CH373 Quiz 4
1/69
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
70 Terms
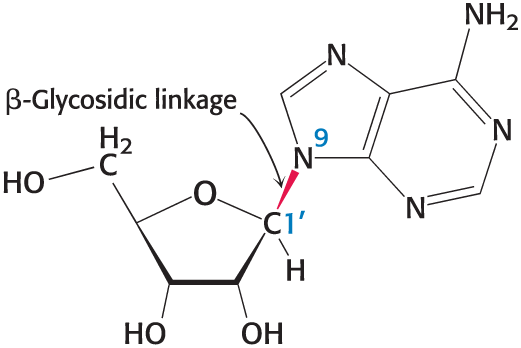
ribose
A five-carbon sugar that forms the backbone of RNA. Ribose is essential for the synthesis of nucleotides.
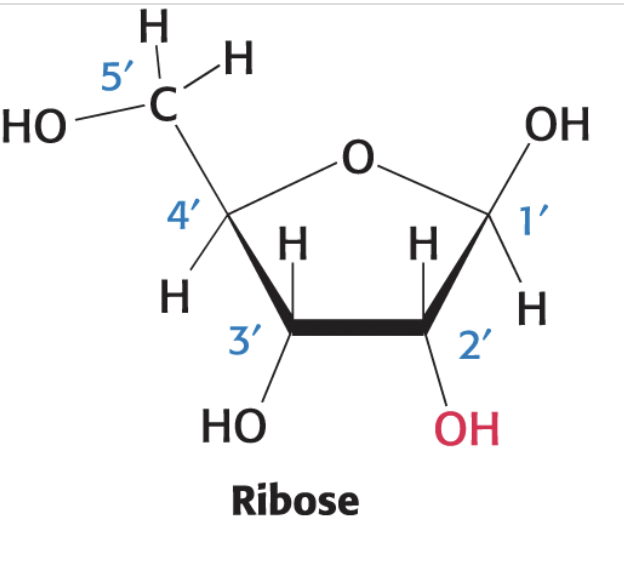
deoxyribose
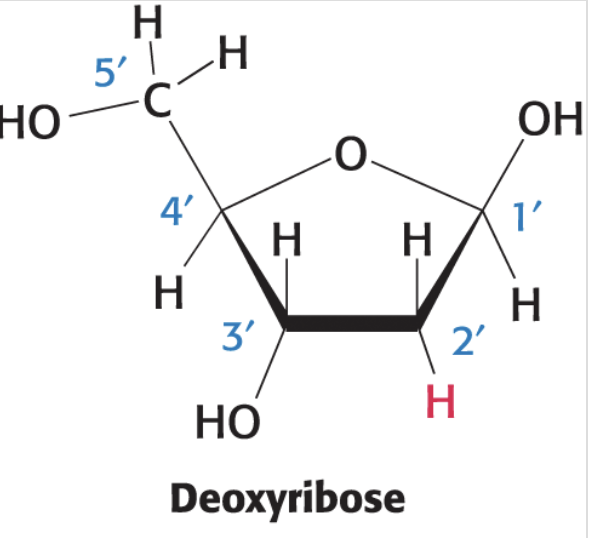
adenine structure
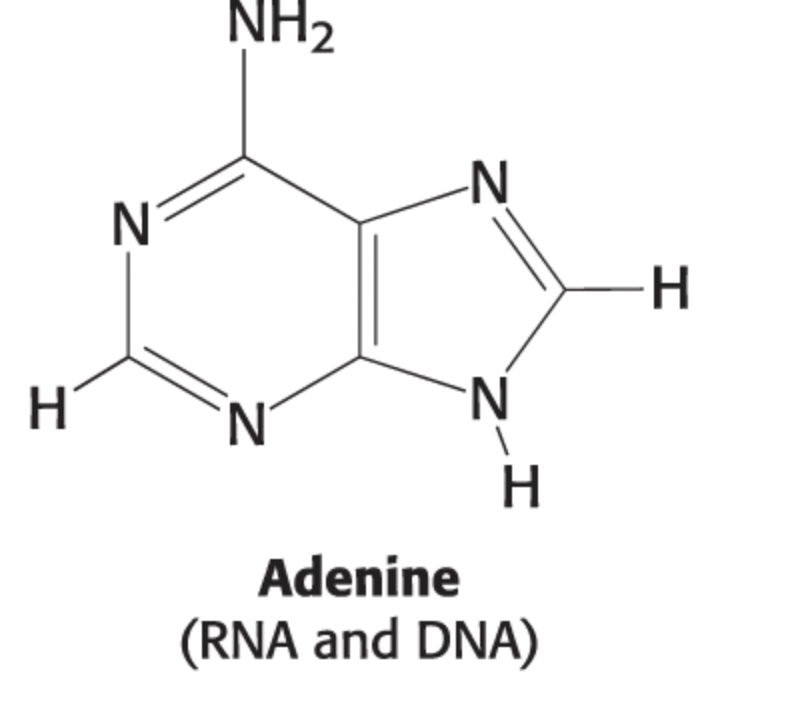
guanine structure
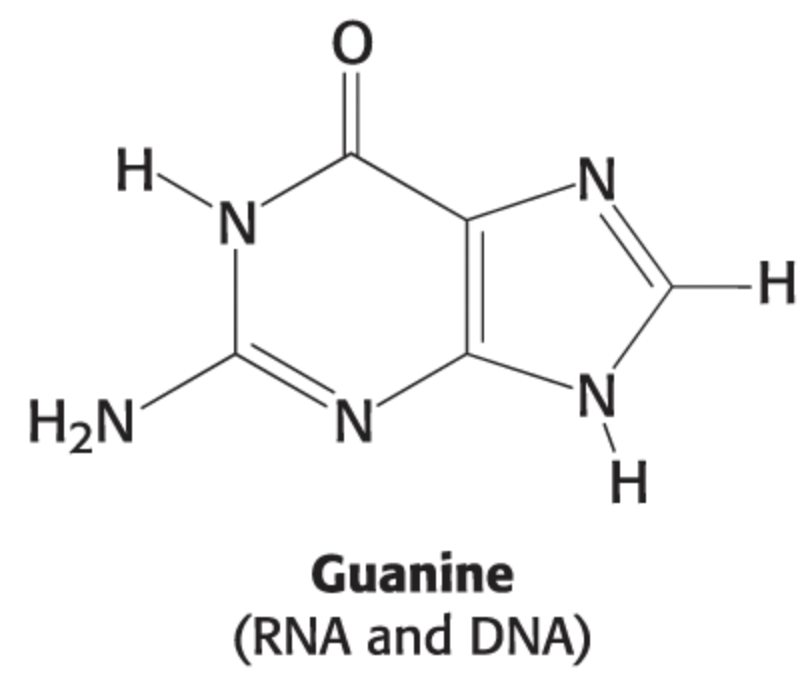
thymine structure
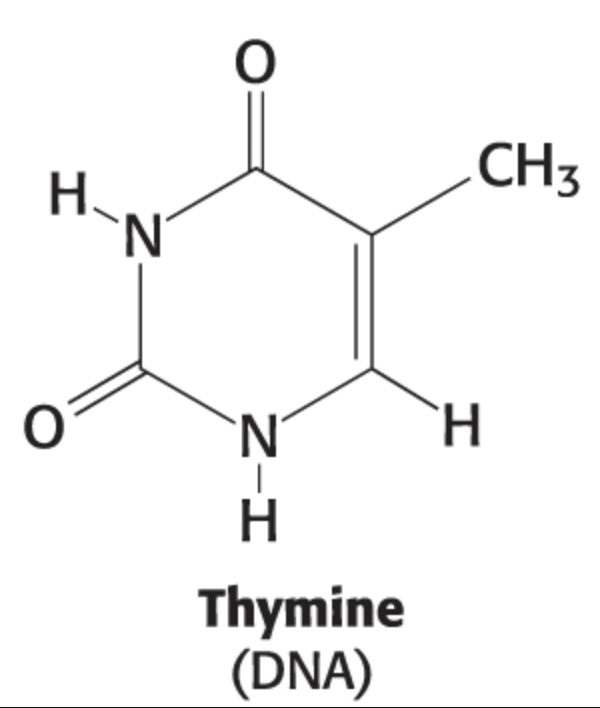
cytosine structure
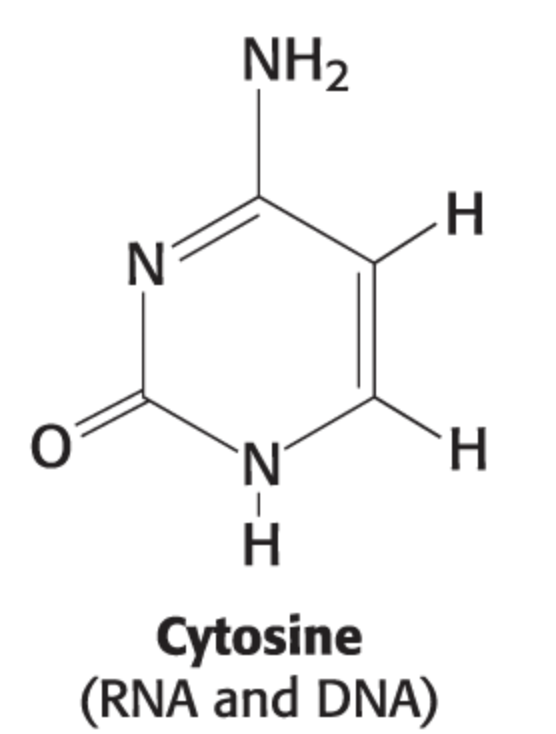
uracil structure
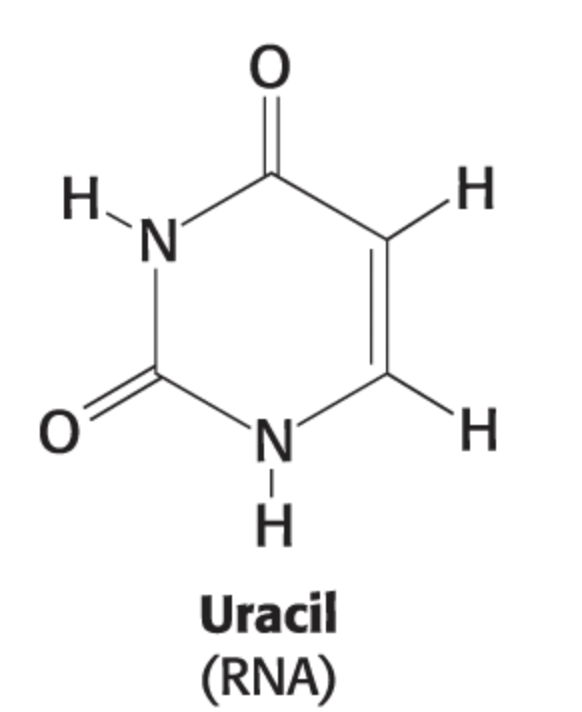
nucleotide vs nucleoside
A nucleotide is a building block of nucleic acids composed of a sugar, a phosphate group, and a nitrogenous base, whereas a nucleoside consists only of the sugar and nitrogenous base, without the phosphate group
Predicting DNA melting temperature (Tm)
higher concentration of C-G = higher melting temp bc they have 2 h-bonds
major vs minor grooves
major groove (24-26) is wider and provides easy access to base pair edges for specific protein binding. The minor groove (12-15) is narrower and less accessible, often used for non-specific structural interactions
A-DNA
right-handed helical structure that is shorter and wider than B-DNA
B-DNA
watsin-crick model
right-handed
Z-DNA
A left-handed double helix in which the backbone phosphates zigzag; can be formed by oligonucleotides with alternating sequences of purines and pyrimidines.
