mmg2040 lecture 24 - rna sequencing
1/7
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
8 Terms
Explain the purpose of RNA-seq
RNA-sequencing: A method to sequence and quantify RNA molecules
Provides a snapshot of the transcriptome (all RNA molecules in a cell at a given time including: m/r/tRNA and noncoding RNA)
Dynamic —> changes conditions, cell types, diseases
RNA shows active gene expression vs. DNA showing potential gene expression
reveals functional output of genome, regulatory changes
Answers:
Which genes are expressed?
How much are they expressed?
Are there alternative transcripts?
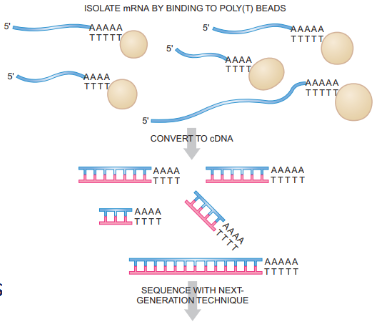
Outline the workflow for RNA-seq
RNA isolation - High-quality RNA is critical
Avoid degradation (RNases!)
Quality check: Integrity + Purity
mRNA enrichment - 2 methods
Poly-A selection - Captures mRNA via poly-A tail
rRNA depletion - Removes abundant rRNA
cDNA synthesis - fragment, reverse transcribe
add adapters + bar codes
Library preparation - PCR amplification, Size selection
Add sequencing adapters
Sequencing - typically short-read sequencing (e.g., Illumina)
Generates millions of reads
Each read corresponds to a fragment of RNA
Data analysis - Reads mapped to reference genome OR transcriptome
Software determines where reads came from
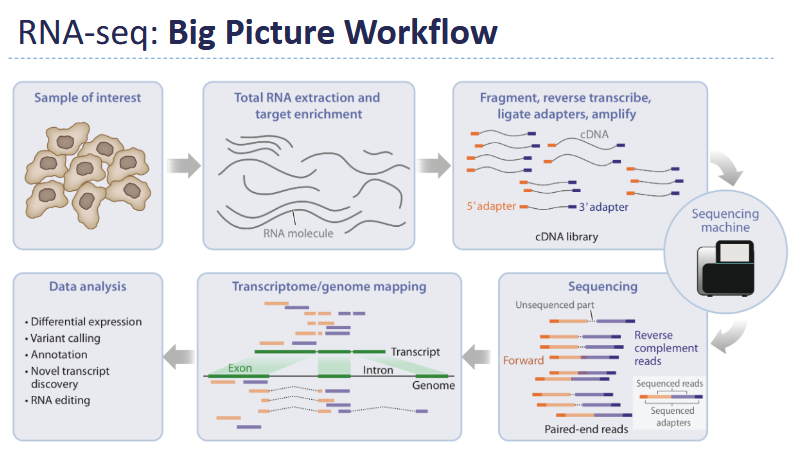
Outline the workflow for scRNA-seq
Key Idea - Measures gene expression in individual cells, not bulk populations
Bulk RNA-seq = average signal
scRNA-seq reveals: Cell-to-cell variation, rare cell types, cell states (e.g., differentiation)
Isolate individual cells - capture RNA from each cell
Microfluidic chip - traps one cell per capture site
Cell lysis, reverse transcription, and amplification within its own chamber
Convert to cDNA - add cell-specific barcodes
Pool and sequence
Assign reads back to original cells
Critical Concept —> Each read includes:
Gene information
Cell barcode → tells you which cell it came from
