protein synthesis, PTM, and transport
1/13
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
14 Terms
tRNA
provides a physical connection between the DNA-RNA genetic code and amino acids
the driving force for protein synthesis; allows for formation of peptide bond to be thermodynamically favorable
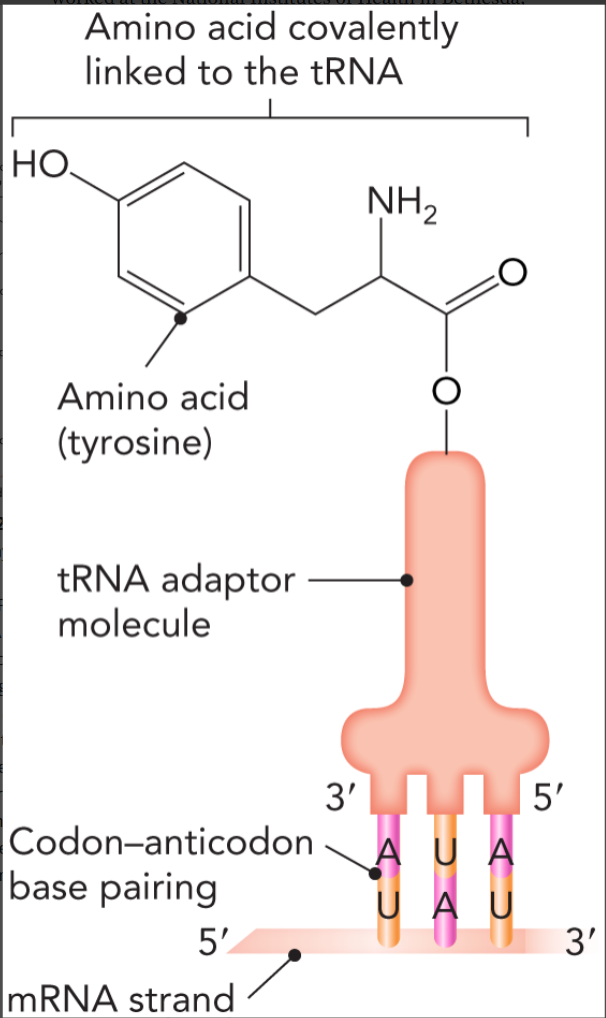
aminoacylation
aminoacyl-tRNA synthetase attaches a specific amino acid to the 3’ acceptor stem of the cognate tRNA (charges the tRNA); coupled to ATP hydrolysis
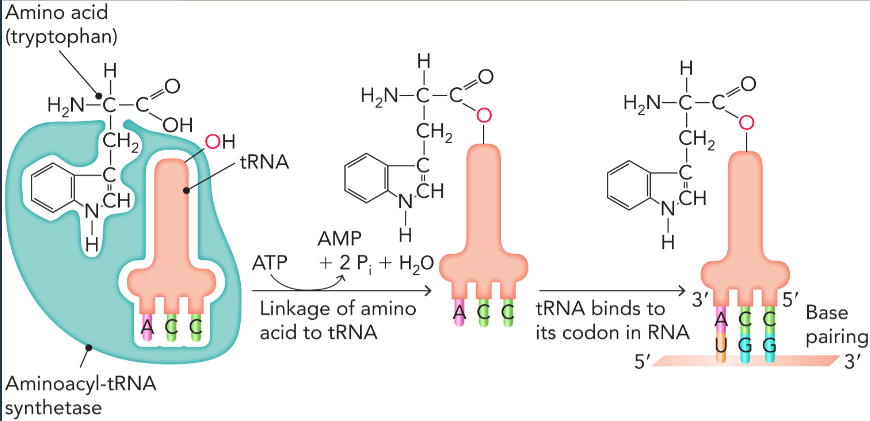
aminoacyl-tRNA synthetase reaction
bond is formed between the carboxyl group of the amino acid and the alpha phosphate of ATP to form an aminoacyl-adenylate