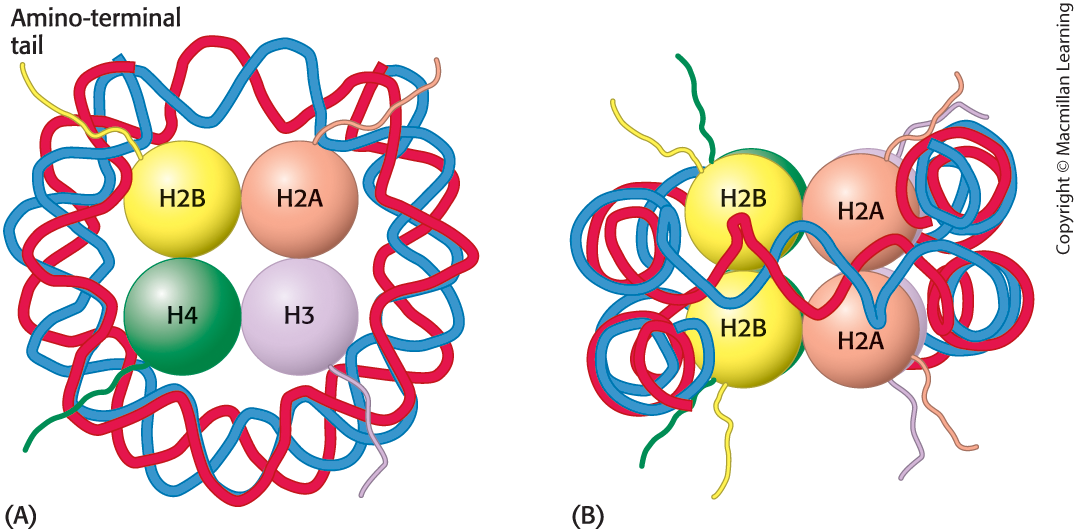
chromatin structure
chromatin consists of nucleosomes, the repeating unit of chromatin that consists of 200 base pairs of DNA and two each of the histones H2A, H2B, H3, and H4.
DNA Polymerase I
Primer removal and DNA repair
5’ —> 3’ and 3’ —> 5’ exonuclease
Removes RNA primers (via exonuclease activity) and replaces them with DNA nucleotides
DNA Polymerase II
Repair
3 > 5 exonuclease
Attachment of bulky hydrocarbons to bases
DNA Polymerase III
Replication - The main enzyme that adds nucleotides in the 5’ to 3’ direction to synthesize new DNA strands
Polymerase IV and Polymerase V
Repair
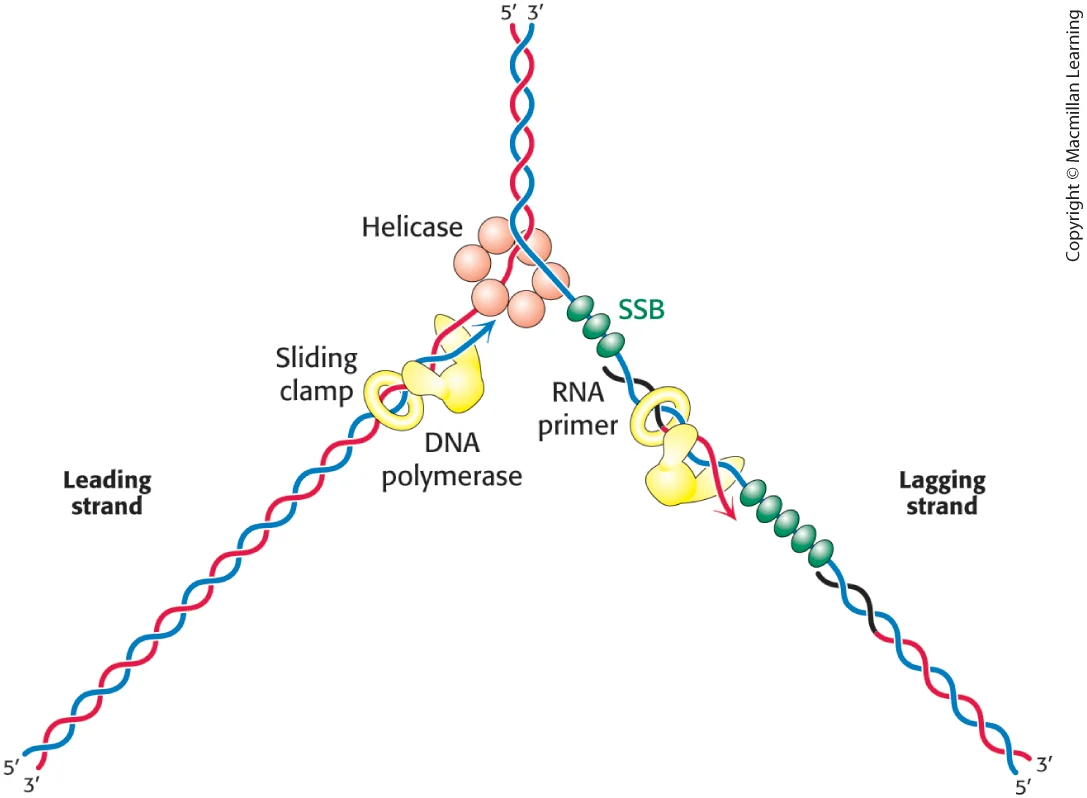
DNA Helicase:
Unwinds the double helix by breaking hydrogen bonds between complementary base pairs, creating a replication fork
DNA Topoisomerase (or Gyrase):
Relieves overwinding/supercoiling strain ahead of the replication fork by cutting and rejoining DNA strands
Single-Strand Binding Proteins (SSBs):
Stabilize the unwound single-stranded DNA, preventing it from reannealing
RNA Primase:
Synthesizes short RNA primers that provide a 3'-OH group for DNA polymerase to begin building the new strand
DNA Ligase:
Joins Okazaki fragments together on the lagging strand by forming phosphodiester bonds.
