BIMM 120_Midterm #2_Saier Milton
1/62
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
63 Terms
What does ATP synthase use to make ATP?
Uses PMF, an electrochemical H⁺ gradient across the membrane, to drive ATP production.
Rotation Direction:
CCW = ATP synthesis. Protons flow down gradient
CW = ATP hydrolysis. ATP drives proton pumping against gradient
F₀ Component:
-Membrane motor that uses proton flow down gradient to drive rotation.
-a: Half-channels that guide protons to/from c-ring.
-c: Forms c-ring; binds and rotates protons
-b: Links F₀ to F₁ and prevents catalytic head rotation.
F₁ Component:
-Cytoplasmic catalytic head that makes ATP
-β: catalytic (make ATP)
-α: structural support
-γ and ε: hydrophilic rotating shaft where γ spins inside α₃β₃ and drives β-subunit conformational to drive ADP + Pi → ATP
Rotor vs Stator:
-Rotor: c-ring + γ + ε rotate together as one unit
-Stator: α, β, a, b, δ hold catalytic head stationary
ATP Synthesis Mechanism:
-Protons flow through F₀ down gradient and bind Asp residues on the c-ring, generating torque that rotates the c-ring and γ shaft inside the stationary β subunits to drives ADP + Pi → ATP.
Comparing Distinct Classes of Rotary ATPases:
-FVAP (Friend Visit Ancient Place)
-F-type ATPases: move 12 H+ to produce 3 ATP with synthesis or hydrolysis, using ion gradient to make ATP
-V-type ATPases: pumps 6 H+ using 3 ATP with hydrolysis. Found in eukaryotic intracellular membranes
-A-type ATPases: move 8 H+ to produce 3 ATP with synthesis. Found in archaea.
-P-type ATPases: Uses ATP to pump ions out of the cell to create acidic outside (low pH/high H+) and basic inside (high pH/lower H+) to build proton/sodium motive force.
Glycolysis:
Net 2 ATP produced to power P-type ATPases to pump H⁺ or Na⁺ and build IMF
Cryogenic Electron Microscopy (Cryo EM):
Enabled detailed structural characterization of ATP synthase.
Flagella:
Powered by IMF, where H⁺ flow into the cytoplasm through the MotA/B proton channel ion selector to generate torque for rotation. Important for surface sensing, adhesion, and biofilm formation
-CCW: Flagella bundle behind bacteria → smooth forward movement (run).
-CW: Flagellar unbundle → tumble (peritrichous) or reverse direction (polar)
Polar vs Peritrichous flagella:
-Polar flagella: At cell ends; enable forward, backward, and tumble motion
-Peritrichous flagella: Distributed over the cell surface. Optimized for chemotaxis (e.g., E. coli, Salmonella).
Flagellar Structure:
-Basal Body: Anchor flagellum and contains motor
-Hook: Connects basal body to filament and transmits torque (HAP1, HAP3).
-Filament: Long, hollow, helical propeller.
-Tip (HAP2): Controls filament growth
-Assembled from tip to base: Basal body → hook → filament using Type III secretion system (T3SS)
Chemotaxis:
Allows bacteria to move toward hydrophilic attractants CCW run and away from hydrophobic repellents CW tumbles using flagella and MCPs. Negative cooperativity makes additional ligand binding harder, allowing gradual receptor responses over wide concentration ranges.
Chemotaxis signaling system:
-My Weird Aunt Yells Zesty Ripe Bananas (MCP, W, A, Y, Z, R, B)
-CheW/A are interactive site & CheR/B control methylation
-MCP (Methyl-Accepting Chemotaxis Protein): Receptor that detect attractants/repellents
-CheW: Link MCP to CheA.
-CheA: Sensor kinase that autophosphorylates CheA to CheA-P
-CheY: Response regulator. Inactive CheY causes CCW run, while active CheY-P causes CW tumble
-CheZ: Dephosphorylase CheY-P, resetting system to CCW run state.
-CheR: Always active methylase (add methyl to MCP) to decrease sensitivity
-CheB: Demethylase (remove methyl from MCP) is activated when phosphorylated by CheA to CheB-P to restore sensitivity.
Fast vs Slow process:
-Fast (Activation): controls movement immediately.
-CheA phosphorylates to CheY-P
(a) More attractant: ↓ CheA and CheY-P → CCW Run
(b) More repellent: ↑ CheA and CheY-P → CW Tumble
-Slow (Adaptation - Methylation system): adjusts sensitivity over time.
(a) Constant high attractant: CheR methylates MCP → receptor desensitization → CheY-P returns to baseline (CCW).