hydrolysis releases pyrophosphate
phosphodiester bond is cleaved to form aminoacyl tRNA and amino acid is transferred to the 2’hydroxyl or 3’hydroxyl of the tRNA acceptor stem
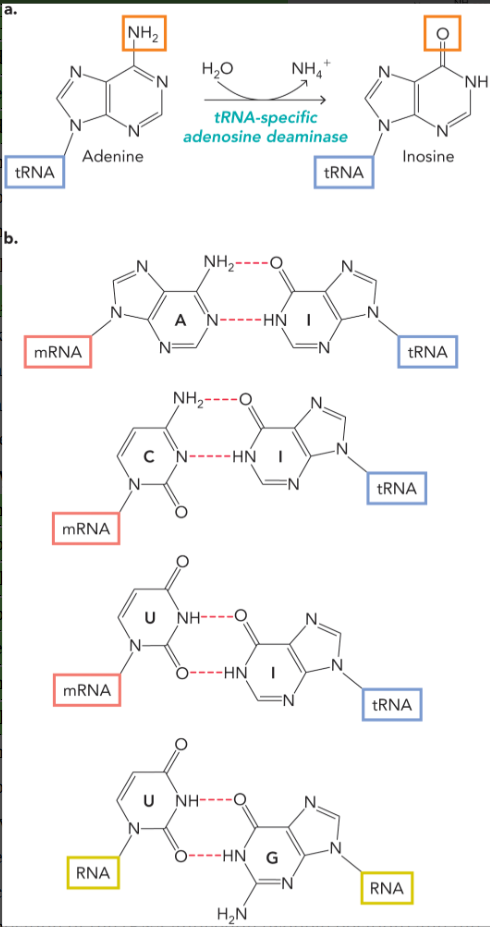
wobble hypothesis
certain nucleotide bases in the 5’-nucleotide of the anticodon could undergo noncanonical hydrogen bonding with the nucleotide base at the 3’ end of the mRNA codon
noncanonical base pairing
A-I
C-I
U-I
U-G
inosine is mostly found in tRNA (very rare in mRNA)
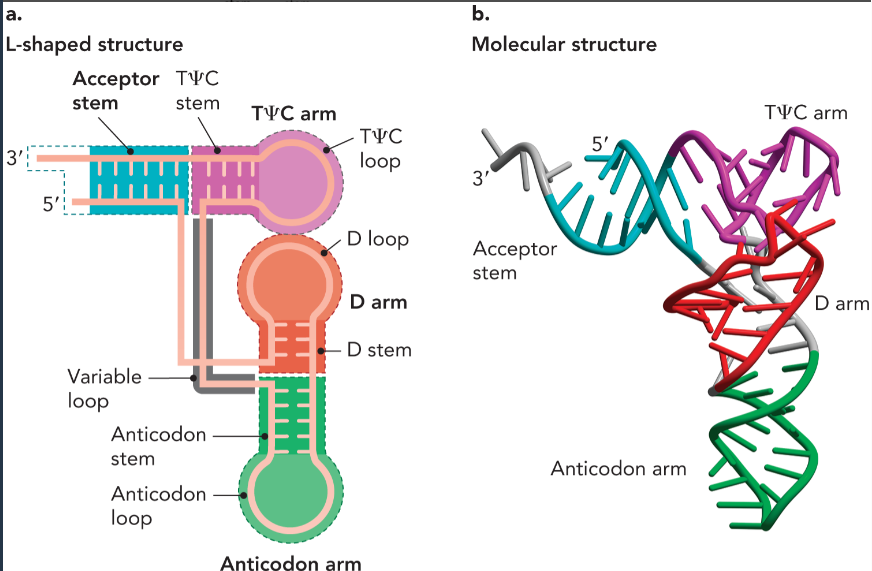
tRNA structural components
acceptor stem- site of aminoacylation
anticodon arm- forms base pairs with corresponding nucleotide bases in mRNA
