Biophys 420 Exam 3
1/328
Earn XP
Description and Tags
Topic 3, Methods of Structure Determination
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
329 Terms
What are the different sample conditions you can use NMR for
solution (micelles), solid state (powder, gel, crystal) cell extracts in vivo, perfused organs (MRI scale)
Why has NMR been used so frequently for membrane structure solving
solving structures is hard when you extract the protein outside of its native environment, ex hydrophobic effect is dependent upon it being in the membrane so removing it causes aggregation
How accurate is NMR structural information
structure determination of protieins and nucleic acids up to 40-50 kDa
How does NMR get structure determination of proteins and nucleic acids up to 40-50 kDA
measuring distances, torsion angles, H-bonds, alignment tensors
How does NMR get structure determination of proteins and nucleic acids above 50 kDA
rapid identification of ligand binding sites, associated conformation changes,
How is NMR useful for protein shuttling / acid base chem
can monitor the change of protons and the environment that they are in directly into the protein
How can you do selective observation of specific sites/components with NMR
isotopic labeling
What are all the purposes of NMR
diverse sample conditions, lots of structural information, selective obervation of sites, macromolecular dynamics on picosecond timescale, reaction kinetics
Frst step of NMR spectroscopy
generate EM radiation
Second step of NMR spectroscopy
send EM through sample, see if EM energy (E = hv) matches energy separation between states in molecules to see if EM will interact and achieve resonance
Third step of NMR spectroscopy
Detect sample induced changes in EM via absorbance, emission, polarization, etc
What is the bore of the NMR machine
cylindrical cavity inside of an NMR machine where the sample is placed
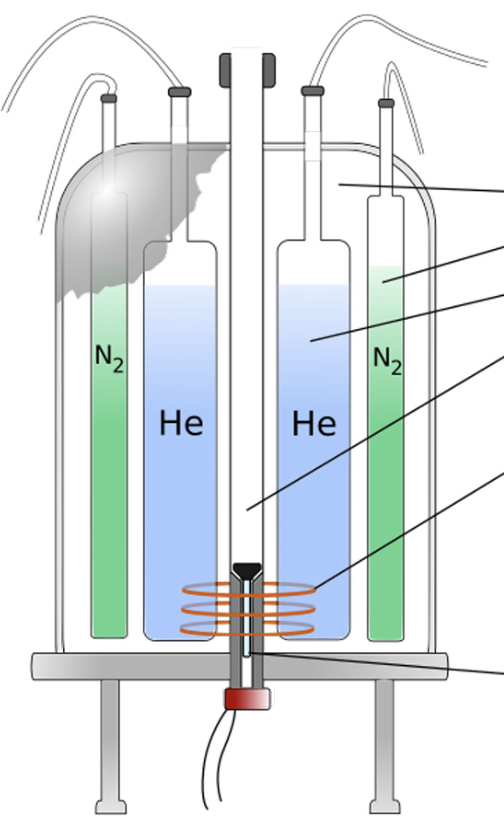
what is the superconducting coil of the NMR machine
uses power and cold temperatures to generate a force that makes metals stick to it
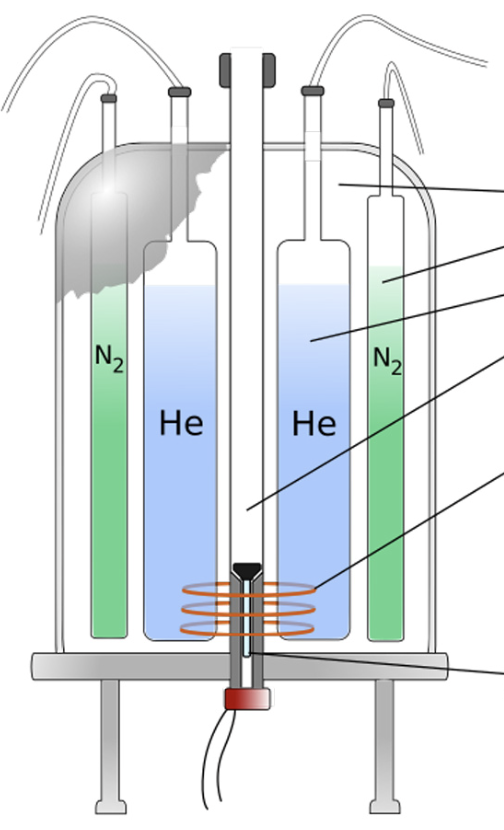
What are the two liquids in an NRM machine
nitrogen, helium
why does the NMR use liquid
cryogenic coolants, maintain the superconducting magnet at low temperatures
What are the shims in the NMR machine
electromagnets used to maintain uniformity/homogeneity of the main magnetic field across the entire sample
Why do superconductors at higher powers need fancier equipment
need to be colder to have the superconducting properties needed
Describe the necessary amount of concentration/sample volume for an NMR machine
high concentration, relatively low volume
What volume advantage does NMR have over x-ray crystallography
requires less sample
Which sample measurement technique requires the lowest sample amount
cryo-EM
How does NMR fix the signal-to-noise ratio problem
repeats experiment multiple times, hundreds of pulses, average the experiment across this
noise averages to 0, signal increases as it averages additively to measurements
What is the NMR formula for signal to noise (S/N) ratio
(number of scans) ^ (1/2)
as # of scans increases, more signal and less noise
What is the change in energy /field ∆E that you’re measuring for NMR based on
quantum property of nuclear spin (I)
What is the purpose of nuclear spin for NMR
Confers a tiny magnetic field (nuclear magnetic moment)
Magnetic moments are randomly oriented without an external field
With an externally applied magnetic field, moments align in specific energy states
Low energy is aligned WITH field and high energy is aligned AGAINST/antiparallel
∆E between these states is proportional to field strength
What does it mean for NMR nuclear spin to be quantized
Nuclear momentum/orientation along the external magnetic field have discrete values ranging from 0-6 (in steps of ½ because of contributing electron spin)
