BSCI331: Final Exam - Intracellular Protein Transport
1/50
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
51 Terms
the movement of proteins between organelles is consistent with ____ among these compartments
topological similarities
topological similarities
compartments with similar membrane orientations

signal patch (3D)
higher order of structure
secondary and tertiary structures of amino acids that originally are not near from each other get brought close to each other

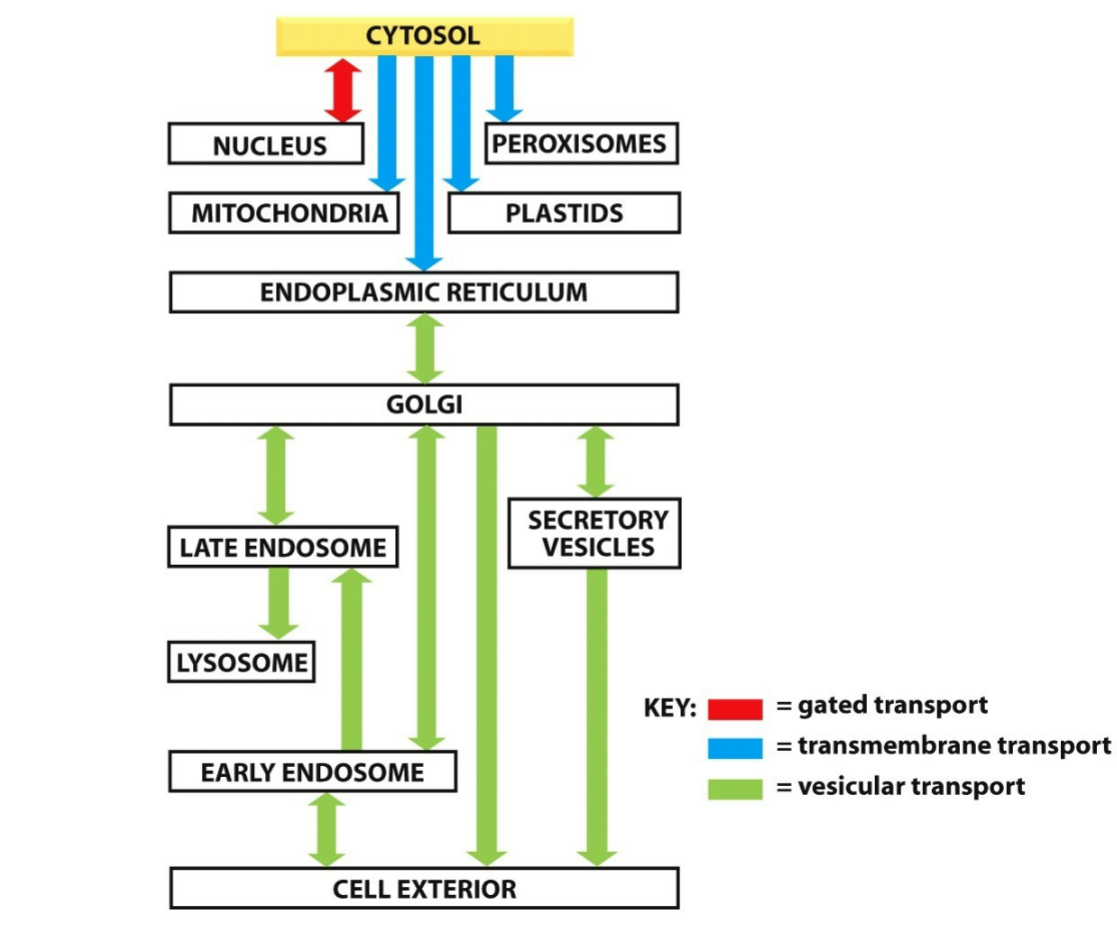
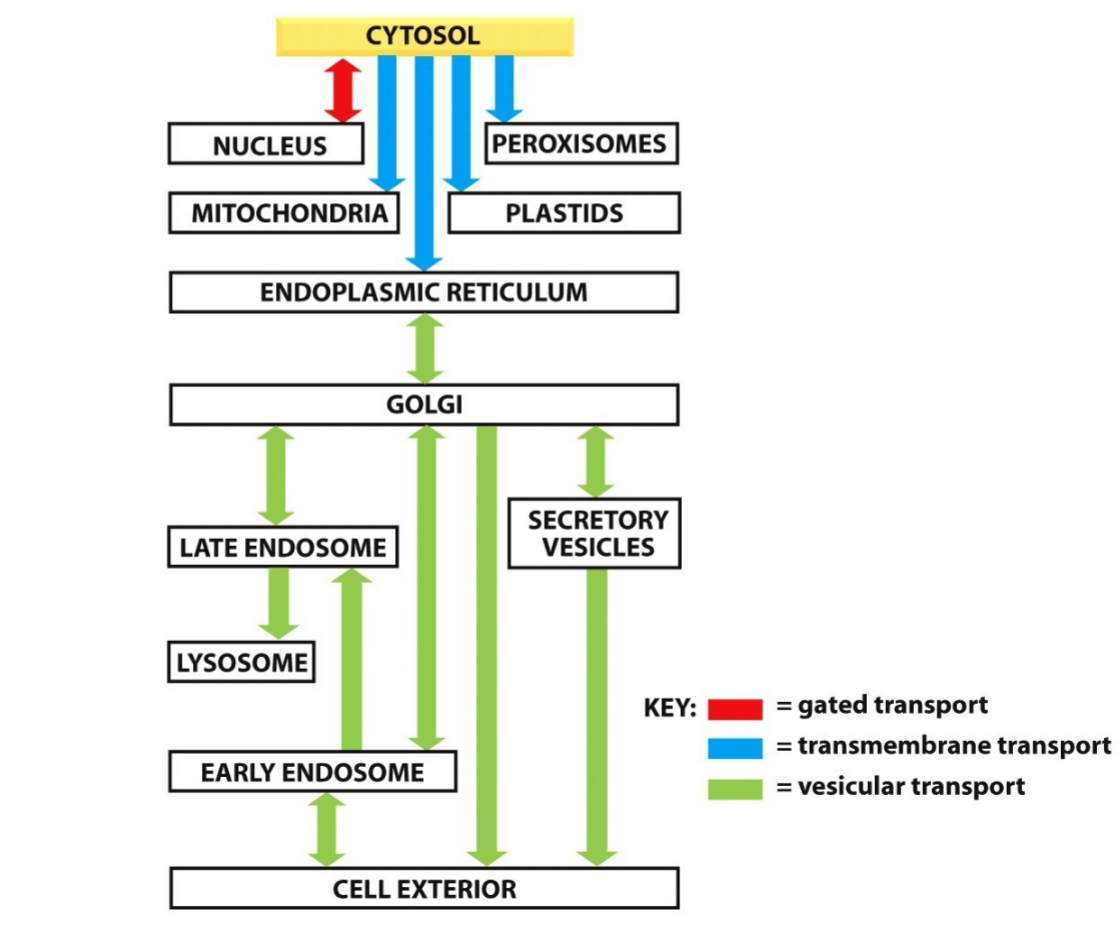
three fundamental mechanisms of the movement of proteins between cellular compartments:
gated transport
transmembrane transport
vesicular transport
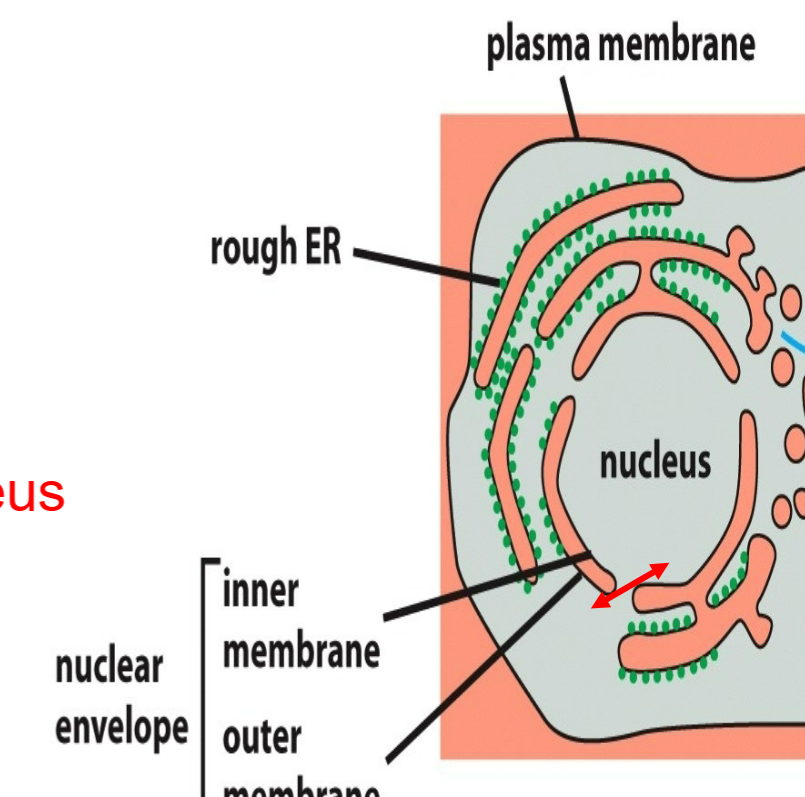
gated transport
protein traffic between the cytosol and nucleus → cytosol to nucleus and/or nucleus to cytosol
occurs through nuclear pore complexes from cytosol to nucleus without stepping off of leaflet
function as selective gates → actively transport specific macromolecules and macromolecular assembles
allow free diffusion of smaller molecules
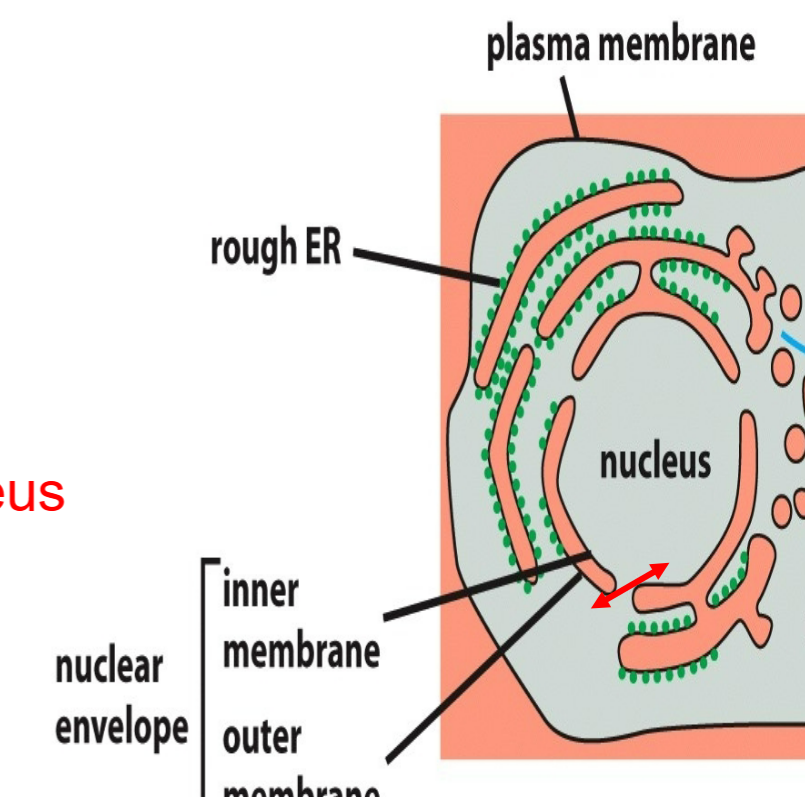
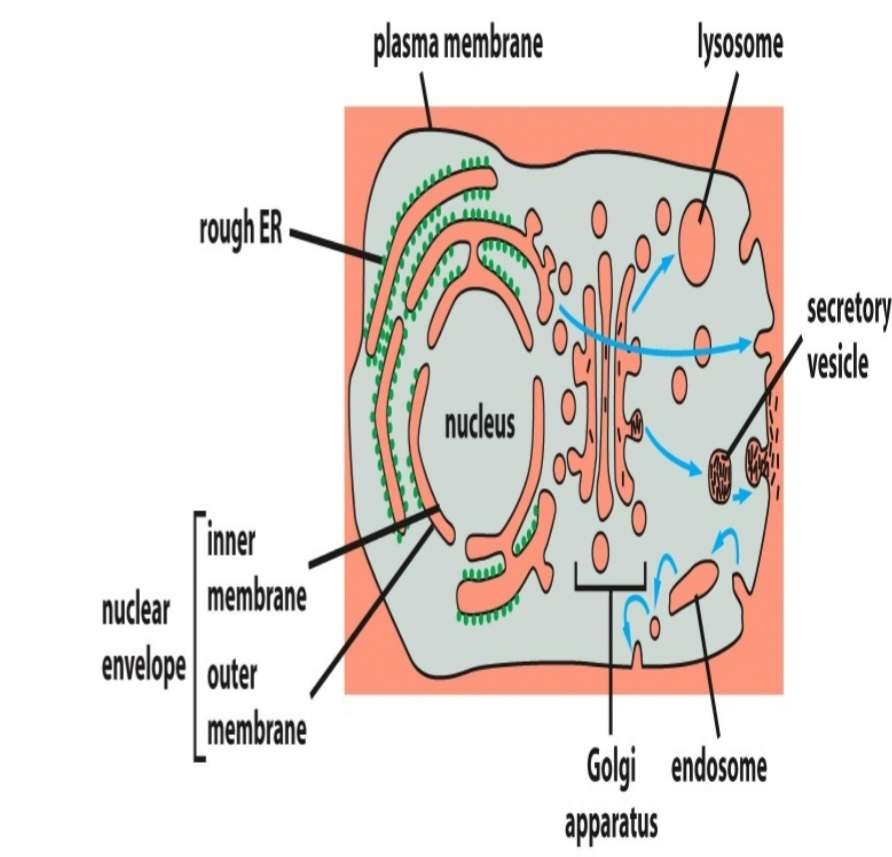
transmembrane transport
thread protein through usage of transporter for proteins to move from one face to another face
protein traffic between the cytosol and an organelle that is topologically different
occurs fthrough membrane-bound protein translocators
transported protein molecule usually must unfold to snake through translocator
ex: cytosol → ER; cytosol → mitochondria
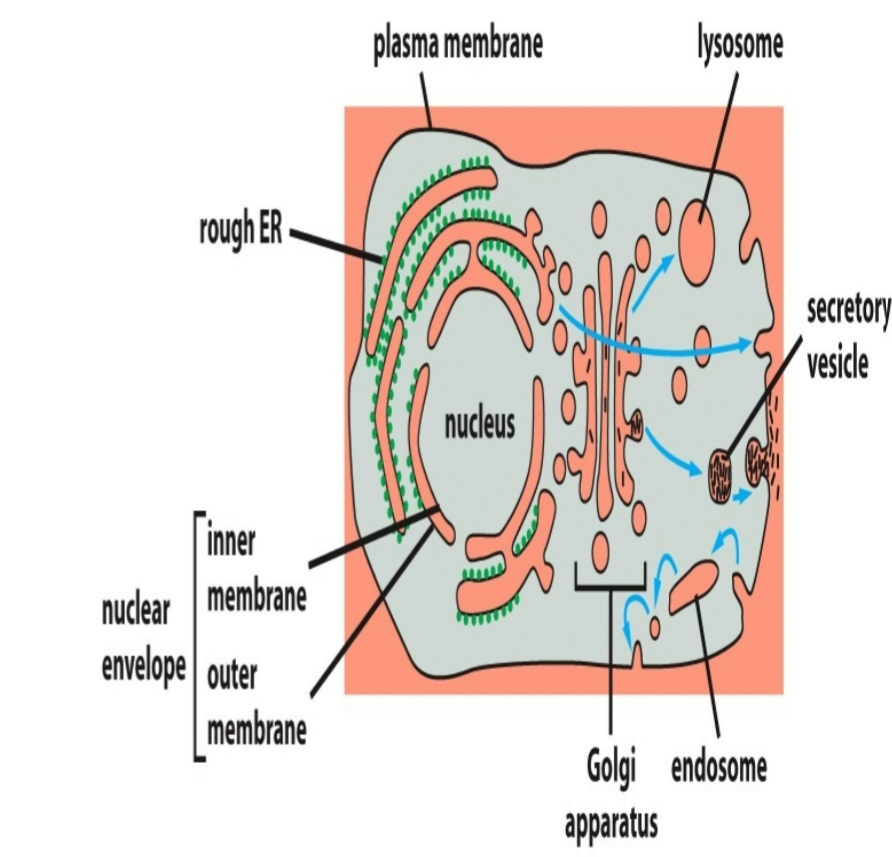
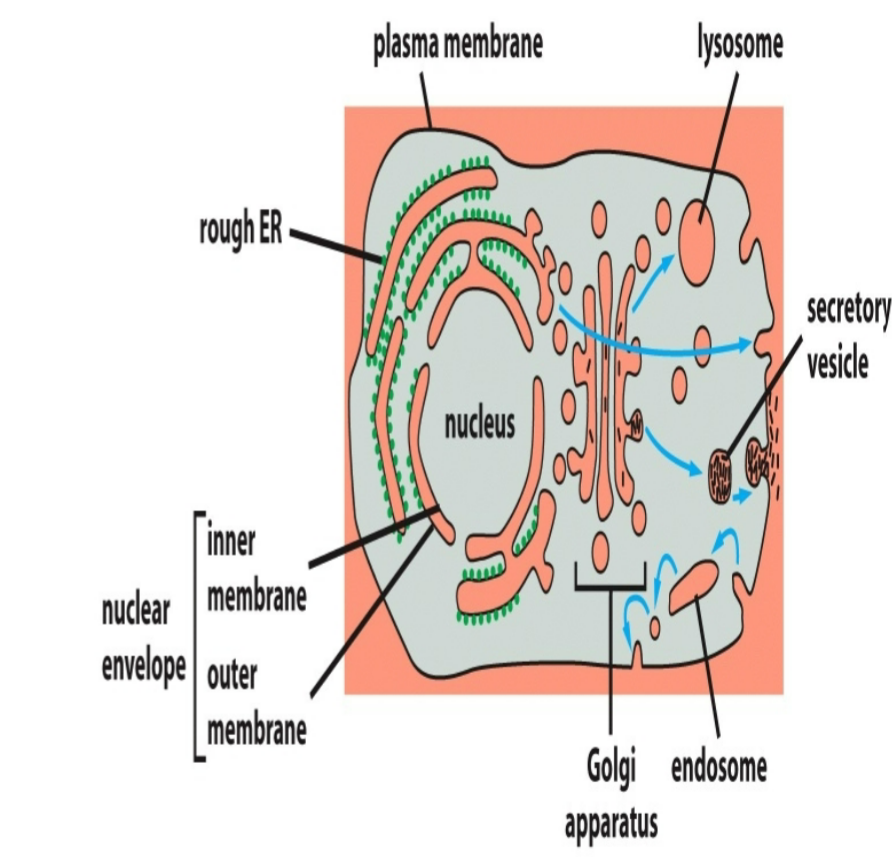
vesicular transport
outside of cell is topologically equivalent to the inside
