BIO 190- Ch #12 Gene Regulation
1/102
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
103 Terms
gene regulation
the ability of cells to control the expression of their genes
"turn genes on or off"
constitutive gene
genes that are always on, their promotors are always open
how do many proteins regulate gene expression
through binding DNA
How must RNA pol be guided to gene promotors
by transcription factors that say "come here"
what happens if a promotor is hidden
then RNA pol can not attach/ find the promotor
how do some transcription factors reveal gene promotors
by changing nucleosome spacing
what do some transcription factors attach to histone proteins to loosen the hold on DNA to reveal promotors
attach functional groups to histones
how do genomes and proteomes differ
all cells have the same genome, but different proteomes
what is cell differentiation
the process by which cells become specialized into particular types by expressing specific genes at specific times
what does hemoglobin do embryonic vs post natal
embryonic: picks up oxygen from uterine environmental/ maternal blood
post natal: picks up blood in lungs
describe the different quaternary composition of hemoglobin in embryonic, fetal, and post natal
embryonic: 2 sigma and 2 zeta (oxygen affinity highest)
Fetal: 2 alpha and 2 zeta (high oxygen affinity)
post natal: 2 alpha and 2 beta (moderate oxygen affinity)
what do most promoters include
a core promoter (essential for the start of transcription)
and regulatory elements (modulate how much, when, and where a gene is transcribed)
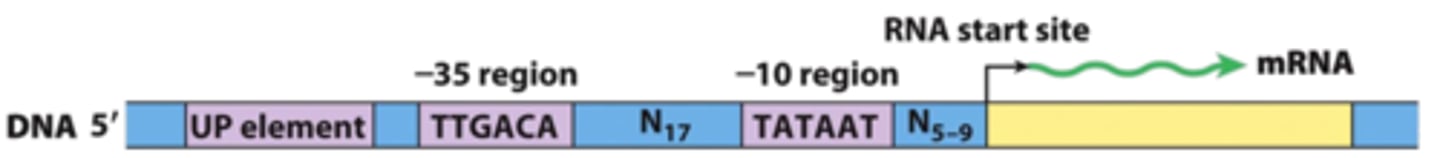
what does the core promoter contain
TATA box and transcription start site
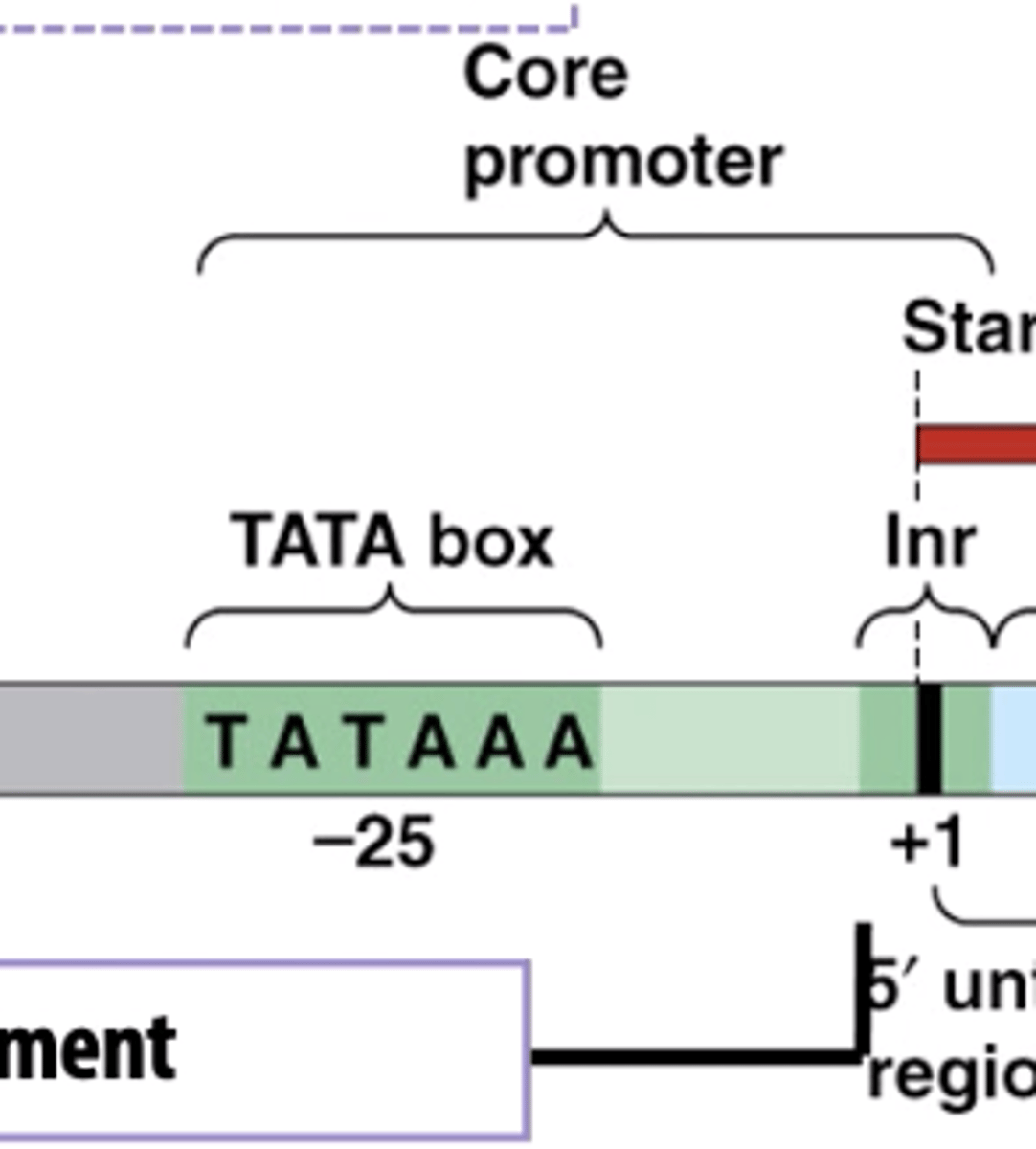
the core promotor alone results in what
a low level basal transcription
The core promoter is enough to start transcription, but only at a very low level. This low level is called basal transcription — it’s the minimum amount of transcription that happens even without help from additional regulatory elements (like enhancers).
what does the +1 mean on the core promotor
refers to the transcription start site (TSS) — the exact spot on the DNA where RNA polymerase II begins transcribing the gene into RNA
+1 site = First nucleotide of the RNA transcript.
transcription factors bind to ______
gene regulatory elements
what are the two types of gene regulatory elements
enhancers and silencers
enhancers vs silencers
enhancers: enhances RNA pol ability to find and bind to promotors
silencers: inhibits RNA pol ability to find/bind to a promotor
what does RNA polymerase II do