Why can’t you use nuceli with an even number of both protons and neutrons for NMR
0 nuclear spin, can’t produce an NMR signal
How is NMR dependence on nuclear particle composition leveraged in studies
atoms that are not NMR active (0 spin state) can be made active by isotopic labeling, ex. C13
How are isotopic labeling methods used to boost signals in NMR
replace inactive nuclei with active nuclei or vice versa to control what is visible and invisible to the NMR
Why is proton NMR so powerful
ton of spin-half isotopes that exist in biological systems, H1 is super abundant. Most carbon is C12 and very little of it is C13.
What are all the types of labeling schemes for NMR
uniform labeling, amino acid specific, site-specific, subunit/domain specific
How does uniform labeling for NMR work
Label as many of the nitrogens or carbons inside a system as possible
How does amino acid/site/subunit specific labeling for NMR work
intein-based protein ligation
Describe the process of uniform labeling for NMR
grow bacteria in regular media to desired OD600
Switch to defined minimal media with a sole source of your desired nucleus ex. N15 ammonium chloride or C13 labeled glucose
Induce expression, bacteria incorporates it into the protein, broadly labels the whole protein
Why isn’t the process of uniform labeling for NMR 100% efficient?
there is some turnover in the bacteria, bacteria degrades some of their own proteins to make new proteins
Describe the process of amino acid/site specific labeling for NMR
load specific tRNAs with amino acids that are labeled
use that in a system to make the protein
more complicated than dumping in isotopically rich media
What is the nuclear magnetic moment formed by
nuclear spin (rotating charge), direction indicated by the right hand rule
What is the magnetic moment formula
μ = 𝛶 𝐼 ℎ
μ = magnetic moment
𝛶 = gyromagnetic ratio
𝐼 = spin
ℎ = Planck’s constant
What is the gyromagnetic ratio
ratio of a particle’s magnetic moment to its angular momentum, specific to each isotope
What happens to μ (magnetic moment) when there is no externally applied magnetic field
μ will be randomly oriented in space, Gaussian of a random species
What happens to μ when there is an external magnetic field applied
Magnetic moment attempts to align to field
Moment is quantized, can only adopt certain orientations
Experiences torque that causes it to wobble about the magnetic field at the Larmor frequency
Measure the vibration that occurs in the system around this
External magnetic field applied around z axis
What is the larmor frequency
rate of wobbling of the magnetic moment around the magnetic field due to torque from the quantized aspect
What is the larmor frequency formula
w0 = - 𝛶 * B0
w0: frequency
𝛶: gyromagnetic ratio
B0: magnetic field strength, closest the magnetic moment can get to the magnetic field
How do nuclei align in the case of an I = ½ nucleus and an external magnetic field B0
nuclei align either up or down relative to the field
up states slightly more populated than down states
Why are the “up” oriented nuclei more popular than the down states
slamming a powerful magnetic field in a particular direction: align with mag field, hit with EM pulse to throw some species up, observe as they decay back down to native speces
in spin you have positive or negative spin states that align with the magnetic field
overall a slight delta in the positive direction because you’re applying a large external field the system
Why is the difference in population of nuclei in up and down states in NMR important
wouldn’t be able to measure anything if they were equal
population difference determines net signal and sensitivity
For a signal to be detected, there must be more nuclei in the ground state than the excited state If the population ratio was 1:1, there would be no net absorption of energy, resulting in no signal.