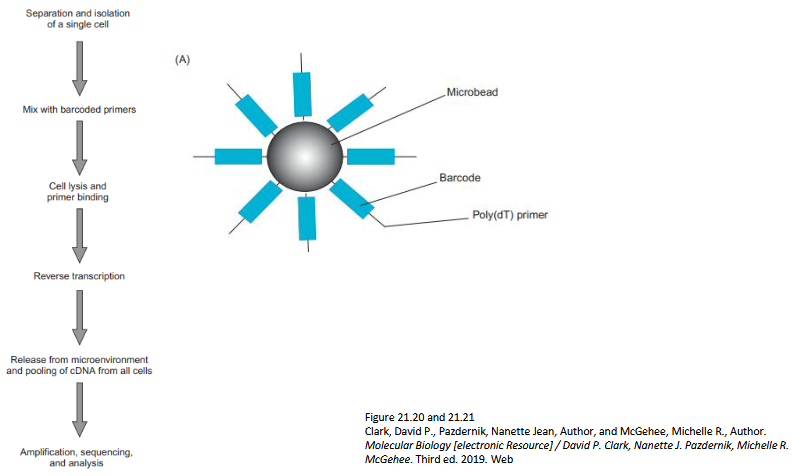
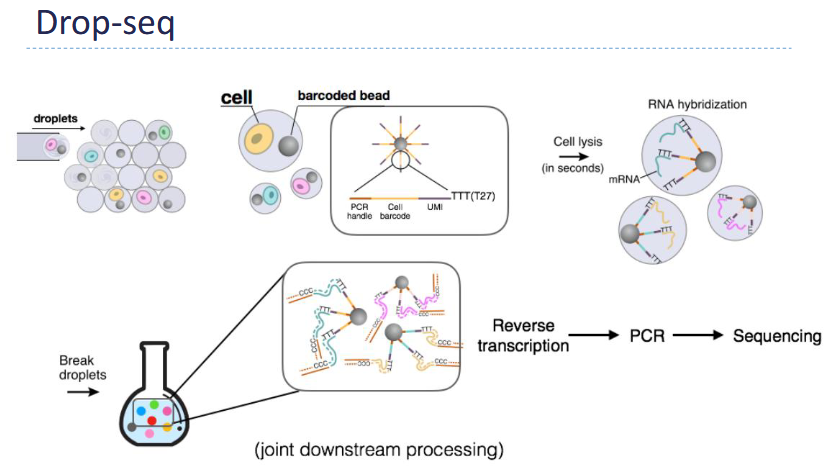
Outline the workflow for Drop-seq
A version of single-cell RNA seq (scRNA-seq)
Core Idea - Encapsulate: 1 cell + 1 bead + reagents inside tiny oil droplets
Inside Each Droplet, a bead carries: Unique barcode + Oligo-dT primers (bind mRNA)
Key Advantage: Thousands of cells processed simultaneously —> High-throughput scRNA-seq
Process:
Cell lyses → mRNA binds bead
cDNA made with barcode attached
All droplets pooled → sequenced together
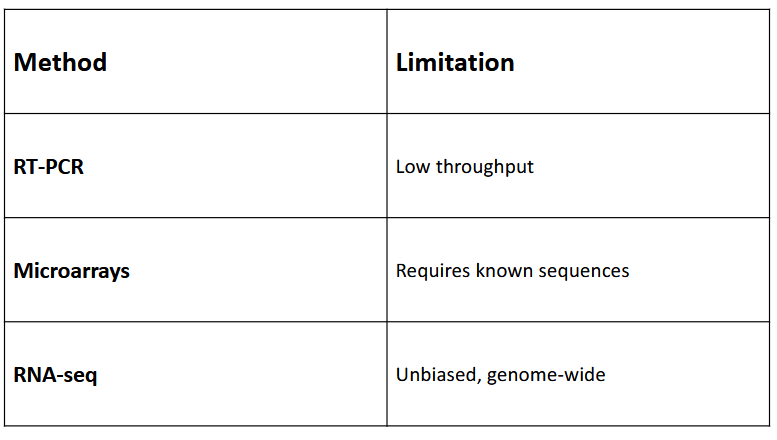
Distinguish RNA-seq from older methods (e.g., microarrays
RT-PCR: low througput
Microarrays: Requires known sequences
RNA-seq: Unbiased, genome-wide
Interpret basic RNA-seq outputs
basic RNA seq: Raw reads are converted into standardized units to compare gene expression across samples
Quantifying Gene Expression - Count number of reads per gene
More reads = higher expression
Normalization needed: Gene length + sequencing depth
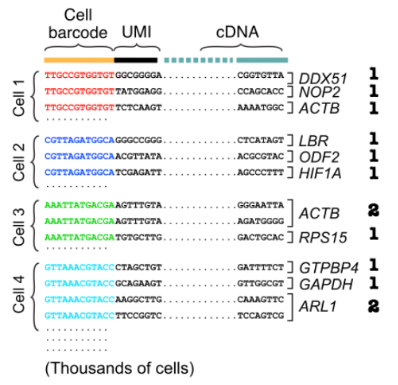
Drop seq: After obtaining sequencing reads consisting of cell barcode, UMI and cDNA
First group reads by cell barcode before aligning cDNA reads
counting unique molecules per cell per gene using the UMIs
estimate the transcript abundances
Describe key applications in research and medicine
Strengths of RNA-seq: Unbiased, Genome-wide, Detects novel transcripts, High sensitivity
Strengths of scRNA-seq: Measures gene expression in individual cells, not bulk
populations —> Bulk RNA-seq = average signal
reveals: Cell-to-cell variation, rare cell types, cell states (e.g., differentiation)
Strengths of drop-seq: Identifies rare cell types, Maps cell differentiation pathways, Reveals heterogeneity in tumors
sc/RNA seq limitations
limitations to rna seq:
Expensive
Requires bioinformatics
Sensitive to RNA quality
Snapshot in time
Limitations to scRNA seq:
Lower RNA capture per cell
More technical noise
Complex data analysis