transcribes genes that encode proteins
what does TF2D bind to
the TATA box and is a common target for activators and repressors
how do activators and repressors commonly exert their regulation of transcription
through affecting the function of general transcription factors
what does it mean by most eukaryotic genes are under combinatorial control
that most expression is regulated by the combination of many factors
what are activators
stimulate RNA polymerase to initiate transcription
what are repressors
inhibit RNA polymerase from initiating transcription
what are modulation
small effector molecules, protein- protein interactions and covalent modifications can modulate activators and repressors
what is chromatin structure
activator proteins promote loosening up the region in the chromosome where a gene is located, making it easier for RNA polymerase to transcribe the gene
push nucleosomes apart so promotor is visible
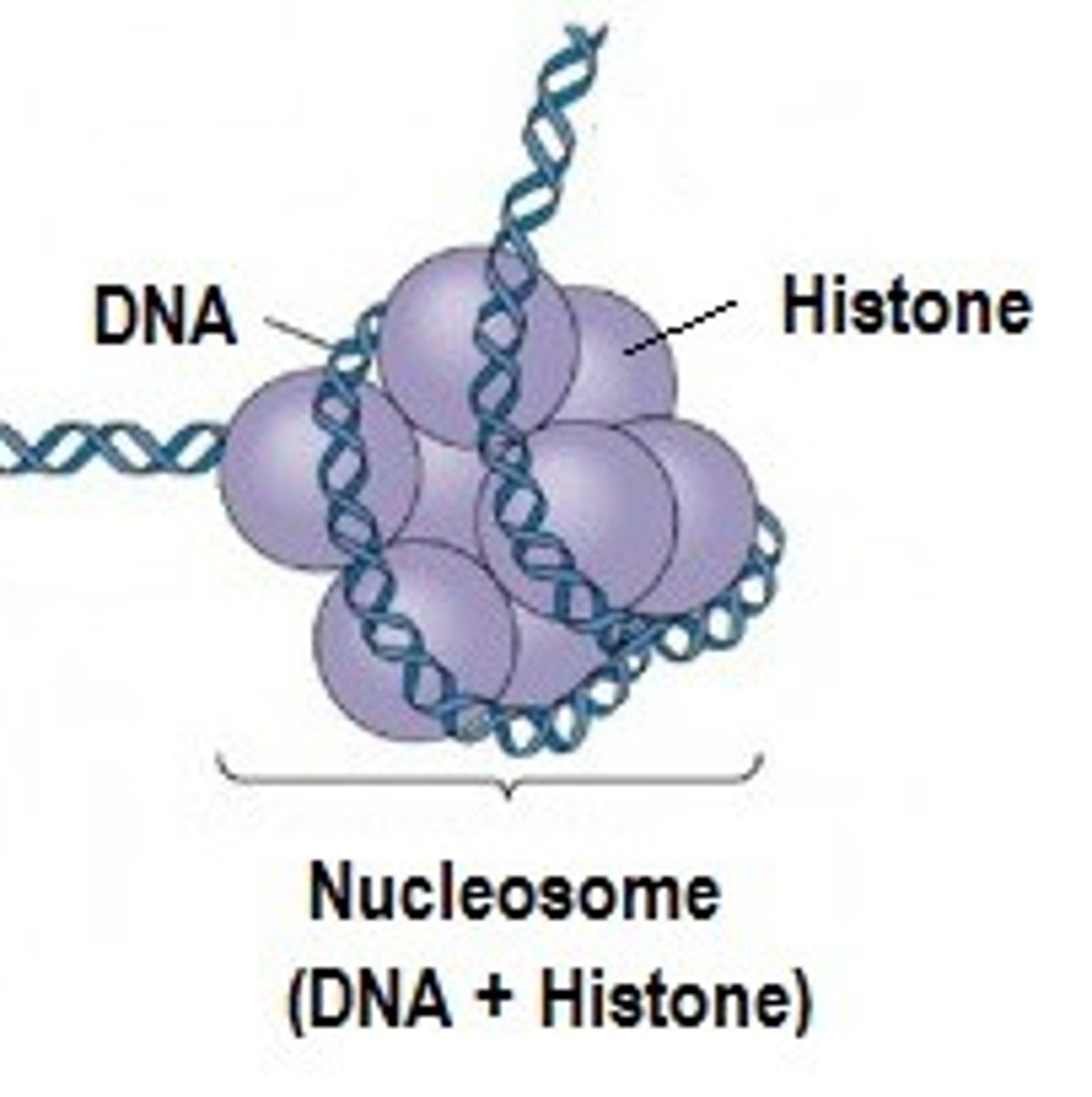
what is DNA methylation
usually inhibits transcription either by blocking an activator protein or by recruiting proteins that inhibit transcription
if eukaryotes do not have operons how do they turn on (express) multiple genes at one time
Multiple RNA polymerase II enzymes can transcribe multiple genes across the genome at the same time.
