D2 cell division, gene expression, water potential
1/47
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
48 Terms
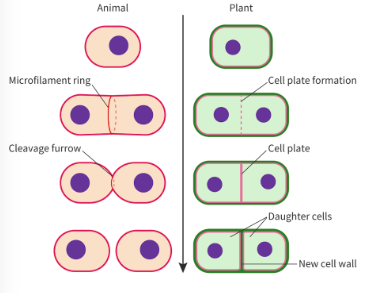
what is cytokinesis? how does it work in animal and plant cells
def: process where cytoplasm of parent cell is split to make 2 daughter cells
animals:
actin/myosin makes contractile ring to pinch cell membrane
forms a cleave furrow (with aid of fluid nature of double membrane)
plants:
need to assemble cell plate (made from fusion of vesicles containing cell wall mateirals)
plate grows outward until fusing with existing cell wall
describe an example of unequal division of cytoplasm in humans
oogenesis
primary oocyte —> 2 rounds of division —> secondary oocyte + first polar body
if fertilisation occurs, secondary oocyte —> cell division —> mature ovum + second polar body
*mature ovum has majority of cytoplasmic contents —> necessary nutrients for early development
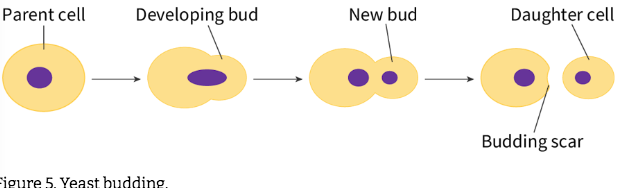
describe budding
unequal cytokinesis
asexual reproduction = outgrowth of a genetically identical daughter cell or ‘bud’ from the parent cell. The bud starts small and grows in size until it forms a fully developed cell that can function independently from the parent cell.
daughter cell is smaller than parent and gets less cytoplasm
parent cell left with small scar
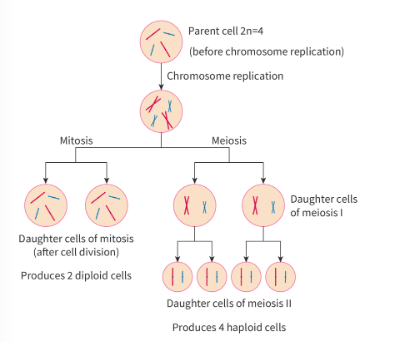
mitosis vs. meiosis functions
mitosis
somatic cells
makes diploid genetically identical daughter cells
for growth or replace damaged cells
meiosis
makes 4 genetically uniquhe daugter nuclei
also beings with diploid cell
has 2 rounds of nuclear division
describe shared features of mitosis/meiosis
need DNA first (in interphase)
both involve movement of chromosomes (microtubule + microtubule motors to move them)
how does DNA look like before and after nuclear division
When the cell is not undergoing nuclear division, the DNA is loosely packed around the histone proteins.
At the beginning of nuclear division the chromatin is supercoiled and condensed by histone proteins. This results in tightly coiled shapes that we recognise as chromosomes. Supercoiling of DNA allows for efficient separation of replicated DNA during nuclear division.
