNA can activate translation
TRUE
- can act in an unconventional manner and activate translation by binding to non-canonical sites in the 5' UTR of target genes
how does miRNA affect cancer-related genes?
can function as oncogenes OR tumor-suppressors!
- repress translation of tumor suppressor genes (miR-21, -10b, -191)
- inhibit the expression of oncogenes and blocks the tumorigenesis process (miR-15a, -16-1, -34a)