can a chromosome have both open and closed regions
yes
describe closed conformation in chromatin structure and an example
a region that is difficult or impossible to transcribe; transcription requires changes in chromatin structure. "constitutive heterochromatin"
ex: when you are awake the sleeping gene is closed in chromatin
describe open conformation in chromatin structure and an example
is accessible to general transcriptional factors and RNA pol II and can therefore be transcribed
facultative heterochromatin
ex: awake gene is open when you are awake
what are chromatin remodeling complexes
protein machines that use the energy of ATP hydrolysis to change the position of the DNA wrapped around nucleosomes
they can move nucleosomes around
describe how nucleosomes reposition
DNA is in a closed conformation --> activator binds to enhancer --> chromatin remodeling --> nucleosome reposition
what are the 2 types of chromatin remodeling (how chromatin structure open up chromatin)
1. Modify histone tails
2. demethylate cytosines in DNA
how does modifying histone tails open up chromatin
Histone acetylation
adds on a acetylation which “loosens” DNA, making genes neutralized and easier to turn on
how does demethylate cytosines in DNA open up chromatin
think of the methyl group as an oil droplet that drags nucleosomes close to each other through hydrophobic attraction which closes the chromatin
demethylation removes that methyl group/ oil droplet groups so the nucleosomes spread and open chromatin
what is a nucleosome free region (NFR)
used for genes we need quickly, they are found at the begging and end of the gene (they don't need to put nucleosomes at promotors) short stretch of DNA that is not wrapped around nucleosomes, meaning it’s free of histones
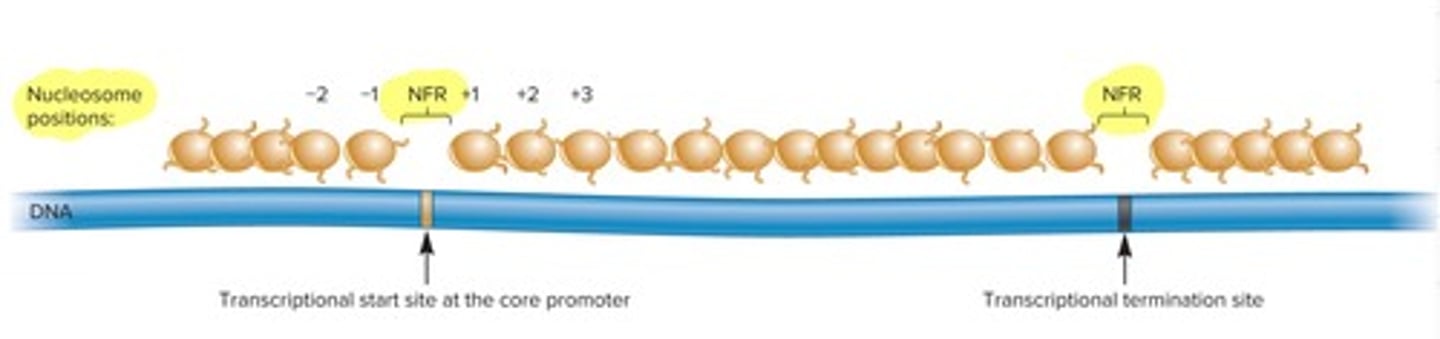
what happens to nucleosomes during transcription elongation
During transcription elongation, nucleosomes are temporarily loosened or rearranged so RNA polymerase II can pass, and then they are reassembled behind it.
does removal of acetyl groups from histone tails decrease or increase gene expression
Removal of acetyl groups from histone tails → decreases gene expression
Acetyl groups neutralize the positive charges on histone tails.
This loosens the interaction between histones and negatively charged DNA, making the DNA more accessible to transcription.
So, when acetyl groups are removed (a process called deacetylation): Histones become more positively charged.
DNA binds more tightly to histones. Chromatin becomes more condensed (heterochromatin) and Transcription factors and RNA polymerase can’t access the DNA as easily.
Removing acetyl groups (histone deacetylation) tightens chromatin and reduces gene expression.
does removal of histones from a region decrease or increase gene expression
Removing histones from DNA increases gene expression by making the DNA more accessible for transcription
Histones package and wrap DNA into nucleosomes, which makes the DNA less accessible to transcription factors and RNA polymerase. Removing histones (even temporarily) from a DNA region: Opens up the chromatin structure
does aceytlation of histone tails decrease or increase gene expression
Histone acetylation opens up chromatin and increases gene expression.
Histone tails have positive charges, and DNA is negatively charged. When acetyl groups are added (acetylation), the positive charges on the histones are neutralized.
This causes histones to bind DNA less tightly, leading to:
Looser chromatin structure (called euchromatin).
Greater access for RNA polymerase and transcription factors.
More transcription of genes in that region.
does a decrease in compaction of histones decrease or increase gene expression
increases gene expression.
Less compacted chromatin (called euchromatin) means:
DNA is more exposed.
RNA polymerase and transcription factors can access promoter and enhancer regions more easily.
This leads to higher levels of transcription.
does an increase in compaction of histones decrease or increase gene expression
decreases gene expression.
When histones compact the DNA more tightly (heterochromatin):
The DNA becomes less accessible to transcription factors and RNA polymerase.
Promoters and enhancers are hidden, preventing gene activation.
As a result, transcription is reduced or silenced.