describe mitosis stages
makes 2 genetically identical diploid daughter cells
prophase
chromatin condenses into chromosomes (already undergone replication, made of 2 genetically identical sister chromatids)
nuclear membrane breaks down
spindle fibres form
plants use MTOC to organise spindle fibres, animals use centrosomes —> MTOCs migrate to each pole of the cell
metaphase
sister chromatids line up on metaphase plate
spindle fibres (bound to centromere of sister chromatids) moves them into position
anaphase
spindle fibres shorten to split centromere and pull sisters apart to either pole
telophase
chromosomes decondense, nuclear membrane reforms
spindle fibres disintegrate
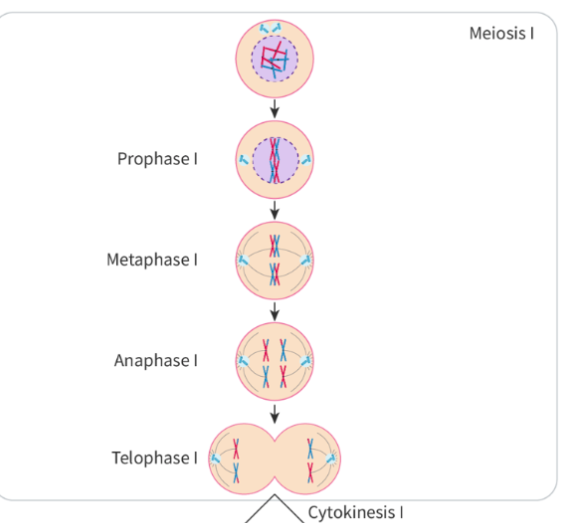
describe steps in meiosis I
produces 4 genetically unique daughter haploid nuclei
is reduction division
Meiosis I
P1:
nuclear mmebrane disintegrates
MTOCs migrate to opposite poles
spindle fibres form
chromatin condenses into chromosomes —> sets of sister chromatids join into a bivalent/tetrad. crossing over may occur (Exchanging equivilant DNA segments between non-sister chromatids)
M1:
spindle fibres attahc to centromeres of homologous chromosomes to pull bivalents to centre of cell
maternal & paternal homologues show random orientation to poles (orientation of maternal is independent of other paternal)
A1:
spindle fibres shorten to separate bivalent
pulls homologous chromoosmes to opposite poles
sister chromatids still connected at centromere
T1:
homologous chromosomes reach poles of the cell and decondense
nuclear membrane forms
spindle fibres breakdown
now, is cytokinesis, and interkinesis (resting)
list steps in meiosis II
Prophase II: During prophase II the DNA recondenses, the nuclear membrane disintegrates, MTOCs migrate to opposite poles and the spindle fibres start to form.
Metaphase II: In metaphase II spindle fibres attach to the centromeres, lining up sister chromatids in the centre of the cell. Sister chromatids show random orientation towards the poles.
Anaphase II: In anaphase II, spindle fibres shorten, splitting the centromere and pulling sister chromatids apart towards opposite poles. Once sister chromatids are separated, they are called chromosomes.
Telophase II: In telophase II the chromosomes reach the opposite poles of the cell and decondense. A nuclear membrane forms around each nuclei.
Following meiosis II, cytokinesis will occur to create four haploid daughter cells.
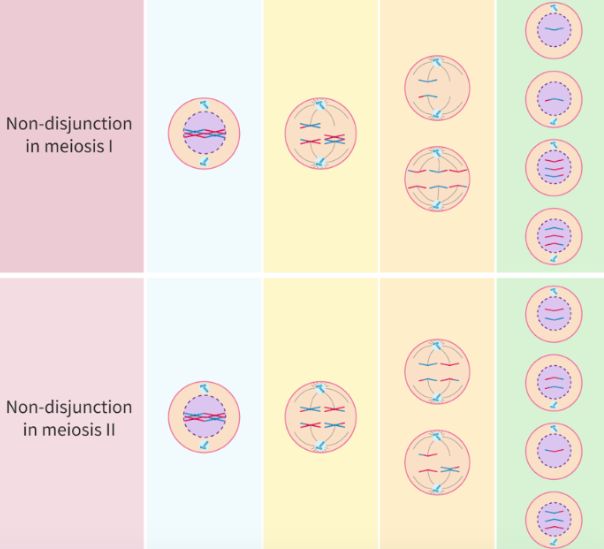
what is non-disjunction caused by? what does it result in?
caused by:
failure of pairs of homologous chromosomes to separate in anaphase I
failure of sister chromatids to separate in anaphase II
leads to gametes with 1 extra or missing chromosome
what is trisomy 21?
a condition that results from an extra copy of chromosome 21. Down syndrome occurs in approximately 1 in every 700 live births worldwide and results in individuals with a range of intellectual and physical disabilities. This includes poor muscle tone, heart defects and delayed development
what is an example of monosomy?