Telomerase:
Extends the telomeres at the ends of linear chromosomes, preventing genetic information loss
this is what shortens as a person ages
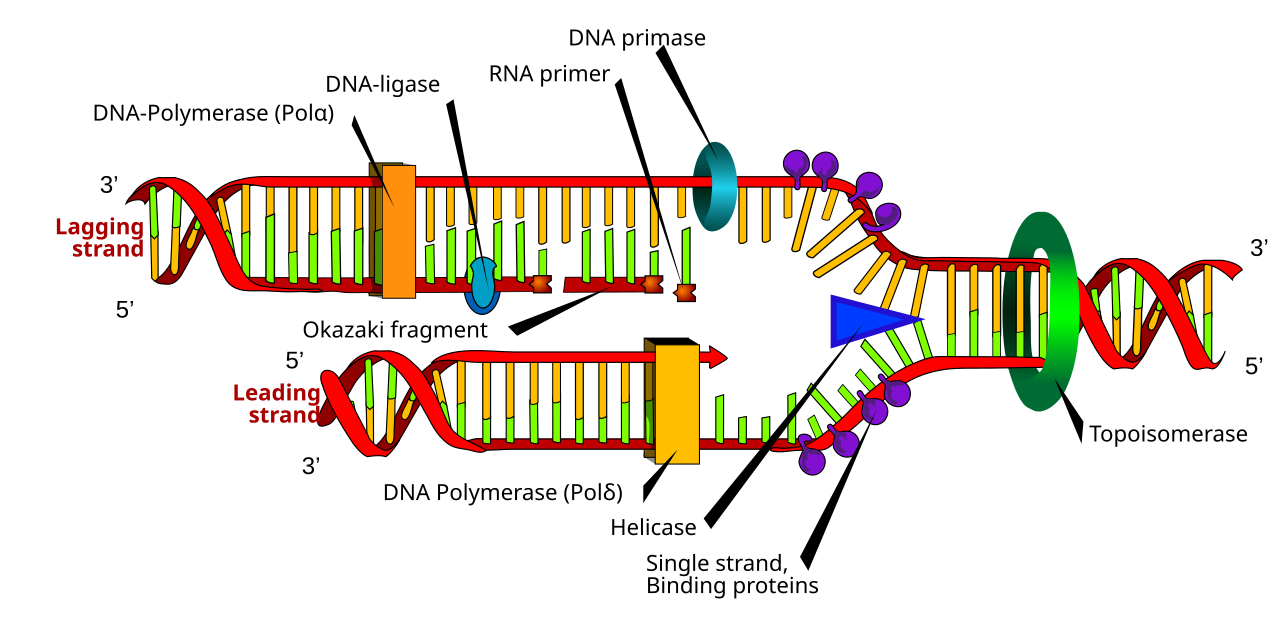
DNA replicaiton diagram
Leading Strand: The strand synthesized continuously in the 5' to 3' direction towards the replication fork by DNA polymerase.
Lagging Strand: The strand synthesized discontinuously in short segments (Okazaki fragments) away from the replication fork.
Initiation of DNA Replication
starts at ORIs
Origin Recognition: Initiator proteins (like DnaA in bacteria) recognize and bind to specific nucleotide sequences known as origins of replication.
Unwinding: Helicase enzymes are recruited to these sites to break hydrogen bonds, unwinding the double helix and forming a replication bubble with two active forks moving in opposite directions.
Primer Loading: DNA primase binds to helicase, forming a primosome, which adds short RNA primers needed to start DNA polymerase activity
Termination of DNA Replication
Fork Convergence: Replication forks move away from the origin, synthesizing new DNA, until they encounter a converging fork or reach the end of the chromosome.
Primer Removal and Ligation: RNA primers are removed, gaps are filled with DNA, and DNA ligase seals the nicked sugar-phosphate backbone, creating continuous strands.
Replisome Disassembly: The replication machinery (replisome) is disassembled.
Resolution of Daughter Strands: In circular prokaryotic chromosomes, Topoisomerase IV separates the interlocked (concatenated) chromosomes. In eukaryotes, topoisomerases handle the untwisting of the DNA.
Modification to the Replication Process in Eukaryotes vs. Prokaryotes
Eukaryotic DNA replication differs from prokaryotic replication in several key aspects:
Multiple Origins of Replication: Eukaryotic chromosomes are linear and larger than prokaryotic circular chromosomes, which allows multiple origins of replication on each chromosome to facilitate faster DNA synthesis. Prokaryotes typically have a single origin.
Complex DNA Packaging: Eukaryotic DNA is wrapped around histone proteins to form nucleosomes, which need to be unwrapped during replication. This adds complexity to the replication process compared to prokaryotes that do not have this level of chromatin organization.
More Enzymes and Proteins: Eukaryotic DNA replication involves additional proteins and enzymes, such as multiple DNA polymerases (e.g., Pol α, β, γ, δ, ε), unlike the single DNA polymerase commonly found in prokaryotes.
RNA Primers: In eukaryotes, RNA primers are later removed and replaced by DNA, requiring complex repair mechanisms; prokaryotes handle this slightly differently.