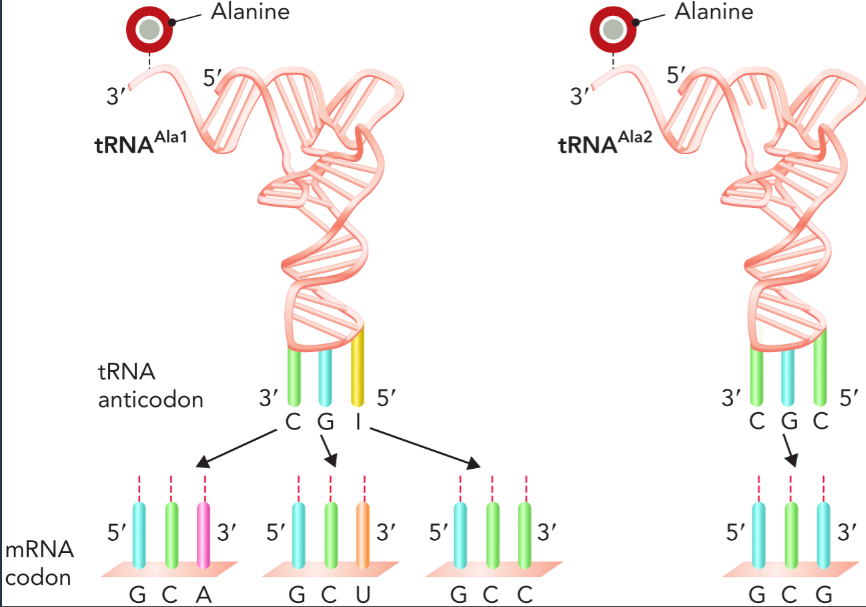
isoacceptor tRNAs
a group of related tRNA molecules that contain the same linked amino acid but bind different codons; can also be a single tRNA that binds to different codons because of the 5’ wobble position in the tRNA
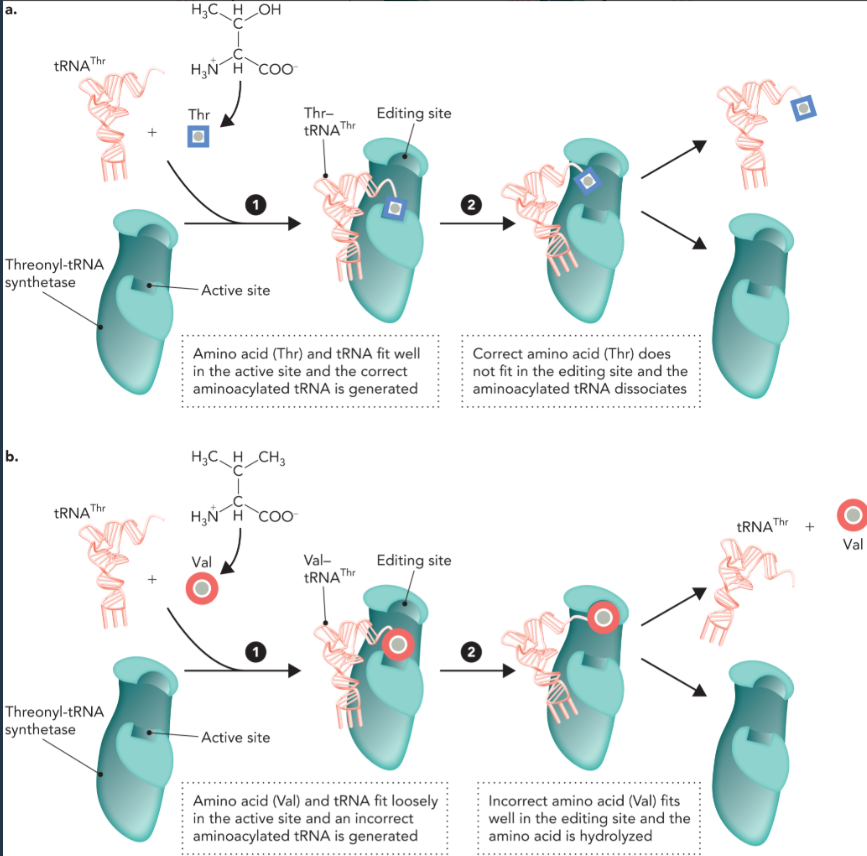
proofreading functions of tRNA synthetases
if the correct tRNA and cognate amino acid bind to the active site in step 1 then aminoacylation occurs in the active site and the aminoacylated tRNA is not hydrolyzed by the editing site in step 2
when a chemically similar but incorrect amino acid binds loosely to the active site with the same tRNA in step 1, then the aminoacylated tRNA is still generated. But the incorrect amino acid is hydrolyzed at the editing site in step 2