(b) Decreasing attractant or increasing repellent: CheA activates & phosphorylates CheB to CheB-P→ demethylates MCP → ↑ sensitivity. → ↓ CheA and CheY-P (return to CCW rotation)
-Note: high deterrent concentration = increased repellent
Fimbriae/Pili:
Short, pilin-based surface structures assembled from base-to-tip (unlike flagella from tip to base via T3SS) that function in DNA transfer, surface adhesion, and gliding/twitching motility.
Swimming vs Swarming:
-Swimming: Individual bacterial movement in liquid using polar flagella and powered by sodium motive force (Na+ MF)
-Swarming: Coordinated surface or viscous motility where transition from swimming causes morphology of cells to elongate and produce many proton-driven flagella.
Bacterial vs Archeal Flagella
-Bacterial flagella: PMF-driven, hollow rotary motors that grow at the tip and control runs/tumble. Mutations causes intact flagella with no chemotact ability
-Archaeal flagella: ATP-powered, non-hollow, and structurally homologous to Type IV pili proteins.
-Structurally and evolutionarily distinct (including filaments) and not homologous.
Non-flagellar movement methods
-Type IV pili (T4P): Drive twitching motility via ATP-powered extension, surface attachment, and retraction PilA forms the pilus, PilB mediates extension, and PilT mediates retraction.
-Centipede or Inchworm Motility: Mycoplasm moves in the direction of a head region, where large surface Gli proteins act like legs at the “neck” to produce gliding motility.
Adventurous and Social Motility:
In M. Xanthus, S-motility is Type IV pili–dependent for coordinated group movement, while A-motility is pilus-independent and driven by polysaccharide secretion.
Thermotaxis:
-Bacterial movement based on temperature changes using MCP.
-Tsr, Tar, Trg are warm sensors, where ↑ temperature = CCW smooth swim and ↓ temperature = CW tumble. Tsr is involved in multiple movement methods
-Tar temperature preference depends on energy/methylation (methylated/high energy = cold sensor; unmethylated/low energy = warm sensor).
-Tap is a cold sensor that biases movement toward cold.
Aerotaxis:
Bacterial movement driven by respiration instead of direct O₂ sensing. Aer sense oxidized FAD = active respiration & reduced FADH₂ = reduced respiration. Tsr sense PMF (H+), where high PMF = high energy and low PMF = low energy. Aer and Tsr act independently
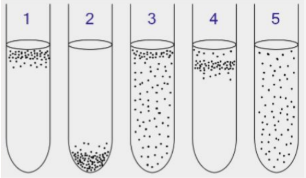
Diagram labeling of bacterial oxygen preferences based on behavior in liquid.
1) Obligate Aerobes: oxygen-dependent and swim toward liquid–air interface
2) Obligate Anaerobes: oxygen-independent and swim deeper down, away from liquid–air interface
3) Facultative Anaerobes: Grow with or without O₂ but prefer O₂, so concentrated near the top but present throughout
4) Microaerophiles require low O₂ and form a sharp band at optimal depth
5) Aerotolerant Microbes: do not use O₂ but tolerate it and are evenly distributed.
Magnetotaxis:
Bacterial movement guided by Earth’s magnetic field using magnetosomes or epigenetically reoriented when environment changes to direct cells toward nutrient-rich, low-oxygen sediments, with MamK (an actin-like ATPase) forming filaments and MamJ anchoring magnetosomes to ensure proper alignment and assembly.
Mechanosensing and surface sensing:
Allow bacteria to detect surface contact via flagella impedance (rotation interrupted) and Type IV pili attachment/retraction, triggering adhesion, biofilm formation, and increased virulence.
Bacterial nanowires:
Conductive Type IV pili (T4P) enable long-range electrons transfer through aromatic AA to insoluble extracellular (outside cell) electron acceptors, reducing them (e.g., Fe₂O₃ → Fe₃O₄) into usable, more soluble forms to enable respiration
Spirochetes:
Opposite rotation of periplasmic flagella and cell body located between the inner and outer membranes drive corkscrew motility effective in viscous environment
Bacteriophages function/structure/isolation:
Bacteria-infecting viruses used in phage therapy due to high specificity, low toxicity, and ability to replicate at infection sites. They consist of a capsid head with genetic material, a tail/sheath for genome injection, and tail fibers for host recognition and are isolated by filtering environmental samples, plating on a bacterial lawn, and purifying plaques.
Phage Infection (Lytic vs Lysogenic):
-Lytic phages: Inject genome, hijack host machinery, replicate, and lyse bacterial cells to cause rapid host death.
-Lysogenic phages: integrate genome into the host chromosome as a prophage and replicated during host cell division without immediately killing the cell with lysis.