Telomere Replication: Eukaryotes utilize telomerase to extend telomeres at the end of linear chromosomes to prevent genetic loss, a feature not applicable to the circular chromosomes of prokaryotes.
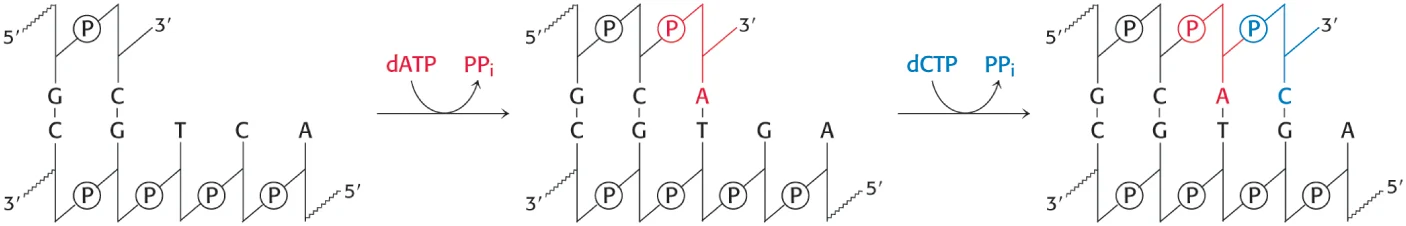
DNA polymerases catalyze the step-by-step addition of … to a DNA strand
deoxyribonucleotides
deoxynucleoside 5’ triphosphate dATP, dGTP, dCTP, and TTP—as well as the Mg2+ ion.
replication fork
trinucleotide repeats
Found in stretches of DNA in which a trinucleotide sequence is repeated many times; these segments of DNA can expand in the course of DNA replication, causing such genetic diseases as Huntington disease.
mutagens
Perturbs the base sequence of DNA and causes a mutation; often chemical but can also be energy sources such as ultraviolet light.
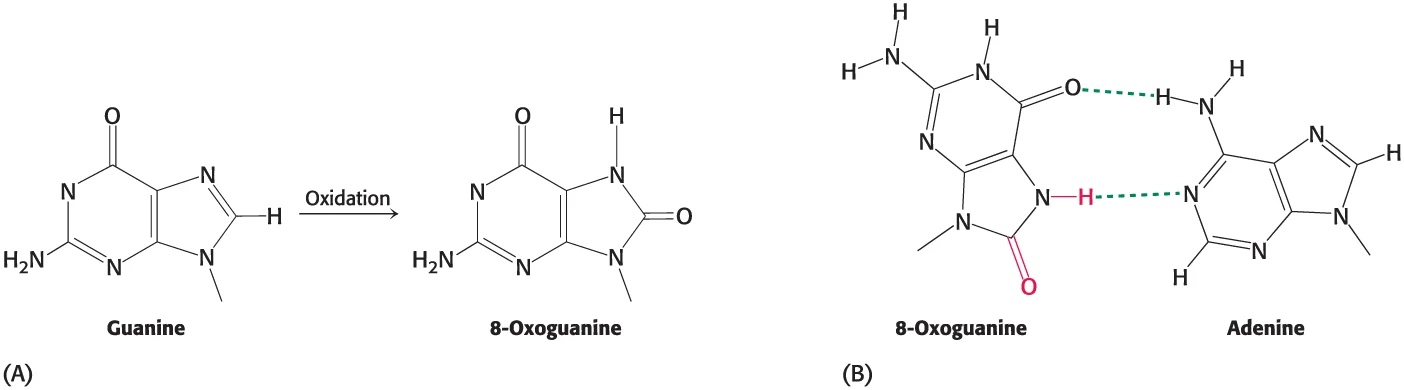
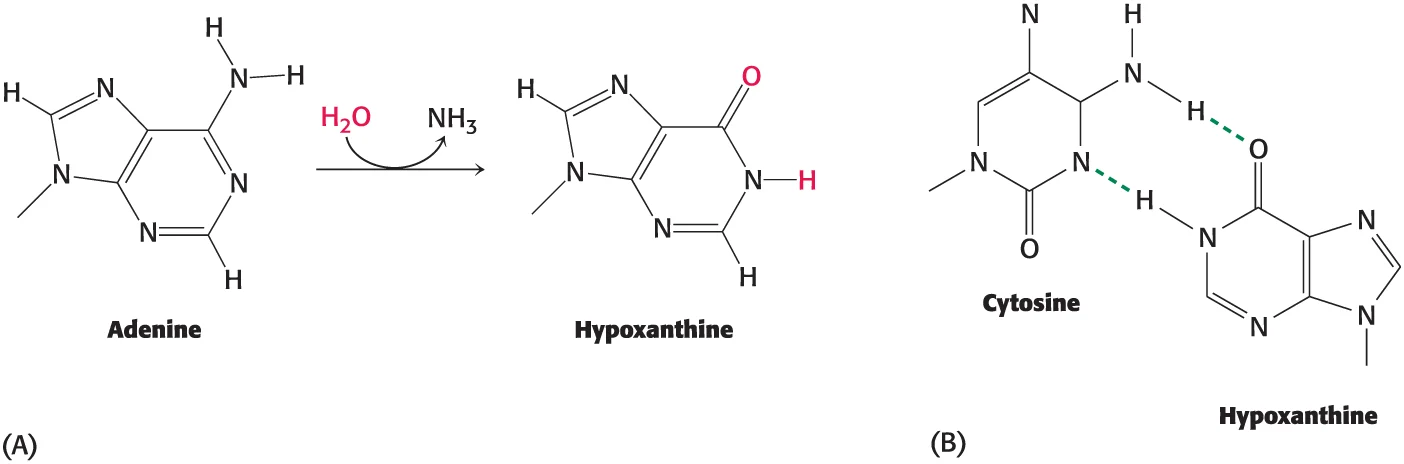
Deamination + Oxidation
Deamination is the removal of an amino group from an amino acid or nucleotide
Oxidation, on the other hand, refers to the addition of oxygen or the removal of hydrogen to or from a molecule (formation of 8-oxoguanine from guanine)
DNA Damage During Replication
Mismatch base pairs: Incorrect nucleotides incorporated by DNA polymerase, leading to mismatches (e.g., G pairing with T).