four key steps of polypeptide synthesis
initiation
elongation
termination
ribosome recycling
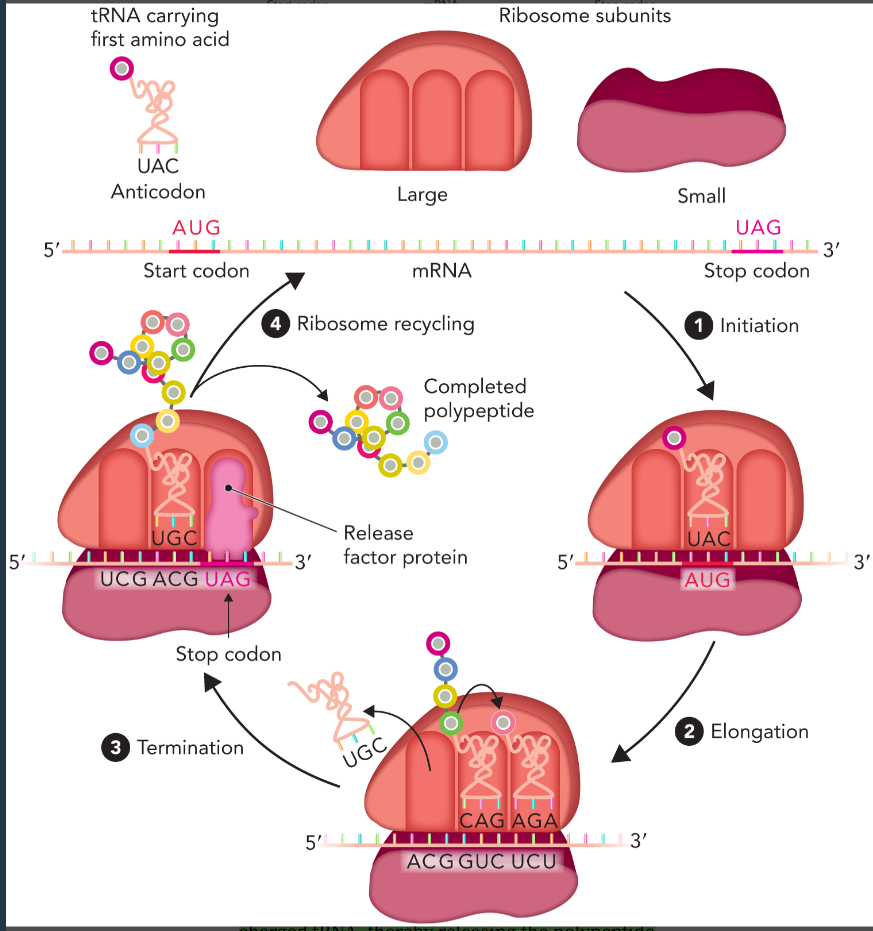
three tRNA binding sites
aminoacyl site (A site)
peptidyl site (P site)
exit site (E site)
A and P are positioned so that the anticodon of a charged tRNA is able to make contact with mRNA
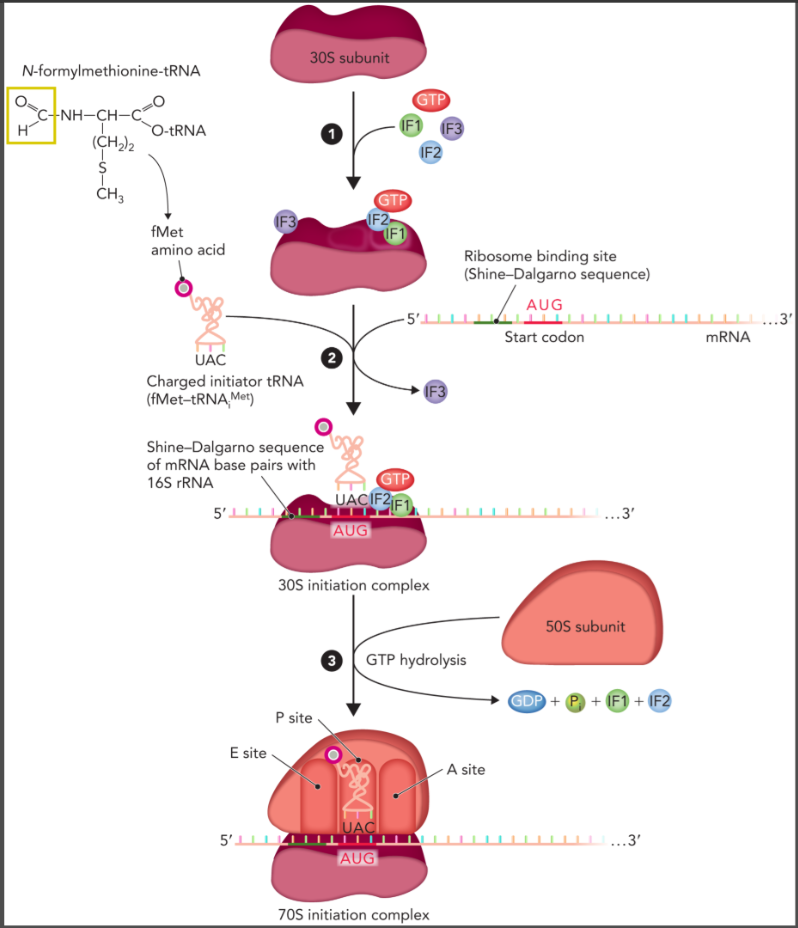
prokaryotic initiation
GTP and three initiation factors bind to the 30S ribosomal subunit
mRNA and initiator tRNA charged with methionine and formylated fMet-tRNAiMet binds to complex with the shine-dalgarno sequence of mRNA aligning with complementary sequences in the rRNA in the 30S subunit