What is the purpose of the radio frequency pulse addition to the populations of up and down states in NMR
some of the population shifted from up to down state
magnetization then relaxed/decayed back to original state
Energy released during decay, measured by reciever
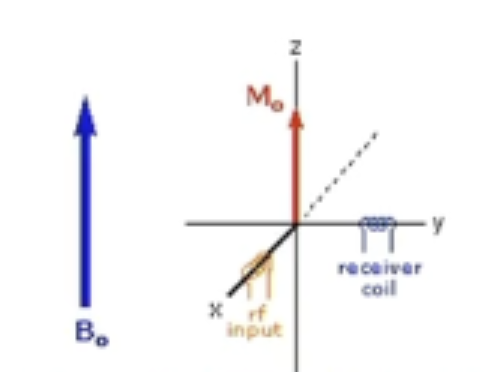
Why is it important that receiver measures in the perpendicular direction in NMR
Receiver is placed perpendicular to the magnetic field to measure perpendicularly
spins are in the XY plane
magnet B0 in the z axis
Receiver coil is perpendicular to transmitter to make sure sensitive receiver not overloaded by high power RF pulse
Otherwise would only see pulse, no change due to magnetic spin about z-axis
Why was it important to standardize NMR?
with more and more powerful magnets, you change the population of states in up versus down
changes effective field strength, magnitude of quantization gets larger
spectra on one instrument less powerful would have different chemical shifts
is the observed frequency in NMR the same as the larmor frequency
depends on local environment of individual nucleus (ex. shielding or exposing), differs slightly from Larmor
What is the chemical shift for NMR
difference in resonance frequencies due to local enviroment
Is the magnetic field at the nucleus the same as the applied magnetic field
no, due to chemical shift
In practice, the chemical shift is defined relative to?
the reference resonance signal
Do NMRs of the same strength have the same chemical shifts?
No, in raw values only have the same shifts when defined relative to a defined 0 point
what is the relation between chemical shift and magnetic field strength
independent of each other
What is an experimental limitation of NMR
relatively insensitive as a technique (super small diff between the populations of up and down magnetic spin states, need lots of signal in order to see anything out of random nucleic noise)
How can you get around NMR insensitivity through sample prep
High concentration, more volume than cryo-EM, relatively low salt
Why do NMR samples need lower salt
ionic strength of solution as you’re applying RF field through it induces currents the stronger the ionic strength is, obscures data by adding random noise and can cook the sample by generating heat with induced current
Why can high concentration of sample of protein be an issue for proteins
some proteins don’t like to be concentrated form aggregates
How can you get around NMR insensitivity through experimental changes
repeats the same experiment multiple times to improve signal to noise ratio
what is the relation between signal to noise ratio and number of scans
proportional to square root of # of scans
Explain NMR with handbell frequency
Nucleus of each atom rings at different frequency depending on size, magnetic moment, overall field strength, applied RF pulse
The ringing of external atoms connected together bleed into each other, can determine where things are in the system but signals can bounce between a lot of diff things
Different ringing frequencies allow you to get different chemical shifts and see individual atoms, not just one massive broad peak
How is carbon frequency different than hydrogen frequency in NMR
Carbon rings at a lower frequency because it’s heavier, bigger, harder to spin. Hydrogen rings at a higher frequency because it’s smaller, easier to disturb
How are fourier transforms used in NMR signals
converts a time domain to a frequency signal (NMR signal/ringing to frequency)
What does the ringing of each atom’s nucleus depend on
its identity, identity of neighboring atoms, overall strength of magnetic field itself
What is the formula to convert frequency to ppm?
chemical shift (ppm) = (resonance frequency of atom - resonance frequency of standard (always 0 ppm) * 10^6 ppm / operating frequency of the magnet
Describe the ease of interpretation 1D NMR for proteins
crowded and overlapped, hard to interpret
Why is NMR harder for bigger molecular weight molecules
more proteins obscured in the center of a species, hard for them to be chemically distinct. wrapped them up, each nuclear spin is shielded and surrounded by diff aromatic residues. Affected by coulombic interactions, salts, hydrogen bonding
What solves the problem of 1D NMR being hard for large molecules?