one side of leaflet or membrane of a compartment faces outside of cell and/or other may face toward cytosol
protein traffic among topologically equivalent organelles
occurs through membrane-enclosed transport intermediates called vesicles
ex ER ←→ Golgi; Golgi ←→Endosomes; Endosomes ←→ Lysosomes; Endosomes ←→ Plasma Membrane
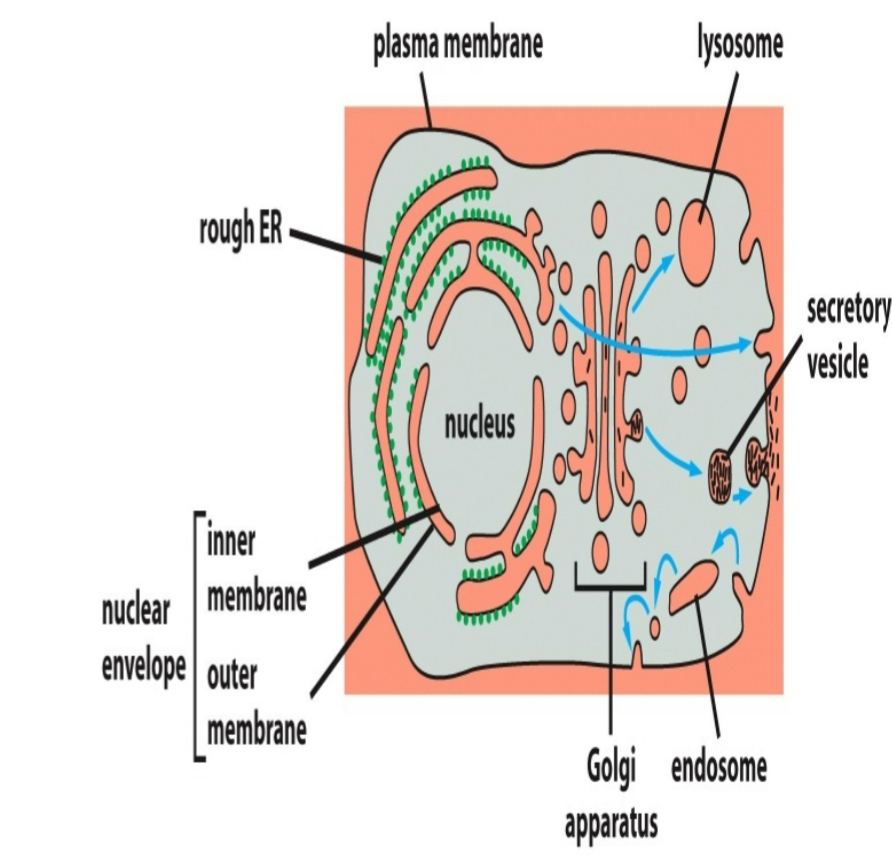
vesicles
bud from one organelle then fused to or from another membrane
merging lipid membranes
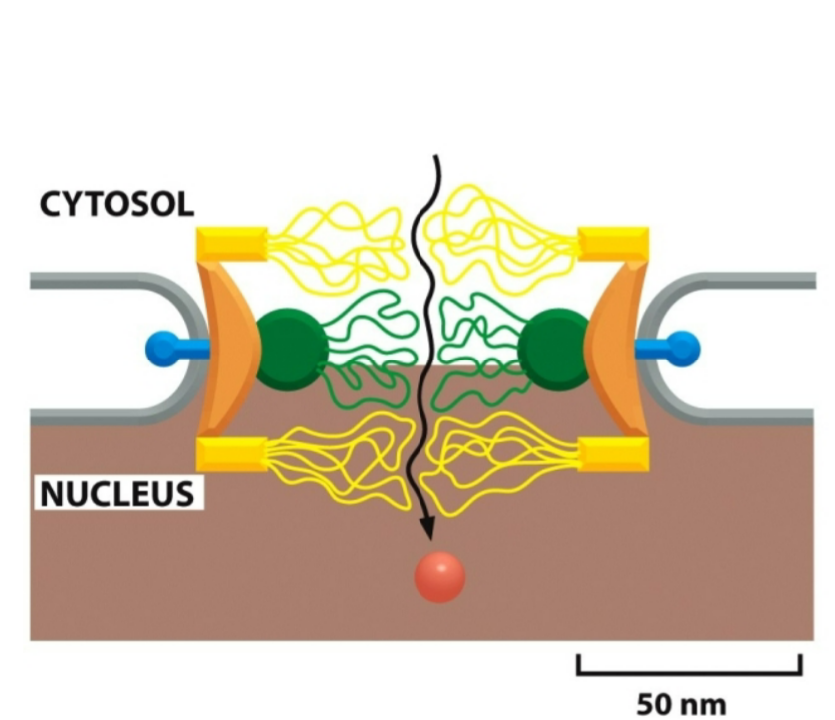
nucleoporins
lining the central pore contain unstructured regions that act to restrict the passage of large macromolecules, only let small molecules through from the cytosol into the nucleus
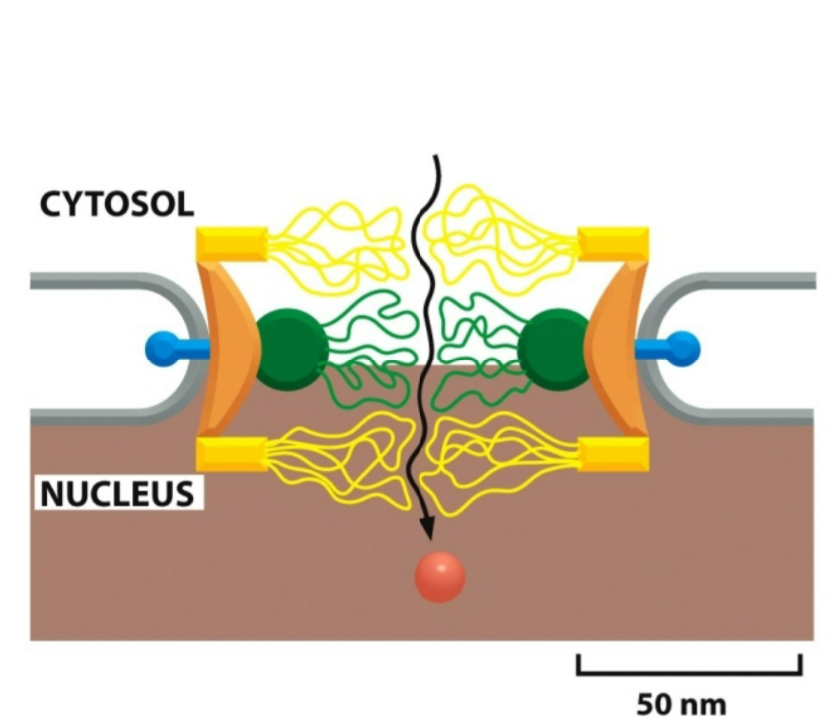

what size do the molecules have to be in order to enter the nucleus by free diffusion to enter through the nucleoporins?
up to 9 nm diameters


what size do these kind of molecules have to be in order to enter the nucleus by active transport to enter through the nucleoporins? what steps are necessary in order for them to enter through the nucleoporins?
upon receiving a signal, channel can open to 30 nm wide
macromolecules where a conformation change has to happen for them to pass through the nucleoporins

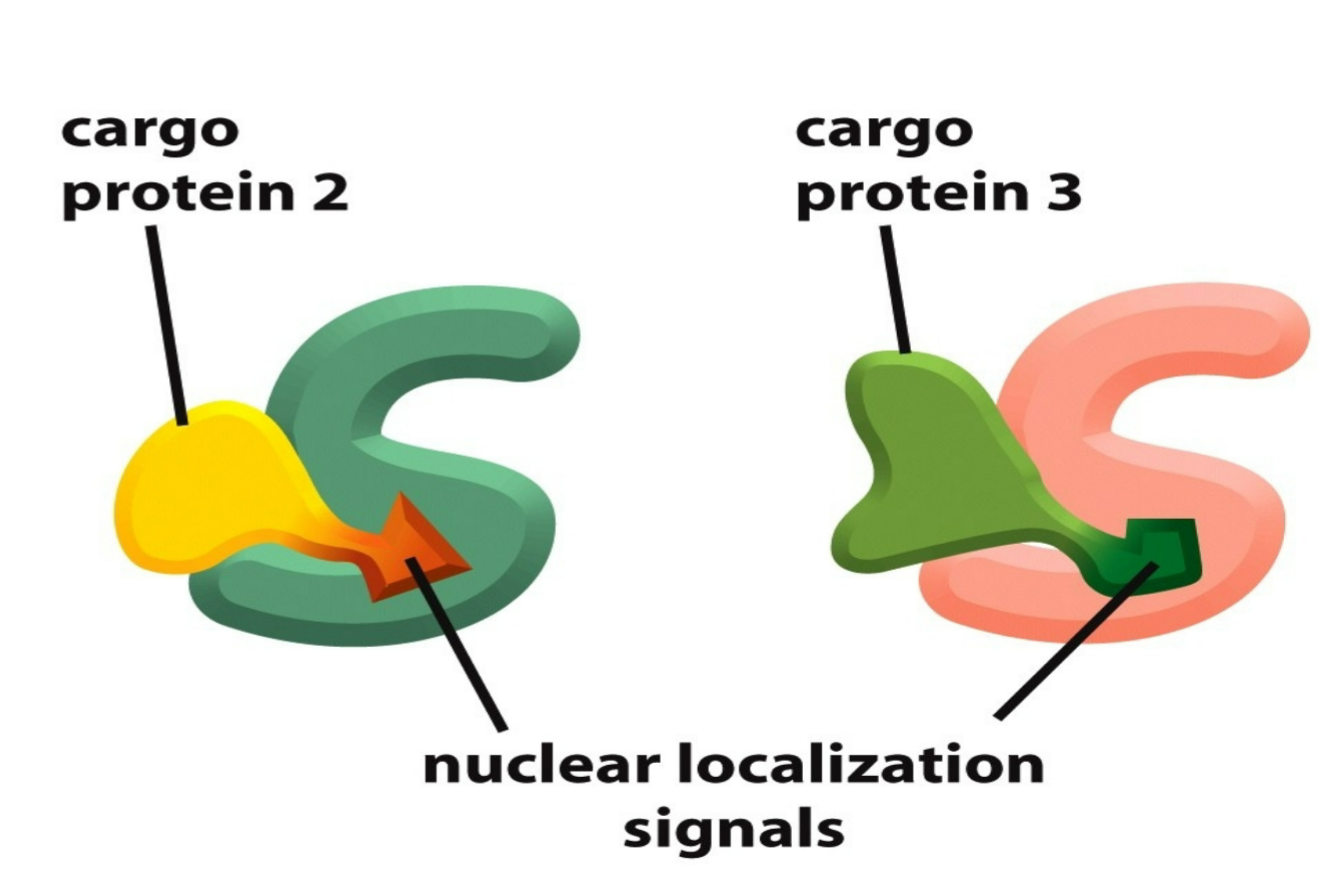
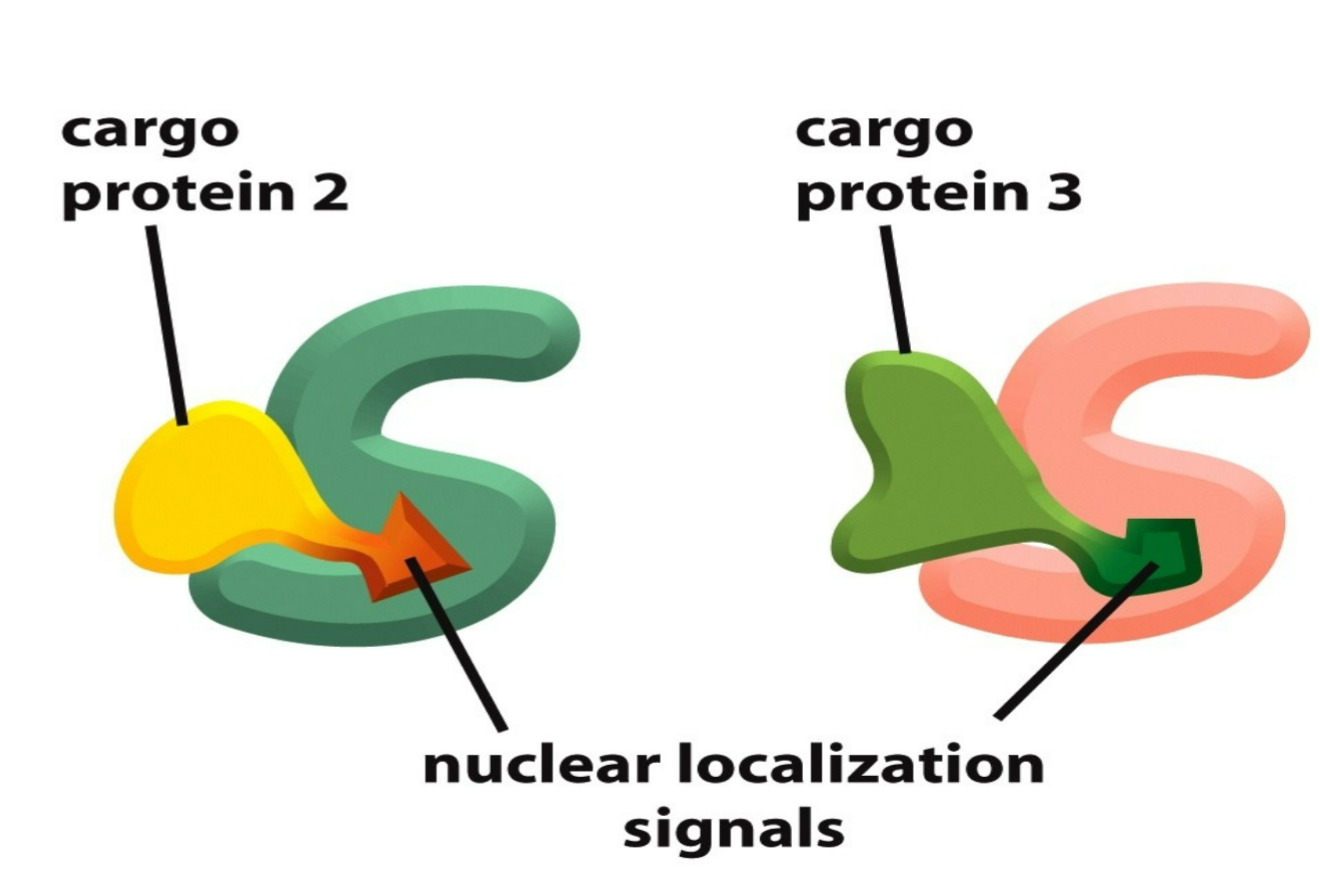
nuclear localization signals (NLS)
appear within end site of cargo which are recognized by nuclear import receptors (aka carrier proteins)
5 or more basic amino acids in a row in nuclear import protein and/or NLS in order to fit as a NLS site
does the import of nuclear proteins through the pore complex increase order in the cell or decrease it that impacts the concentration of specific proteins in the nucleus?