DNA is associated with proteins to form
chromatin
a ______ is composed of DNA wrapped around an octamer of histone proteins
nucleosome
An activator can increase transcription by attracting a _____ to the region.
histone acetyltransferase
removal of acetyl groups from histones result in a _________ in gene expression
decrease
addition of -COCH3 groups to histones amino terminal tails results in a _______ gene expression
increase
regulation of transcription is efficient however ..
it requires a fair amount of time to drive effects of cell function
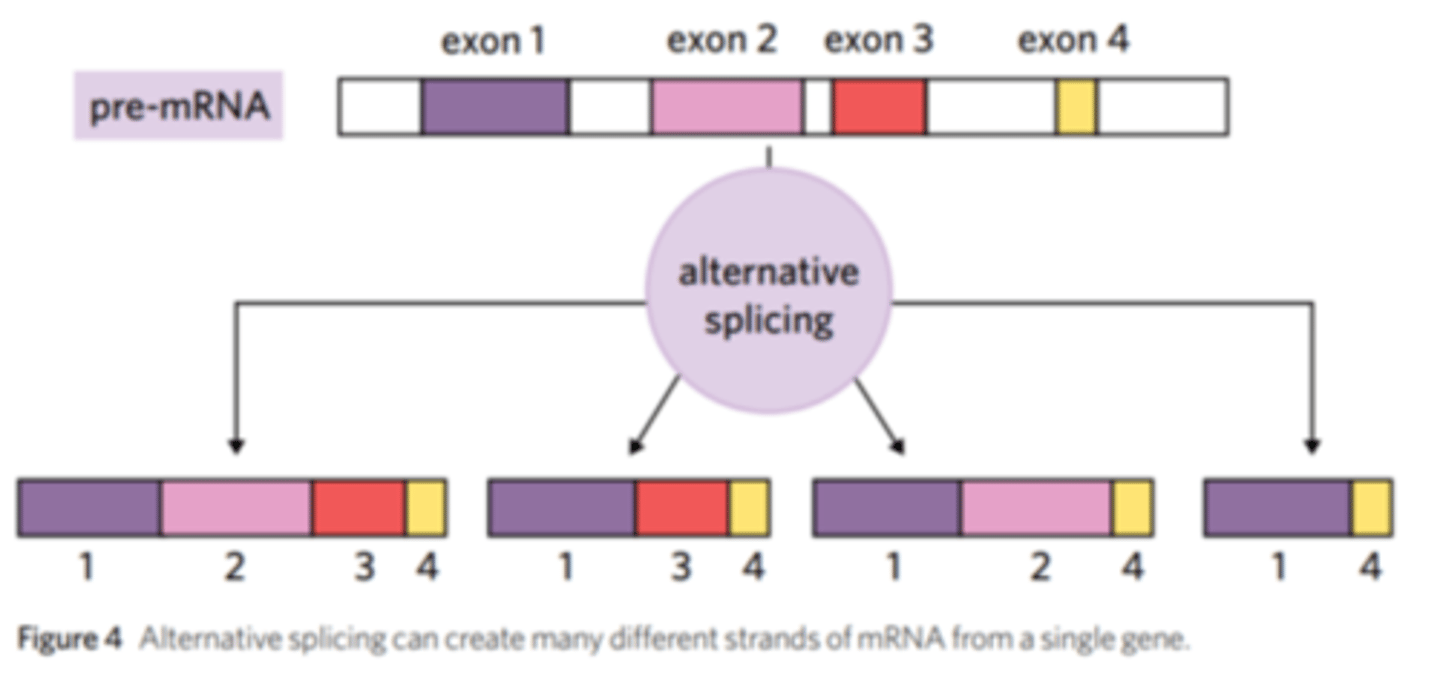
what is alternative splicing and when does it occur
occurs during post translational action
allows an organism to use the same gene to make different proteins at different stages of development
Alternative splicing is a process in eukaryotic gene expression where a single pre-mRNA can be spliced in different ways to produce multiple different mRNA transcripts, and therefore different proteins, from the same gene.
faster regulation can be achieved at the stages of pre-mRNA _______ & ________
splicing and translation
what is ferritin
storage form of iron (iron ion sponges)
how are ferritin levels controlled
by holding its mRNA in standby until needed
what does the active iron regulatory protein do
stops ferritin when iron levels are low you don't want a lot of it (inhibits translation)
what happens when iron levels are high
iron regulatory protein binds to iron causing a conformational change that releases it from the iron regulatory element and translation proceeds.
