CH30: Protein Biosynthesis (1)
1/31
Earn XP
Description and Tags
pt 1
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
32 Terms
what is translation
the process of biosynthesis - named bc the 4-letter nucleic acid alphabet is translated into the 20-letter protein alphabet
in what direction is mRNA decoded in? in what direction is protein synthesized in?
5’—3’ direction 1 codon at a time
protein is synthesized in the amino (5/N))-to-carboxyl (3/C) direction
for the following, name which terminus it can be found at
free amino group
free carboxyl group
first aa added
addition of aa
free amino group — N term
free carboxyl group — C term
first aa added — N term
addition of aa — C term
what reaction does polypeptide peptide bonds undergo?
dehydration synthesis (condensation) where carboxyl group of one aa reacts with amino group of another and releases a water mc
what is a codon and why is accurate recognition of codons important
three coding bases on the mRNA template (5’—3’)
accurate recognition is important bc it is required for the fidelity of protein biosynthesis (translation)
accuracy with which genetic code is translated to precise aa seq
what is the function of tRNA in terms of protein biosynthesis
functions as adaptor molecules between a codon & amino acid
what is an anticodon
portion of the tRNA that base pairs with codon (3’—5’)
how can each tRNA sequence be arranged and what is its structure?
2。structure can be arranged in a cloverleaf pattern which 50% nucleotides are base-paired
3D structure is L-shaped
each tRNA is a single chain w/ 73-93 ribonucleotides
what kind of unusual bases do tRNAs contain?
methylated/demethylated derivatives of A, U, C, and G
methylation increases the hydrophobic nature of the base
in tRNA, five groups of bases are not base-paired, but they participate in what interactions?
H-bonding
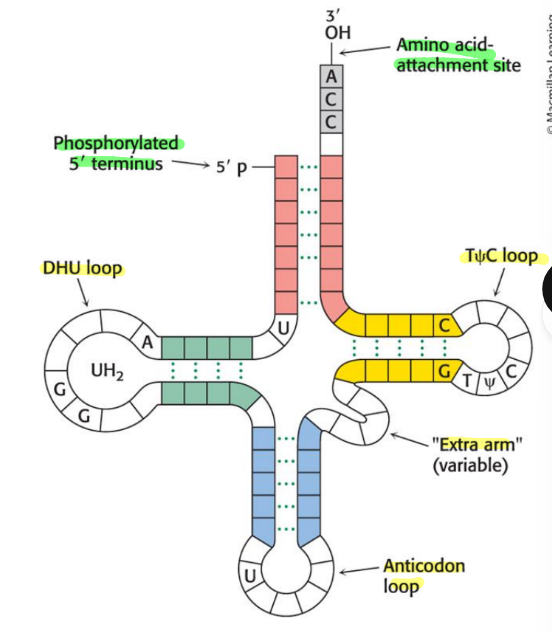
Draw and label the following on a tRNA
3’ CCA terminal region
TψC loop
extra arm
DHU loop
anticodon loop
acceptor stem
amino acid attachment site
acceptor stem is the red region and CCA region
t loop on right
dhu loop on left
extra arm lower of t loop
anticodon loop on bottom
what forms the L-shape tertiary structure of tRNA & why’s it important?
four double-stranded regions of tRNA stack
the acceptor stem, T-arm, anticodon arm, and DHU arm
it allows ribosomes to interact with both the CCA terminus & anticodon loop at the same time
what happens to the 5’ end of a tRNA and what is its 5’ terminal residue?
phosphorylated
usually a pG
where exactly is an activated amino acid attached to on the tRNA?
a hydroxyl group of adenosine in the CCA region of the acceptor stem
what couples amino acids to tRNAs
ester linkages
why can some tRNA molecules recognize more than one codon?
because of wobble in base-pairing
wobble: steric freedom in the pairing of the third base of the codon
what is redundancy or degeneracy of the genetic code? what is the wobble hypothesis?
redundancy: indicates that recognition of the 3rd base of a codon is sometimes less discriminating than the other two
wobble hypothesis: predicts binding of anticodons to codons
what codons must be recognized by different tRNAs
codons that differ in either of their first two bases
what codon and anticodon position does wobble occur?