GTP hydrolysis allows binding of the 50S subunit and initiation factors are released to form 70S initiation complex
fMet-tRNAiMet is bound in the P site
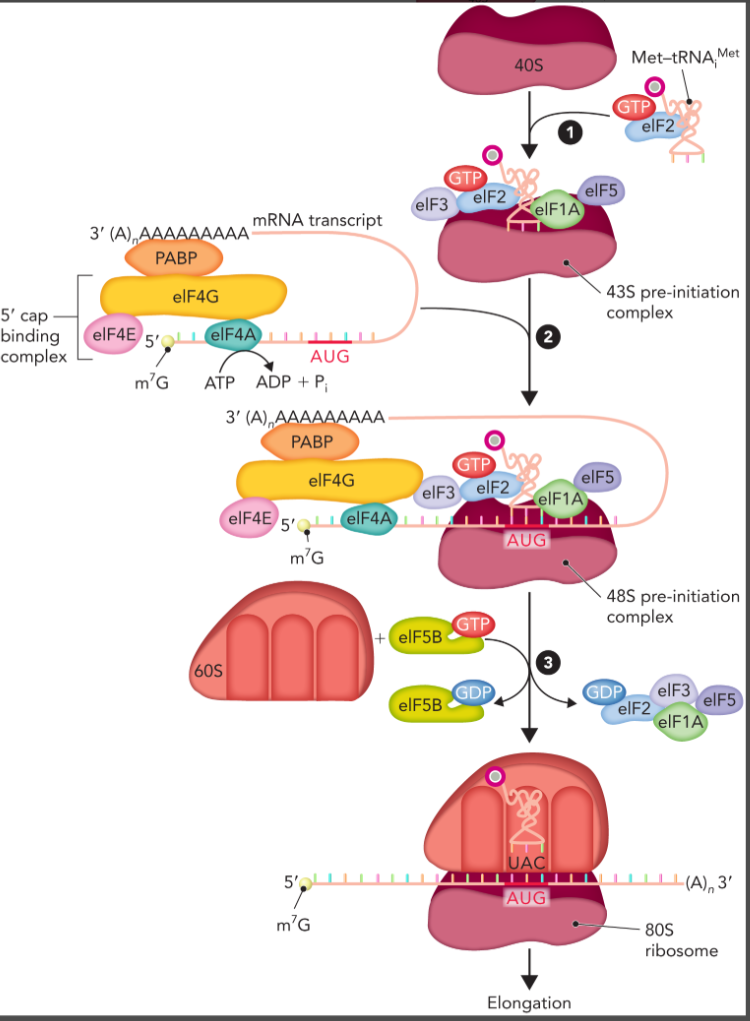
eukaryotic initiation
43S pre-initiation complex is formed between 40S subunit and a complex consisting of initiation factors, GTP, and the aminoacylated Met-tRNAiMet
the mRNA transcript with a RNA binding proteins associated with the 5’ cap and poly(A) tail associates with the 43S to form the 48S pre-initiation complex
the 60S subunit associates resulting in the exchange of initiation factor proteins and formation of the fully assembled 80S complex