Phage therapy:
uses lytic phages (sometimes engineered/CRISPR-enhanced) to specifically kill pathogenic bacteria and disrupt biofilms, requiring understanding of host range and phage diversity. Pharmacokinetics (PK): phage behavior in the body & Pharmacodynamics (PD): how phages interact and kill bacteria
Pseudomonas aeruginosa (sue-dough-know-miss, aero-gin-nosa):
Uses Type IV pili for surface sensing, where increased intracellular cAMP decreases flagellar rotation due to surface impedance and activates translation of virulence factors, and relies on OprM efflux pumps for antibiotic resistance
Bacterial Virulence Factors (Adhesins, Invasins, Serum Protectant, Cytotoxins & Proteases. Capsules):
-Molecules that help bacteria survive and cause disease.
-Adhesins: Help attach to host cells surface carbohydrates, often at fimbrial tips but non-fimbrial exist
-Invasins: Help enter host cells and avoid immunity
-Serum Protectants: Block complement/innate immunity killing
-Cytotoxins & Proteases: Damage host cells/tissues to aid infection
-Capsules: Prevents immune recognition and limits antibiotic penetration.
Secretor (Sec) system:
Exports hydrophilic virulence proteins from the cytoplasm across the hydrophobic membrane to temporarily unfold to cross and then refold outside the cell to function
Gut microbiome:
Diverse internal microbial ecosystem that produces short-chain fatty acids (SCFAs) for energy, suppresses pathogens, regulates immunity, and ferments indigestible food using microbial niches.
Microbial niche (Fermenter, Pathogen suppressors, Mucus degraders, Nitrogen cyclers. Secondary metabolite producers):
-Specific gut microenvironment shaped by nutrients, oxygen, pH, and host interactions
-Fermenters → break down carbs/fiber to produce SCFAs. Example of mutualistic relationship
-Pathogen suppressors → compete for nutrients + produce antimicrobials
-Mucus degraders → break down gut mucins
-Nitrogen cyclers → affect nitrogen metabolism & gut pH
-Secondary metabolite producers → make vitamins/neurotransmitters that influence gut-brain communication
Mutualism vs Commensalism vs Pathogenicity
-Mutualism: Microbe and host benefit.
-Commensalism: Microbe benefits and host is unaffected.
-Pathogenicity: Microbe benefits at host’s expense (causes disease).
Helicobacter pylori (pie-lore-ride):
Stomach pathogen causing chronic inflammation and increase risk of ulcers (uhl-sir) and gastric cancer.
Inflammatory Bowel Disease (IBD):
1. Crohn’s disease: Affect any GI region and all intestinal wall layers.
2. Ulcerative colitis: Limited to the colon and affects only the innermost lining.
Factors affecting the gut microbiome:
1. Antibiotics & Medications – Disrupt balance and allow harmful overgrowth.
2. Lifestyle & Environment – alter microbiome composition.
3. Diet: High-fat shape gut microbial
(a) Probiotics = beneficial microbes.
(b) Prebiotics = nutrients that feed beneficial microbes.
4. Western Diet – Promotes inflammation, chronic gut disease, and insulin resistance.
5. Gut-Brain Axis – Communication between the ENS (gut) and the CNS (brain) where disruption affects mood, sleep, and pain.
Pap system:
A chaperone-usher system in E. coli that assembles fimbriae by stabilizing pilus subunits in periplasm and inserting them through an outer membrane usher to extend the pilus outward.
Molecular tropism:
Concept that a single molecular structure or mechanism can produce different disease outcomes.
Persistence vs Resistance vs Tolerance
Persistence: Dormant cells that survive antibiotics and regrow.
Resistance: Genetic changes that allow growth despite antibiotics, increasing MIC.
Tolerance: Bacteria die at a very slow rate in antibiotics with no MIC change.
Two Types of Persistent Bacteria:
-Type I Persisters: Environmental stresser
-Type II Persisters: Occur randomly at low frequency.
Mechanisms of Persistence
1. Toxin–Antitoxin (TA) System: Stress-activated system via ppGpp when in high concentration to slow growth
2. SOS Response: DNA damage repair response (e.g., UV/oxidants) that generates single-stranded DNA.
3. Quorum Sensing: Cell-density signaling that promotes persister formation.
Where persister cells are found and how to kill them:
-Found in hostile environments, biofilms, intracellular regions, and granulomas (grand-new-low-miss)
1. Direct-killing: Using membrane-targeting antibiotics, antimicrobial peptides, or phage therapy
2. Drug Combinations: Use antibiotics plus metabolic/environmental modifiers to improve killing.
Key Idea of Antibiotic History:
Antibiotic resistance evolves quickly once selective pressure is applied
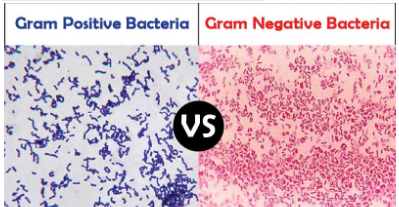
Gram positive vs Gram negative bacteria
-Gram-positive: Thick peptidoglycan cell wall with no outer membrane.
