Genetics Ch 12: Prok Transcription
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
__________ genes = transcribed and translated
aka ________-coding genes
90% of genes
__________ genes = transcribed, but not translated (function as RNA)
ex: ribosomes, spliceosomes, signal recognition particle, telomerase
structural genes = transcribed and translated
aka protein-coding genes
90% of genes
non-structural genes = transcribed, but not translated (function as RNA)
ex: ribosomes, spliceosomes, signal recognition particle, telomerase
Bacterial gene sites for transcription
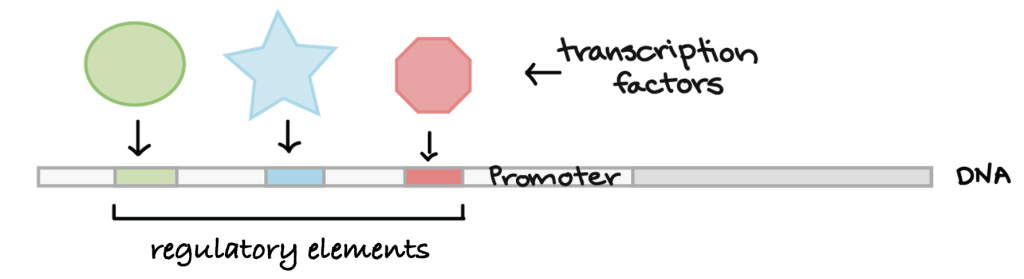
__________ = sites for binding regulatory transcription factor proteins
increases or decreases rate of transcription
__________ = site for RNA pol binding = beginning of transcription
__________ = site that signals end of transcription
__________ = site for ribosome binding to mRNA in bacteria
ribosome scans RNA for start codon
translation beings here
in eukaryotes, ribosome binds to _________ cap in mRNA
Bacterial gene sites for transcription
regulatory elements = sites for binding regulatory transcription factor proteins
increases or decreases rate of transcription
promoter = site for RNA pol binding = beginning of transcription
terminator = site that signals end of transcription
ribosome-binding site = site for ribosome binding to mRNA in bacteria
ribosome scans RNA for start codon
translation beings here
in eukaryotes, ribosome binds to 7-methylguanosine cap in mRNA
codons are on ______ and anticodons are on ______
codons are on mRNA and anticodons are on tRNA
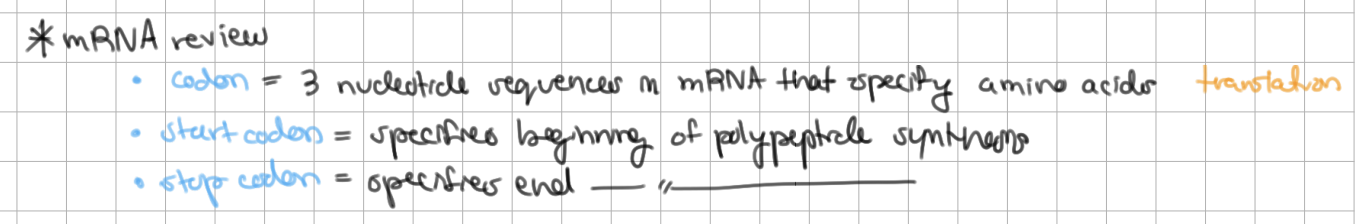