increases order in the cell → consumes energy due to a decrease in entropy
Ran
small GTPase found in both cytosol and nucleus
required for both nuclear import and export systems
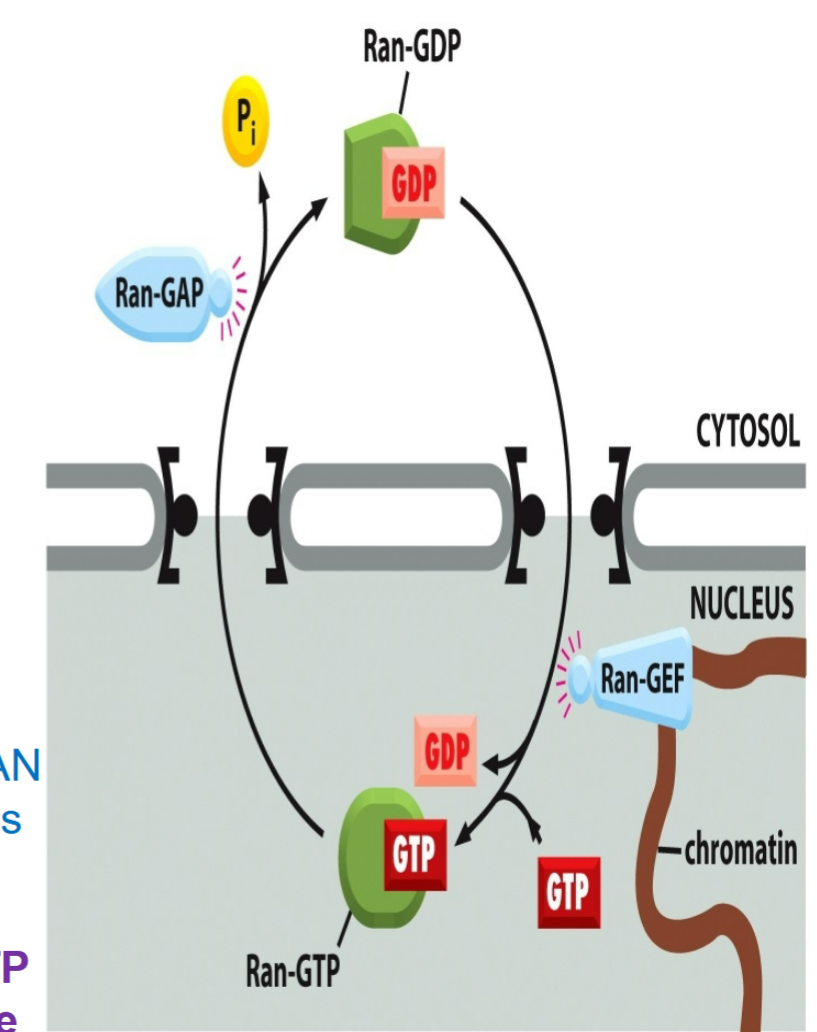
RAN, like other GTP-binding proteins, exists in two states:
one with GTP attached (“on” state)
one with GDP attached (“off” state)
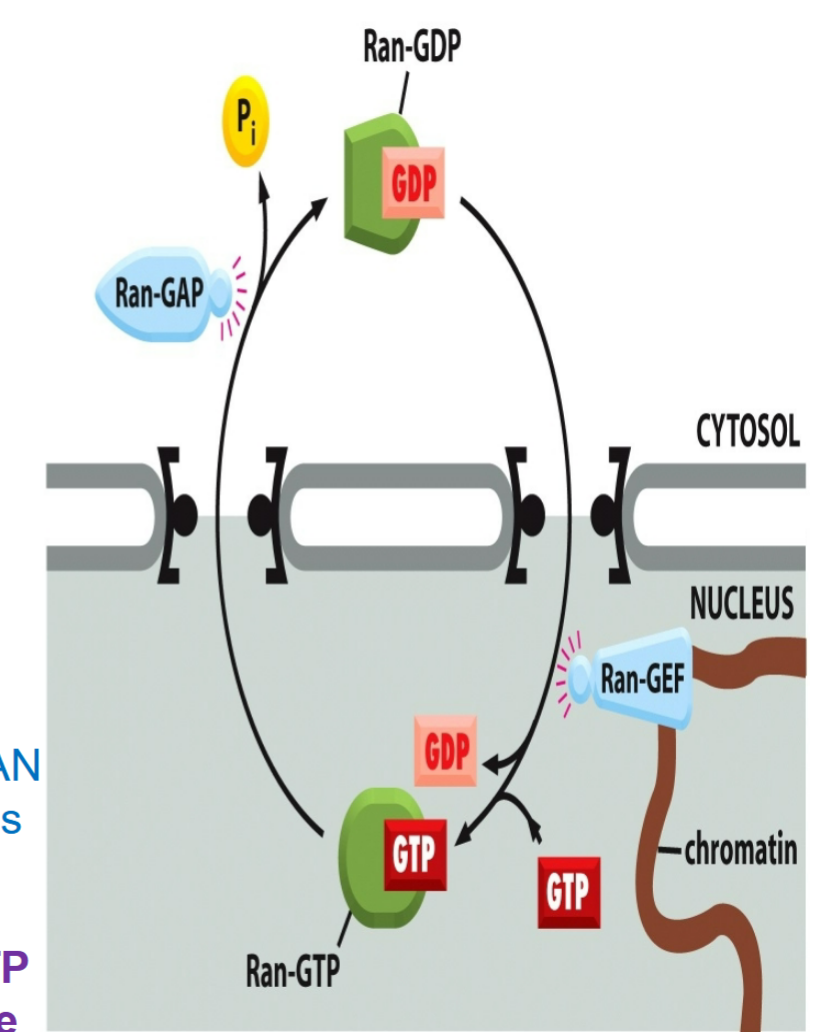
RAN-GEF (RAN guanine exchange factor (GEF))
nuclear protein, binds to chromatin inside of cell
catalyzes binding of GTP to RAN inside the nucleus
RAN-GDP found in cytosol
RAN-GAP (RAN GTP-ase activating protein)
cytosolic protein, outside of cell
activates hydrolysis of GTP attached to RAN to hydrolyze to GDP
creates gradient of RAN-GTP across nuclear pore with more RAN GTP inside nucleus than the outside
RAN-GTP found in nucleus
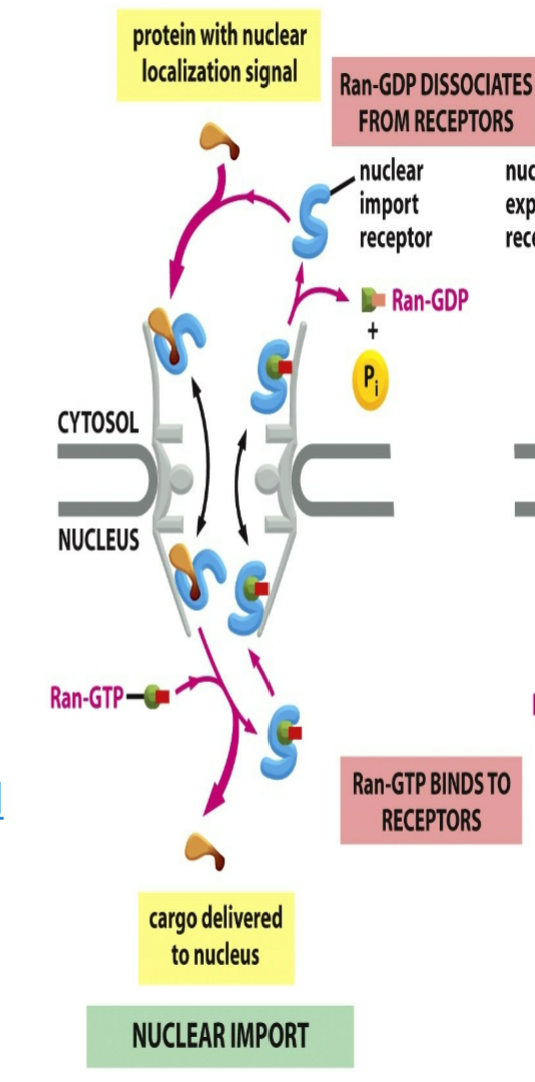
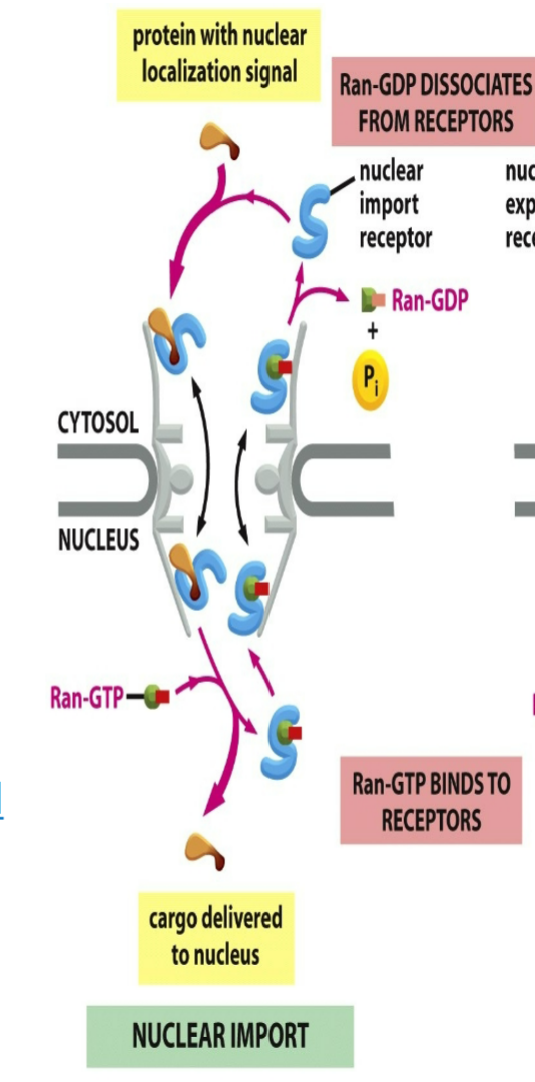
role of nuclear import receptor when RAN binds:
unicellular and mutually exclusive → when RAN-GTP binds to nuclear import receptor then cargo protein won’t be able to bind
cargo protein will bind to import receptor from the cytosol to be allowed to enter inside of nucleus → cargo lets go where RAN-GTP comes and binds to import receptor → enter out of cytosol and gets hydrolyzed into RAN-GDP which allows cargo protein to bind to it again
binds to import receptor after they diffuse through nuclear pore and into nucleus
causes them to release their cargo proteins, which therefore accumulate inside the nucleus
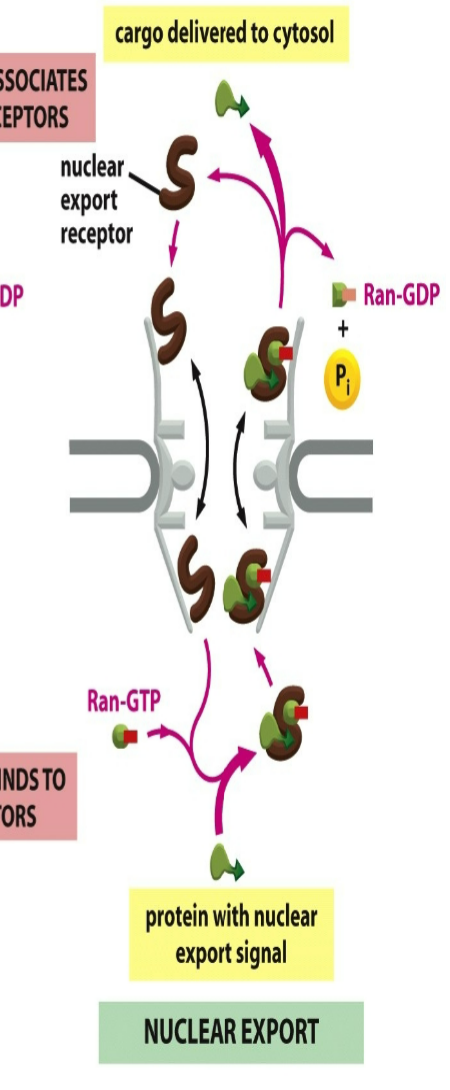
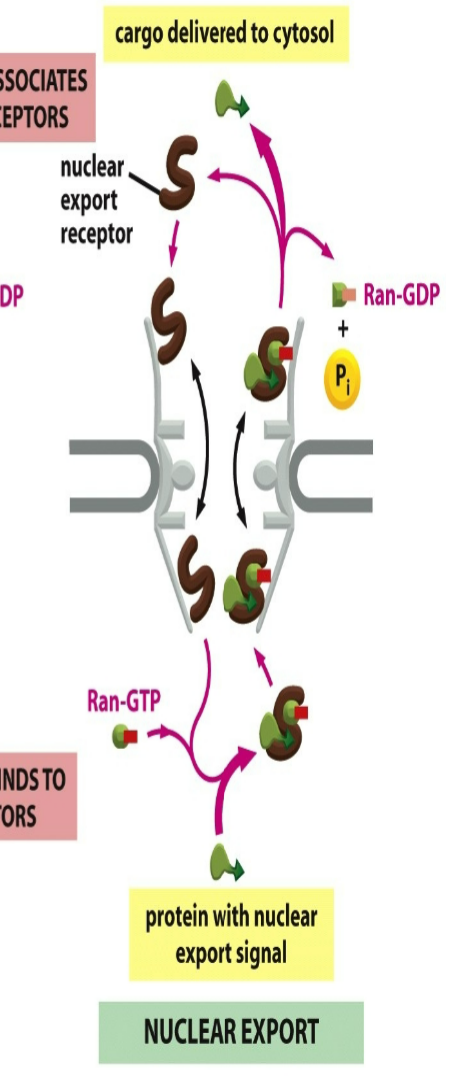
role of nuclear export receptor when RAN binds:
not mutually exclusive → would need both RAN-GTP and cargo protein binded to export receptor to carry reaction
nuclear export receptor needs to be naked to enter in cytosol → both cargo protein and RAN-GTP both bind on either end sites of receptor → goes back out into cytosol and gets hydrolyzed into RAN-GDP which releases RAN-GTP and cargo protein
RAN-GTP has opposite effect on export receptors, causing them to bind their cargo
then diffuse through pore into the cytosol
in mitochondrial transport, what are mitochondrial proteins?
first fully synthesized as precursor proteins in cytosol and then translocated into mitochondria
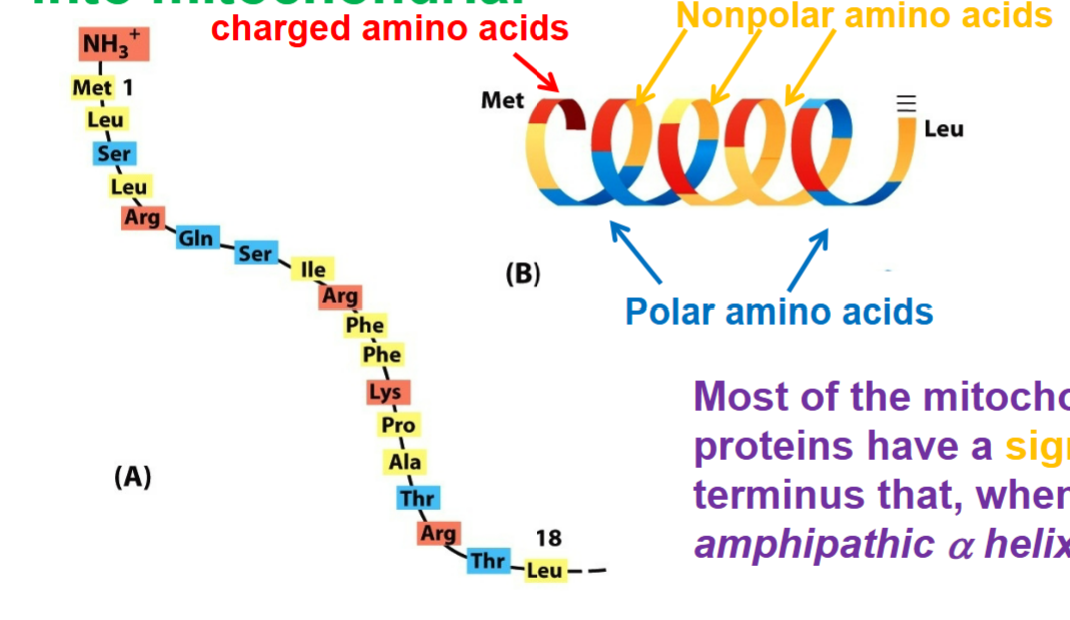
requires secondary structure, rely on chemical nature of amino acids and not specific amino acids
AMPHIPATHIC nature → one side or face has charged residues clustered, while other side has uncharged residues clustered (represent as signal sequences)
most of the mitochondrial precursor proteins have signal sequence at their N terminus that, when folded forms an amphipathic alpha helix