-Gram-negative: Thin peptidoglycan layer with an outer membrane containing LPS. Ex: e-coli has inner & outer periplasmic space
Antimicrobial Resistance (AMR):
The ability of microbes to survive and grow despite antimicrobial treatment due to human or environmental selective pressure.
Horizontal Gene Transfer (HGT)
-Rapid spread of resistance genes between bacteria.
1. Conjugation: Direct cell-to-cell DNA transfer via pilus.
2. Transformation: Uptake of free environmental DNA.
3. Transduction: DNA transfer via bacteriophages.
Mechanisms of Antimicrobial Resistance (AMR)
-Enzymatic Degradation of Antibiotics: Produce β-lactamases to degrade β-lactam ring of antibiotics.
-Reduced Antibiotic Uptake: by decreasing porin size or expression to limit drug entry.
-Active Efflux: Pump antibiotics out of the cell.
-Target Modification: Mutations alter binding sites to prevent antibiotic binding.
-Alternative Metabolic Pathways: Bypass inhibited pathway to maintain the same function.
-Biofilm Formation & Quorum Sensing: increasing resistance and persistence.
Ways to combat Antimicrobial Resistance (AMR):
Reduce selective pressure through responsible antibiotic use, develop new antibiotics, and phage therapy
Biofilms:
sessile microbial communities embedded in a self-produced EPS matrix that provides structural protection and antimicrobial resistance in response to stress and regulated by cyclic-di-GMP and quorum sensing, with organized surface and core cells to enhance persistence.
Stages of Biofilm Formation
1. Reversible attachment
2. Irreversible attachment
3. Microcolony formation
4. Dispersal
eDNA (extracellular DNA):
DNA released from lysed cells to reinforce biofilm structure and facilitates horizontal gene transfer (HGT)
Surface vs Core Cells:
-Surface cells: High oxygen and nutrients, aerobic, metabolically active, and produce EPS.
-Core cells: Low oxygen and nutrients, anaerobic, grow slowly, and prone to persistent state.
Bacterial capsules:
Sticky polysaccharide layers that aid attachment and evade immunity by blocking antibody binding.
Chaperonin vs Proteasome vs DnaJK:
-Chaperonins: Protein complexes that assist folding by providing a protected environment to prevent aggregation but do not determine final structure.
-Proteasomes: ATP-dependent complexes that degrade misfolded proteins. Eukaryotes use ubiquitin tagging, while bacteria use other proteases.
-DnaJK: Chaperonin system that initiates protein folding. If folding fails, proteins are tagged for degradation and recycled.
Protein folding pathway in E. coli:
-Overview: 1) Unfolded protein → DnaK-DnaJ-GrpE → Partial folding 2) GroEL-GroES → Folded protein
-GroEL: Double-ring barrel where folding occurs.
-GroES: Cap structure that binds to GroEL w/ ATP to enclose chamber and allow proper folding
“-some” in bacteria (Cellulosomes, Carboxysomes, Degradosomes/Exosomes):
-Often indicates breakdown/hydrolysis
-Cellulosomes: Extracellular complexes that degrade cellulose into sugars.
-Carboxysomes: Protein shells that concentrate bicarbonate (HCO₃⁻) & CO₂ for fixation.
(a) RuBisCo: Catalyzes CO₂ fixation
(b) Carbonic anhydrase: Converts HCO₃⁻ to CO₂, to increase local concentration
-Degradosomes (prokaryotes) / Exosomes (eukaryotes): Degrade RNA into nucleotides and maintain RNA turnover.
Sulfur Granules:
Store sulfur in photosynthetic and chemolithotrophic bacteria for later redox energy use.
Gas Vesicles:
Air-filled structures that allow aquatic bacteria and archaea to float.
Protein compartments:
Proteins can self-assemble into distinct cytoplasmic compartments without membranes or shells.
Circadian clock of cyanobacteria:
-Regulates daytime photosynthesis and nighttime nitrogen fixation.
-KaiA: Promotes KaiC phosphorylation.
-KaiB: Inhibits KaiC phosphorylation.
-KaiC: Autokinase and autophosphatase
-CikA: Dephosphorylates RpaA to reset clock
-SasA: Competes with KaiB to bind KaiC → RpaA phosphorylation.
-RpaA: Transcription factor controlling circadian gene expression.
-Day: KaiA activates KaiC → KaiC autophosphorylates.
-Night: KaiB binds phosphorylated KaiC → inhibits KaiA.