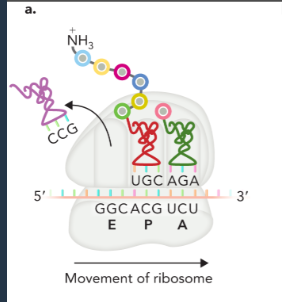
elongation
similar in prokaryotes and eukaryotes
six steps that requires the hydrolysis of 2 GTP, one to promote binding of the AA-tRNAAA to the A site, and one to facilitate translocation of the ribosome in the 3’ direction
uses elongation factors
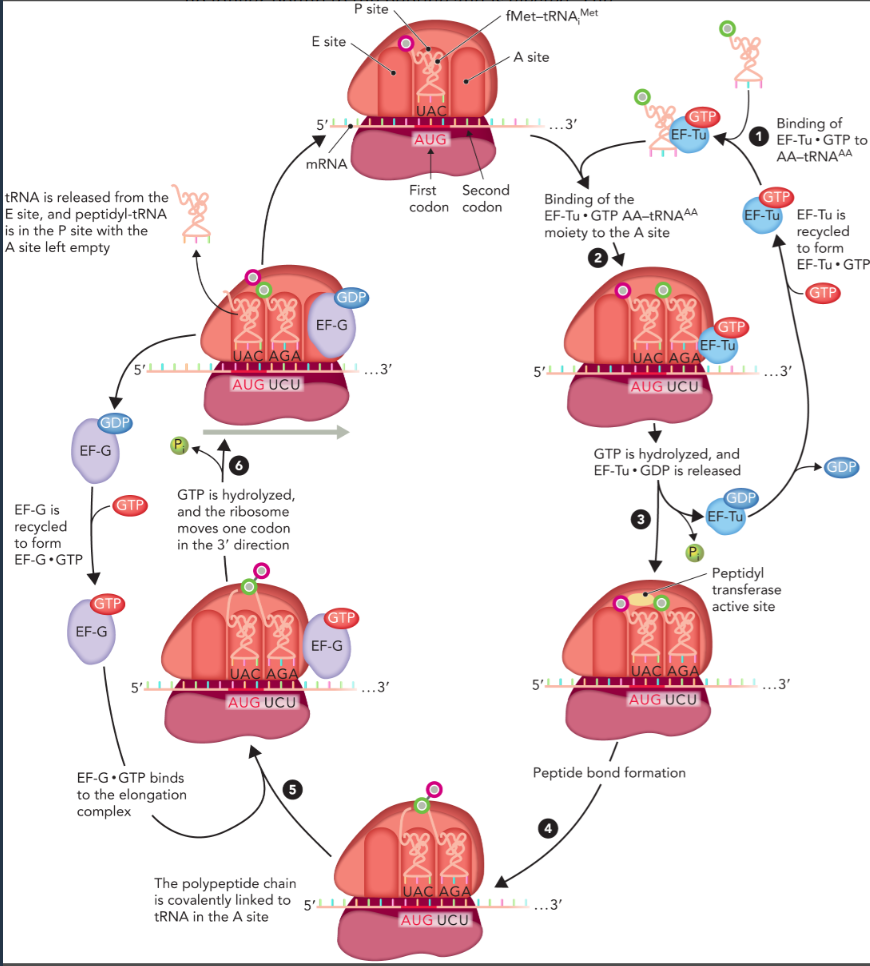
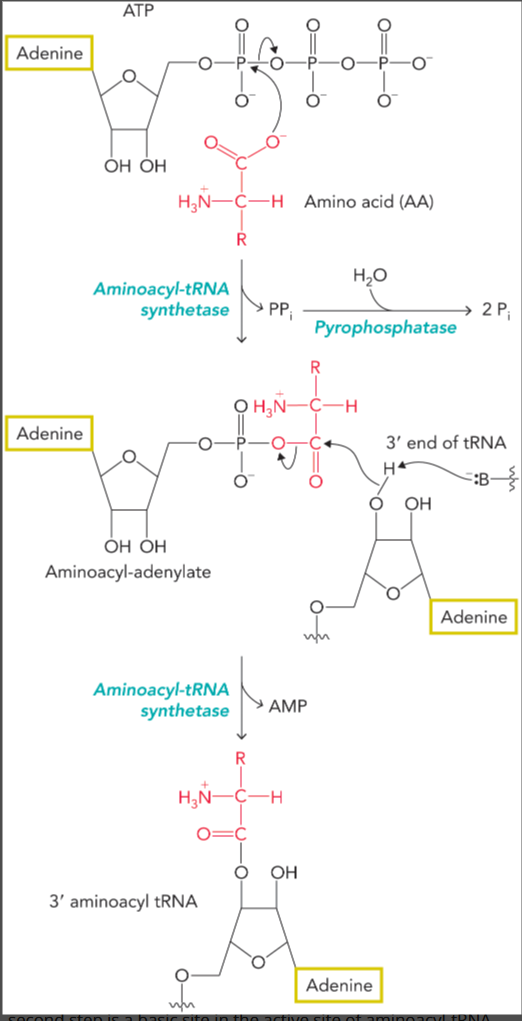
peptidyl transferase reaction
catalyzed by the ribozyme activity of rRNA
the nucleophilic amino group of the amino acid bound to A site tRNA attacks the electrophilic carbonyl carbon of the P site tRNA
rearrangement results in transfer of the former P site amino acid or polypeptide to the amino acid on the A site tRNA