CSN BIO 251 LAB 7-11 Practical #2
1/160
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
161 Terms
Non-ionizing, short, high energy wavelength radiation (from 4 - 400 nm).
UV radiation
UV-A (longest wavelength)
UV-B (280-315 nm)
UV-C (100-280 nm, most damaging, kills 99.9% of all pathogens)
3 types of UV radiation
Caused by prolonged exposure to UV radiation at wavelengths of 254-260 nm.
DNA damage
Molecular lesions formed by UV radiation that kink the DNA strand, preventing replication (thymine-thymine most common).
Pyrimidine Dimers
A repair system activated by major DNA damage that removes pyrimidine dimers and restores DNA. If system is overwhelmed, the bacteria dies.
SOS system
Time length of exposure
Wavelength (260 nm is most damaging)
Distance from the source
Barriers
Factors affecting UV radiation lethality
Acid-soluble proteins that protect DNA and enzymes.
Can withstand UV radiation. Biofilms also provide protection.
Bacterial Endospore
They repair damaged DNA within endospores.
What is the role of enzymes in bacterial endospores regarding UV damage?
Chemical metabolites produced by fungi and bacteria to treat infections.
Natural antibiotics
The first commercial antibiotic discovered.
Penicillin
Alexander Fleming
Who discovered penicillin?
Entirely produced inside laboratories (ex. Sulfonamide).
Synthetic antibiotics
Antibiotics produced chemically in the lab by modifying the R side chain of natural antibiotics; Used if parent (original) antibiotic is unsuccessful (ex. Penicillin).
Semi-synthetic antibiotics
Bactericidal antibiotics kill bacteria, while bacteriostatic antibiotics inhibit their growth.
Bactericidal vs. Bacteriostatic antibiotics
Narrow spectrum antibiotics target specific bacteria.
Broad spectrum antibiotics target multiple types.
Narrow spectrum vs. Broad spectrum antibiotics
Target a specific Gram class of bacteria.
Vancomycin - active against Gram-positive bacteria.
Polymyxin - active against Gram-negative bacteria.
Narrow spectrum antibiotic
Target both Gram types of bacteria (positive and negative).
Tetracycline - target both Gram-positive and Gram-negative bacteria but remove normal microbiota (found in colon), allowing superinfections.
Broad spectrum antibiotic
1. Inhibit cell wall synthesis
2. Inhibit protein translation
3. Disrupt cell membrane
4. Inhibit DNA/RNA synthesis
Antibiotics Mechanisms of Action
The antibiotic targets bacteria and not humans (very specific, ex. Penicillin)
High Selective Toxicity
The antibiotic can cause more serious side effects in humans like polymyxin B (not very specific for bacterial cell membranes). Used externally only.
Low Selective Toxicity
Bacteria swabbed over entire agar to form a uniform growth.
Bacteria lawn
A method used to determine the in vitro sensitivity of antibiotics or the antibiotic resistance of bacteria.
Incubation 37°C for 16-18 hours.
Kirby Bauer Method
Mueller-Hinton II agar
pH 7.2 - 7.4
Agar thickness of 4 mm
Standardized variables for Kirby-Bauer test
The diameter of the zone of inhibition in millimeters is compared to a standardized chart.
Susceptibility rating determination in Kirby-Bauer test
A quantitative technique for determining antimicrobial susceptibility using a predefined gradient of antibiotic concentrations on a plastic strip.
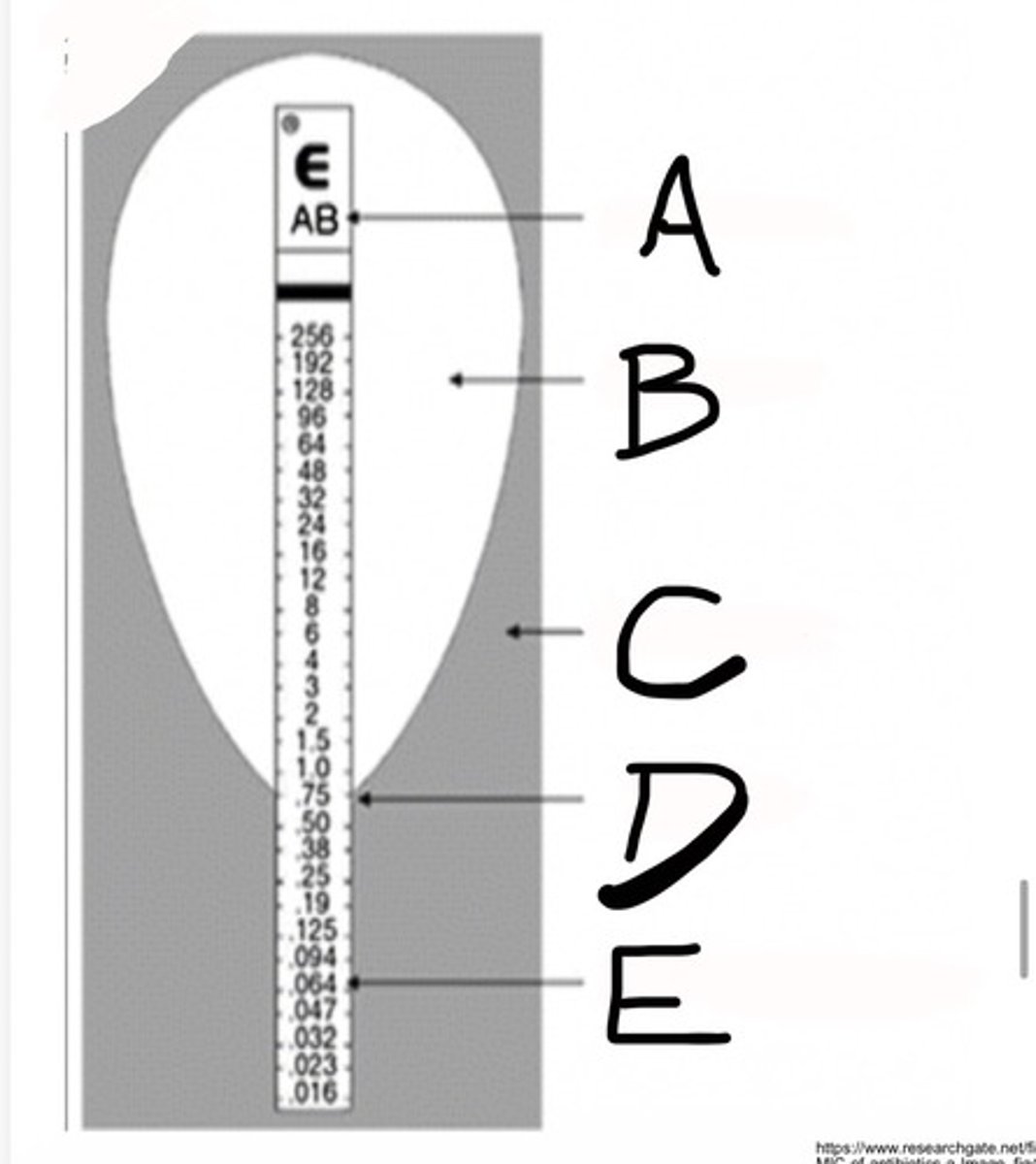
E-TEST
Indicates the lowest concentration of an antimicrobial agent that prevents visible growth of bacteria.
Minimum Inhibitory Concentration (MIC)
Code for Antibiotic
What is A?
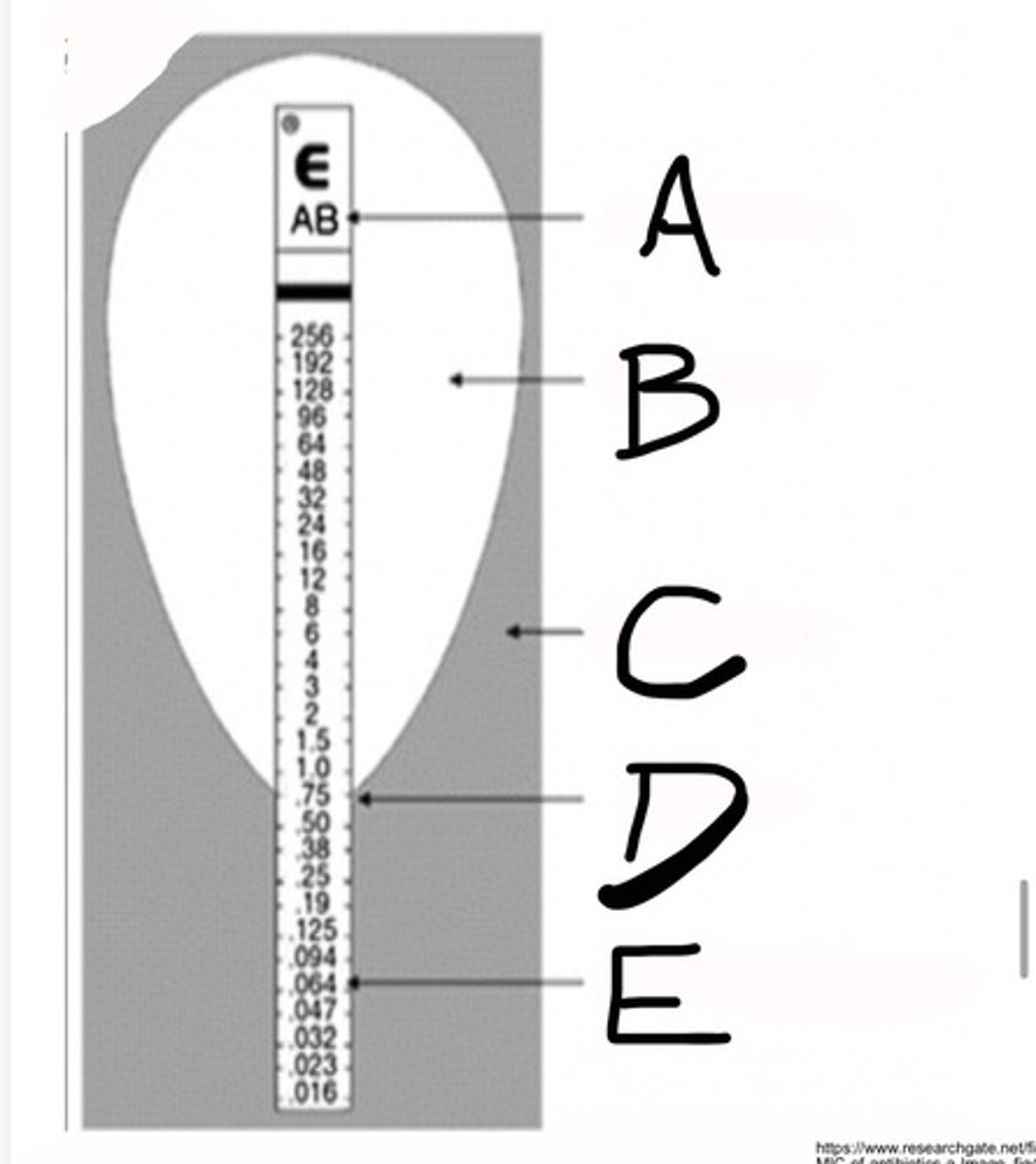
Inhibition Ellipse
What is B?
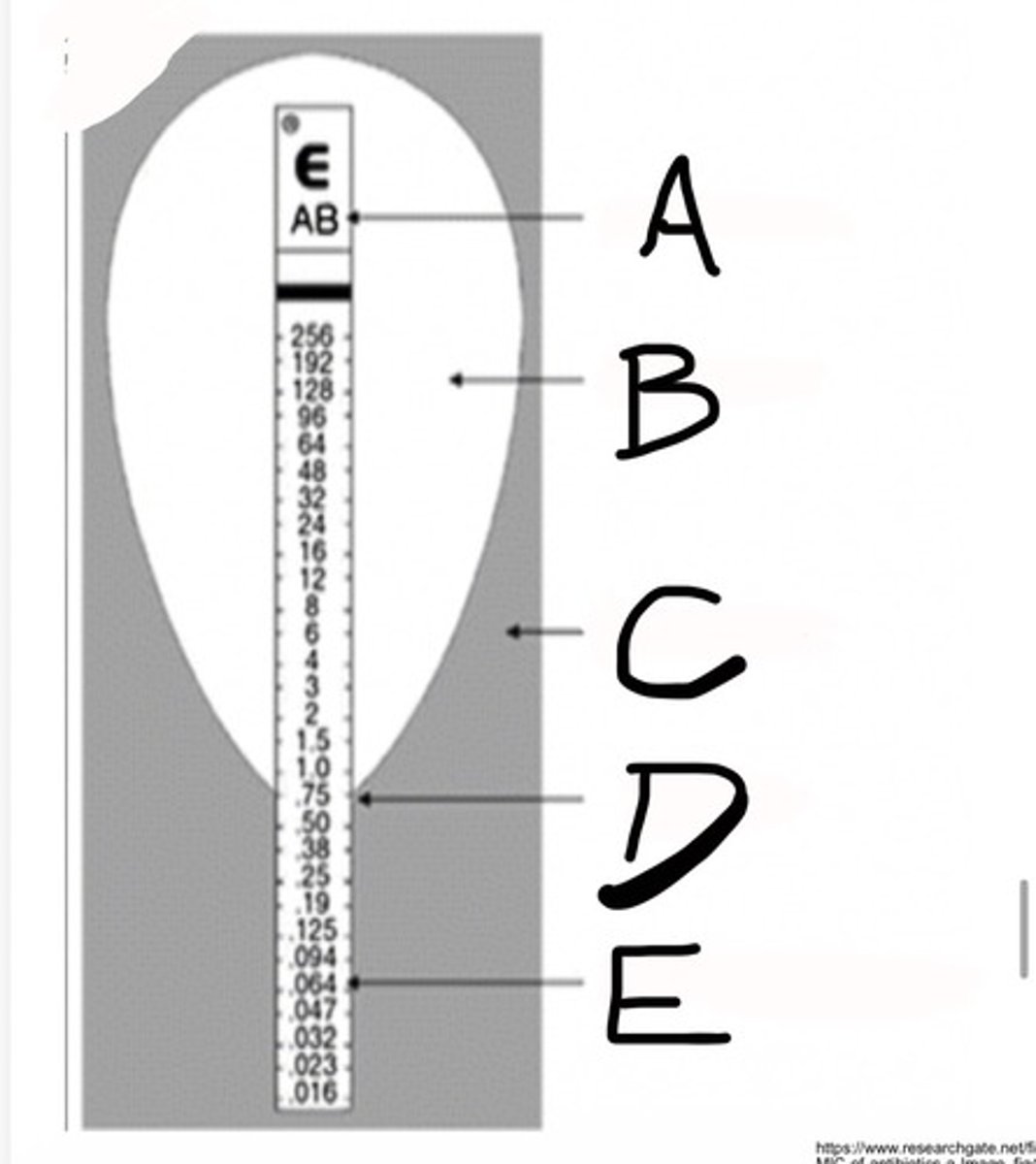
Bacterial Growth
What is C?