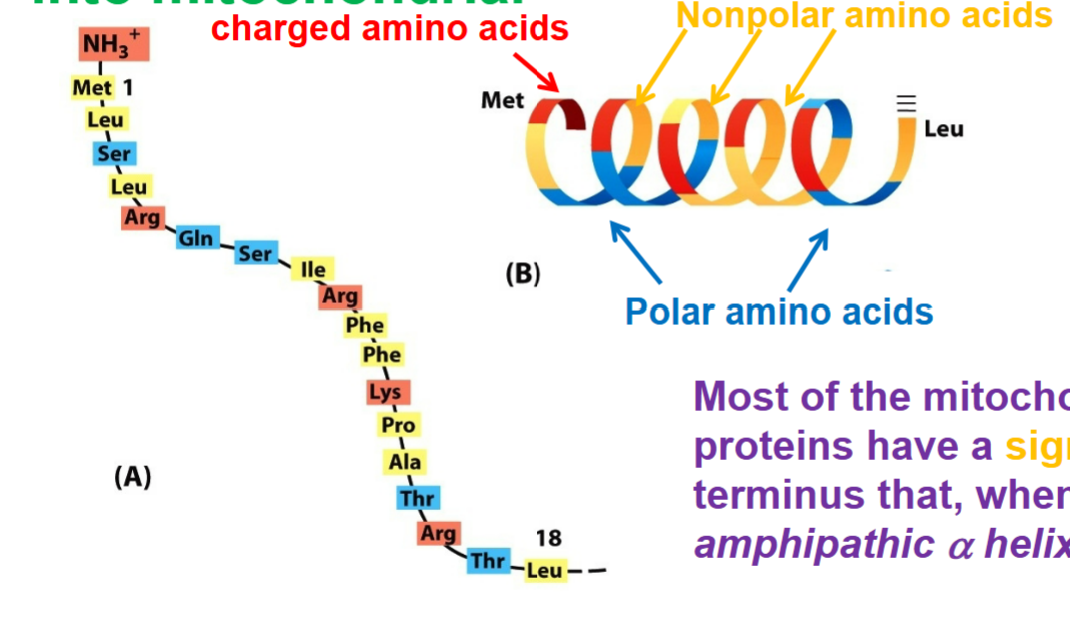
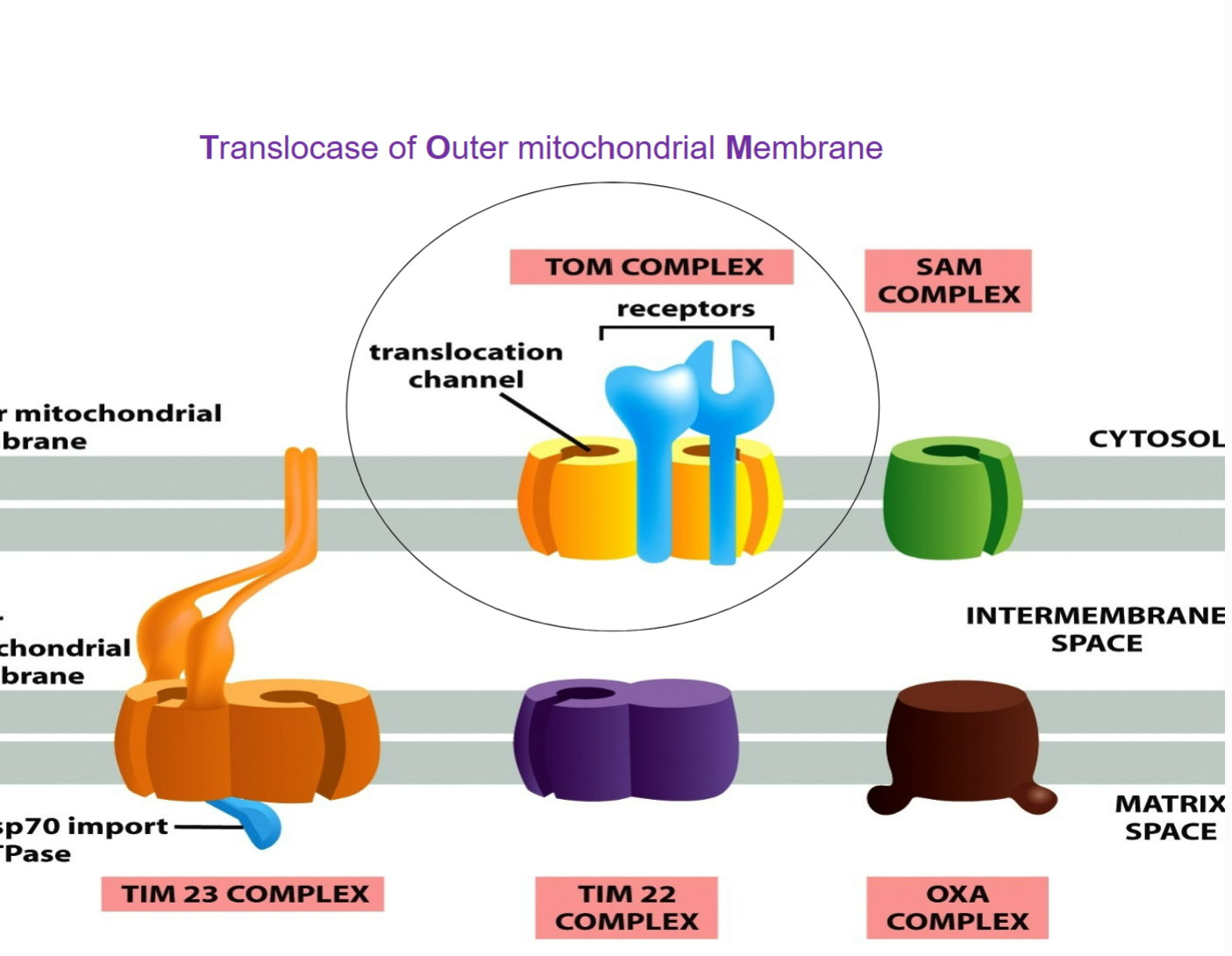
Protein translocation across mitochondrial membranes is mediated by multi-subunit protein complexes that function as protein translocators:
TOM complex (translocase of outer mitochondrial membrane)
two TIM complexes (translocase of inner mitochondrial membrane) → TIM23 and TIM22
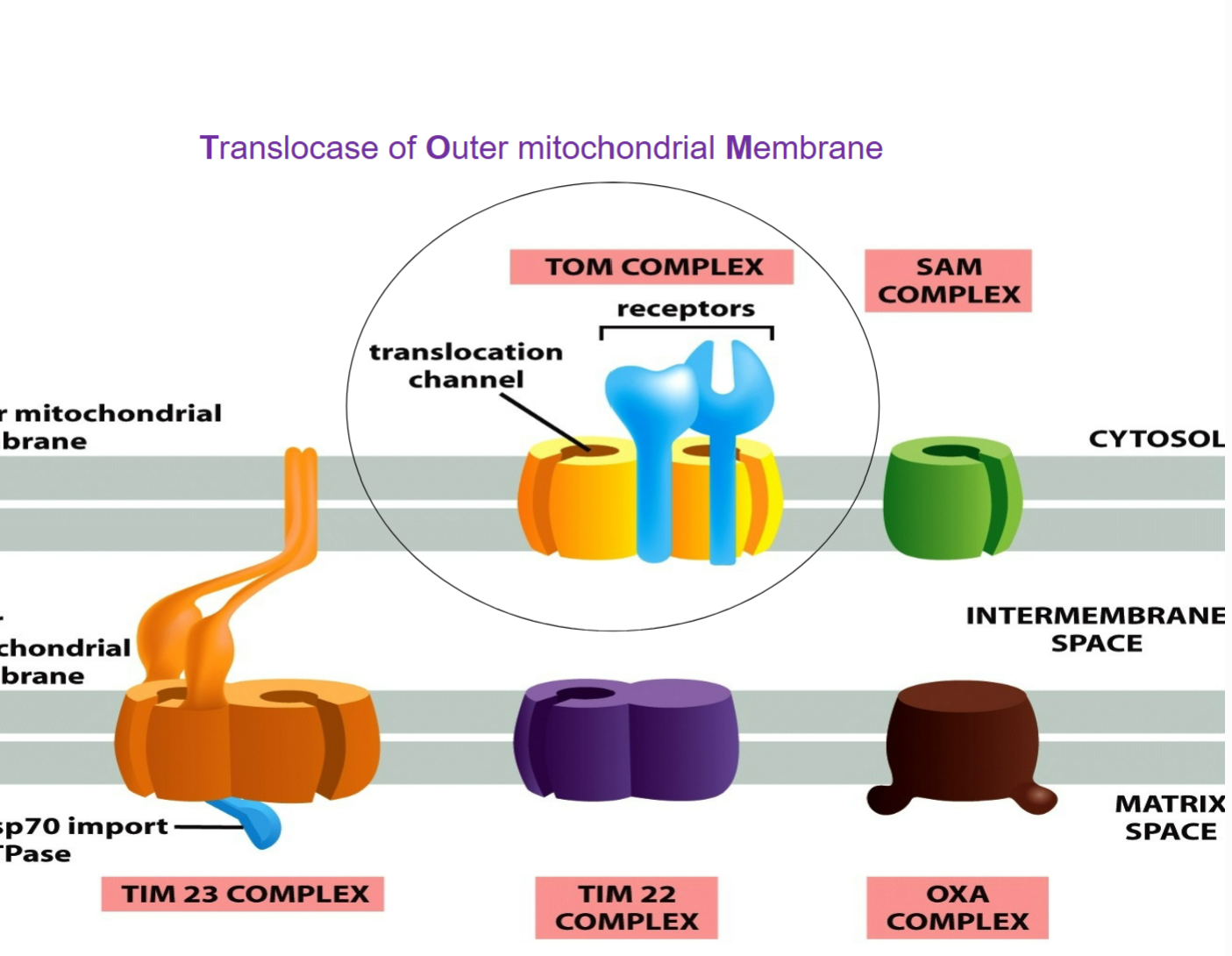
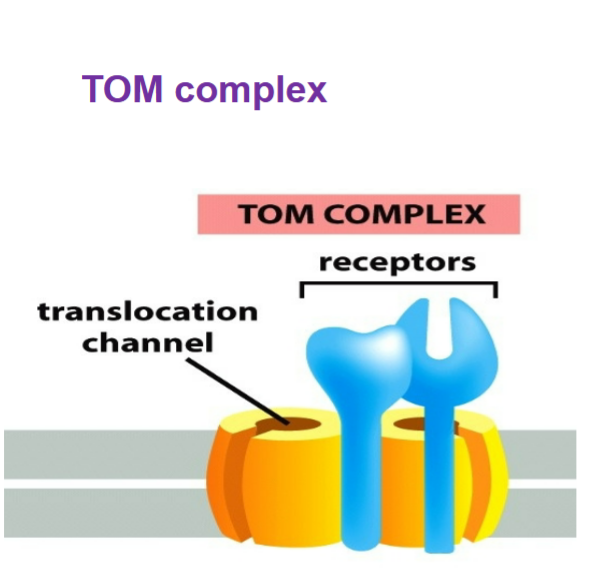
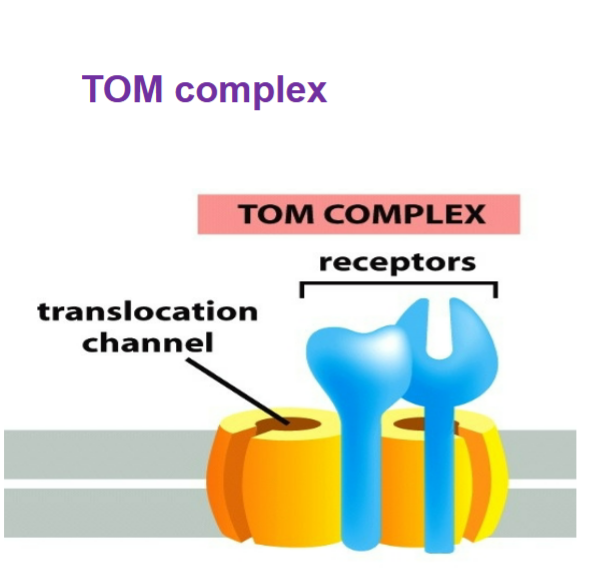
TOM complex (translocase of outer mitochondrial membrane)
ALL nucleus-encoded mitochondrial proteins must first enter via TOM → must be UNFOLDED in order to enter through
helps insert transmembrane proteins into outer mitochondrial membrane, gatekeeper of mitochondrion
transmembrane proteins with a B (beta)-barrel structure are transferred to the SAM (functions sorting and assembly) complex for proper folding
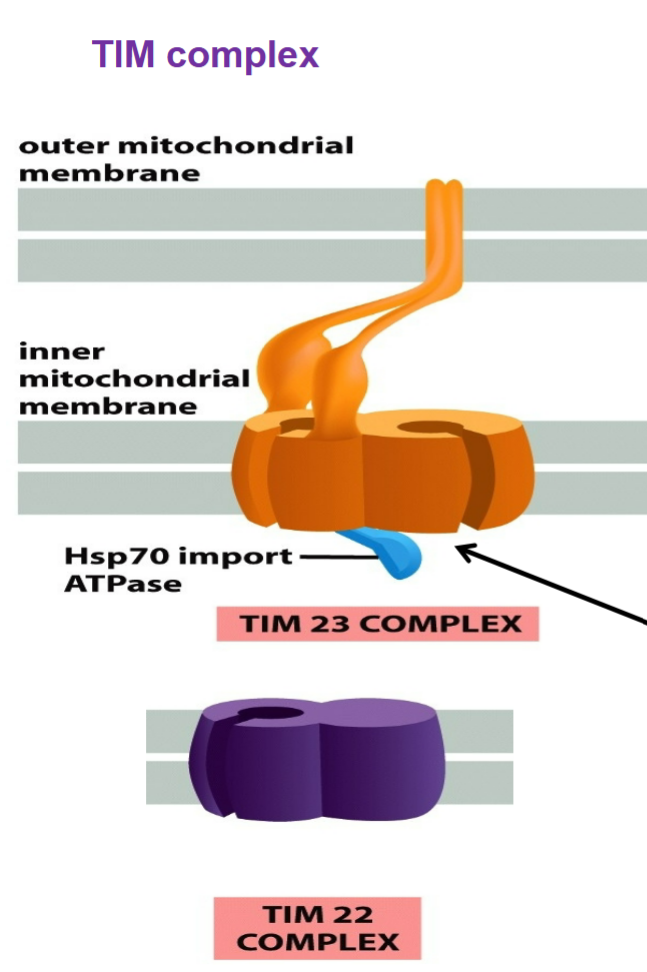
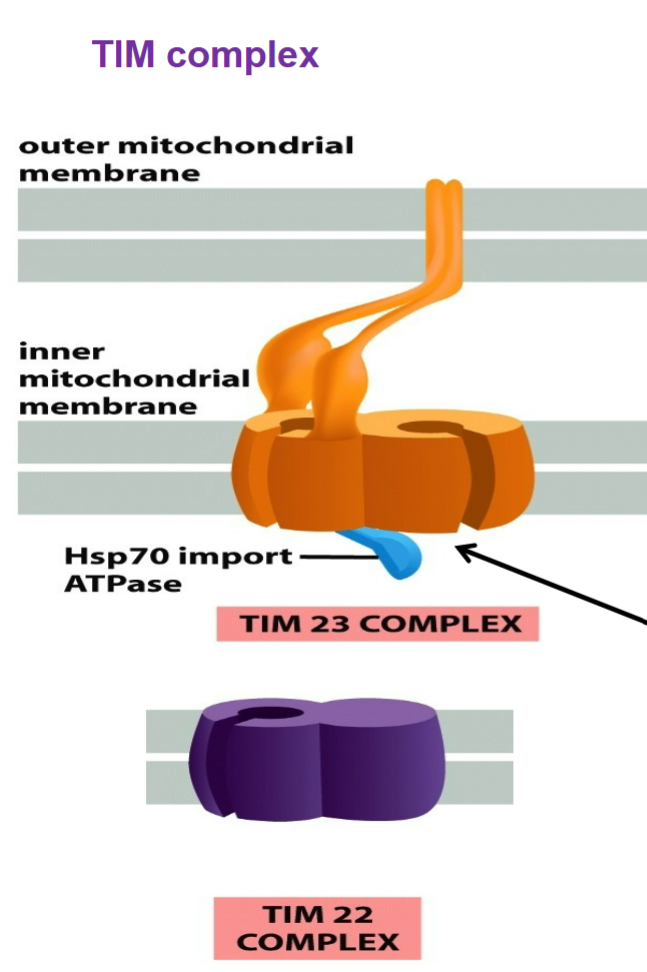
TIM complex (translocase of inner mitochondrial membrane)
TIM23 spans both outer and inner mitochondrial membranes → has extension from inner to outer membranes that allow TIM and TOM to interact for proteins to enter through matrix and threaded through series
Transports: soluble proteins into MATRIX; membrane proteins into inner mitochondrial membrane
import Hsp70 import ATPase complex binds to and pulls proteins through TIM23 channel → chaperone protein used to clear out unstable/unfolded proteins, pulls protein across inner membrane channel by hydrolyzing ATP, increase order of cell from unfolded to fold into 3D shape
does the protein have to be folded or unfolded to go through the TOM complex?
unfolded because it needs to fit in order to thread through and then later folding up again

Hsp70 ATP-ase chaperone
newly synthesized (precursor) mitochondrial proteins in cytosol are surrounded by protein-folding chaperones that prevent them from aggregating
mitochondrial versions of these chaperones also exist and help these precursor proteins fold into 3D structures once they enter the mitochondria
part of TIM complex and known to be most common chaperone
instead of assisting in folding protein it prevents folding to happen → ensure proteins don’t fold properly to allow them to cross through mitochondrial membrane
need to hydrolyze ATP to unfold proteins

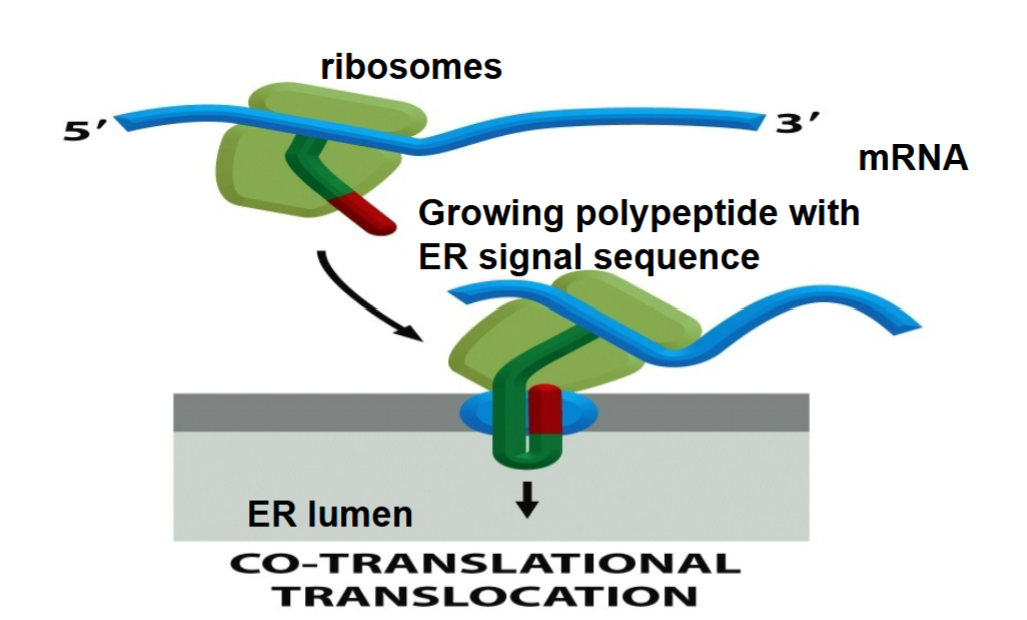
Co-translational translocation
imported into ER as they are being synthesized at the same time
helps to save much effort in unfolding protein
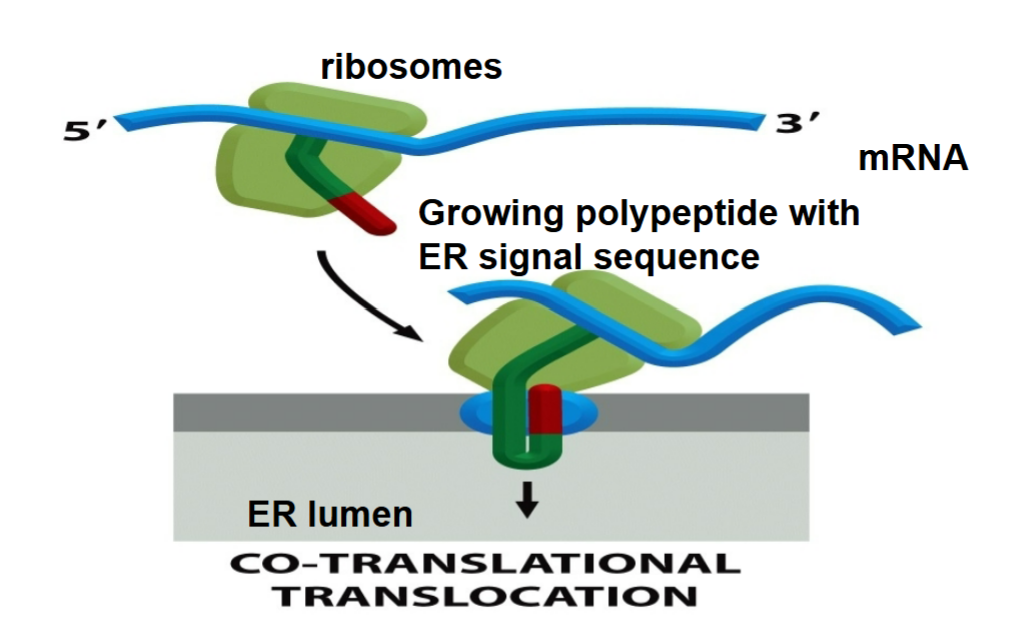
What types of proteins require co-translational translocation?