Insertion/Deletion mutations: Strand slippage causing the addition or removal of nucleotides.
Single-Strand Breaks (SSBs): Occur due to fork stalling, collision with transcriptional machinery, or unrepaired nicked DNA.
Double-Strand Breaks (DSBs): Arise when a replication fork encounters a SSB and collapses, resulting in highly deleterious DNA fragmentation.
DNA Damage After/Independent of Replication (Spontaneous/Exogenous)
Base Oxidation: Reactive oxygen species (ROS) damage bases, commonly forming 8-oxo-7,8-dihydro-2′-deoxyguanosine (8-oxo-dG).
Deamination: Loss of amine groups, such as cytosine converting to uracil, causing point mutations.
Depurination/Depyrimidination: Hydrolysis of the glycosidic bond, leaving an "AP site" (apurinic/apyrimidinic).
Alkylation: Addition of alkyl groups to bases, often by toxic compounds.
Bulky Adducts: Addition of large chemical groups, such as benzo[a]pyrene from cigarette smoke, which distort DNA structure.
Pyrimidine Dimers (UV Damage): UV light induces covalent bonds between adjacent thymine or cytosine bases, causing physical blocks to replication.
DNA Crosslinking: Covalent linkages between two strands (interstrand) or within one strand (intrastrand)
DNA repair
protein MutS (or homolog MSH) scans DNA and binds to the distorting mismatch
MutL (or MLH/PMS) is recruited to the complex, activating an endonuclease (MutH in E. coli).
Exonuclease enzymes remove a segment of the daughter strand containing the incorrect base, and DNA polymerase fills the gap
DNA ligase ligates ends together
Importance of DNA Methylation in Repair
protein MutH binds to these hemimethylated sites and only nicks the unmethylated (daughter) strand, ensuring the error is removed rather than the correct template
so basically the methylated strand is the correct one
Endonucleases
initiate repair by cutting out damaged sections or recognizing mismatches
internal to DNA
Exonucleases
Following endonuclease cleavage, exonucleases "nibble" or degrade the damaged DNA strand starting from the cut site. They also proofread during DNA replication, removing mispaired bases
initiation of transcription
RNA polymerase (with a sigma factor in bacteria, or general transcription factors in eukaryotes) recognizes and binds to specific consensus sequences in the promoter region, such as the TATA box
also unwinds helix
elongation of RNA transcript
RNA polymerase moves along the DNA template strand, synthesizing the RNA by adding ribonucleoside triphosphates (NTPs) complementary to the DNA template. This addition occurs in the 5' to 3' direction, with RNA polymerase unwinding the DNA ahead of it and rewinding it behind as the RNA transcript is formed
template strand
is the 3' to 5' DNA strand that RNA polymerase uses as a guide to synthesize messenger RNA (mRNA) in the 5' to 3' direction
noncoding strand or antisense
Explain and compare the two ways in which transcription can be terminated
Rho-independent (Intrinsic) Termination
DNA contains a region rich in Cytosine (C) and Guanine (G) followed by a stretch of Adenine (A) nucleotides. When transcribed, the C-G rich RNA region folds back on itself to form a stable hairpin loop structure causing release
no ATP
Rho-dependent Termination
Rho binds to a specific, naked sequence on the nascent (growing) RNA chain called the Rho utilization (rut) site. The Rho protein, a ring-shaped ATPase/helicase, travels along the RNA transcript towards the RNA polymerase
requires ATP
RNA Polymerase I
transcribes rRNA (ribosomal RNA)
RNA Polymerase II
transcribes mRNA (messenger RNA).
RNA Polymerase III
transcribes tRNA (transfer RNA).
Compare and contrast DNA and RNA polymerases
DNA Polymerase
DNA replication
deoxyribonucleotides (dATP, dTTP, dCTP, dGTP)
requires primer (Requires free 3’ OH)
requires helicase/topoisomerase
RNA Polymerase
transcription
Ribonucleotides (ATP, UTP, CTP, GTP)
Intrinsic helicase activity
de novo synthesis
BOTH synthesize new strands in the 5’ → 3’ direction
Compare and contrast the structural features of DNA and RNA, and explain how these similarities and differences affect replication and transcription.
Double-Strand Stability: The double helix allows for high-fidelity replication where one strand serves as a direct template for the other
INTACT genome
Single-Strand Flexibility: RNA is produced as a single strand by RNA polymerase using one DNA strand as a template, facilitating its role as a temporary message.