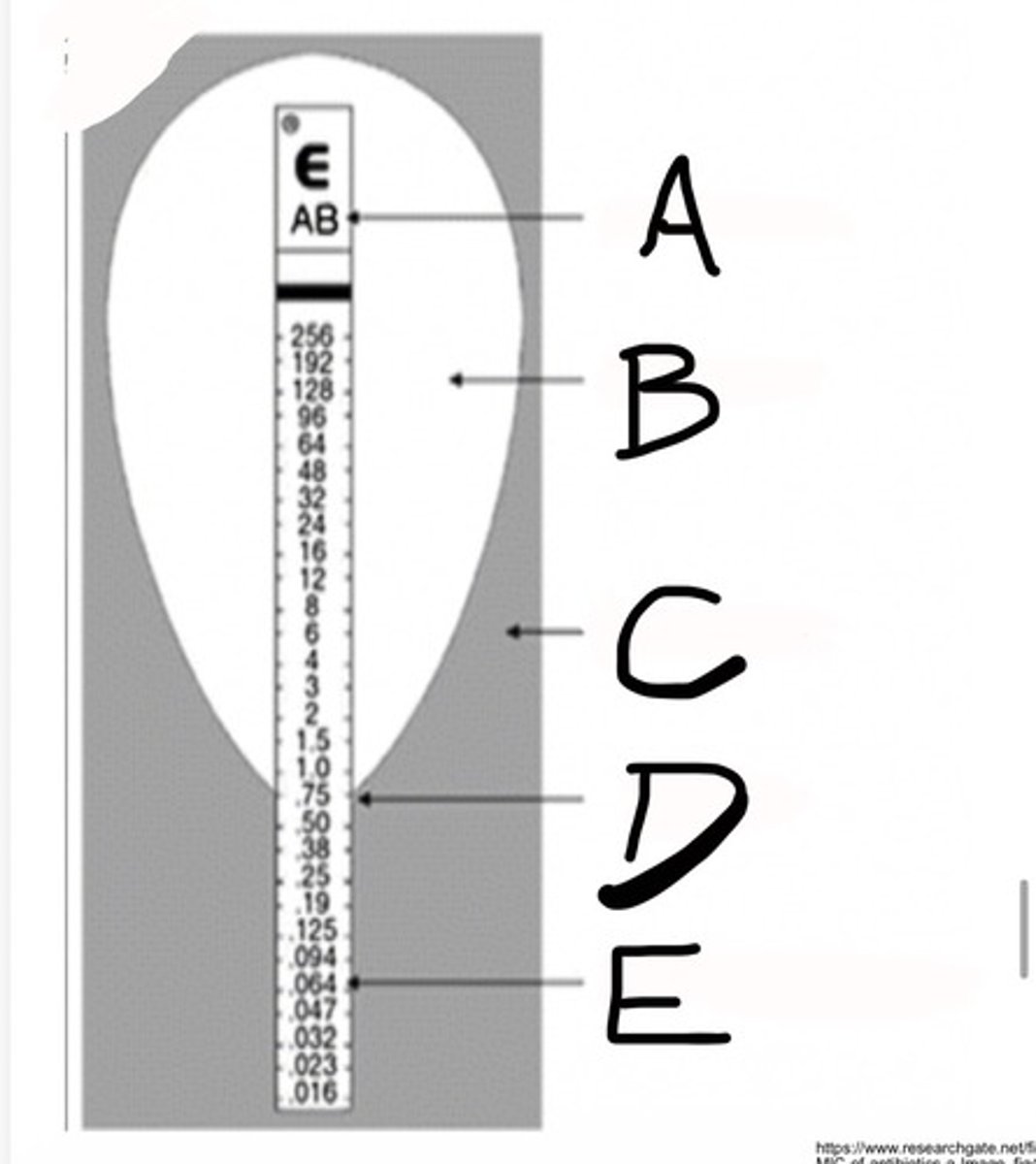
MIC (ug/ml)
What is D?
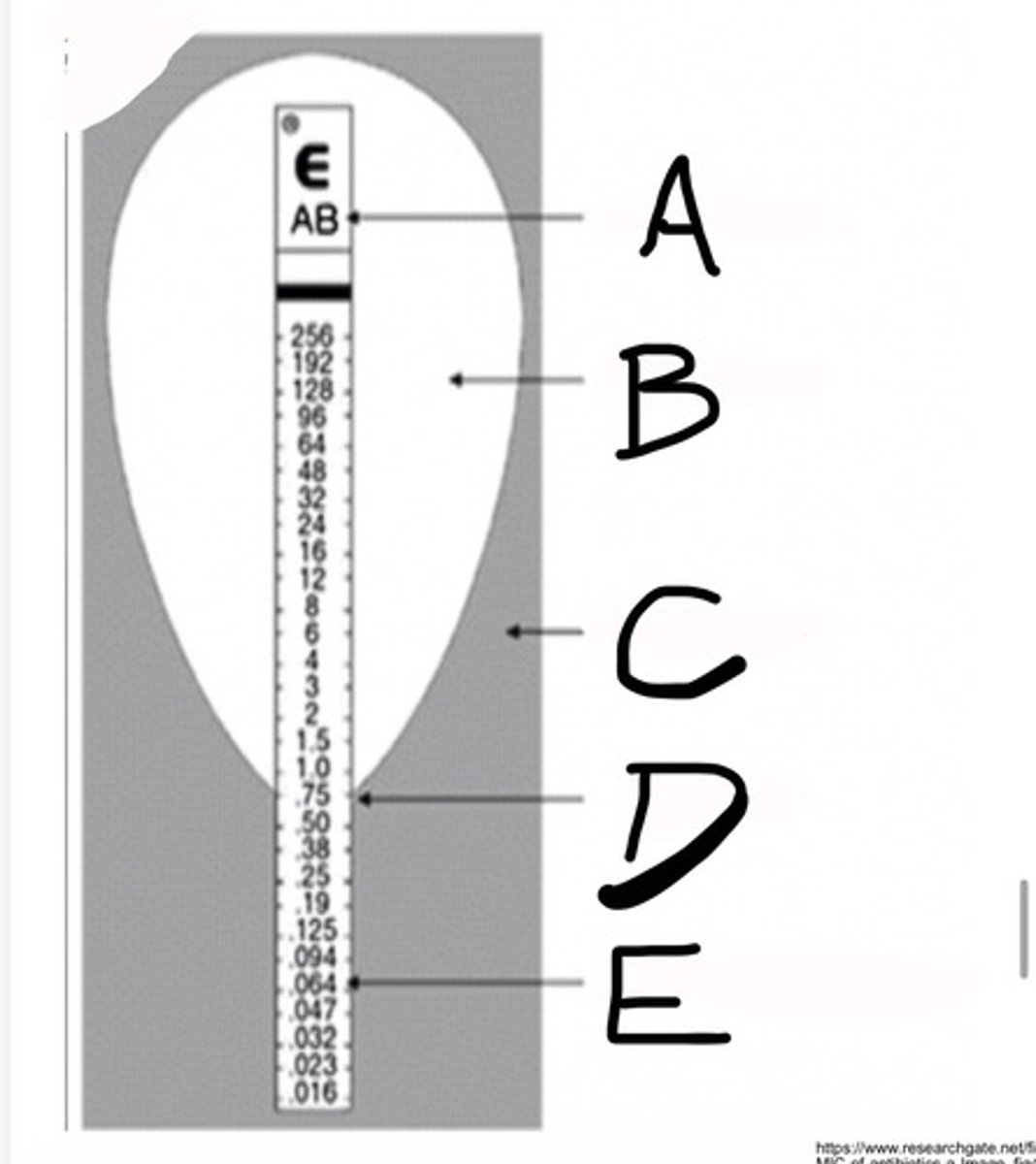
Etest MIC predefined gradient
What is E?
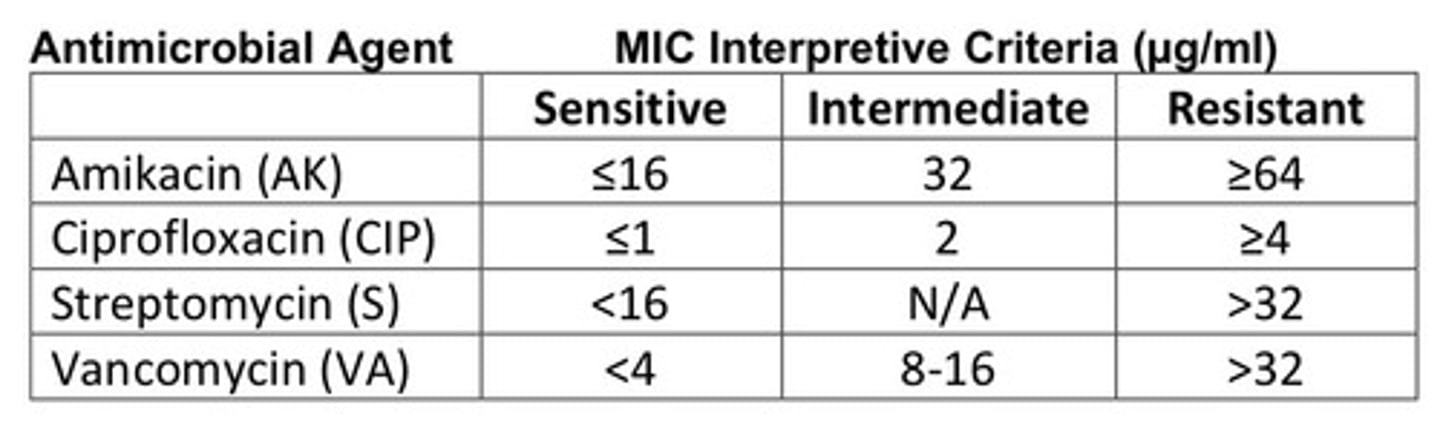
MIC Interpretation
Study this chart
The lowest concentration of an antibacterial agent required to kill a bacterium over a fixed period.
Minimum Bactericidal Concentration (MBC)
The process of transferring genetic information from one donor bacterial cell to a recipient cell through a sex pilus.
Bacterial conjugation
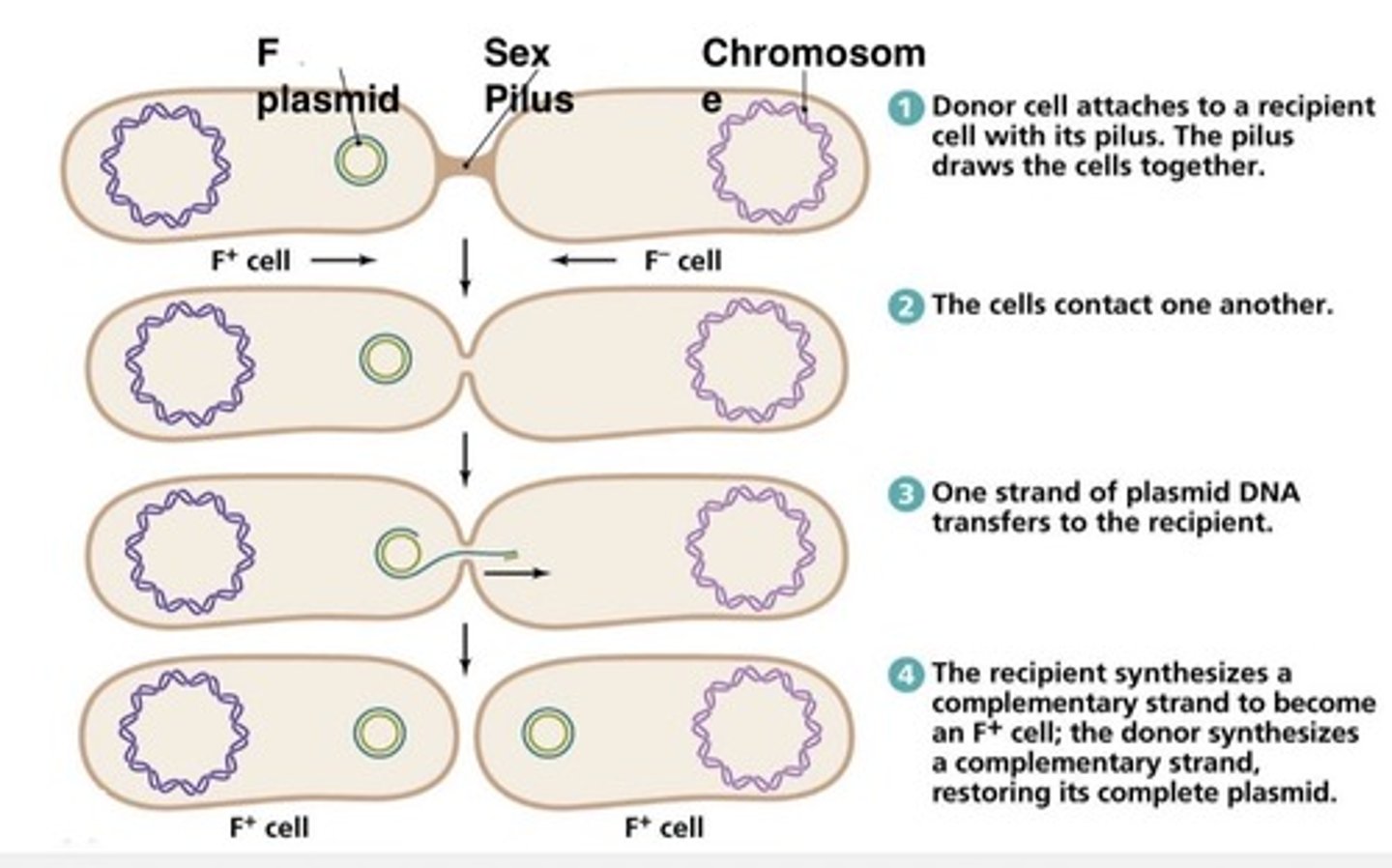
Small, circular, double-stranded DNA molecules that exist outside of the bacterial chromosome and can carry antibiotic resistance genes.