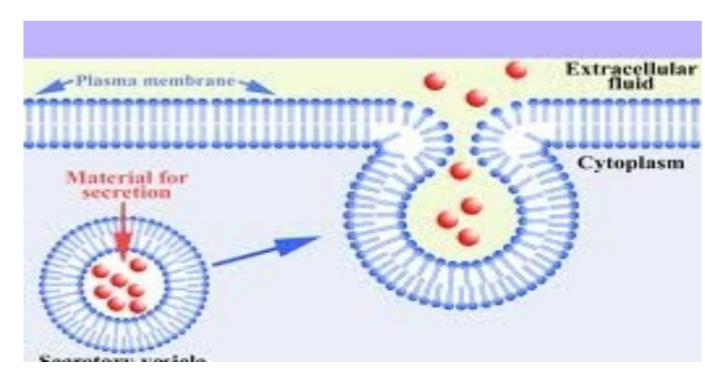
water soluble (non membranous) proteins destined to: localize to the lumen of any non nuclear organelle (ER, Golgi, lysosomes, etc.); be secreted out of the cell (e.g. hormones) → found in ER first, don’t need protein translocator to secrete proteins since ER lumen topologically similar to extracellular fluid
transmembrane proteins destined to: localize to the membrane of an organelle (some nuclear membrane, plasma membrane, ER, Golgi, lysosomal, etc.); located inside membrane or plasma membrane → translocated in ER
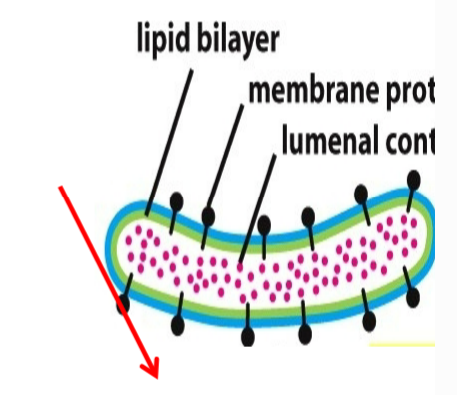
water soluble (non membranous) proteins
localize to the lumen of any non nuclear organelle (ER, Golgi, lysosomes, etc.)
be secreted out of the cell (e.g. hormones) → found in ER first, don’t need protein translocator to secrete proteins since ER lumen topologically similar to extracellular fluid
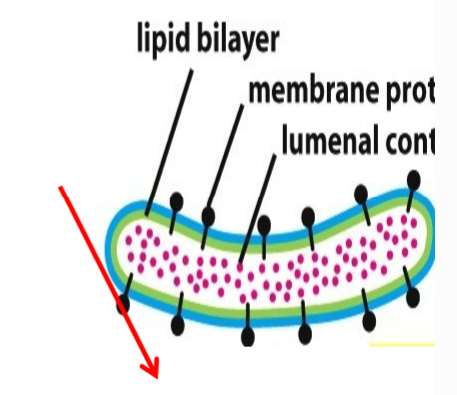
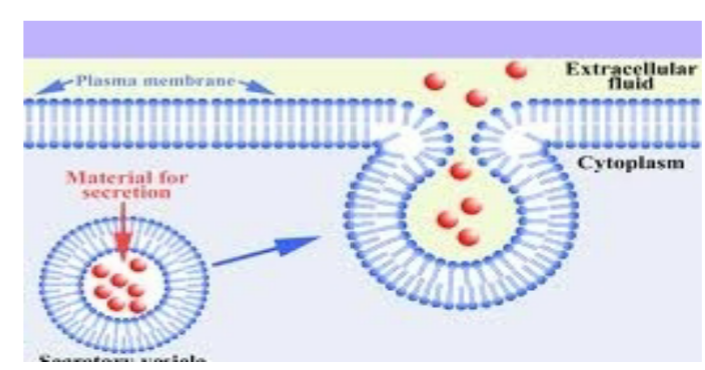
transmembrane proteins
localize to the membrane of an organelle (some nuclear membrane, plasma membrane, ER, Golgi, lysosomal, etc.)
located inside membrane or plasma membrane → translocated in ER
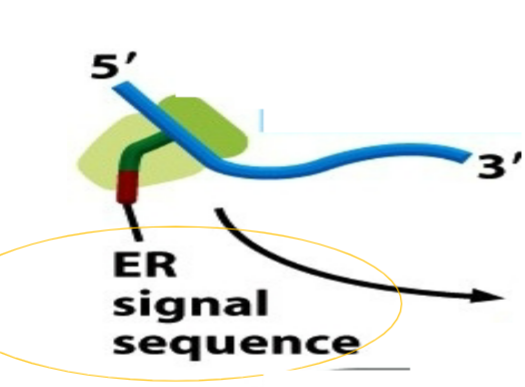
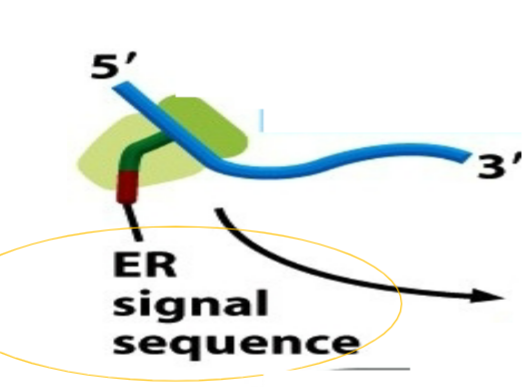
ER signal sequence
ALL proteins requiring co-translational translocation possess a ER signal sequence
defined from chemical nature and based on 5 or more basic amino acids (similar to NLS or nuclear localization signalling)
vary somewhat in sequence but all are: n-terminal, hydrophobic → contain 8 or more hydrophobic/nonpolar amino acids (about the width of one lipid bilayer of ER)
signal sequences recognized by a signal recognition particle (SRP)
recognized by an SRP receptor in the ER membrane
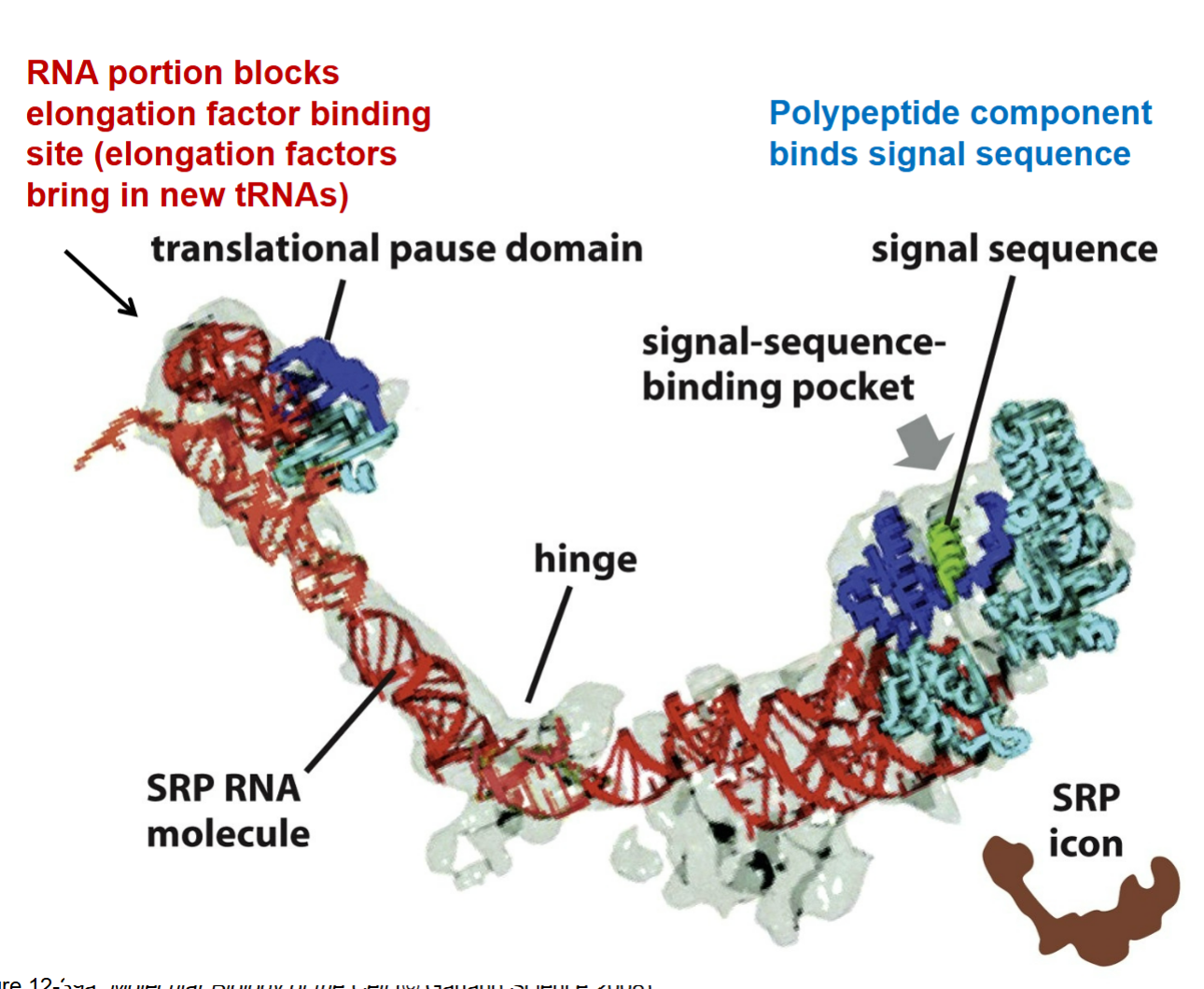
SRP (signal recognition particle)
non-coding RNA
recognizes ER signal sequences in co-translational translocation
complex proteins in that they contain both RNA and polypeptide components
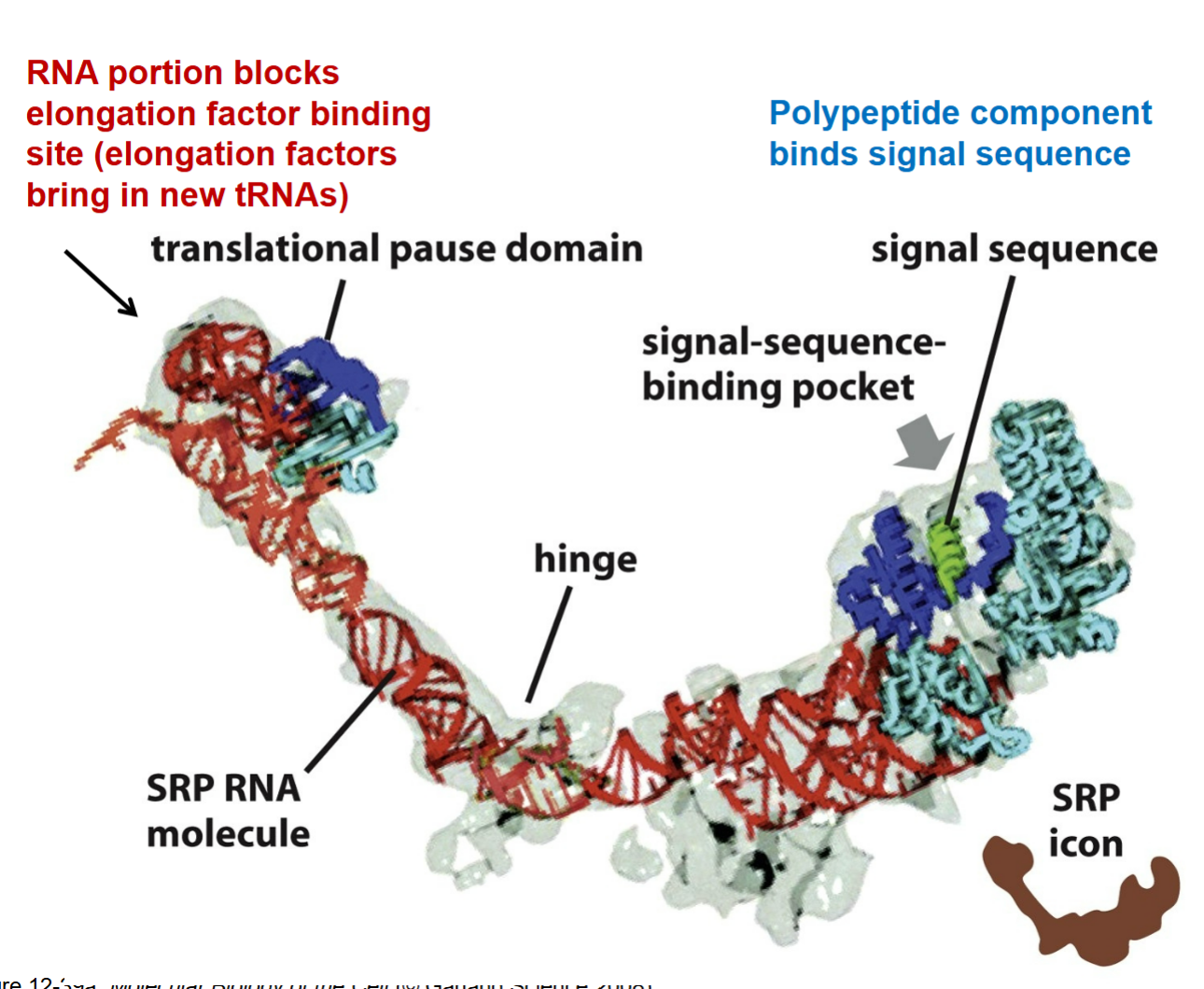
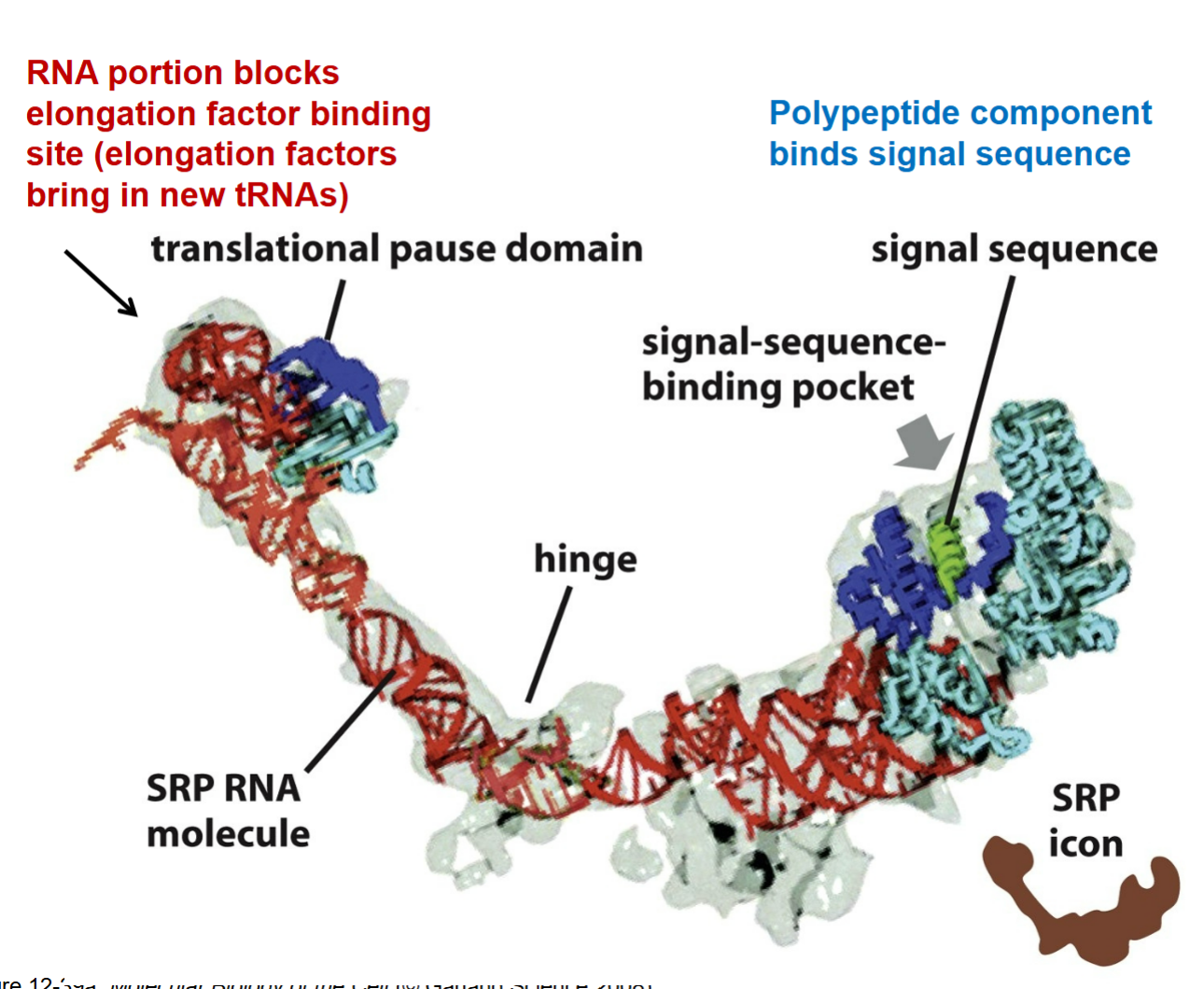
how does a SRP play a role for the SRP RNA molecule?