bacterial mRNA are __________ = one mRNA → many polypeptides
each mRNA can have many start/stop codons
polycistronic
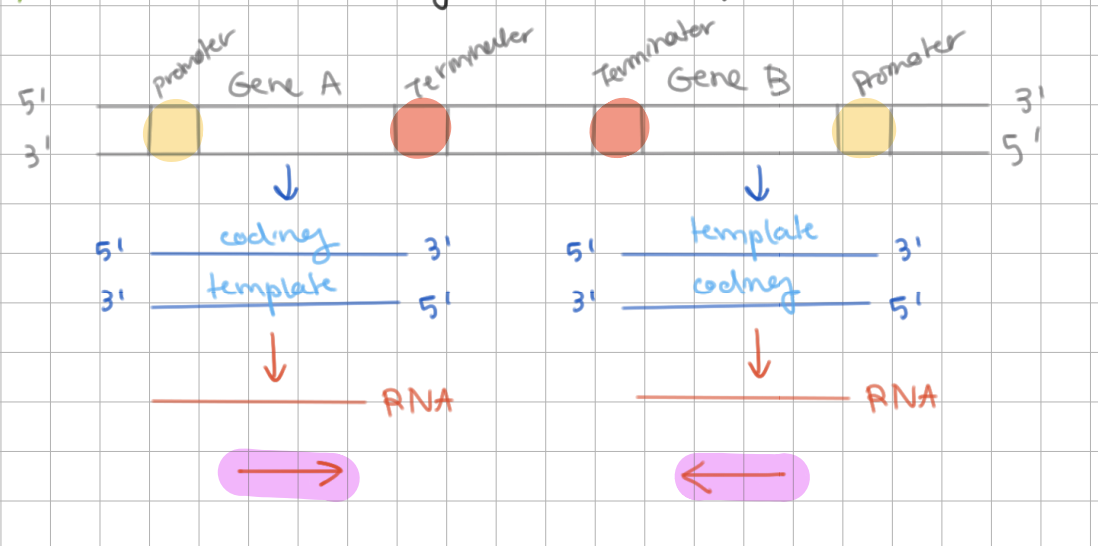
nontemplate/template strand = transcribed strand, complementary to to mRNA
nontemplate/template strand = not transcribed, identical to mRNA
template strand = transcribed strand, complementary to to mRNA
nontemplate strand = not transcribed, identical to mRNA

template = sense/antisense = coding/noncoding
nontemplate = sense/antisense = coding/noncoding
template = antisense = noncoding
nontemplate = sense = coding

promoters = promote gene expression
direct location for start of transcription
upstream/downstream where transcription begins
in other words, at __’ of nontemplate strand
contain numbering system relative to transcription start site
promoters = promote gene expression
direct location for start of transcription
upstream where transcription begins
in other words, at 5’ of nontemplate strand
contain numbering system relative to transcription start site
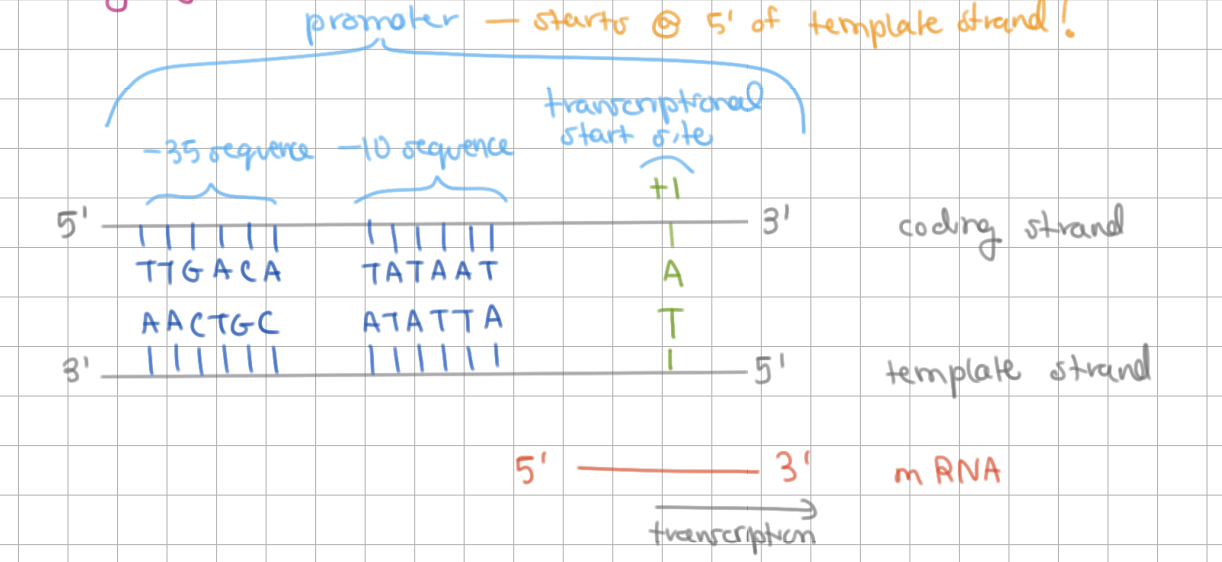
__________ = common sequences
most common are _____ and _____ sequences
results in high/low transcription
if promoter deviates from consensus sequence → high/low transcription
consensus sequence = common sequences
most common are -35 and -10 sequences
results in high transcription
if promoter deviates from consensus sequence → low transcription
__________ = catalyzes synthesis of RNA
RNA pol
RNA pol
core enzyme = ________
holoenzyme = ________
RNA pol
core enzyme = ɑ2ββ’⍵
holoenzyme = ɑ2ββ’⍵σ
contains core enzyme + sigma factor
Transcription contains 3 stages:
Initiation
Elongation
Termination
Initiation of Transcription (1/4)
RNA pol holoenzyme binds _____ to DNA
scans DNA for _______ region
Initiation of Transcription (1/4)
RNA pol holoenzyme binds loosely to DNA
scans DNA for promoter region
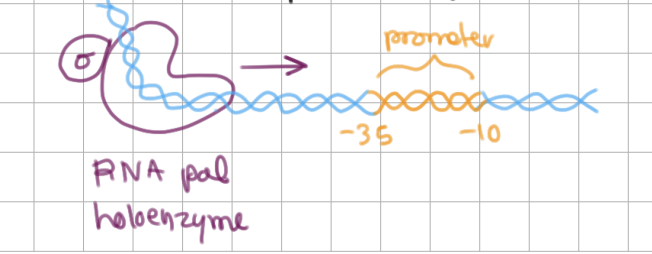
Initiation of Transcription (2/4)
________ recognizes promoter region by -10 and -35 sequences
contains ____________ and involved in ________ binding
RNA pol binding to promoter → _____ complex
Initiation of Transcription (2/4)
sigma factor (σ) recognizes promoter region by -10 and -35 sequences
contains helix-turn-helix and involved in tighter binding
RNA pol binding to promoter → closed complex
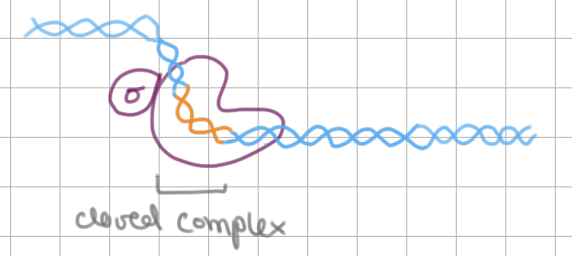
Initiation of Transcription (3/4)
TATAAAT box in ___ region is unwound → _____ complex
AT bonds can be easily separated
Initiation of Transcription (3/4)
TATAAAT box in -10 region is unwound → open complex
AT bonds can be easily separated
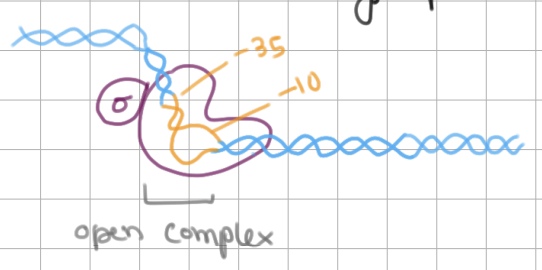
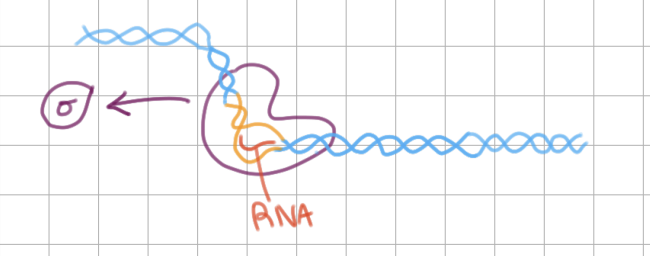
Initiation of Transcription (4/4)
short RNA strand made in open complex
_______ is released
Initiation of Transcription (4/4)
short RNA strand made in open complex
sigma factor is released
Elongation during Transcription (1/3)
open complex is about ___ bases long
rate of RNA synthesis = ___ nucleotides/second
Elongation during Transcription (1/3)
open complex is about 17 bases long
rate of RNA synthesis = 43 nucleotides/second
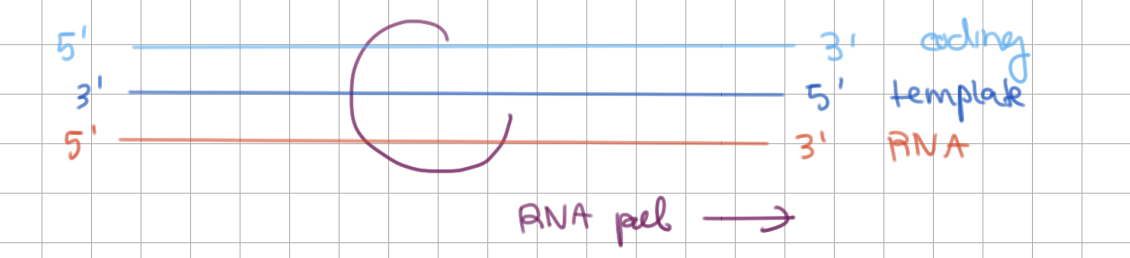
Elongation during Transcription (2/3)
RNA pol synthesizes RNA from __’ → __’
RNA pol slides along template from __’ → __’