Plasmids
Genes providing bacteria resistance to antibiotics.
Antibiotic resistance genes
The transfer of genetic information from a parent cell to a daughter cell as a result of binary fission.
Vertical transmission
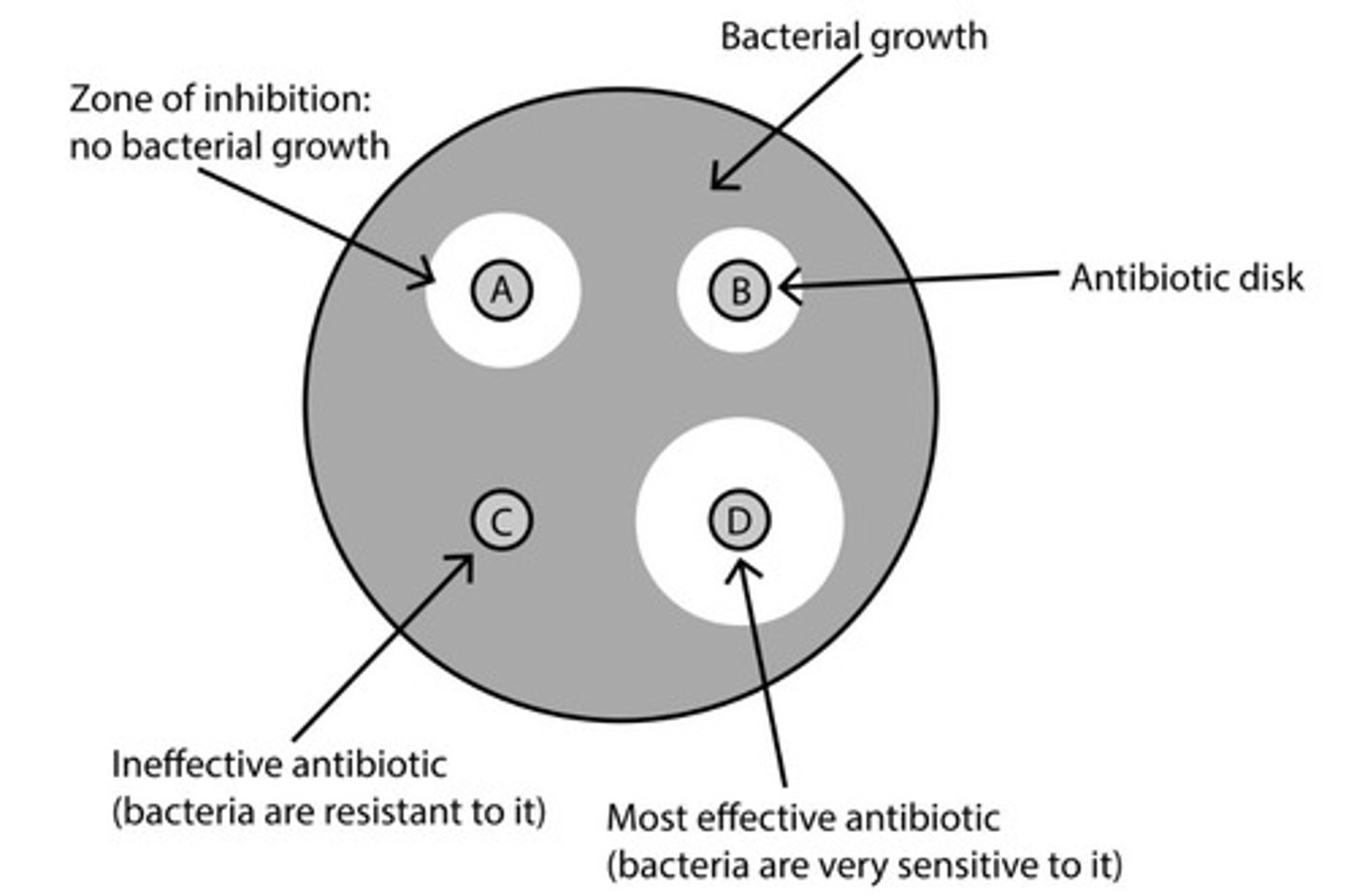
Zone of inhibition
The area around the antibiotic disk in which no bacteria grow, indicating susceptibility to the antibiotic.
What is this?
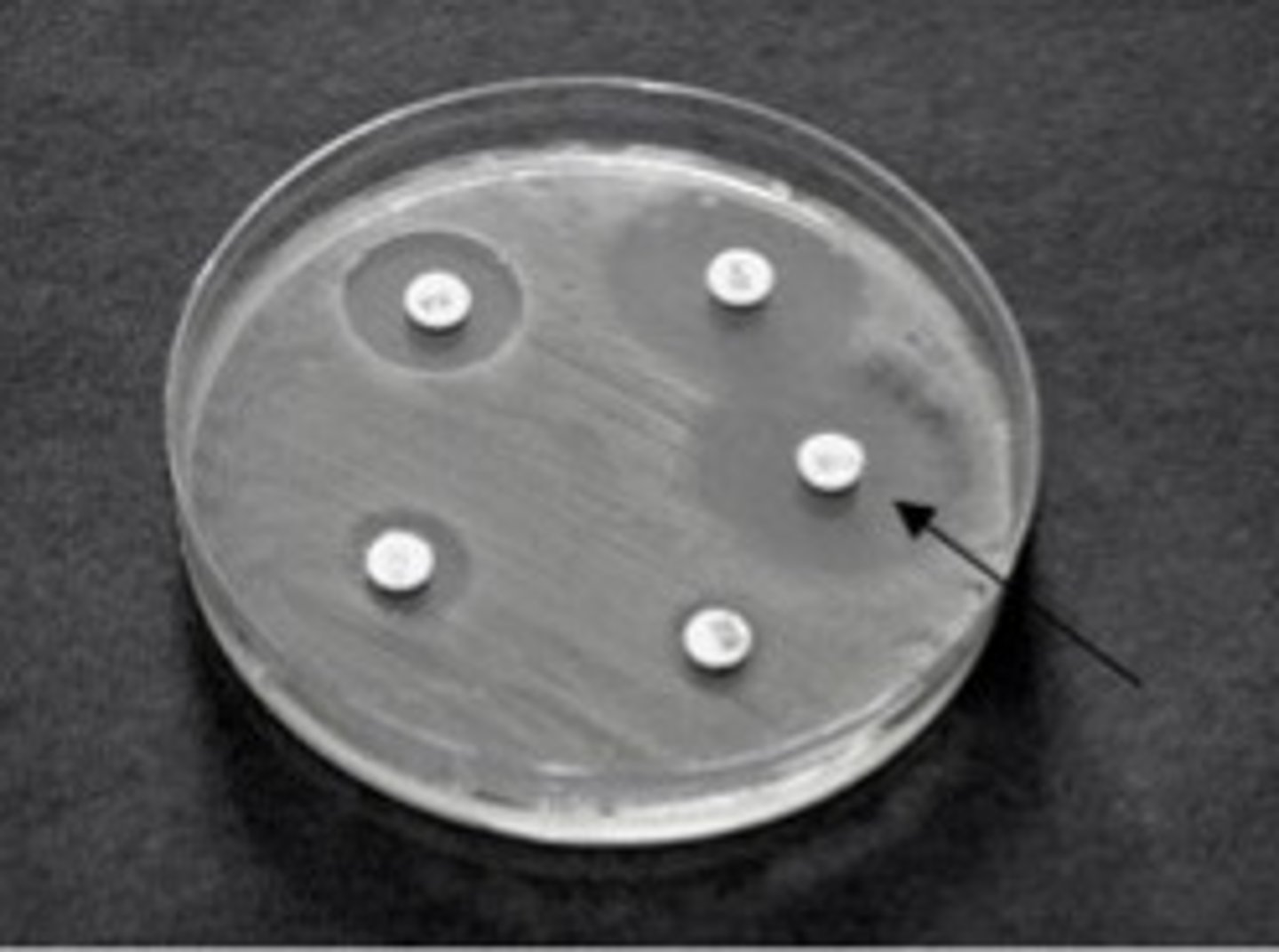
Sensitive - when a bacterium is killed or inhibited in growth.
Resistant - when a bacterium is not susceptible to an antibiotic.
Intermediate - when a bacterium gains resistance and another concentration may be necessary.
Susceptibility Classification
Results for Antibiotic Sentitivity
A form of horizontal transfer that cells use to incorporate novel DNA into their genome.
Transformation
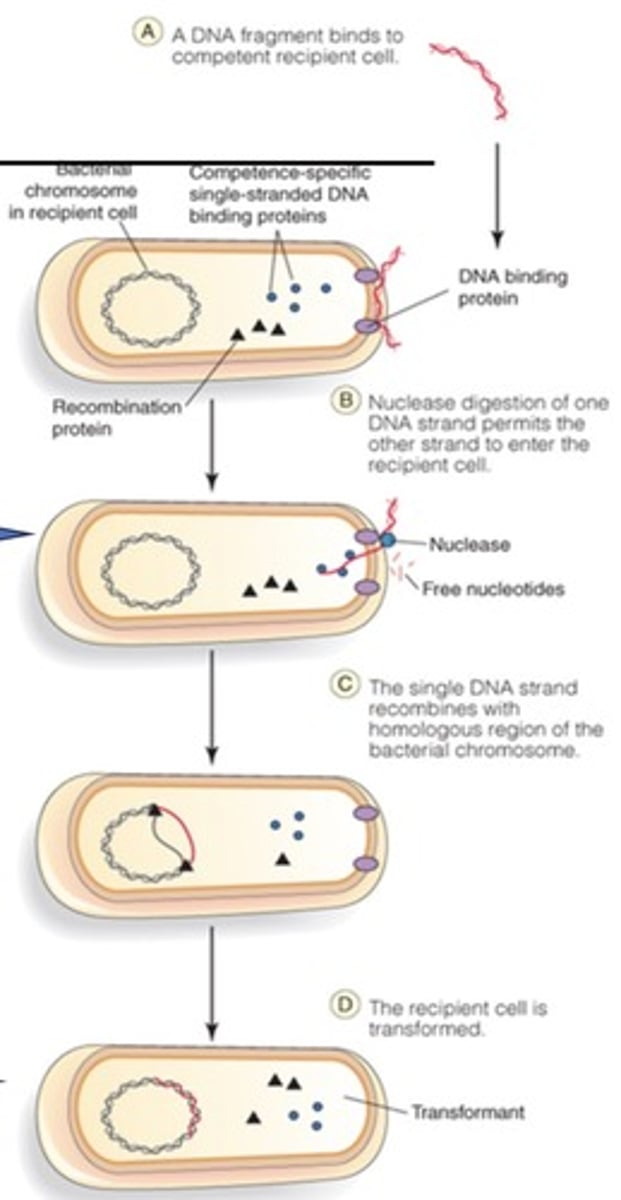
1% of cells in a natural population that have expressed a protein system on their surface which allows them to move intact ssDNA into the cells.
Competent
A family of Gram Negative rods that ferment glucose.
Can be normal gastrointestinal microbiota, opportunists (E. coli), and pathogens(Salmonella, Shigella).
These bacterial pathogens can cause diarrhea, dysentery, hemolytic uremic syndrome,UTI, pneumonia.
Pathogens can release toxin that cause illness and can be fatal.
Many normal flora can cause nosocomial (hospital acquired) infections and are often resistant to multiple antibiotics.
Enterobacteriaceae
Allows us to determine the presence of enzymatic activity because some enzymes cause changes in the pH of the media.
Differential Media
1. Sulfide-Indole-Motility (SIM) medium
2. Triple Sugar Iron (TSI) agar
3. Simmon's Citrate Agar (SCA)
4. Urea agar
Differential Tests
Sulfide, Indole, Motility
Differential Media: SIM
Some bacterial reduce sulfur directly or can remove sulfur from the amino acid cysteine to produce H2S.
Media contains ferrous ammonium sulfate that combines with H2S gas to form ferric sulfide, a black precipitate.
Sulfide
Some bacteria have the enzyme tryptophanase that hydrolyzes the amino acid tryptophan to produce Indole and pyruvate.
Detected with Kovac's reagent (red).
Indole
Semi-solid media
Some bacteria move via flagella
Can be seen moving away from stab line.
Motility
Enzyme that breaks down the amino acid tryptophan to indole, pyruvic acid, and ammonia.
Tryptophanase
Differentiates bacteria based on sugar fermentation and sulfur reduction.
Triple sugars: Glucose, Sucrose, Lactose
Ferrous sulfate (Iron sulfate)
Differential Media: Triple Sugar Iron (TSI) Agar
SCA tests for citric acid utilization
Media contains sodium citrate as the sole carbon source
Also has ammonium phosphate as a single nitrogen source and bromthymol blue (pH indicator)
Organisms with the enzyme citrase are able to utilize the sodium citrate for energy to convert the ammonium phosphate to ammonia
Differential Media: Simmon's Citrate Agar (SCA)
Differentiates bacteria based on the presence of the enzyme urease.