Prokaryotic Promoters
-10 (Pribnow) & -35 boxes
Direct binding by singma factor
Polycistronic (multiple genes)
Eukaryotic Promoters
Core (TATA), Proximal, Distal (Enhancers)
Packaged in chromatin/nucleosomes
single gene
List the differences between prokaryotic and eukaryotic gene expression
Prokaryotic gene expression occurs in the cytoplasm, allowing simultaneous transcription and translation (coupled), whereas eukaryotic gene expression is separated, with transcription in the nucleus and translation in the cytoplasm
mRNA modifications
5' Capping: A modified guanine nucleotide (7-methylguanosine) is added to the 5' end, aiding in ribosome binding and protecting the transcript
3' Polyadenylation: A tail consisting of many adenine nucleotides is added to the 3' end, increasing mRNA stability and regulating lifespan in the cytoplasm
RNA Splicing: Introns (non-coding regions) are removed, and exons (coding regions) are joined, sometimes creating different mature mRNA molecules through alternative splicing
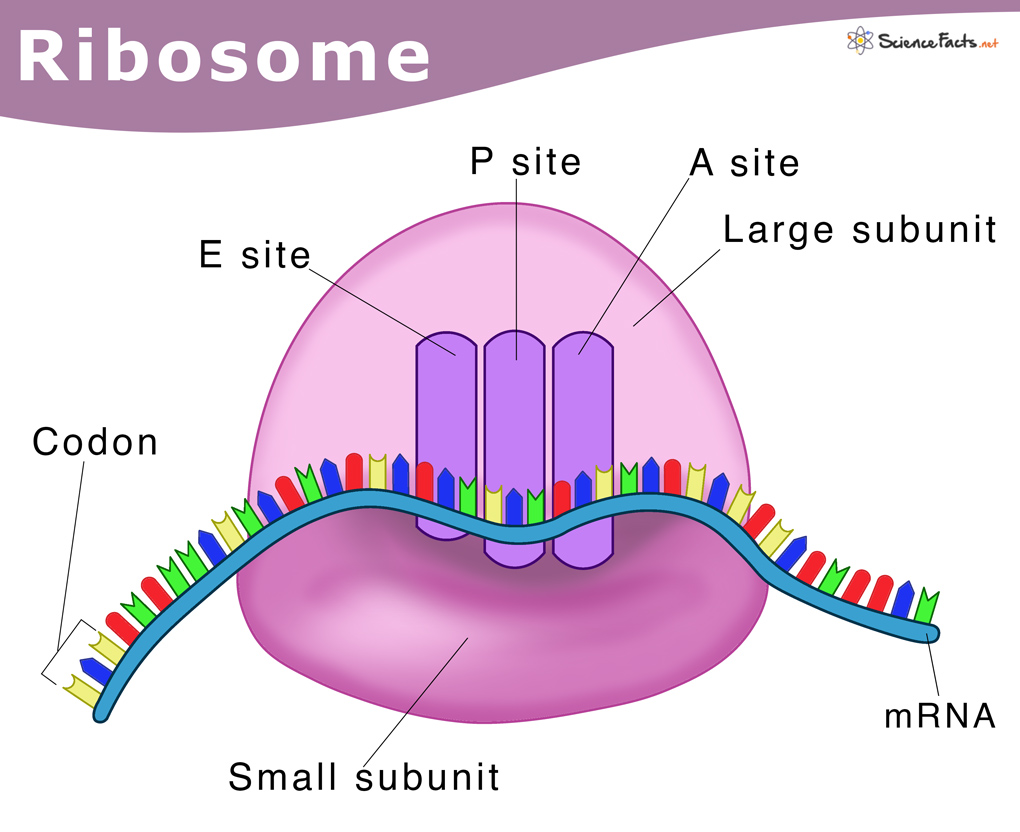
ribosome structure
The small subunit (e.g., 30S) binds to mRNA and ensures fidelity by matching tRNA anticodons to mRNA codons.
The large subunit (e.g., 50S) contains the catalytic site that links amino acids together.
3 tRNA binding sites
A Site (Aminoacyl): Binds incoming tRNA with the next amino acid.
P Site (Peptidyl): Holds the tRNA attached to the growing polypeptide chain.
E Site (Exit): Releases the deacylated (empty) tRNA
tRNA structure
anticodon loop for codon recognition, a 3' acceptor stem for amino acid attachment, and structural loops (D and TC) for ribosomal binding
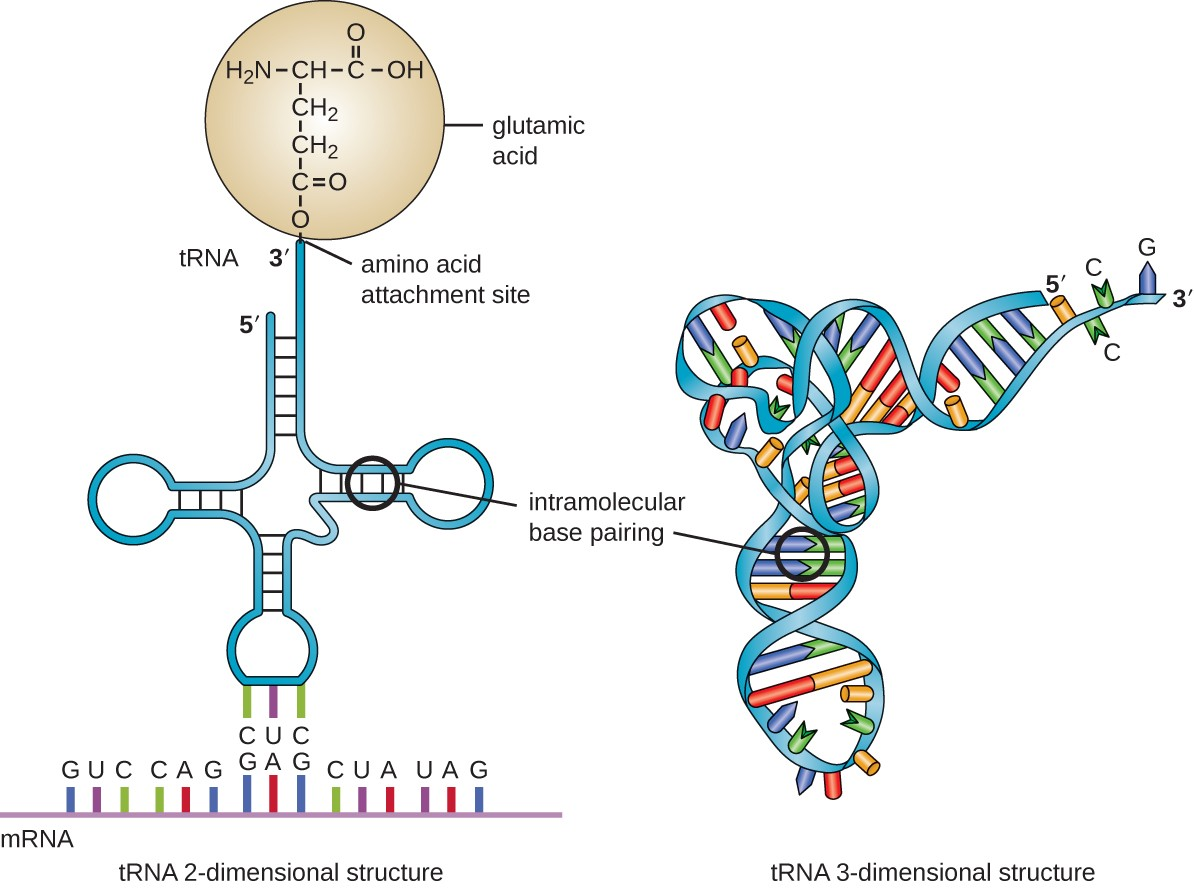
Describe how tRNA synthetases achieve specificity
precise binding of the anticodon loop and acceptor stem, followed by proofreading via hydrolytic active sites that remove mischarged amino acids
wobble hypothesis
the third base of an mRNA codon (the 3' base) can form non-standard, flexible base pairs with the first base of the tRNA anticodon (the 5' base).
This allows a single tRNA to recognize multiple synonymous codons, explaining the degeneracy of the genetic code and reducing the total number of tRNAs needed for translation
translation factors
Translation Factors (Protein factors)
Initiation Factors (IFs/eIFs): Help assemble the ribosome on the mRNA.
Elongation Factors (EFs/eEFs): Assist with adding amino acids to the chain.
Termination Factors (RFs/eRFs): Recognize stop codons to release the polypeptide.
GTP must be hydrolyzed
Describe how polymerization of the polypeptide chain is achieved