red part → RNA portion end that blocks elongation factor binding site (elongation factors bring in new tRNAs); binds import channel to preven bringing in new tRNAs into ribosome and stops translation by allowing recognition that a protein belongs in the ER which signals to stop translation
brings ribosomal component over to ER to interact with SRP receptor from the blue part which is protein end and hydrophobic that allows interaction to come together to stop translation
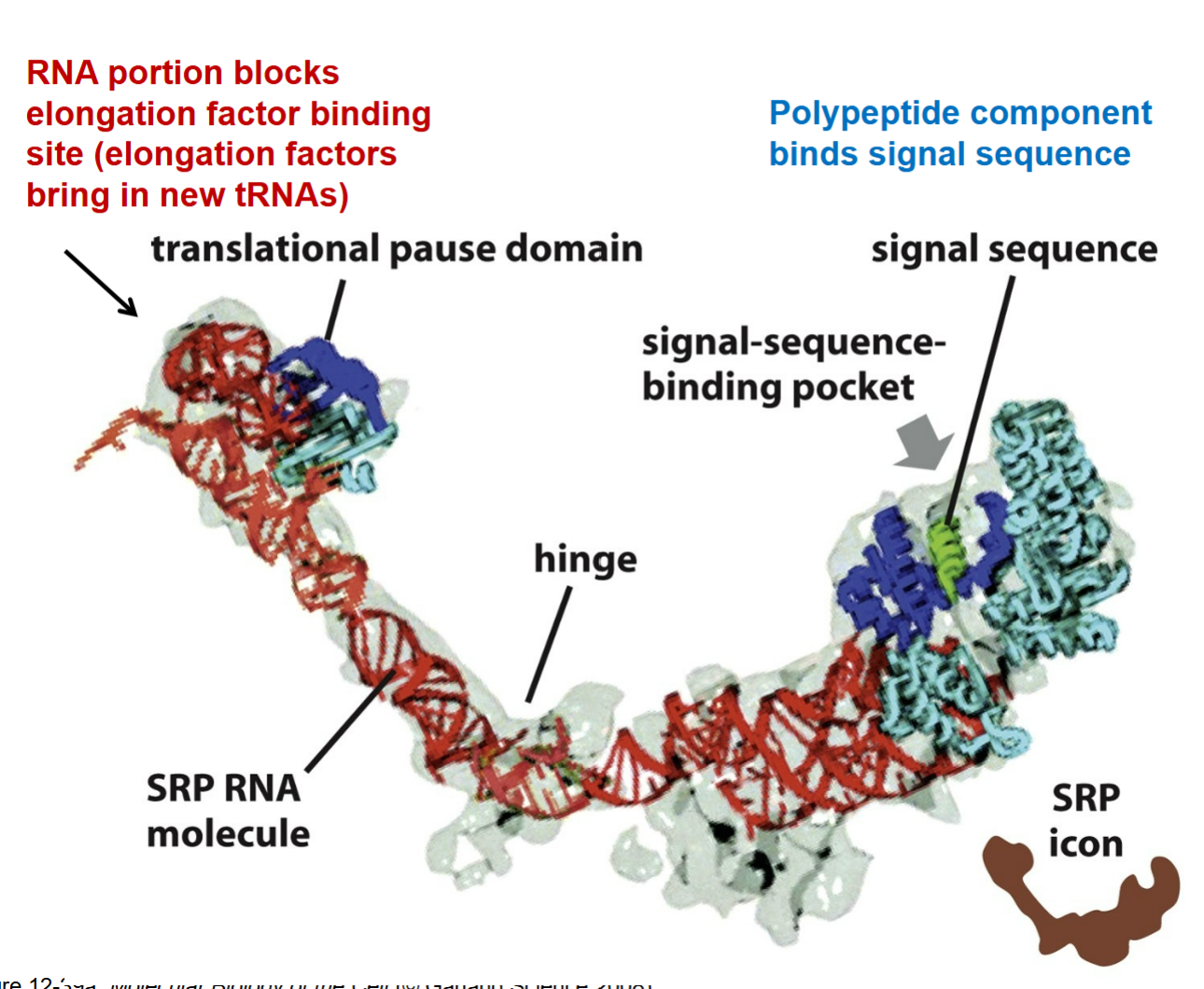
would you consider the SRP to be a ribozyme?
no! because it doesn’t have enzyme characteristics to speed up a reaction
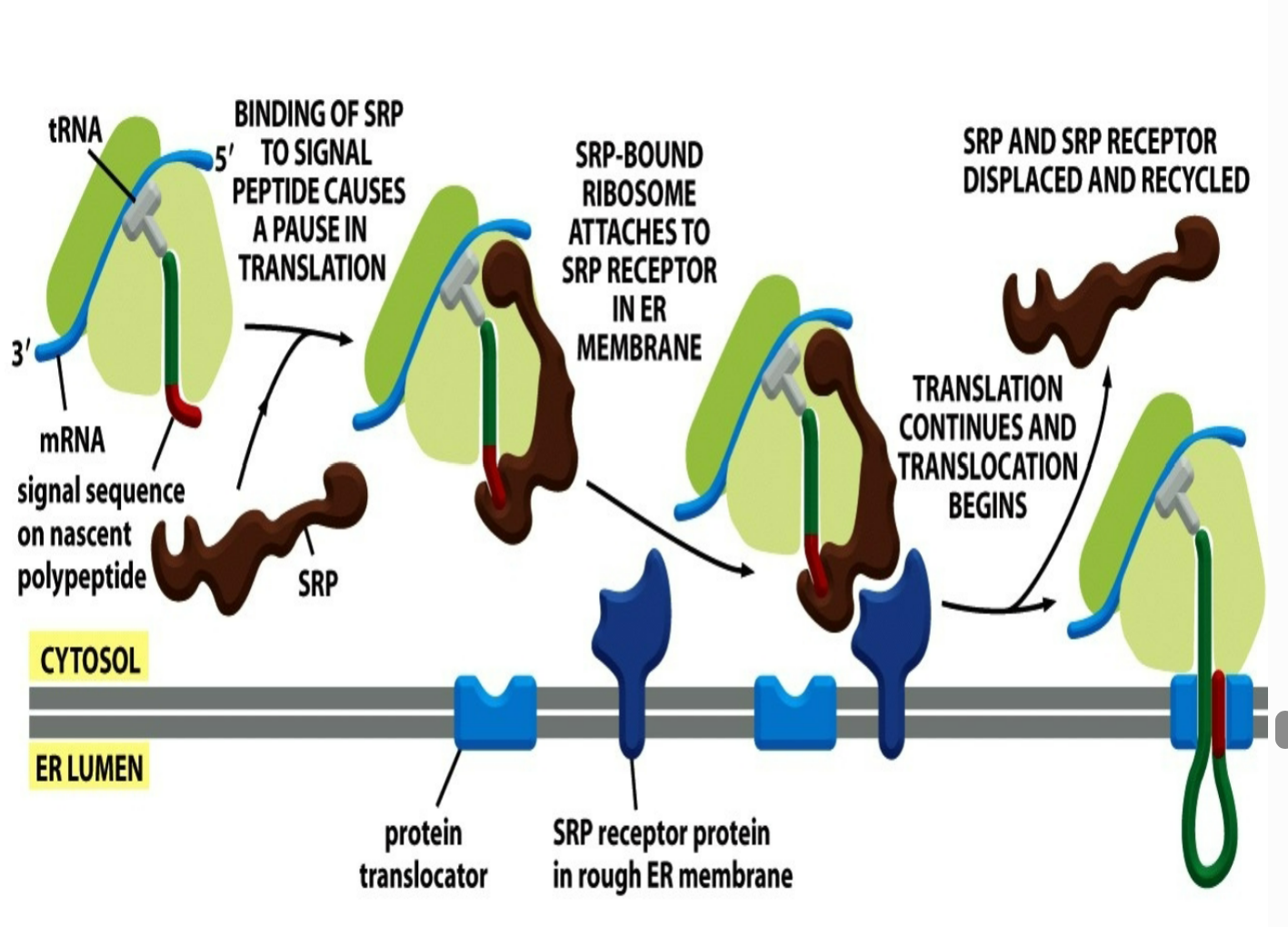
how does the mechanism for SRP look like when it comes to recognizing the protein being in the ER in order to pause translation?
binding of SRP to signal peptide causes a pause in translation where the ER signal sequence gets recognized and binded to SRP
SRP-bound ribosome (signal sequence attached to SRP) ataches to SRP receptor in rough ER membrane (receptor is known to recognize and help form rough ER)
once binded to SRP receptor protein in rough ER membrane → translation continues and translocation begins
SRP and SRP receptor gets released and displaced and recycled and signal sequence gets entangled with protein translocator
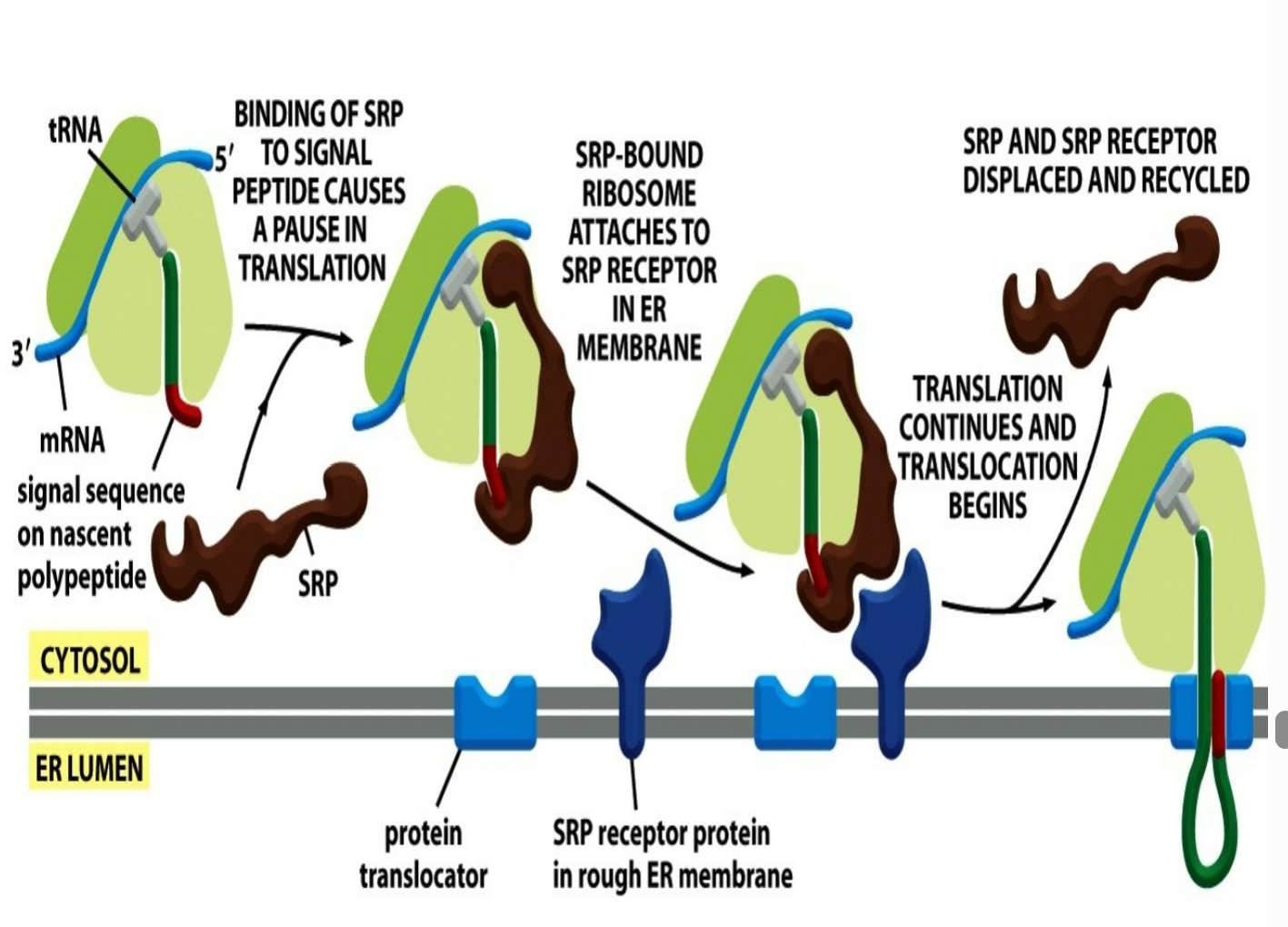
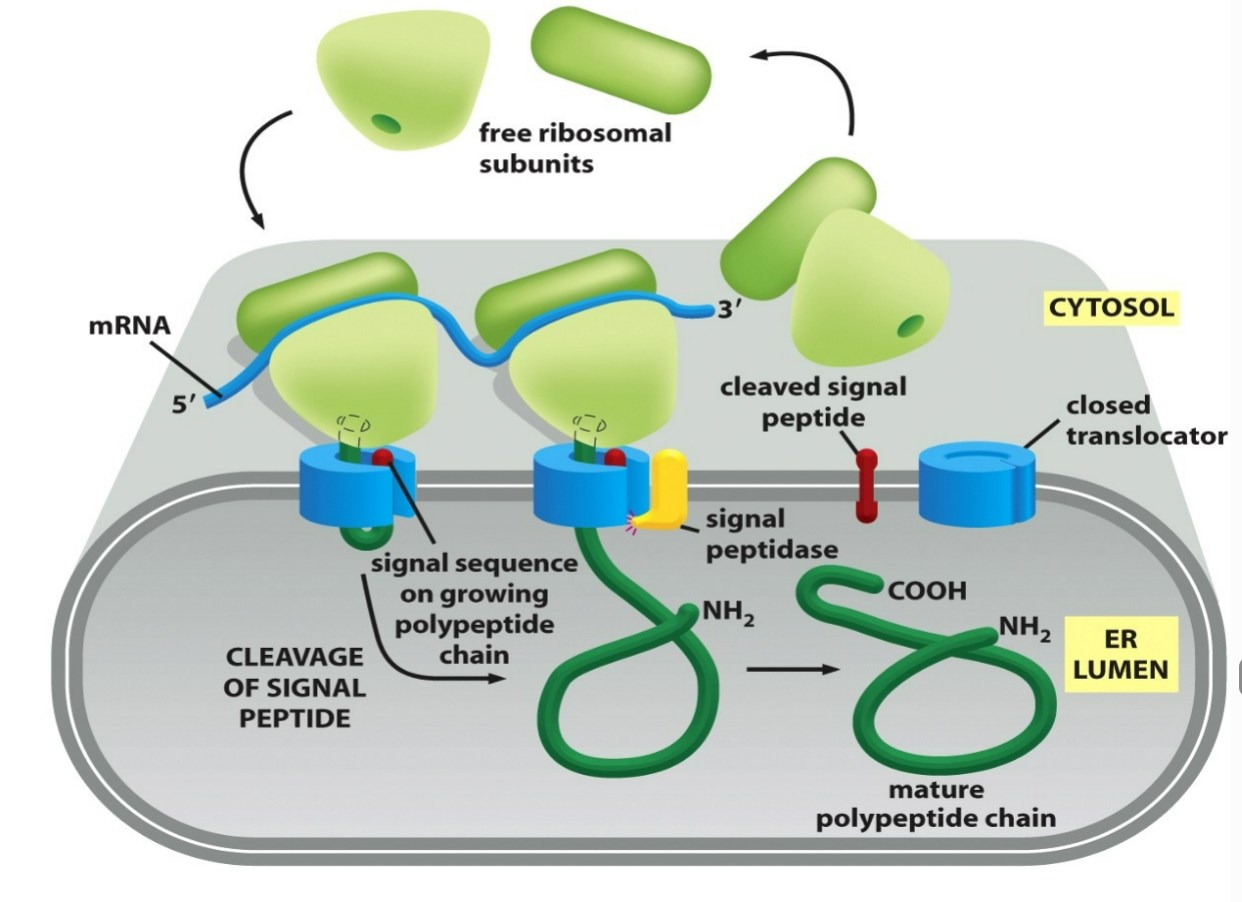
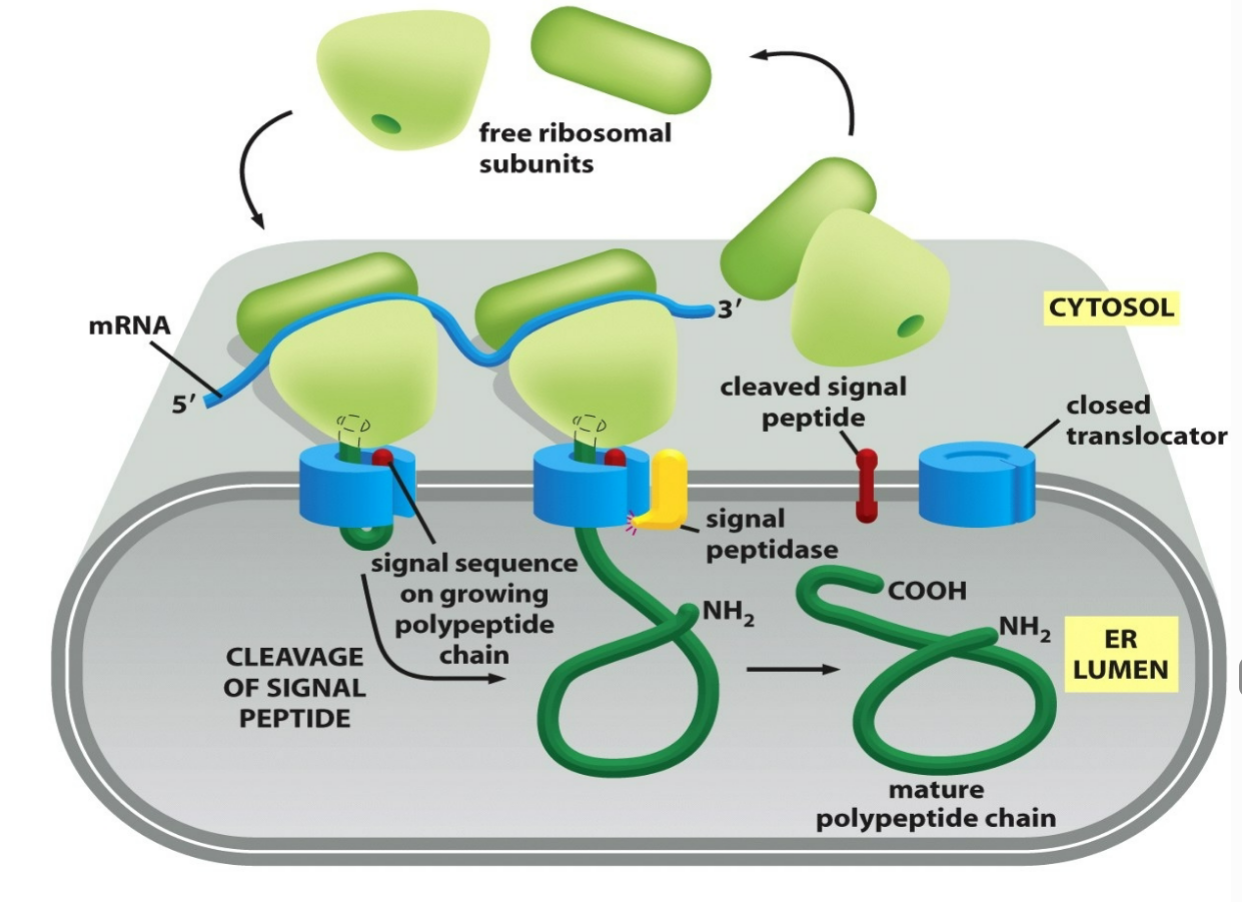
what happens to the ER signal sequences for water soluble proteins following translocation?