Bacteria are small and typically can not perform endocytosis.
Urease hydrolyzes certain amino acids and urea to form ammonia.
Differential Media: Urea Agar
Enzyme that changes urea into ammonia and carbon dioxide
Urease
Helps differentiate many members of the Enterobacteriaceae based on color.
Chromagar
It facilitates the transfer of genetic material from the donor to the recipient bacterial cell.
Sex pilus
The transfer of genetic information between bacteria through methods like conjugation, transformation, transduction; Also uses transposons.
Horizontal gene transfer
To ensure that the water is safe to drink (potable) and free from microbial contamination.
What is the primary purpose of testing public drinking water?
Parasites (e.g., Cryptosporidium, Giardia)
Viruses (e.g., Norovirus)
Bacteria (e.g., Salmonella, Shigella, E. coli, Vibrio cholera, Campylobacter, Leptospira)
Pathogens found in contaminated drinking water
Pathogens in water, some of which are resistant to chlorine and can cause gastrointestinal outbreaks.
Fecal Contamination
Bacillus that indicate fecal contamination and the possible presence of gastrointestinal pathogens; Ferments lactose.
Gram-Negative
Rod shaped
Facultative Anaerobes
Coliform Bacteria
Rapid'E.coli 2 Agar
Eosin Methylene Blue (EMB)
Hektoen Enteric Agar (HEA)
MacConkey Agar (MCA)
Selective-Differential Media used for screening water
Detects E. coli and other coliforms by identifying two enzymatic activities:
β-D-Galactosidase (GAL; pink)
β-D-Glucuronidase (GLUC; blue).
Rapid'E.coli 2 Agar
Rapid'E.coli 2 Agar
Coliform (GAL+/GLUC-) = blue-green colonies
E. coli (GAL+/GLUC+) = violet-pink colonies
What type of media is this?
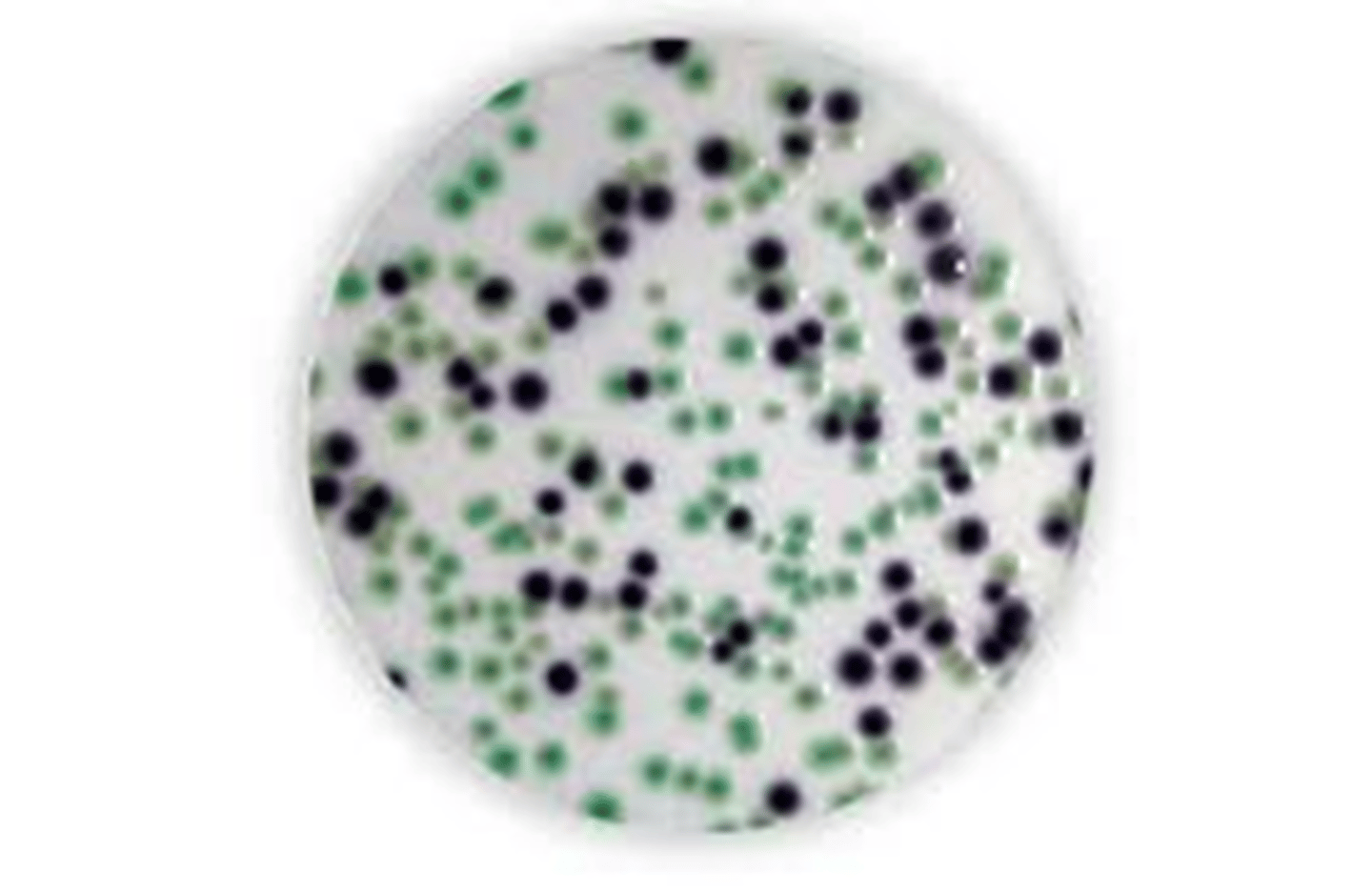
Contains two dyes: eosin Y and methylene blue
Methylene blue inhibits Gram-positive bacterial growth.
Eosin Y is an indicator that reacts with the acid produced by lactose fermenters.
Eosin Methylene Blue (EMB)
Eosin Methylene Blue (EMB)
Coliforms present
E. coli produce a striking metallic green sheen due to excessive acid production in addition to turning the media to a pink-dark purple color.
What type of media is this?
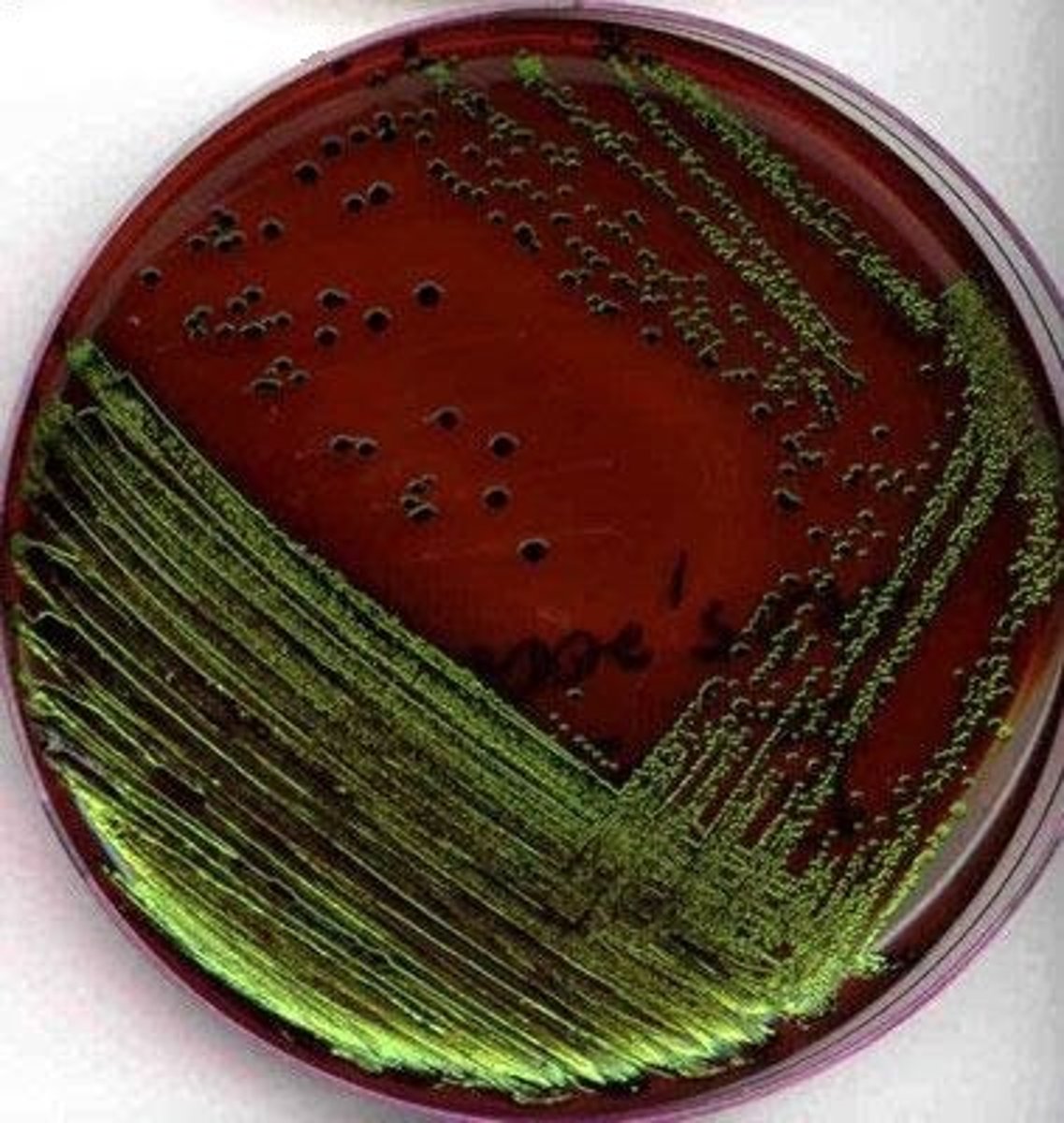
Eosin Methylene Blue (EMB)
Salmonella; Non-coliforms
If there is no lactose fermentation, the agar remains red in color. This means that no coliforms are present, but there are non-coliforms growing in the media.
What type of media is this?
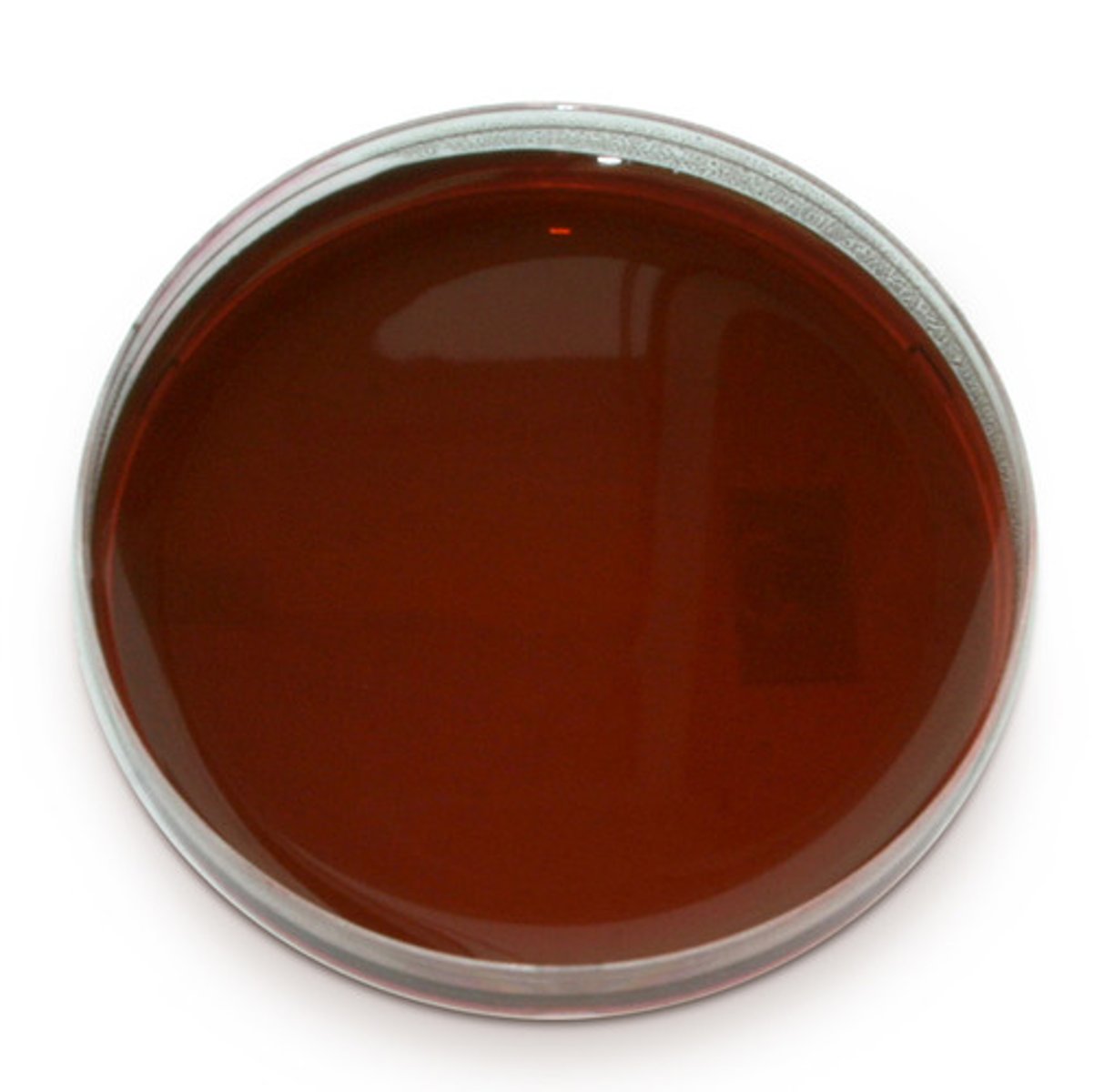

Differentiates Salmonella and Shigella from other enteric bacteria.