Polypeptide chain polymerization is achieved through dehydration synthesis (condensation), where the amino group (NH2) of one amino acid joins the carboxyl group (COOH) of another, forming a peptide bond and releasing a water molecule (H2O)
ribosome proofreading
Aminoacyl-tRNA arrives at the ribosome's A-site within a ternary complex containing elongation factor Tu (EF-Tu) and GTP
The ribosome's decoding center (in the 30S subunit) monitors the codon-anticodon pairing. If the match is correct (cognate), it induces a structural change (lock-in) that triggers GTP hydrolysis on EF-Tu.
If the tRNA is incorrect (near-cognate), it does not bind tightly. The incorrect tRNA is often ejected before GTP is hydrolyzed
Following GTP hydrolysis, EF-Tu changes its conformation and releases the tRNA.
GTP HYDROLYSIS
elongation and termination in ribosome
Elongation
Decoding: A tRNA with an anticodon binds to the mRNA codon in the ribosome's A site, aided by elongation factors.
Peptide Bond Formation: The ribosome (peptidyl transferase) links the amino acid from the A site to the polypeptide chain on the P site.
Translocation: The ribosome moves one codon forward; the empty tRNA moves to the E (exit) site and is released, moving the peptide-carrying tRNA to the P site.
Termination
Stop Codon Recognition: A stop codon (UAA, UAG, UGA) enters the A site.
Release Factors: Release factors bind to the A site instead of a tRNA.
Polypeptide Release: The completed polypeptide is detached from the P-site tRNA, and the ribosomal complex dissociates
Describe the role of signal peptides for secretory proteins
Signal peptides are short (16–30 amino acids) N-terminal sequences that act as "zip codes" to direct newly synthesized proteins toward the secretory pathway
signaling to the ER for packaging or the membrane for transport etc.
translation of secretory protein VS from proteins that are not secreted into the cell
Secretory proteins are translated on the rough endoplasmic reticulum (ER) via cotranslational translocation, guided by a signal peptide and signal recognition particle (SRP) to the ER membrane.
secretory proteins are inserted into the ER lumen, cleaved, and folded in an oxidizing environment, followed by Golgi transport
cytosolic proteins translated on free ribosomes
Restriction enzymes
Types of Restriction Enzymes
Type I: Cleave randomly away from the recognition site and are multifunctional.
Type II: Cut within or near their specific recognition sequence; used most often in laboratories.
Type III: Cut outside the recognition sequence and are large enzymes.
Type IV: Recognize modified, typically methylated, DNA.
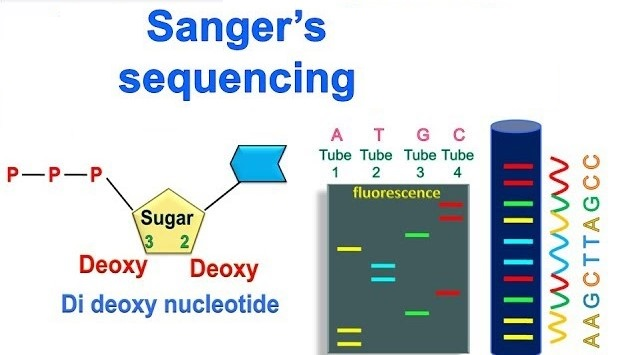
Sanger sequencing
Template DNA is mixed with a primer, DNA polymerase, normal nucleotides (dNTPs), and fluorescently labeled dideoxynucleotides (ddNTPs).