What level of regulation do eukaryotes possess that prokaryotes do not?
transport RNA out of the nucleus
which of the following is not involved in the control of gene expression in eukaryotes
transcription
modification of mRNA
translation
post- translation
replication
replication
is this a valid reason for a cell to regulate gene expression;
made additional cells of the same type in response to demand
true
is this a valid reason for a cell to regulate gene expression;
synthesize enzymes to metabolize a particular nutrient
true
is this a valid reason for a cell to regulate gene expression;
keep a gene product available under all conditions
false, not regulation
is this a valid reason for a cell to regulate gene expression;
execute a specific program of development
true
is this a valid reason for a cell to regulate gene expression;
stop synthesis of a cellular component when there is enough available in the cell
true
is this a valid reason for a cell to regulate gene expression;
synthesize mRNA for every gene in the genome at the same time
false
the most common point of gene regulation in bacteria is
transcription
DNA methylation in many eukaryotic organisms usually cause
decreased transcription levels
what does TF2d do
recruits RNA pol to bind
where does TF2D bind
TATA box
what do activators do
stretch/ move nucleosomes apart in 2 ways
1. acetylate histone tails
2. demythaltate DNA cytosines
for active eukaryotic genes the core promotor is found at a nucleosome free region (NFR) this NFR is needed so that
activators can promote the formation of a pre-initiation complex
is a enhancer a DNA element or protein
DNA element
is a repressor a DNA element or protein
protein
is a silencer a DNA element or protein
dna
is a activator a DNA element or protein
protein
is RNA pol a DNA element or protein
protein
is a promotor a DNA element or protein
DNA element
the iron regulatory protein binds to the iron regulatory element when iron levels are _____ and _____ translation of ferritin mRNA
low and inhibits
when histone tails are unacetylated is chromatin most likely to be open or closed
closed
how do organisms benefit from gene regulation
Gene regulation helps organisms adapt, conserve energy, develop properly, stay healthy, and survive in changing environments.
combinatorial control
the way multiple regulatory factors work together to control the expression of a single gene in eukaryotic cells.
how does alternative splicing increase protein diversity
by allowing a single gene to produce multiple different mRNA transcripts, which are then translated into different proteins.
what are regulatory transcription factors in bacteria
proteins that bind to regulatory sequences in DNA are frequently used to change levels of gene expression
what are the 2 main types of regulatory transcription factors in bacteria
repressors and activators
describe the difference between repressors and activators in bacteria
repressors: transcription factors that exert a negative control and decrease expression
activators: transcription factors that exert a postive control and increase transcription
describe the location in reference to the promotor of repressors vs activators in bacteria
repressors are downstream/ in front of the promotor so they can block RNA pol binding
activators are behind RNA pol and tell it to go
What are operons?
Groups of bacterial genes that share one promoter. allows bacteria to keep genes off until they are needed
eukaryotes do not have operons
what is an example of operons in bacteria
Ecoli have genes that metabolize glucose on all the time but they also have genes that use other fuel sources like lactate to but they only express those when needed
where does the repressor bind to in bacteria
on the operator (lacO)
lacZ
encodes B-galactosidase (part of lac operon )
lacY
encodes lactose permease (part of lac operon )
lacl
encodes the lac repressor
in a bacteria what happens in the absence of lactose
the lac repressor binds the operator and prevents RNA polymerase from transcribing the structural genes (lacZ, lacY, and lacA)
what does the lac repressor do
binds to the operator and blocks in front of RNA pol
what happens when lactose binds to the lac repressor
it changes its shape and function so now RNA pol does not have a road block and can generate mRNA
if lactose is around then the ________ is not bound
repressor
what happens in the presence of lactate
there is a transcription of genes
lac operon is under ___________- control when glucose is low
positive
what is the activator of the lac operon
CAP (catabolite activator operon)
CAP can only bind to a CAP site when bound by ______
cAMP
high cAMP = _____ glucose
Low
When glucose levels are low, cells increase the production of cyclic AMP (cAMP). This is especially important in bacteria like E. coli during the lac operon regulation:
Low glucose → ↑ cAMP
High cAMP binds to CAP (catabolite activator protein)
low glucose = __ ATP and ______ADP
ADP gets converted to _____________ = high ______
low ATP and high ADP
ADP gets converted to cyclic AMP = high cAMP