Bile salts = Gram-positive
Hektoen Enteric Agar (HEA)
Hektoen Enteric Agar (HEA)
Yellow-pinkish orange = presence of coliform
Blue-green = no coliforms but non-coliforms growing
Ferric ammonium citrate > H2S gas > Ferrous sulfide = Black (Salmonella)
What type of media is this?
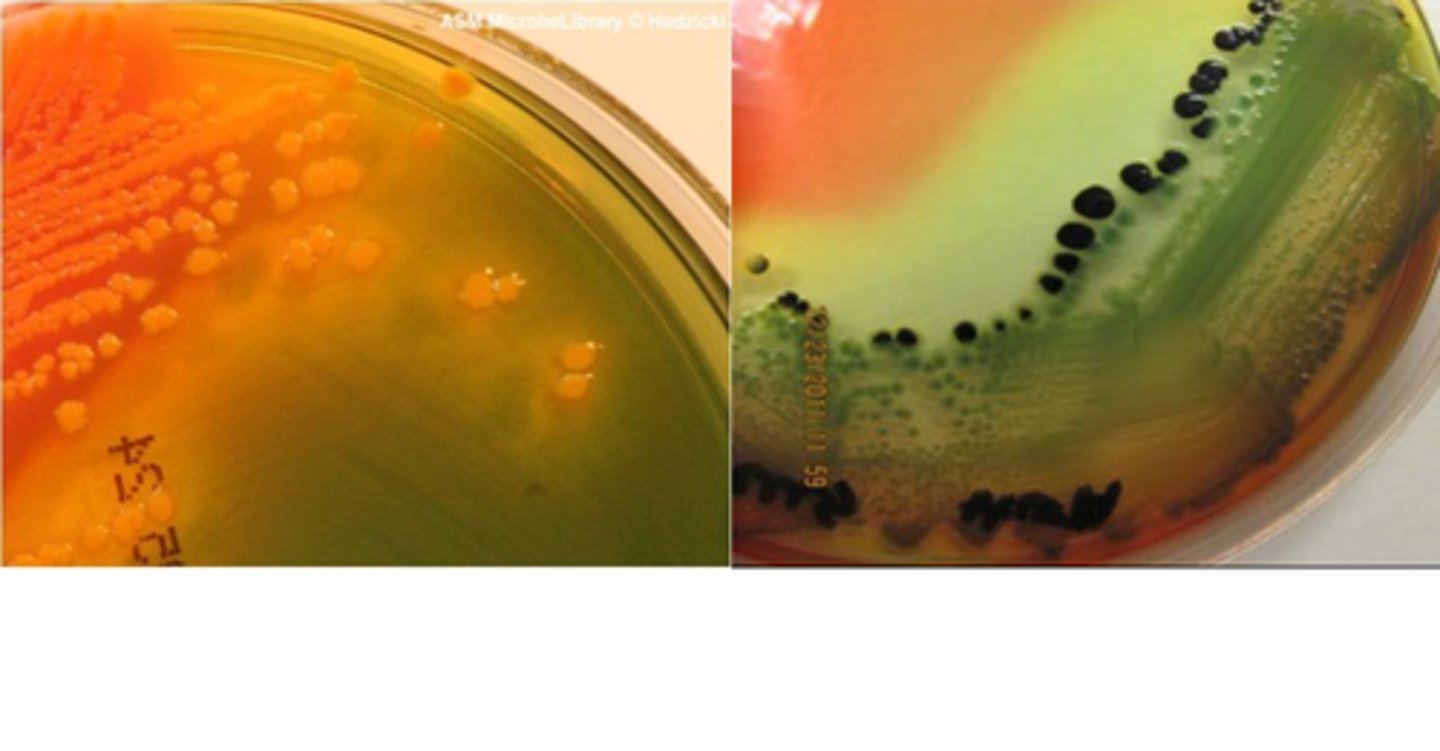
Differentiates Salmonella/Shigella from other enterics. MCA contains bile salts and crystal violet which inhibits Gram-positive growth.
MacConkey Agar (MCA)
MacConkey Agar (MCA)
Pink = E. coli (coliforms present)
Fermentation occurs
What type of media is this (left)?
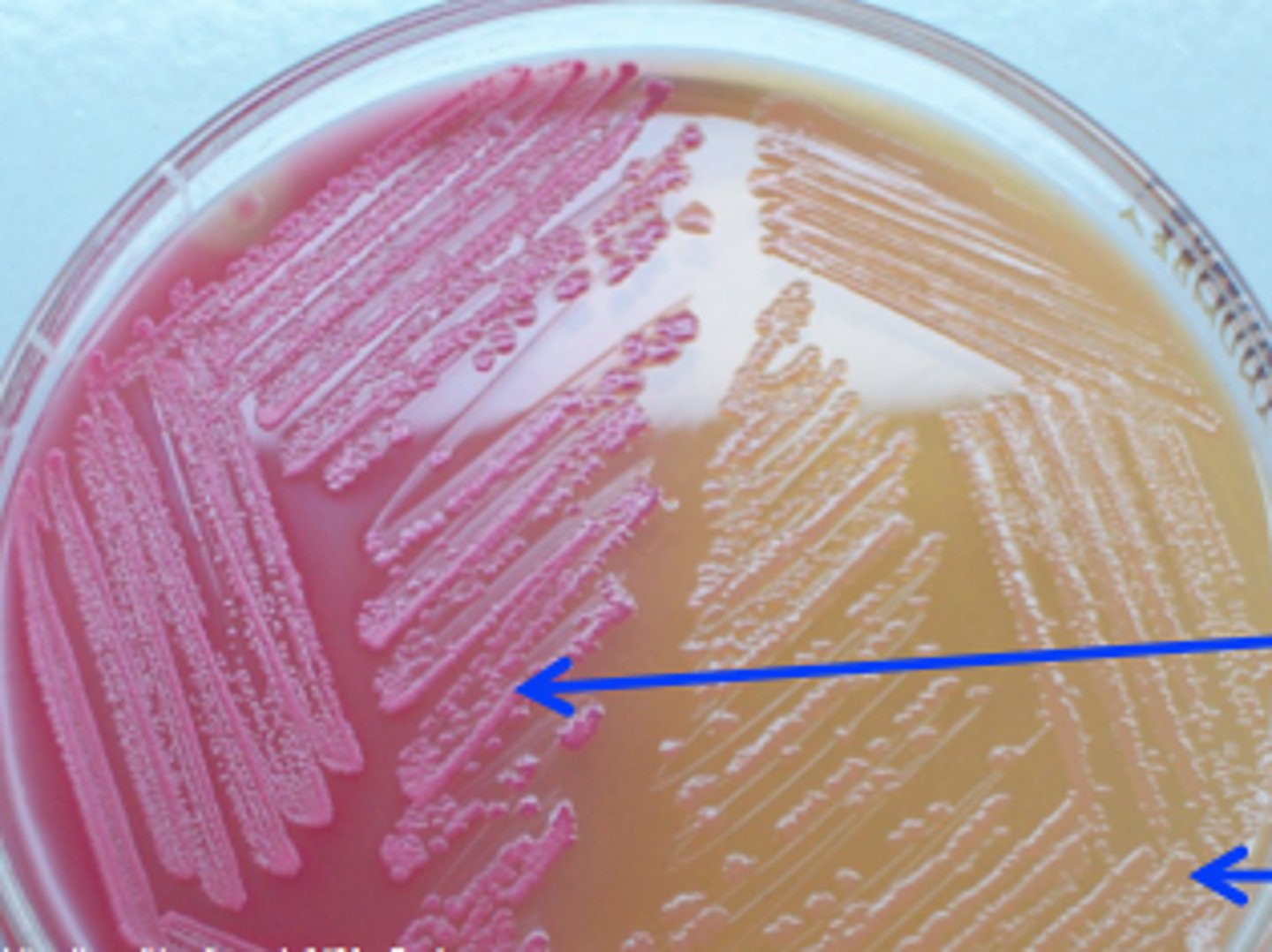
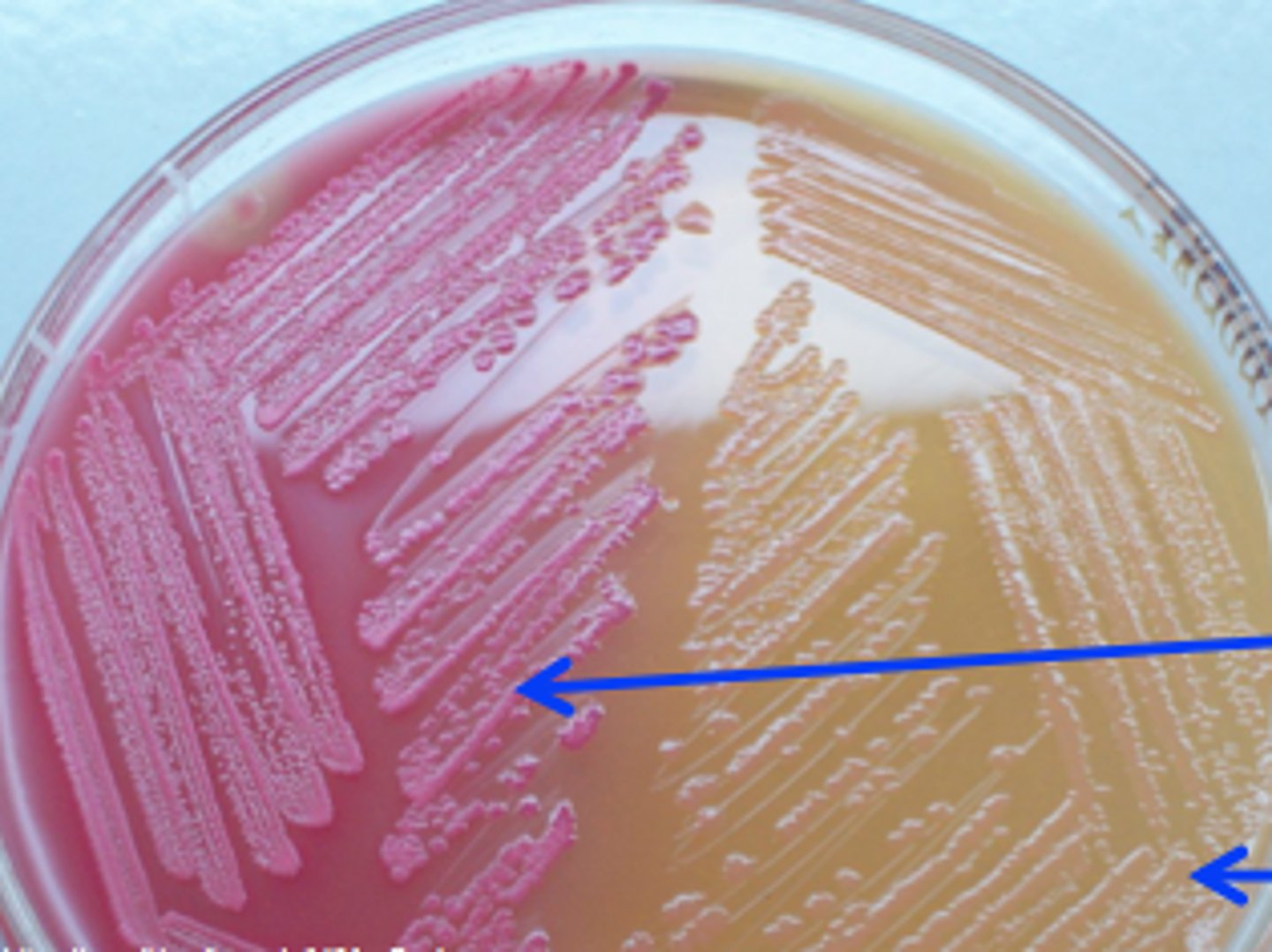
MacConkey Agar (MCA)
Colorless = Proteus Vulgaris (no coliforms present but non-coliform growing)
What type of media is this (right)?
A process where bacteriophages mistakenly package host DNA and transfer it to another bacterial cell during infection.