codon position: 3rd base
anticodon position: 1st base
what are the allowed pairings at the 3rd base of the codon according to the first base of anticodon (using hypothesis)
C
A
U
G
I (inosine)
note that I - inosine purine base pairs with cytidine, uridine or adenosine
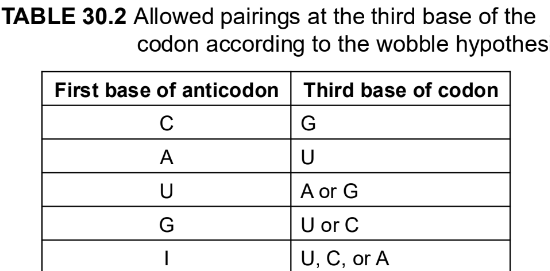
using wobble rules, how many codons can each anticodon recognize?
anticodon 5’ base = G
anticodon 5’ base = I
1: 2 (third base can be U or C)
2: 3 (third base can be U, C, or A)
how many min different tRNAs are required to read these codons?
Gly codons:
GGU, GGC, GGA, GGG
2
since the 3rd base is wobble, we need to find anticodons that pairs with U, C, A and G
an anticodon with first base of G can read GGC and GGU
an anticodon with first base U can read GGA and GGG
an anticodon with first base I can read GGU, GGC, GGA
is the formation of a peptide bond between free amino acids favorable?
it is not thermodynamically favorable
how are amino acids activated? where does it occur and what does it form? what catalyzes this?
formation of ester-linkage
happens between carboxyl group of aa & either the 2’ or 3’ hydroxyl group of the terminal adenosine of tRNA
forms an aminoacyl tRNA (charged tRNA)
aminoacyl-tRNA synthetases catalyze aa activation (adenylation)
what is the first step of aa activation?
formation of aminoacyl adenylate (aminoacyl-AMP)
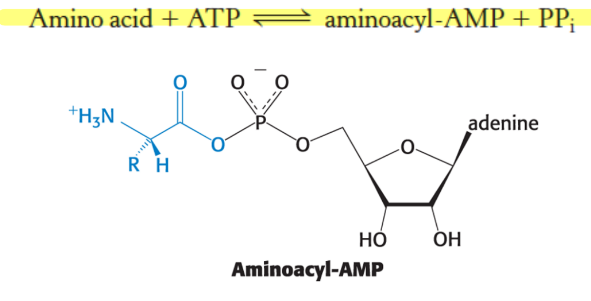
what is the second step of aa activation? what is the sum of activation & transfer steps? what is this reaction driven by?
next step: aminoacyl group of aminoacyl-AMP is transferred to a particular tRNA to form aminoacyl-tRNA
aminoacyl-AMP + tRNA —> aminoacyl-tRNA + AMP
sum:
amino acid + ATP + tRNA —> aminoacyl-tRNA + AMP + PPi
reaction is driven by hydrolysis of pyrophosphate
what is consumed in forming the ester linkage of aminoacyl-tRNA and what is consumed in driving reaction forward?
one ATP equivalent for each step
what does it mean when the aminoacyl-AMP intermediate doesn’t dissociate from the synthetase
the same aminoacyl-tRNA synthetase catalyzes both steps of the reaction
why are these enzymes essential for maintaining the accuracy of translation? and how do they avoid coupling to the wrong aa? what about Val and Ser?
each aminoacyl-tRNA synthetase is specific for a given amino acid
=
to avoid coupling to wrong aa, threonyl-tRNA synthetase has a zinc ion at active site that binds to amino & hydroxyl group of Thr
Val is similar in overall structure to Thr but lacks hydroxyl group, so it’s not joined to tRNA Thr
Ser is occasionally linked to tRNAThr bc of presence of hydroxyl group
how are aminoacyl-tRNA synthetases the true translators of genetic code?
they assign a particular amino acid to a specific tRNA
some synthetases recognize their tRNA partners on the basis of their anticodons
synthetases may recognize other aspects like loops rich in unusual bases of tRNAs
how does CCA arm extend and how does this aminoacyl-tRNA synthetase interact?
extends into the zinc-containing activation site
enzyme interacts with the acceptor stem of tRNA & anticodon loop
T/F tRNAs have multiple recognition sites for aminoacyl-tRNA synthetases
true