monosomy X (turner’s syndrome)
normally, females have XX, males have XY
non-disjunction happened in mother/father sex chromosomes not separating properlI
a gamete is missing an X chromosome
offspring only get 1 X chromosome from a parent
what are 3 ways meiosis results in genetic diversity
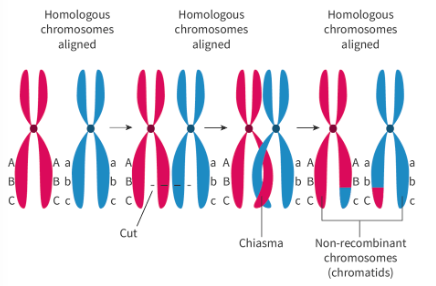
crossing over
in prophase I:
may occur several times in the same bivalent
occurs at different places each time meiosis occurs
happens anywhere along chromosome
sister chromatids no longer genetically identical —→ gamete inherits diff. combinations of alleles
random orientation/independent assortment
in metaphase I or II:
homologous chromosomes line in equator and are orientated randomly (position don’t matter)
random fertilisation
the randomness of which egg/sperm fuse
define cell proliferation
the process of cellular division and replication
for growht, cell replacement, etc
what are 3 stages in cell proliferation
growth
cell proliferation increases cell number and organism size & complexity
plants: growth occurs in meristems —> cells at tip remain differentiated, so cells behind them can specialize and differentiate into specific cell types
humans: cell division happens every 24 hours in early embryonic development
cell replacement
ex. skin cells
tissue repair
cells divide and migrate to site of injury
what are. 3stages in cell cycle
interphase (G1, S, G2)
G1: in cytoplasm, cell grows in size and carries out regular metabolic fx, mito and chloroplasts replicate through binary fission, cell doubles in size
G2: in nucleus, DNA replication
G2: in cytoplasm, cell continues growing, synthesizes microtubles and proteins needed for cell/nuclear division, checks that DNA replication was accurate
mitosis
cytokinesis
how is cell cycle controlled?
by cyclins
a family of proteins that regulate cell cycle
they bind and activate CDKs (enzymes that phosphorylate proteins for cell cycle)
only active in some stages of cell cycle (need threshold)
ex. cyclin E binds to CDK before S-phase, activating DNA replication
a mutation can occur in ____ or _____ which are genes involved in the cell cycle
proto-oncogenes
genes coding for proteins promoting cell growth
a mutation results in uncontrolled division
tumour suppressor genes
genes coding for proteins that slow down/prevent cell division
when mutated, leads to uncontrolled cell division
contrast 2 types of tumours
benign
abnormal growth of cells
non cancerous
grows slowly
has well defined borders
malignant
lacks well defind border
can spread through metastasis in bloodstream/lymphatic system (forms secondary tumour that’s difficult to treat)
formula for mitosis
mitotic index
measures the % of actively dividing cells in a population
from 0-1
actively dividng cells w/visible chromosomes/total number of cells
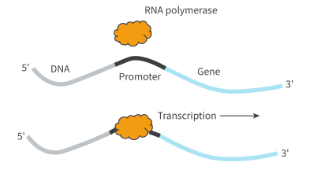
what are promoters?
noncoding regions of DNA where RNA polymerase binds to
typically loacted at 5’ end of a gene
allows initiation of transcription
what do transcription factors do?
proteins that influence gene expression (Affects interaction of RNA polymerase and promoter)
encourages RNA polymerase to bind to promoter or blocks it
the availability of it affects how easily RNA polymerase binds to promoter
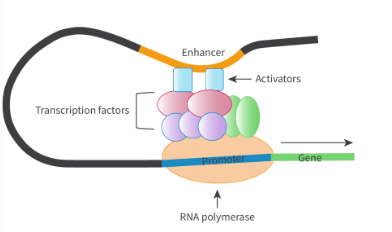
what is enhancer? what does it work with
regions of DNA that regulate when and to what extent a gene. is expressed
noncoding
unlike promoters, enhancers can be located far away from genes
works with activator proteins (Type of transcription factor), where binding forms a complex that interacts with promoter region
can multiple genes transcribed at the same time?
yes, since 1 enhancer can interact w/multiple promoters
what is operon
a collection of genes with the same promoter
these genes are usually transcribed together
what determines how long an mRNA persists?
usually, the longer = translated more
will eventually be broken by nucleases
lifespan depends on:
chemical modifications (guanine cap at 5’, poly-A tail, all increases stability of mRNA)
short tails are likely to be translated since they are vulnurable to degradation —> can vary the length to vary how much each protein is made
presence of stabilising proteins (interfears with nucleases to extend lifespan, as well as help binding of protective proteins)

define epigenesis
the process where cells/organisms develop from an undifferentiated zygote through interactions between DNA and environmental factors
basically, how stem cells differentiate into different tissues
environmental factors can modify gene expression factors without changing DNA sequence (by activating/silencing genes) to alter phenotype
the study of how behaviour and environment affects gene transcription
describe DNA methylation
methylation of cytosine in a promoter represses transcription —> prevents binding of transcription factors
results in silencing of genes
level may depend on age, diet, etc
describe histone protein structure
has core domain + tail
tails (N-terminus) protrude outwards
has net positive charge, mainly in the tails
helps attract DNA to keep it together
describe the different structures that nucleosomes can be in based on level of transcription
heterochromatin
when DNA is supercoiled around histones
DNA less accessible to RNA poly.