Transduction
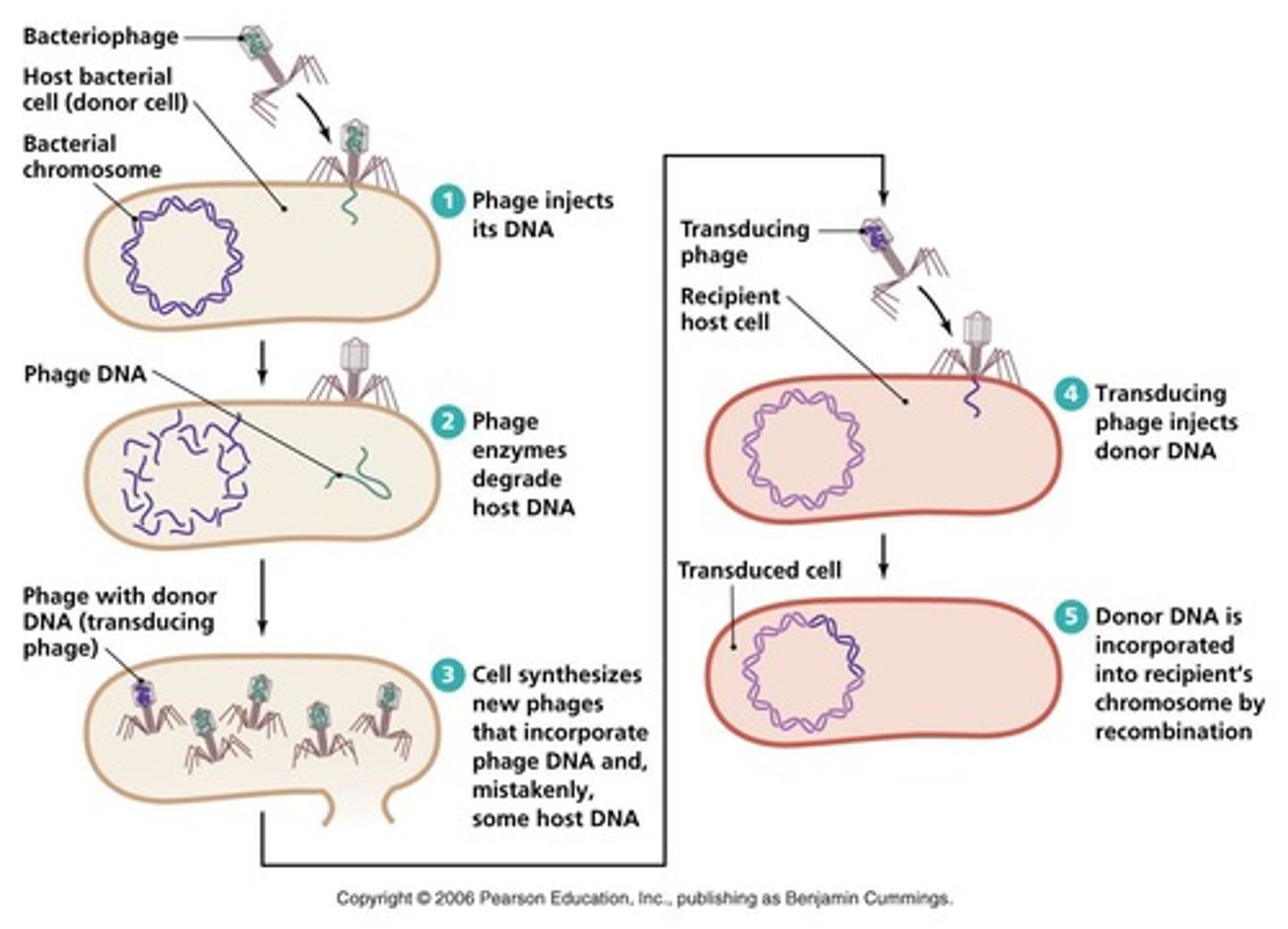
Viruses that infect and replicate inside bacteria.
Bacteriophages
The phage injects its DNA into the bacterial cell, uses the host's machinery to replicate, and then lyses the cell to release new phages.
Bacteriophage - Lytic Infection
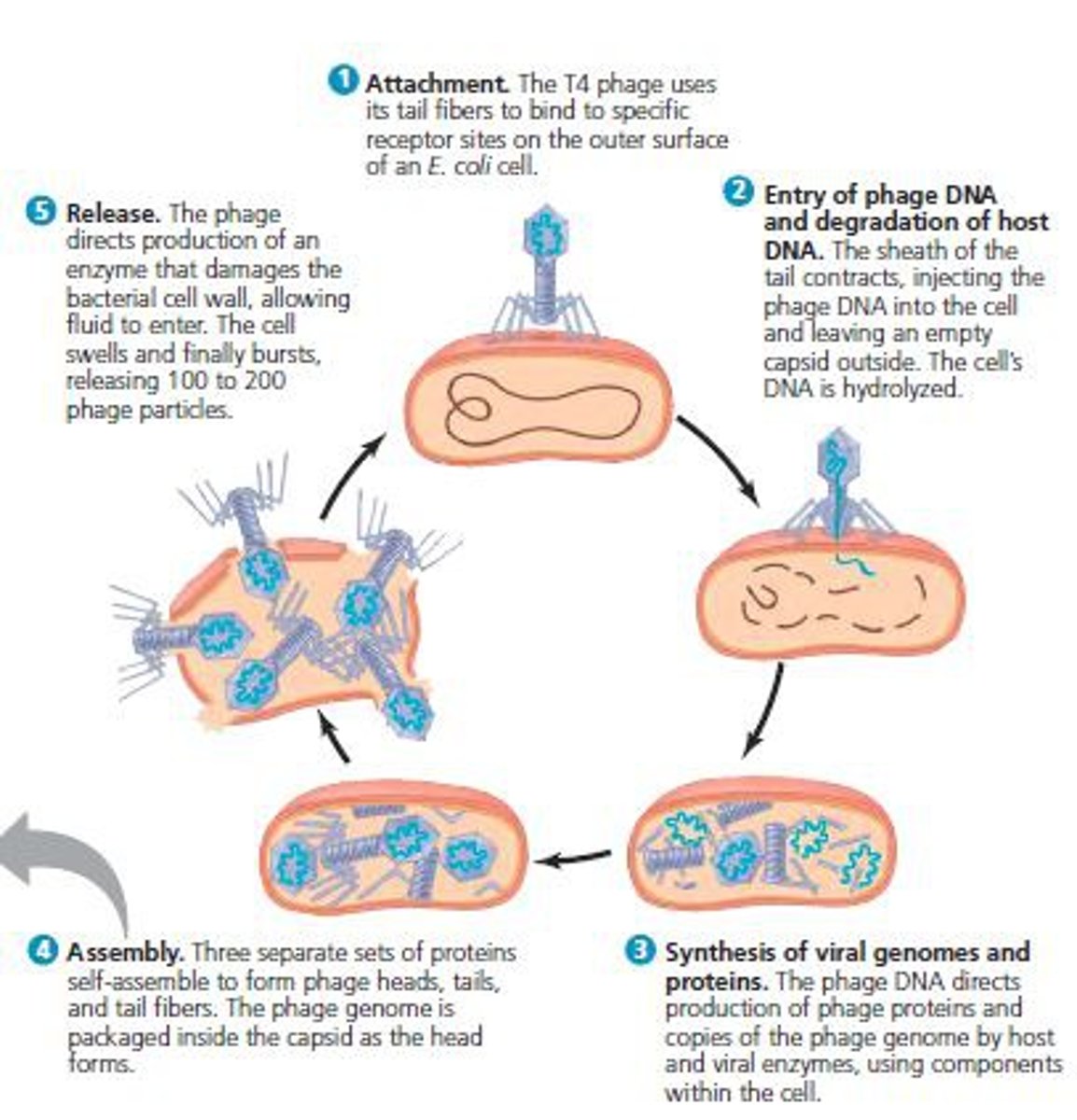
Dormant viral DNA that is incorporated into the bacterial chromosome during lysogenic infections.
Prophage
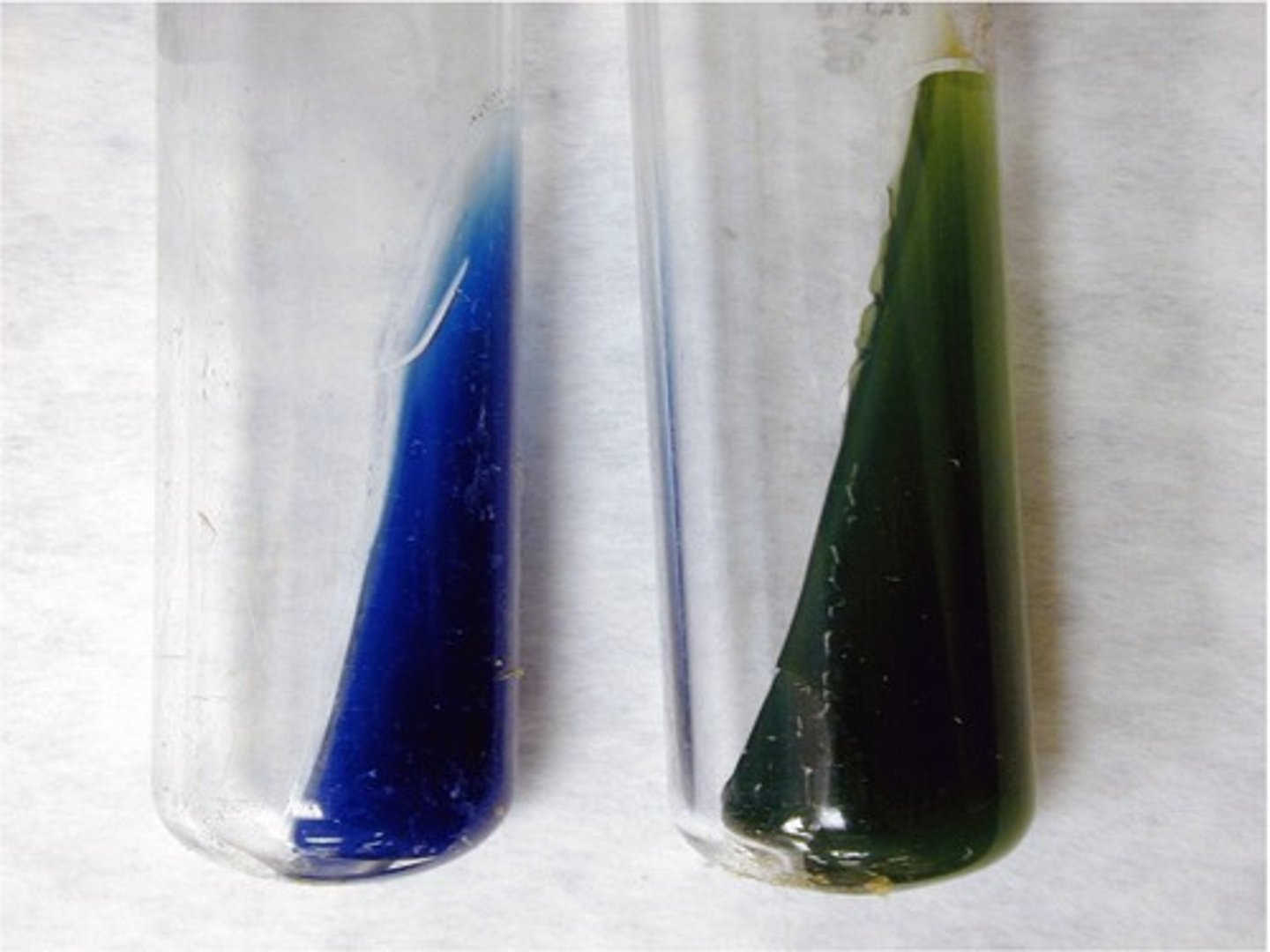
Simmon’s Citrate Agar
Sodium citrate, which bacteria with citrase enzyme can use to produce ammonia.The agar changes from green > blue due to an increase in pH; Ammonia produced (bromothymol blue).
Blue = positive citrate
Green = negative citrate
What is this?

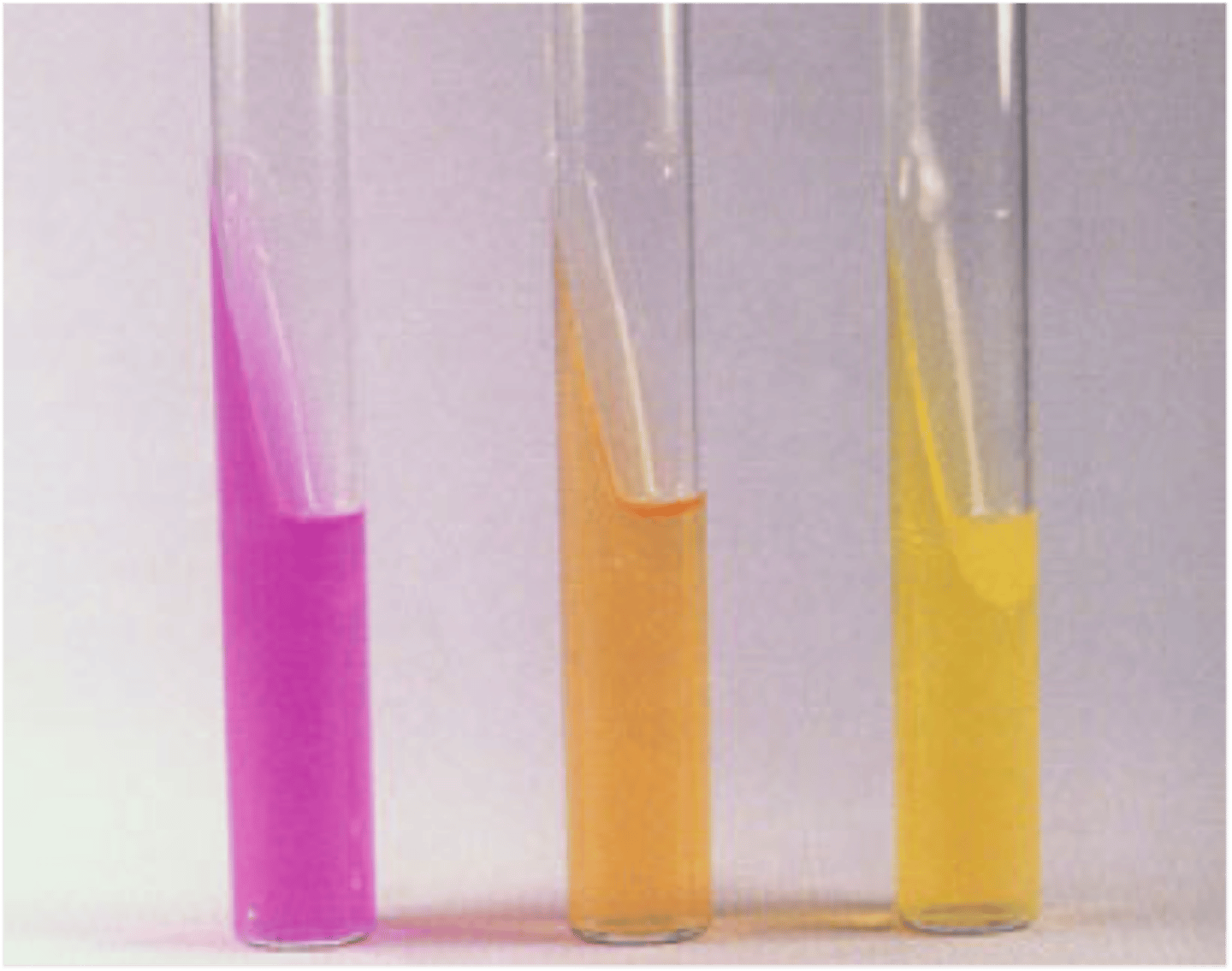
Urea Agar
Hydrolyzes urea, releasing ammonia and carbon dioxide, which raises the pH.