ER signal sequences will be cleaved off by a signal peptidase following translocation
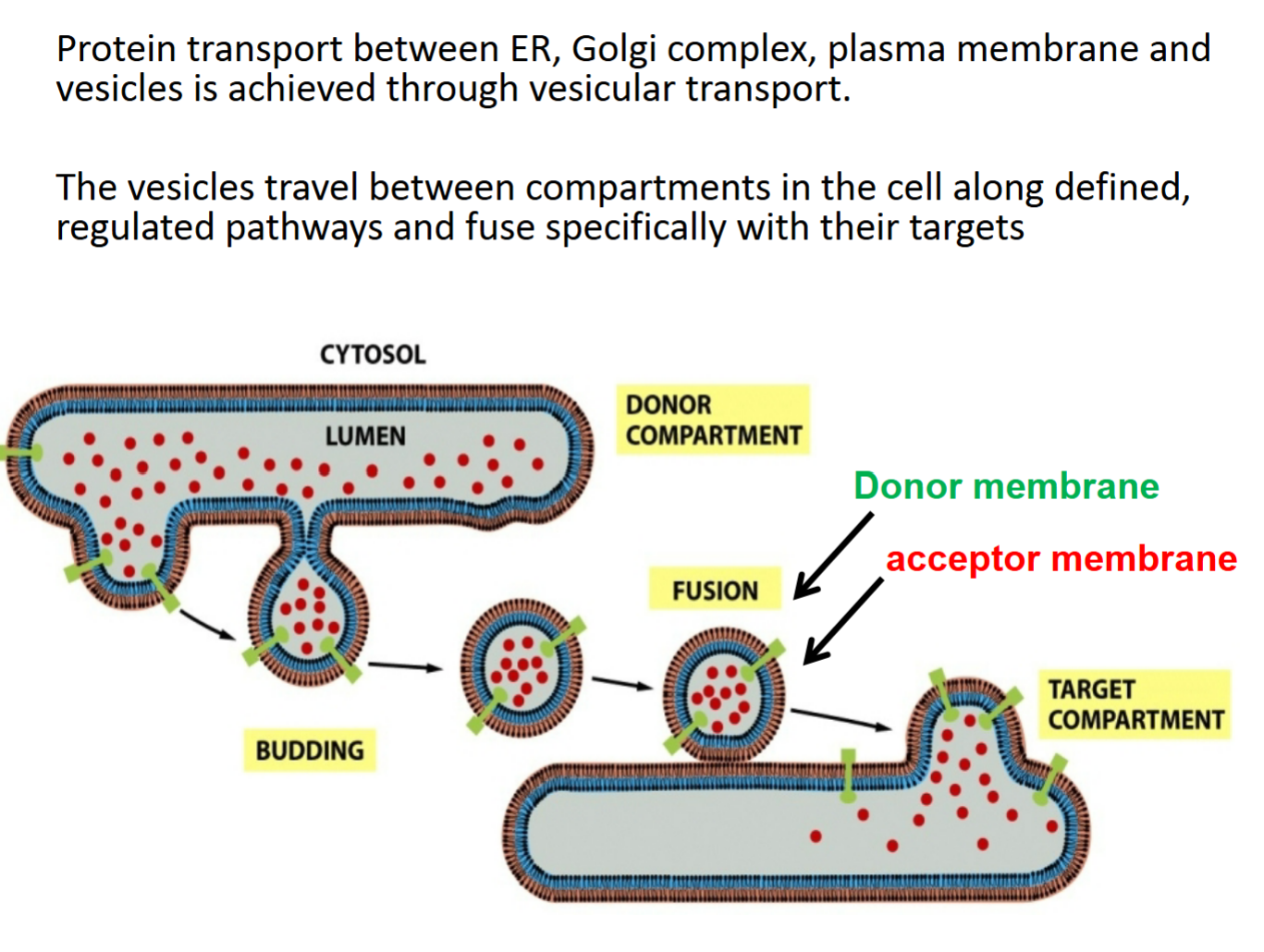
vesicular transport
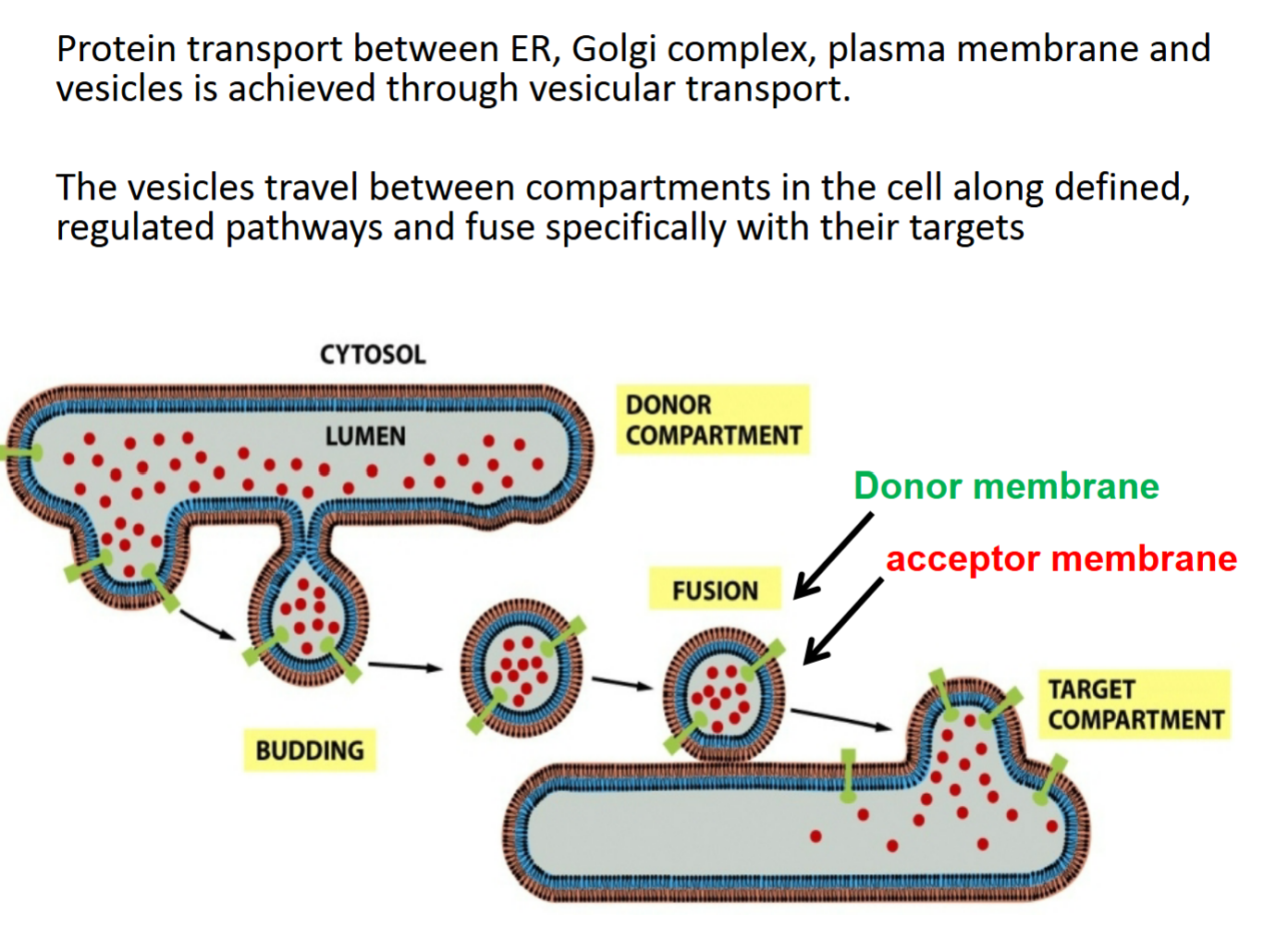
protein transport between ER, Golgi complex, plasma membrane and vesicles achieved through vesicular transport
vesicles travel between compartments in cell along defined, regulated pathways and fuse specifically with their targets
protein will NEVER travel across membrane
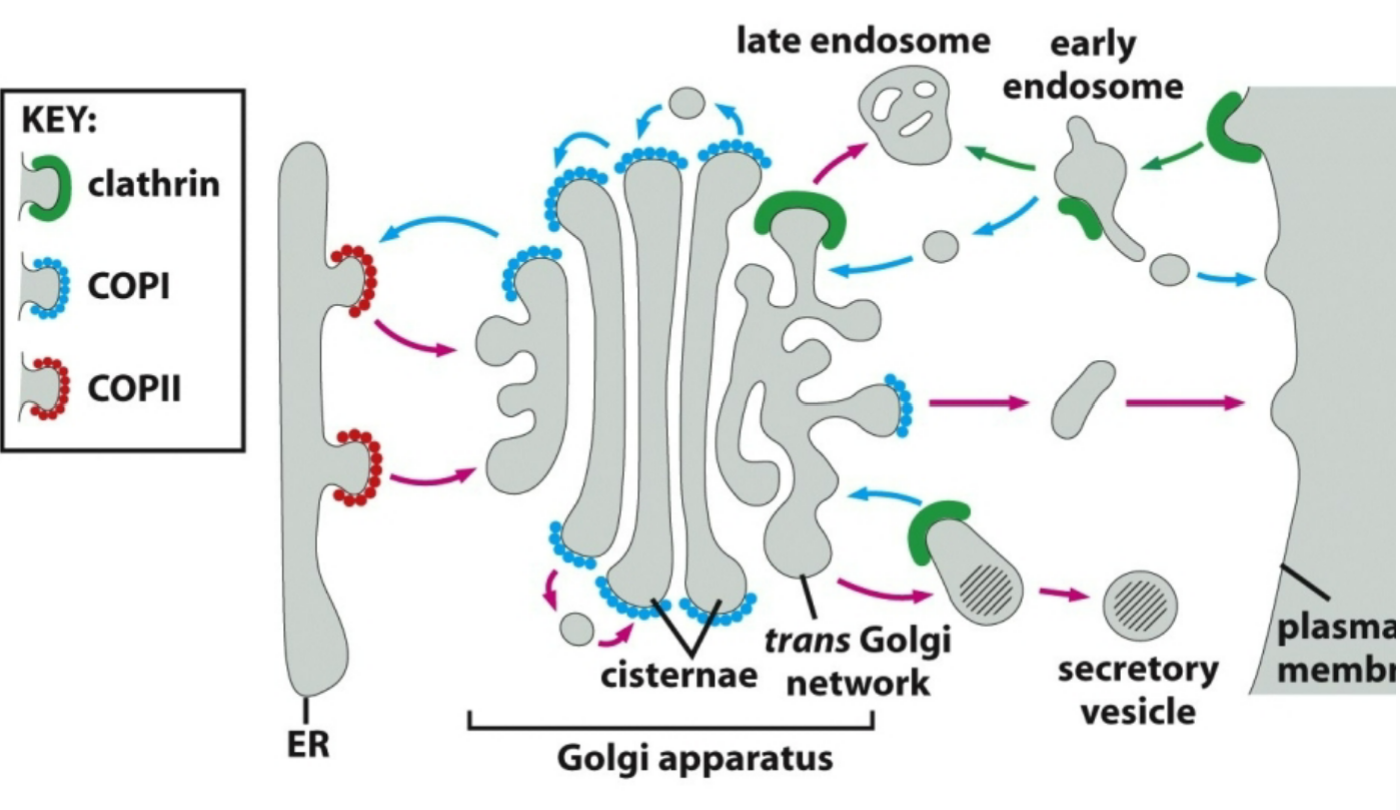
vesicular sorting depends on the assembly of a special protein coat formed at specific locations along a given donor compartment. what are those three protein coats that are involved?
COPII: coats from ER to golgi vesicles
COPI: coats from golgi to ER, golgi to plasma membrane (secretory vesicles), and within golgi
Clathrin: coats from plasma membrane to endosomal systems
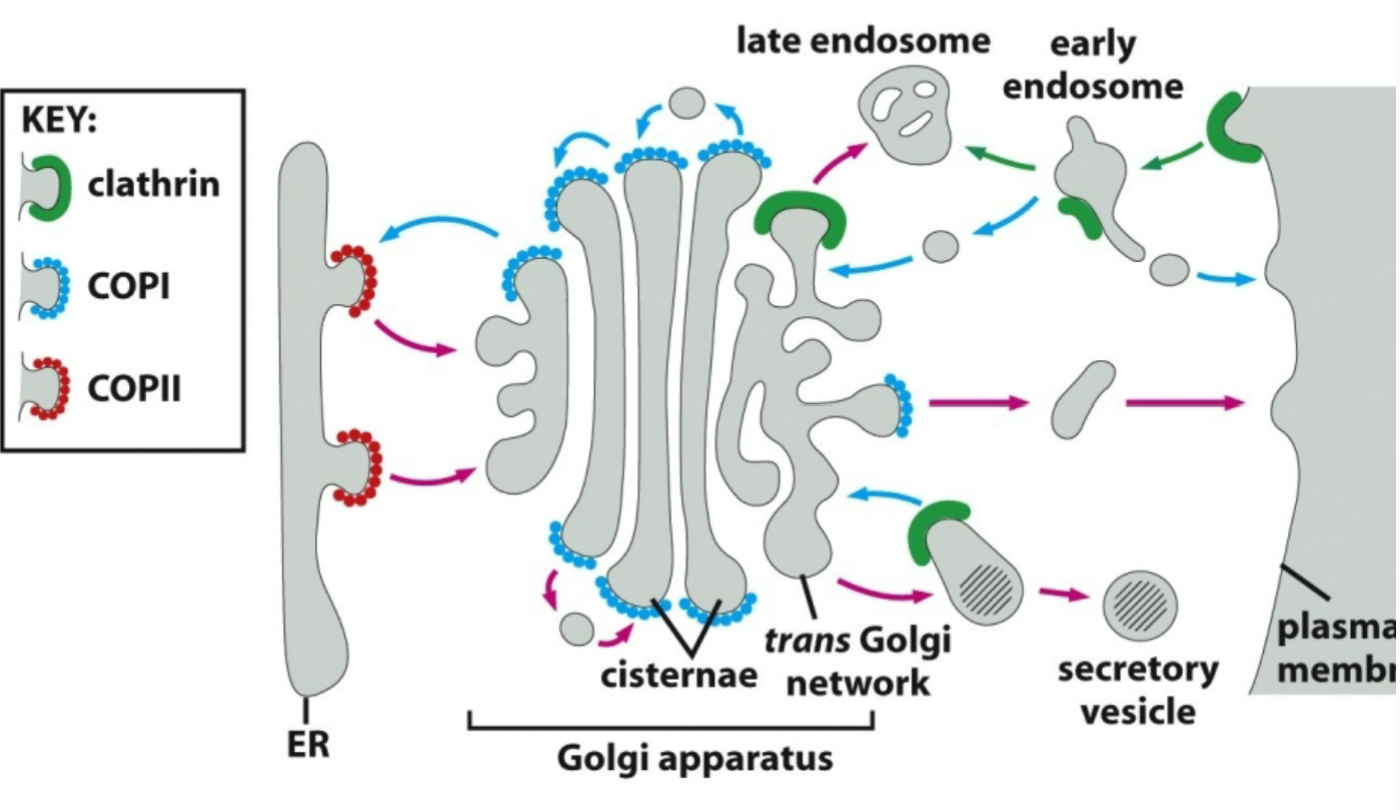
coat proteins represent initial step in vesicle formation
transport vesicles bud off as coated vesicles that have a distinctive cage of proteins (defined as membrane being distorted to create a "bulge” then releases to later fuse like a bubble) covering their cytosolic surface
before the vesicle fuses with target membrane, the coat is discarded → allow two cytosolic membrane surfaces to interact directly and fuse
transport vesicles bud off as coated vesicles that have a distinctive cage of proteins covering their cytosolic surface. what are cage of proteins?
distorts membrane to create “bulge” then releases to then later fuse into a bigger bubble
different coat proteins are involved in transport between different organelles. what are these three?