less transcription
euchromatin
when DNA is loosely packed
more accessible to RNA polymerase
higher transcirption
what are ways histone tails can be modified?
acetylation
decreases charge of histone protein, reducing attraction between histone and DNA
easier for RNA polymerase to access DNA
increased gene expression
methylation
of AA in histone tails
represses/activates trancsription
both alter nucleosome structure
define epigenetic inheritance
def: inheritance of non-genetic information that can influence gene expression and phenotypic traits
for this to occur, epigenetic changes (DNA methylation, histone modificaiton) must occur in germline cells
list 3 environmental effects on gene expression
exposure to air pollution
affects DNA methylation
diet
high folic acid = increased methylation
temperature
the sex of reptiles
what are epigenetic tags?
tags like methyl groups that regulate gene expression
after fertilisation, most tags are removed
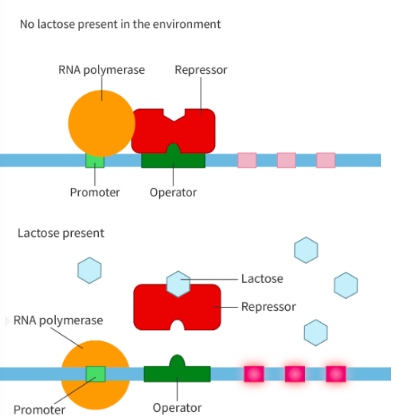
discuss effect of lactose on lac operon expression
lac operon = cluster of 3 genes in bacterial DNA coding for proteins for lactose digestion
bacteria use lactose as a backup energy source
when lactose absent:
lac repressor binds to operator region
prevents attachment of RNA polymerase to lac operon promoter
represses transcription
when present:
lactose bindst o repressor, detaching it from promoter, allowing RNA polymerase to bind and transcribe it
discuss effect of tryptophan on expression of bacterial genes
it affects tryptophan operon = a cluster of 5 genes in bacterial DNA, necessary for synthesis of tryptophan AA
when tryptophan absent:
RNA polymerase binds to tryptophan operon to transcribe genes for tryptophan
bacteria can produce tryptophan when not available
when present:
tryptophan binds to repressor protein, which binds to generator region of operon
inhibits transcription fo operon
when can water dissolve the substance
when force of attraction between ions and water is greater than force of attraction between oppositely charged ions
hyper vs. hypotonic solution
hyper = high solute concentration (high osmotic concentration)
hypo = low solute concentraiton (low osmolarity)
water flows from…
hypotonic —> hypertonic until isotonic
what happens if cell is put in hypertonic solution:
will be net movement of H2O out of cell
cell shrinks (crenated or plasmolysis)
what happens if cell is placed in hypotonic solution
will have net movement of water into cell
cell swells
animal cells may lyse
how to determine isotonic [solute] of a plant tissue?
meausre % change in tissue mass and length of plant tissues placed in different solution concentrations
same mass/length of plant tissue is put in different solute concentrations
then, removed and patted dry
data is collected
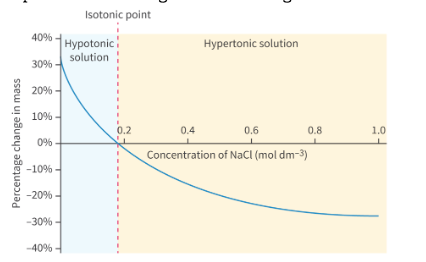
discuss H2O movement on plants
influx of H2O = accumulates turgor pressure, making cytoplasm exert pressure against cell wall. however, the cell is turgid since the wall present busting
efflx of H2O = plant cell plasmolyses
define water potential
def: measure of the pot. energy of water/unit volume H2O, measured in kPa. pure water has 0kPa
water potential is defined by what 2 things
solute potential
ability for H2O to move given the concentration of solutes
the more solutes in a solution, the less free water (less potential to move)
pressure potential
pressure that H2O exerts on cell membrane
can be positive (increase water pot. when exerted outwards from cell) or negative (xylem suction)
negative water pot means..
harder for water to move
water moves by osmosis….in terms of water pot.
from high water pot to low water pot.
describe what happens when plants placed in hypotonic solution in terms of water pot.
solute pot of tissue is more negative than the solution
H2O moves into tissue (less negative water pot. —> more negative)
the additional water in tissue cells tell them to appply turgor pressure against cell wall
thus, the pressure pot. is positive, since pressure inside cell is higher than outside cell