Pink = Positive (Alkaline)
Orange = Uninoculated or Negative
Yellow = Negative
(NH2) 2CO + 2 H2O (Urea) > Urease > CO2 + H2O + 2 NH3 (Carbon Dioxide, Water, Ammonia)
What is this?

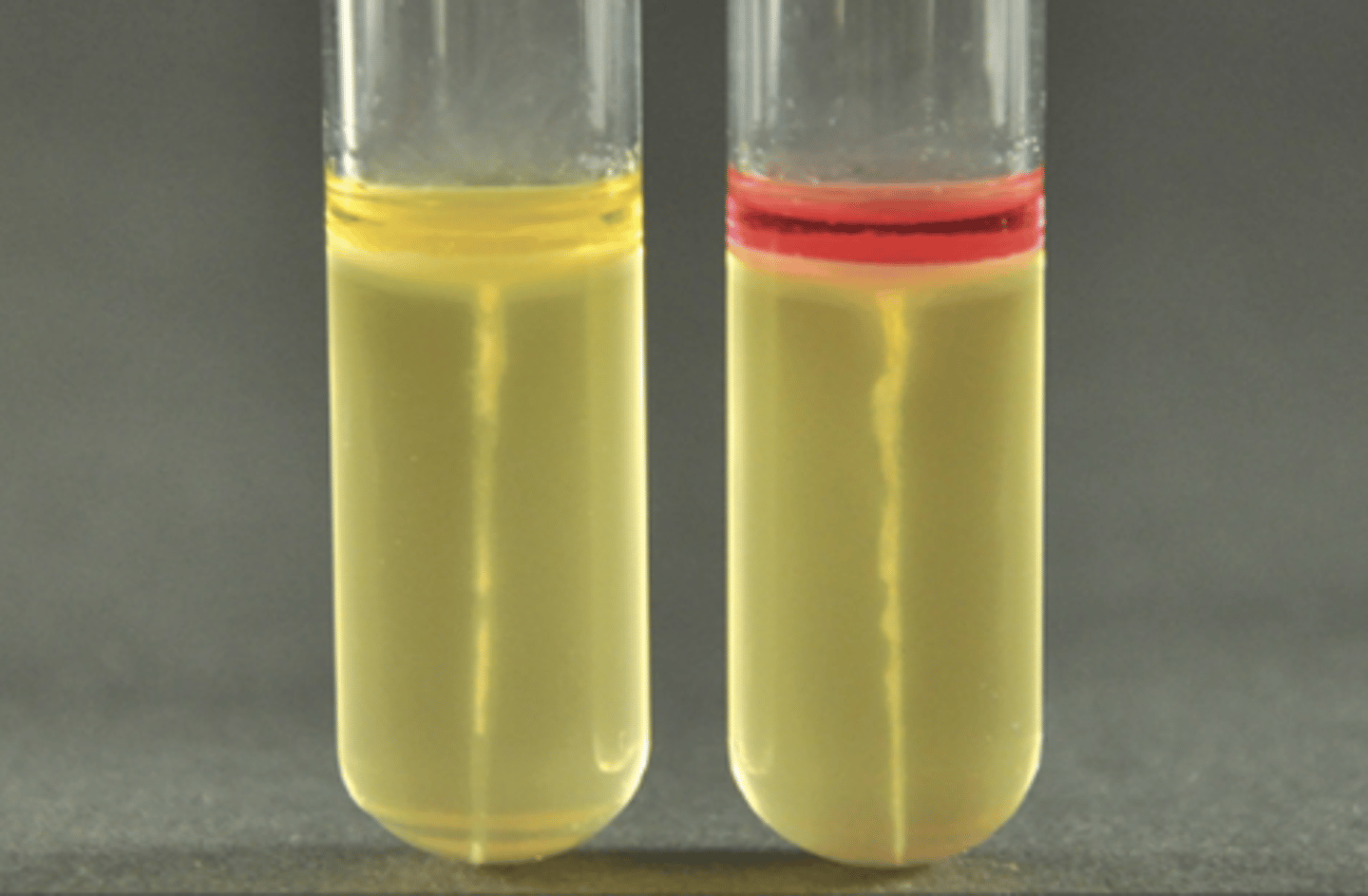
Sulfide-Indole-Motility (SIM) Medium
Tests for sulfur reduction, tryptophan metabolism, and motility.
Yellow = Negative
Black = Positive (H2S gas present)
What is this?
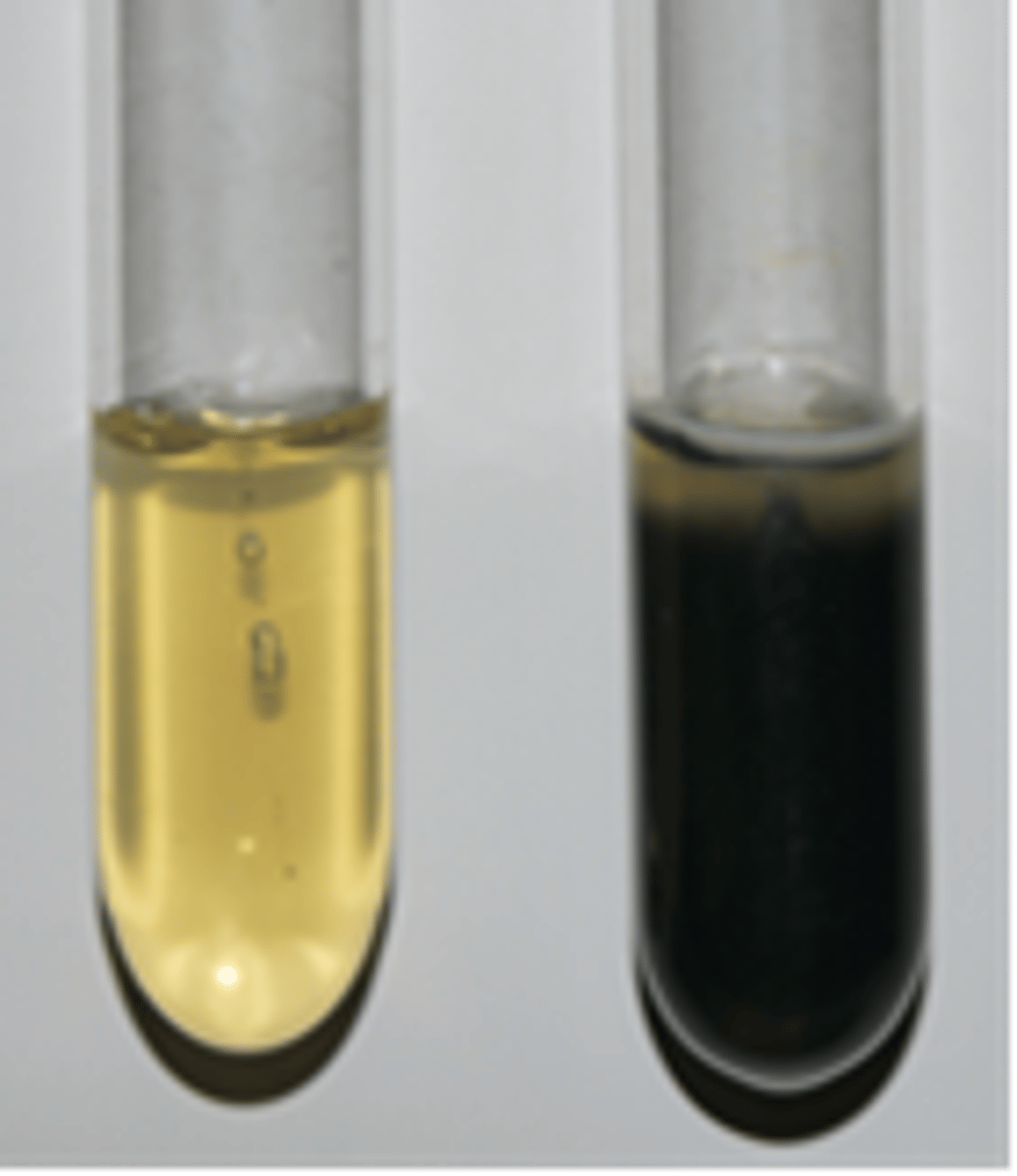

It indicates that the bacteria are motile.
What does cloudiness extending from the stab line in SIM Medium indicate?
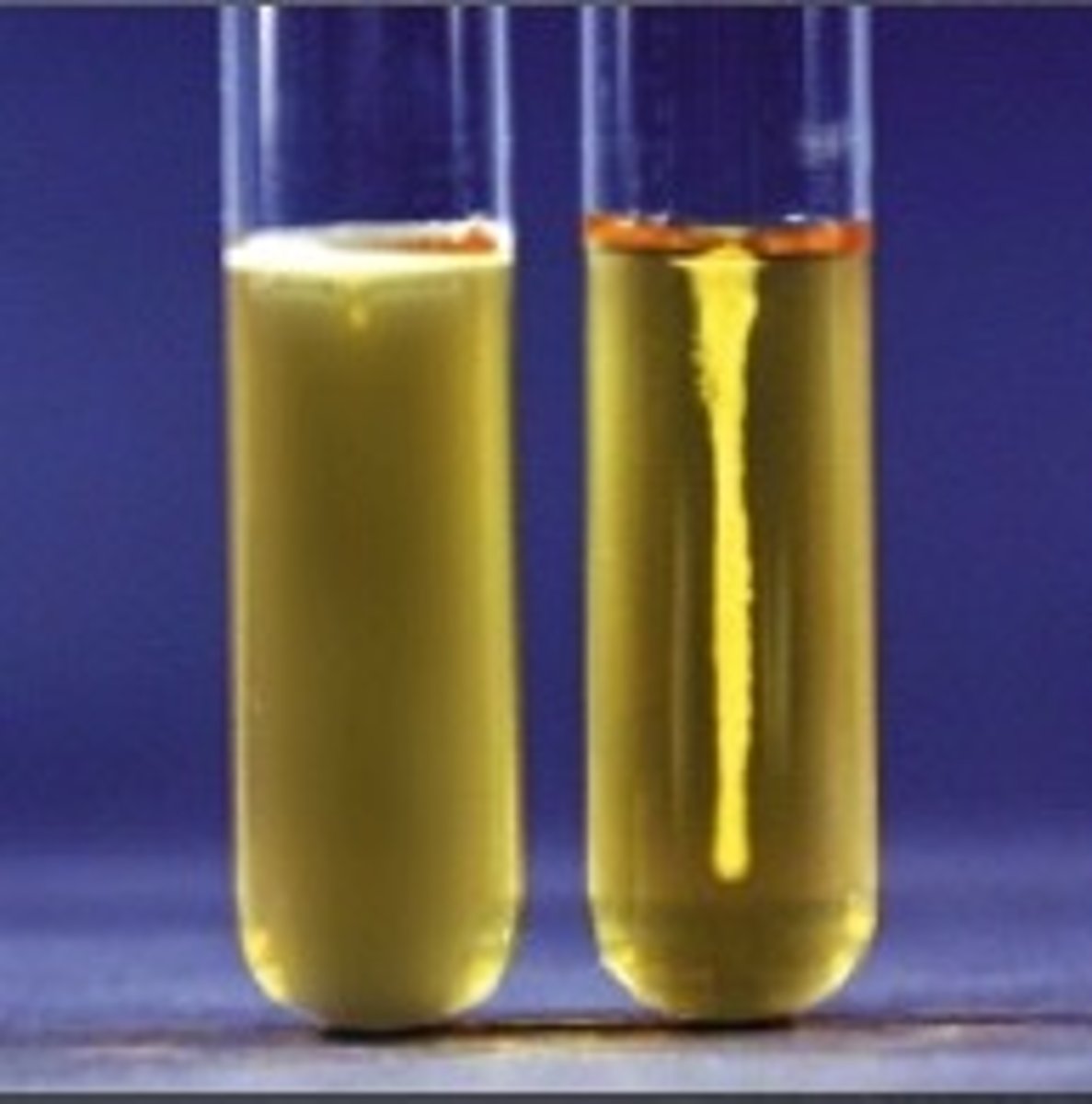

SIM Medium
Pink = Indole positive
Yellow = Indole negative
What is this?

Added to SIM to detect indole; contains DMABA; reacts with indole to form rosindole (cherry red compound).
Kovac's Reagent
Some bacteria have the enzyme, tryptophanase, which hydrolyzes the amino acid, tryptophan. This hydrolysis produces indole and pyruvate.
Tryptophan Metabolism (Indole)
Triple Sugar Iron (TSI) Agar
The bacteria fermented glucose and either sucrose or lactose, producing acid.
Yellow = Acidic pH (A)
Pink-red = Alkaline pH (K)
What are these?