COPII → coats ER to golgi vesicles
COPI → coats vesicles moving from golgi to ER, golgi to plasma membrane (secretory vesicles), and within the golgi
clathrin → transport to/from and within plasma membrane to endosomal system (endocytosis)
how does coat assembly and vesicle stability work?
adapter proteins bind to membrane proteins and recruit coat proteins → often bind to cargo receptors - transmembrane proteins that bind soluble cargo proteins for transport
how is coat assembly controlled?
coat recruitment GTPases control coat assembly:
monomeric GTPases → regulate many steps in vesicular traffic
Sar-1 → regulates COPII assembly
Arf protein → regulate COPI and clathrin assembly
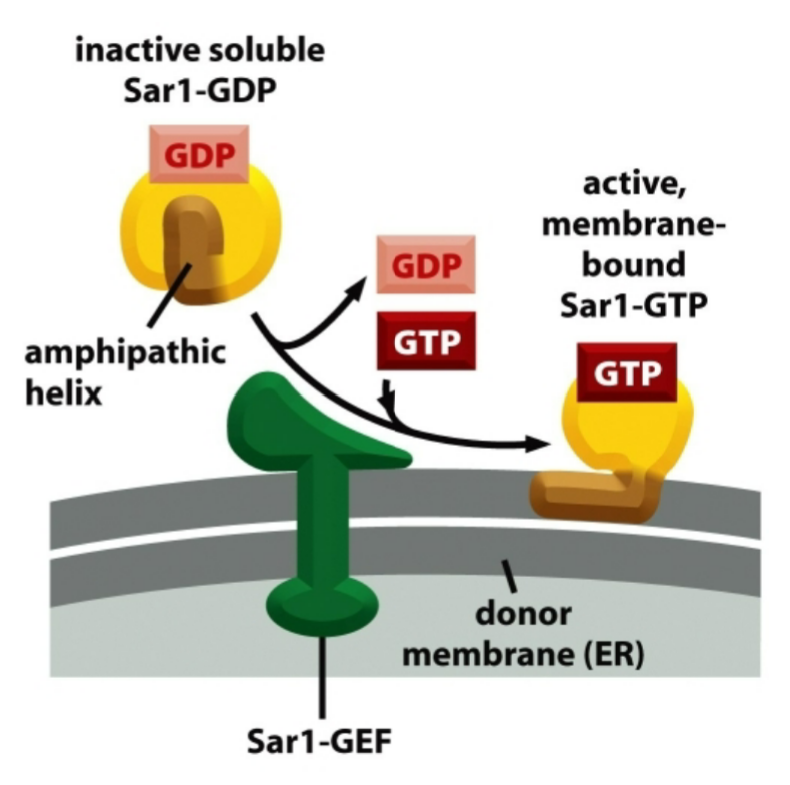
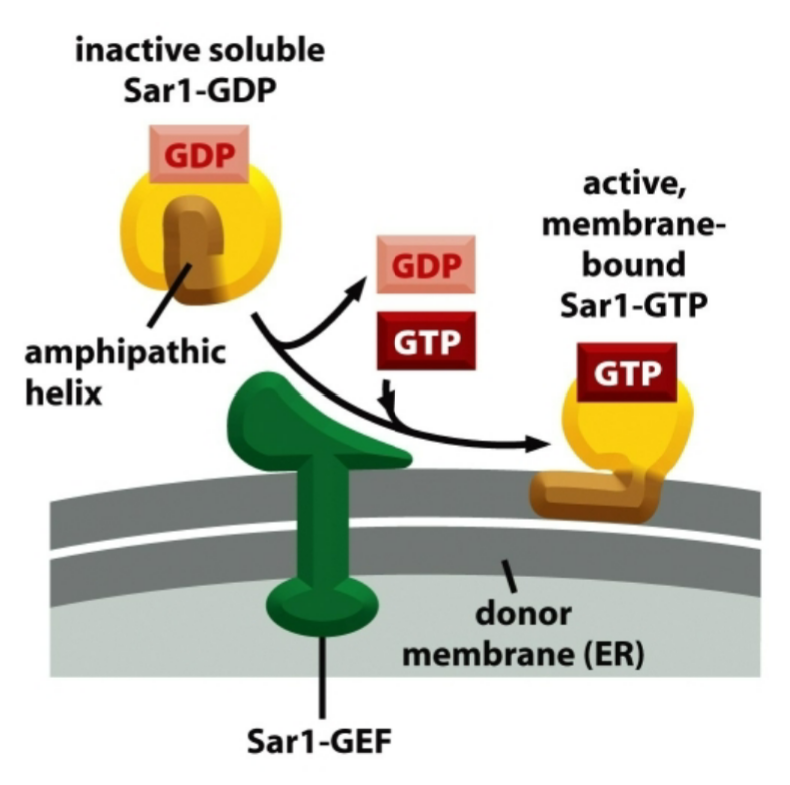
Sar-1 mechanism
Sar1-GDP = cytosol = inactive
Sar1-GTP = ER membrane-bound = active
active Sar1-GTP then promotes the assembly of coat complexes
GTP hydrolysis causes coat disassembly after budding
recognition of donor vesicles by acceptor membranes is controlled mainly by two classes of proteins. what are they?
SNAREs and Rabs
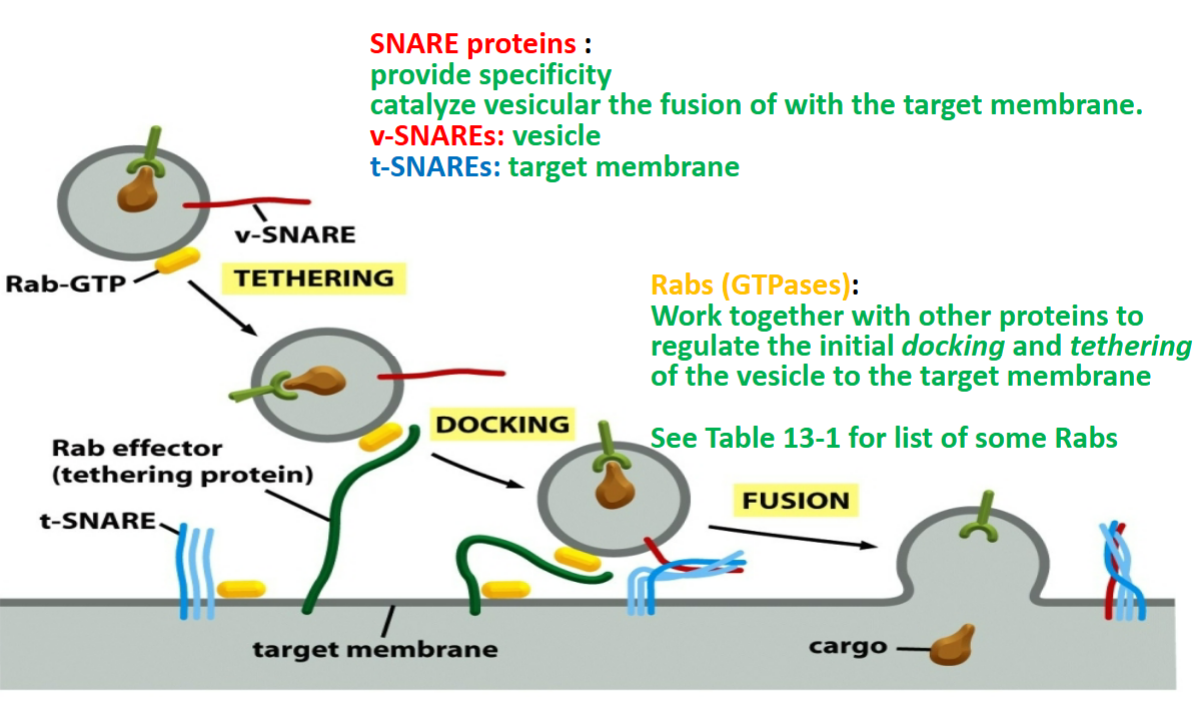
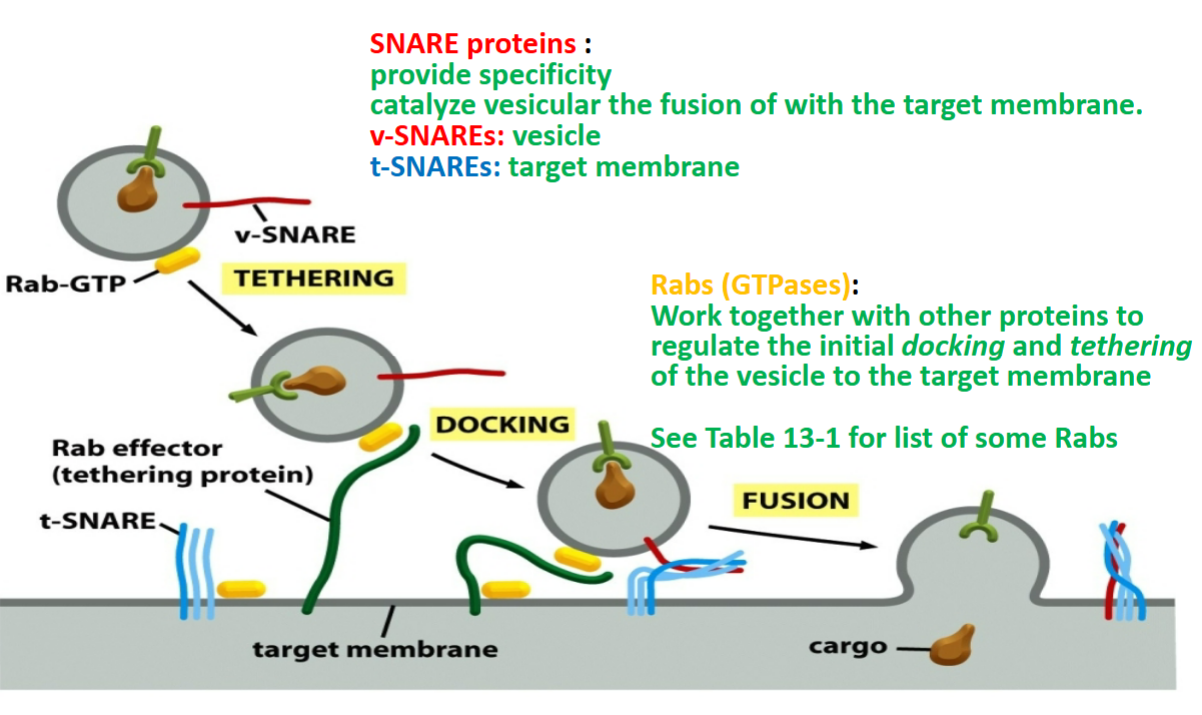
SNAREs
proteins that traps vesicle and causes fusion by snapping together of target membrane
provide specificity
catalyze vesicular the fusion of with target membrane
two kinds:
v-SNAREs → vesicle, thin and unfolded that threads through bubble to maintain structure, must have Rab-GTP binded
t-SNAREs → target membrane, within target membrane that entangles with v-SNARE to allow vesicle bubble to fuse along with target membrane where Rab-GTP lets go once vesicle fuses
Rabs
GTP-binding protein (GTP-ases)
recognizes to reach right membrane
work together with other proteins to regulate initial docking and tethering of the vesicle to the target membrane
tethering protein of Rab effector grabs or binds to Rab-GTP to allow v-SNARE and t-SNARE to entangle and allow vesicle bubble to fuse into membrane
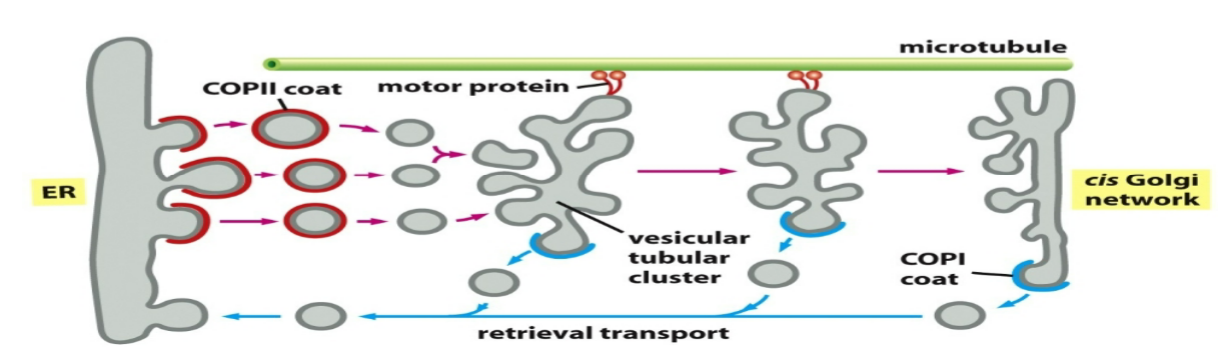
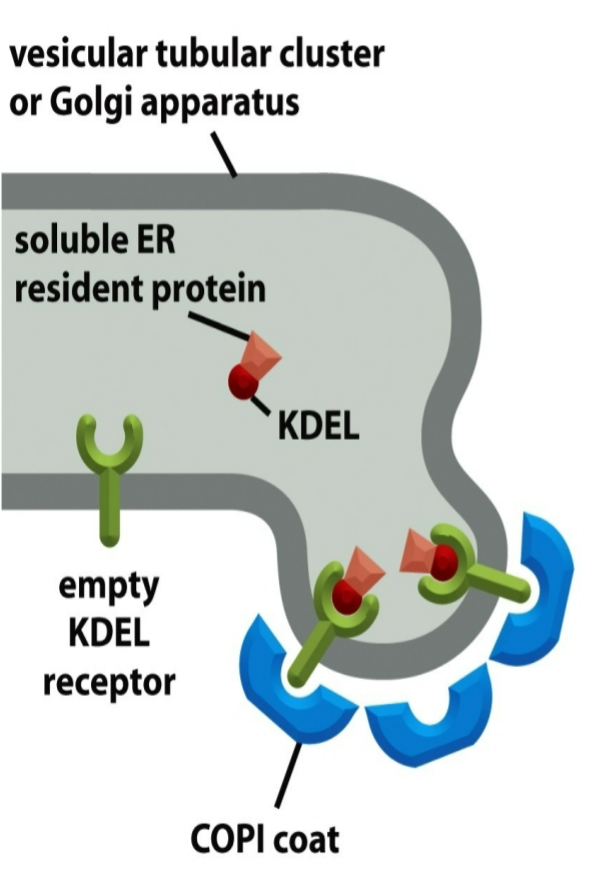
vesicular tubular clusters
transport vesicles leaving the ER fuse together to form intermediate compartments called vesicular tubular clusters
clusters travel towards cis Golgi via motor proteins on microtubule tracks and generate coated vesicles going back to the ER (COPI coat) - retrograde transport
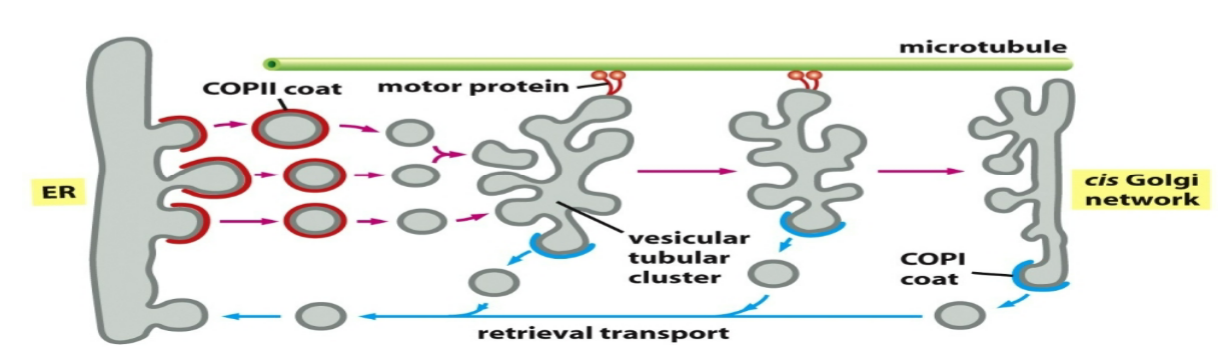
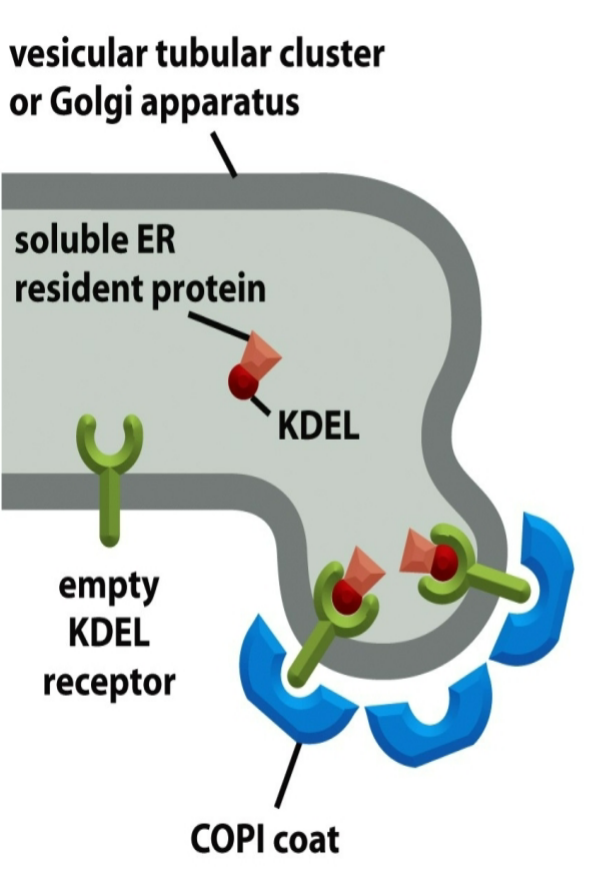
ER retrieval signals
membrane ER resident proteins: retrieval signals in cytosolic tails, recognized by COPI coat proteins
soluble ER resident proteins: retrieval signals within their structure, bind to receptors (ex: KDEL sequences → cannot have chemical nature altered)