Elongation during Transcription (2/3)
RNA pol synthesizes RNA from 5’ → 3’
RNA pol slides along template from 3’ → 5’
Elongation during Transcription (3/3)
precursors = __________
__________ is released
Elongation during Transcription (3/3)
precursors = nucleoside triphosphates
pyrophosphate (PPi) is released
Both strands may / may not be used as template
Both strands may be used as template
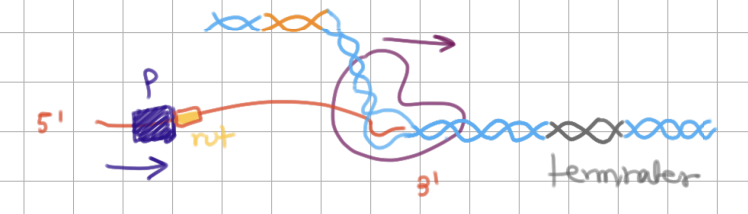
Rho-dependent termination = requires ⍴ (1/3)
______ protein binds to ____ site on RNA and moves to 3’ end
Rho-dependent termination (1/3)
⍴ (rho) protein binds to rut site on RNA and moves to 3’ end
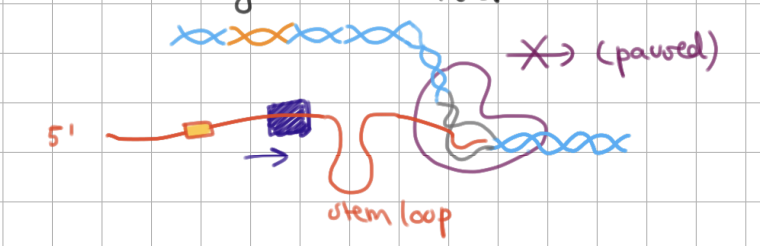
Rho-dependent termination (2/3)
RNA pol transcribes _____ loop (causing RNA pol to _____) and goes to terminator
Rho-dependent termination (2/3)
RNA pol transcribes stem loop (causing RNA pol to pause) and goes to terminator
Rho-dependent termination (3/3)
During pause, rho can reach open complex to separate ________
Rho-dependent termination (3/3)
During pause, rho can reach open complex to separate RNA-DNA hybrid

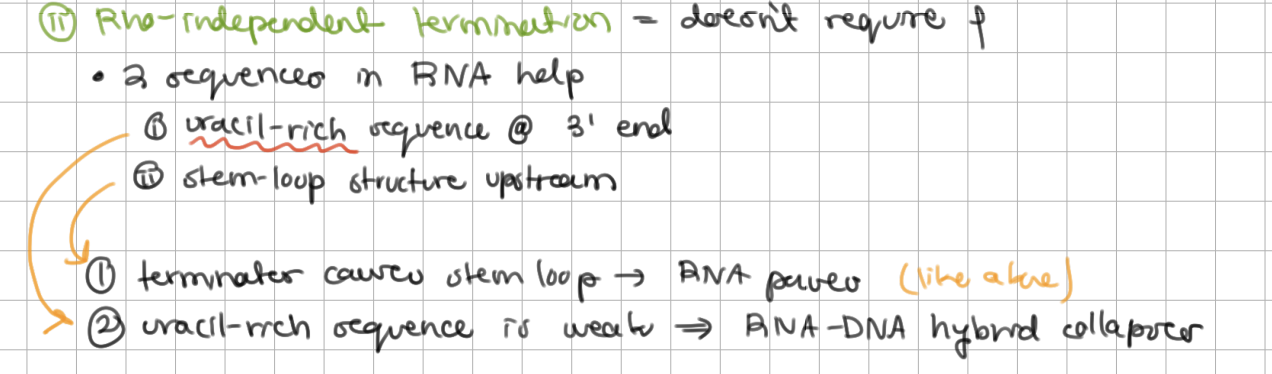
Rho-independent termination = does not require ⍴
RNA contains ______-rich sequence at 3’ end
Like rho-dependent termination, terminator causes stem loop so RNA pol pauses
______-rich sequence is weak => RNA-DNA hybrid collapses without rho
Rho-independent termination = does not require ⍴
RNA contains uracil-rich sequence at 3’ end
Like rho-dependent termination, terminator causes stem loop so RNA pol pauses
uracil-rich sequence is weak => RNA-DNA hybrid collapses without rho