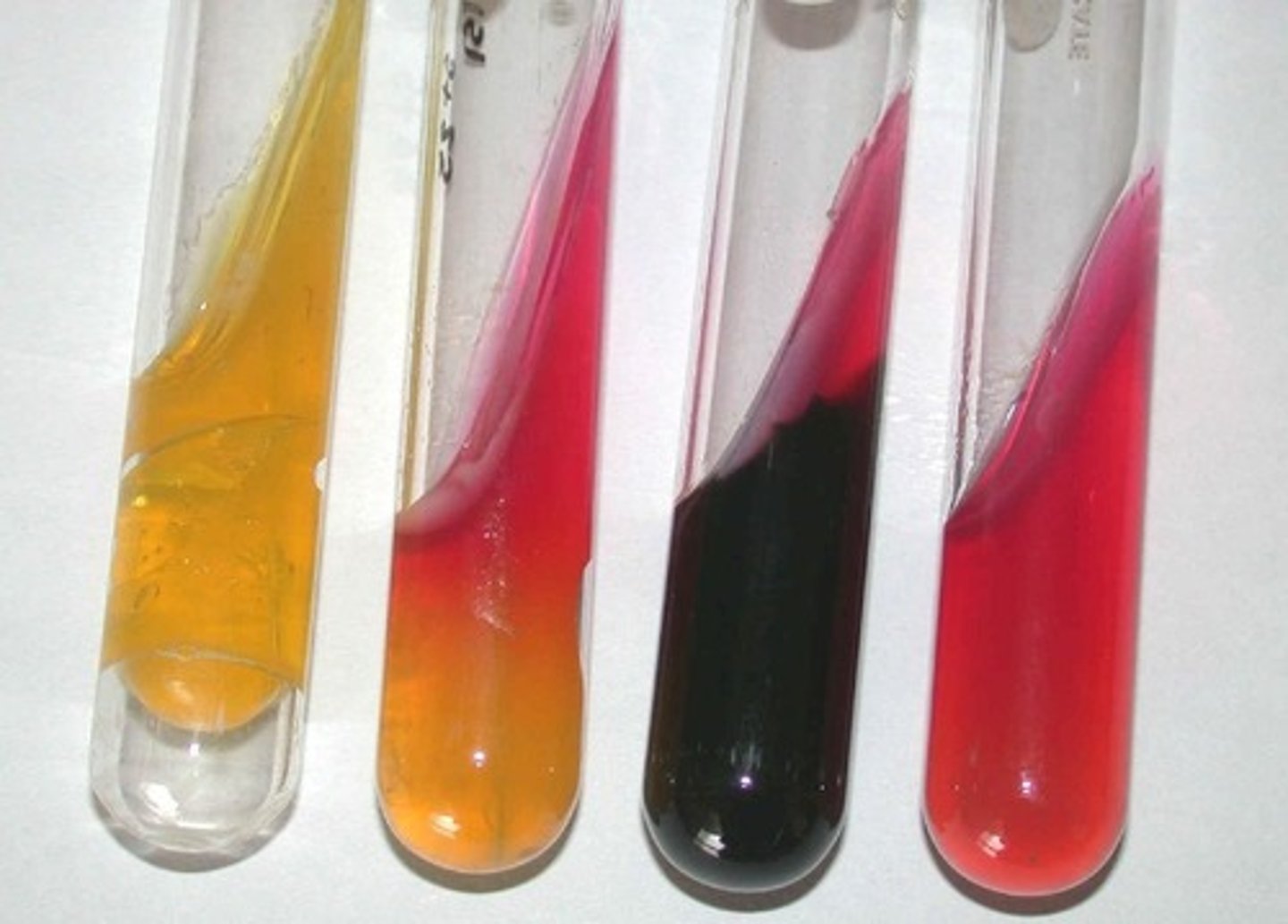

Yellow slant and butt in TSI Agar (A/A)
The bacteria fermented the glucose and either the sucrose or lactose (or both), resulting in a yellow slant and butt. Pockets = gas.
What is this?

Pink-red slant and yellow butt in TSI Agar (K/A)
The bacteria fermented glucose but could not use sucrose or lactose, leading to peptonization.
What is this?
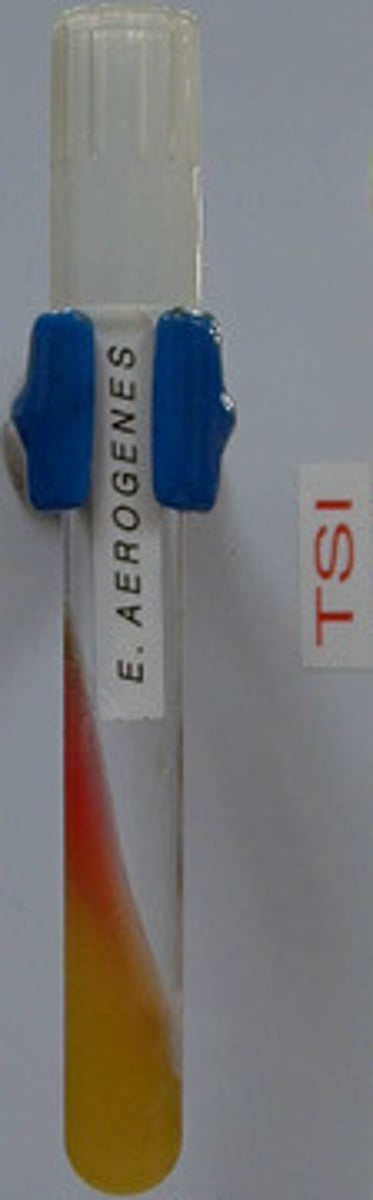
Red slant and butt in TSI Agar (K/K)
The bacteria did not ferment any sugars and only used peptones; alkaline reversion/conversion
What is this?

Indicates that bacteria reduced sulfate during anaerobic respiration, producing H2S gas.
Black medium in TSI Agar
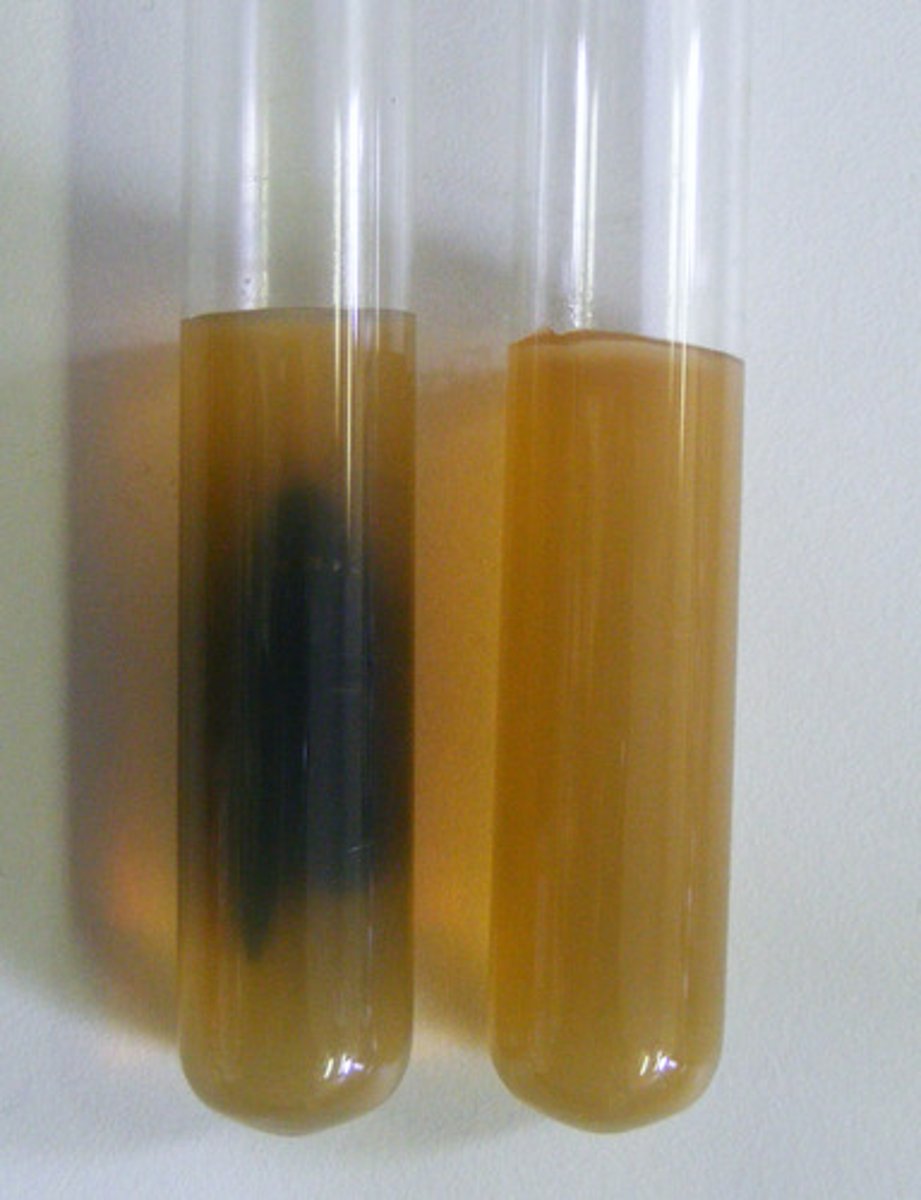

pH indicator in TSI Agar that changes color with acidity.
Phenol Red
Occurs when the bacteria initially rapidly ferment all the sugars, turning the media yellow. After depleting the sugars, the bacteria then use the amino acids for food and the media turns red.
Alkaline reversion/Conversion
Small, non-resistant colonies that grow around a central colony due to beta-lactamase diffusion.
Satellite colonies
Toxic metabolic product of cellular respiration (ex. hydrogen peroxide).
Reactive Oxygen Species (ROS)
Is pyogenic (pus-forming) and can cause abscesses and systemic diseases.
Staphylococcus Aureus
MRSA causes thousands of deaths annually in the U.S. and is resistant to common antibiotics.
Methicillin-Resistant Staphylococcus Aureus (MRSA)
Is typically not hemolytic on blood agar, it is halotolerant and generally ferments mannitol. It is coagulase negative.
Staphylococcus Saprophyticus
Is non-hemolytic on blood agar, halotolerant, does not ferment mannitol, and is coagulase-negative.
Staphylococcus Epidermidis
May be found in gastrointestinal tract. It's an opportunistic and is associated with nosocomial infections.
Enterococcus Fecalis
Coagulase, which coagulates blood plasma.
What enzyme is produced by Staphylococcus aureus that aids in its pathogenicity?
Is an enzyme that breaks down hydrogen peroxide (H2O2) into water and oxygen (H2O + O2).
Staphylococci species = catalase-positive
Catalase
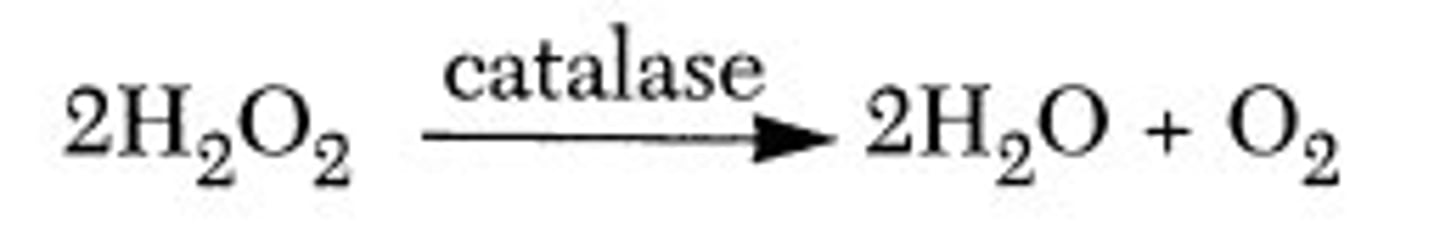
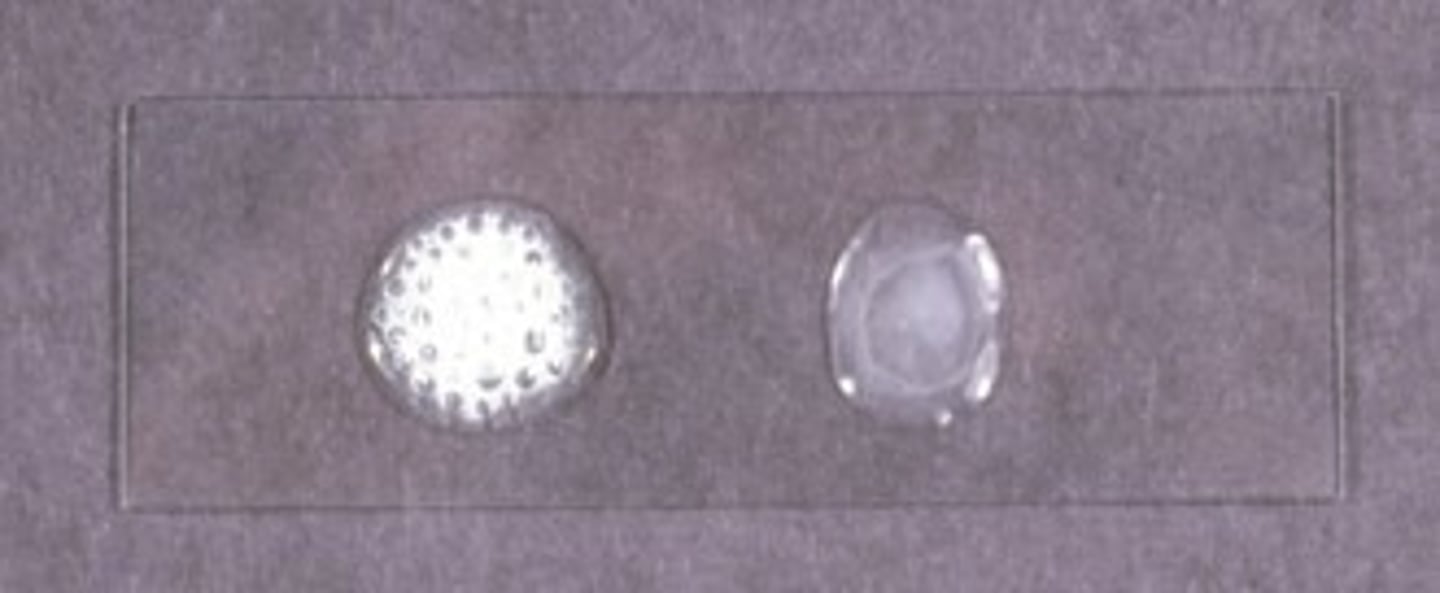
Bubbles = Catalase-positive
No bubbles = Catalase-negative
Catalase test