Multidimensional NMR, to separate out the peaks
How does multidimensional NMR connect 1D peaks
Interaction between nuclei observed in the 1D spectra, ex. J-coupling or NOE
atoms close to each other have similar properties when hit with an EM pulse, can be correlated
in HSQC, what does the 2D NMR take advantage of
takes advantage of coupling between atoms because they are connected by chemical bonds and shared within space
in 2D NMR, magnetization only happens if:
two atoms are connected via covalent bond (J-coupling). Probing one to ring transfers ringing to the other atom to make it ring too
close in space (<5 A)
What is coupling in NMR
when 2 nuclei are coupled, the activity of one nucleus affects the activity of the other nucleus
How is 2D NMR visualized
stacked plots and contour plots
Describe what a stacked plot looks like
2D surface with peaks emerging from the plane, wherever there is a peak, there is a correlation between those two atoms
Why is there a bunch of signals along the center diagonal in stacked NMR plots
that represents atoms that are physically the same, always going to be correlated with each other
what is a key advantage of a stacked plot
easy to see intensity differences between peaks, strengths of the interactions between atoms
What is the most common way to visualize 2D NMR data
contour plots
Why are contour plots more commonly used than stacked plots
stacked plots are complicated and hard to interpet
What does a contour plot look like
topographic map, different rings indicate steepness of the peak
How does weak peak visualization differ between stacked and contour plots
You can see more of the weaker peaks in a contour plot, whereas they’re hidden behind larger peaks in the stacked plot
Why is HSQC (proton nitrogen) the most common experiment for protein structure determination
relatively fast to acquire, 10 mins for 500 micromolar, cheap (15-N labeling)
What does HSQC (proton nitrogen) experiment allow for
fingerprinting, comparing where positions of amide/peptide bond and nitrogen containing side chains.
Why is HSQC called fingerprinting
you can get overall relative position of a lot of the atoms in a species
What are the benefits of 15-N labeling in HSQC
Eliminate aliphatic aromatic protein peaks
Only observe 15N1H pairs
15N chemical shift provides a second dimension that spreads out peaks with the same proton shifts that would otherwise overlap, side chains are further spread apart in spectra than if you just took two proton spectra and correlated them
What are the applications of H-15N HSQC experiment
rapid identification of folded proteins and monitoring ligand binding
How is HSQC NMR used for identifying protein folding
Chemical shift dispersion or range of many protons is much worse in unfolded proteins than in folded proteins, can qualitatively assess folding of recombinant protein fragments. Useful for intrinsically disorderd regions, transcription factors, or other instances with lots of structural flexibility.
Why is there a broadening of ppm shifts in unfolded proteins
most of the protons and nitrogens will see the same environment, surrounded by a large bulk of solvent with the same forces and interactions
Why is the ppm shifts in folded proteins more narrow
specific hydrogen bonds, things become more specific in a system
If you have a large broad peak, the protein is likely to be ?
unfolded
If you have distinct peaks that are more spread out, the protein is likely to be ?
folded
Why is being able to rapidly identify folded proteins so helpful for intrinsically disordered regions
you can monitor the folding process, can quantify the peaks going from very broad to sharpening up to individual peaks that start to separate out better than initial broad peaks
How is HSQC NMR used for monitoring ligand binding
monitoring physical position of atoms in your ligand assuming ligand or protein is labeled in some fashion
titrate ligand into 15-N labeled protein, monitor chemical shift cahnges
measures Kd by plotting shift vs ligand concentration
can map which residues bind to ligand if you know what resonance belongs to what residue
timescales important
How does the timescale analysis component of ligand binding monitoring work for HSQC NMR
P L complex is in rapid exchange with P + L
As it comes together you can tell whether it’s diffusing together or snapping together
What dictates the PL complex’s exchange with P + L
∆w
What happens if Kex chemical exchange rate is > ∆w chemical shift difference
avg 𝛿 is observed, one averaged peak
What happens if Kex chemical exchange rate is < ∆w chemical shift difference
two resonances are observed, not averaged
How can you figure out what resonance belongs to what residue for HSQC NMR?
Make a mutation and look for what resonance disappeared/moved. But super time consuming and not feasible.
Use a third frequency as a third dimension, needs uniformly labeled protein with both a 15N and 13C atom. Time consuming, needs to be repeated to assign peaks
What is 3D NMR
Triple resonance-based assignments, based on comparing data acquired from pairs of 3D NMR experiments
What do all 2D or higher spectra require for the nuclei they link?
Scalar coupling or NOE interaction between the nuclei that they link, in order to identify peaks
in 3D NMR, what is each peak associated with
3 chemical shifts (amide hydrogen, nitrogen, carbonyl)