Spring 2026 Biomedical Science Exam 2 material
1/155
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
156 Terms
agents that damage the DNA and cause mutations
- ionizing rays
- UV rays (*UVC rays)
- reactive oxygen species
- chemicals in the environment
- chemicals used in chemotherapy
UVC rays
UV-C from the sun does not reach the earths surface because it is absorbed by ozone layer
- Most of UV-A and some UV-B do reach earth's surface and can cause DNA damage
reactive oxygen species
chemical reactive molecules that contain oxygen; e.g., oxygen ions and peroxides
- produced during cellular metabolism
chemicals in the environment
hydrocarbons in cigarette smoke (tar, benzene)
- plant and microbial products such as aflatoxins
- (metabolites can intercalate DNA) produced by moldy peanuts
catagoeies of mutations
- substituiton
- transition
- transversion
- insertions
- deletions
- frame shift mutations
substitution mutation
a mutation that exchanges one base for another
transition mutation
One pyrimidine is changed to the other pyrimidine, or one purine is changed to the other purine
transversion mutation
Substitution of a purine with a pyrimidine, or substitution of a pyrimidine with a purine
insertions mutation
mutations in which extra base pairs are inserted into a new place in the DNA.
deletions mutation
mutations in which a section of DNA is lost or deleted.
frame shift mutations
Since protein coding DNA segment is divided into codons which is a triplet (3 letter code), insertions and deletions can alter the reading frame.
serine structure
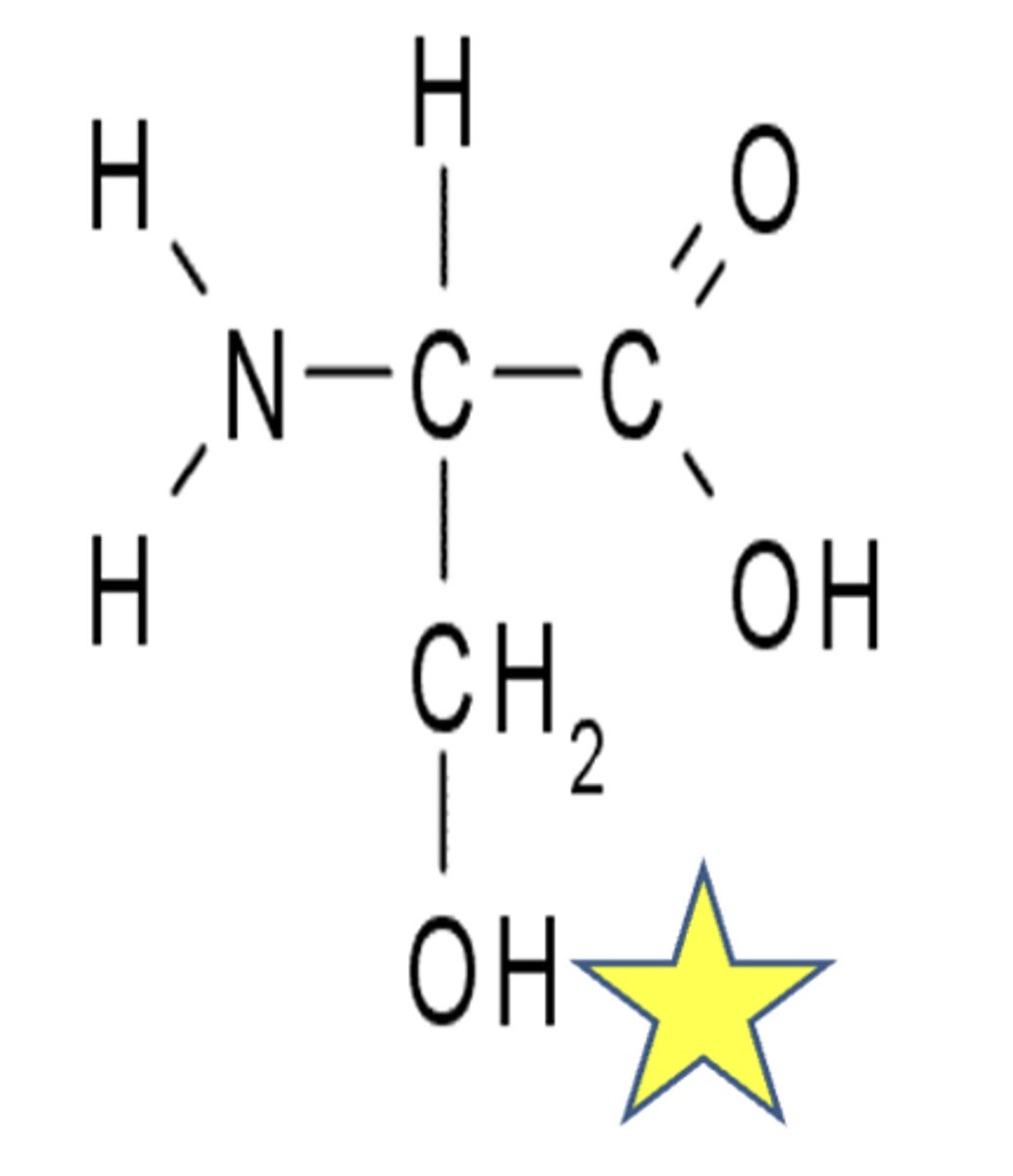
threonine structure
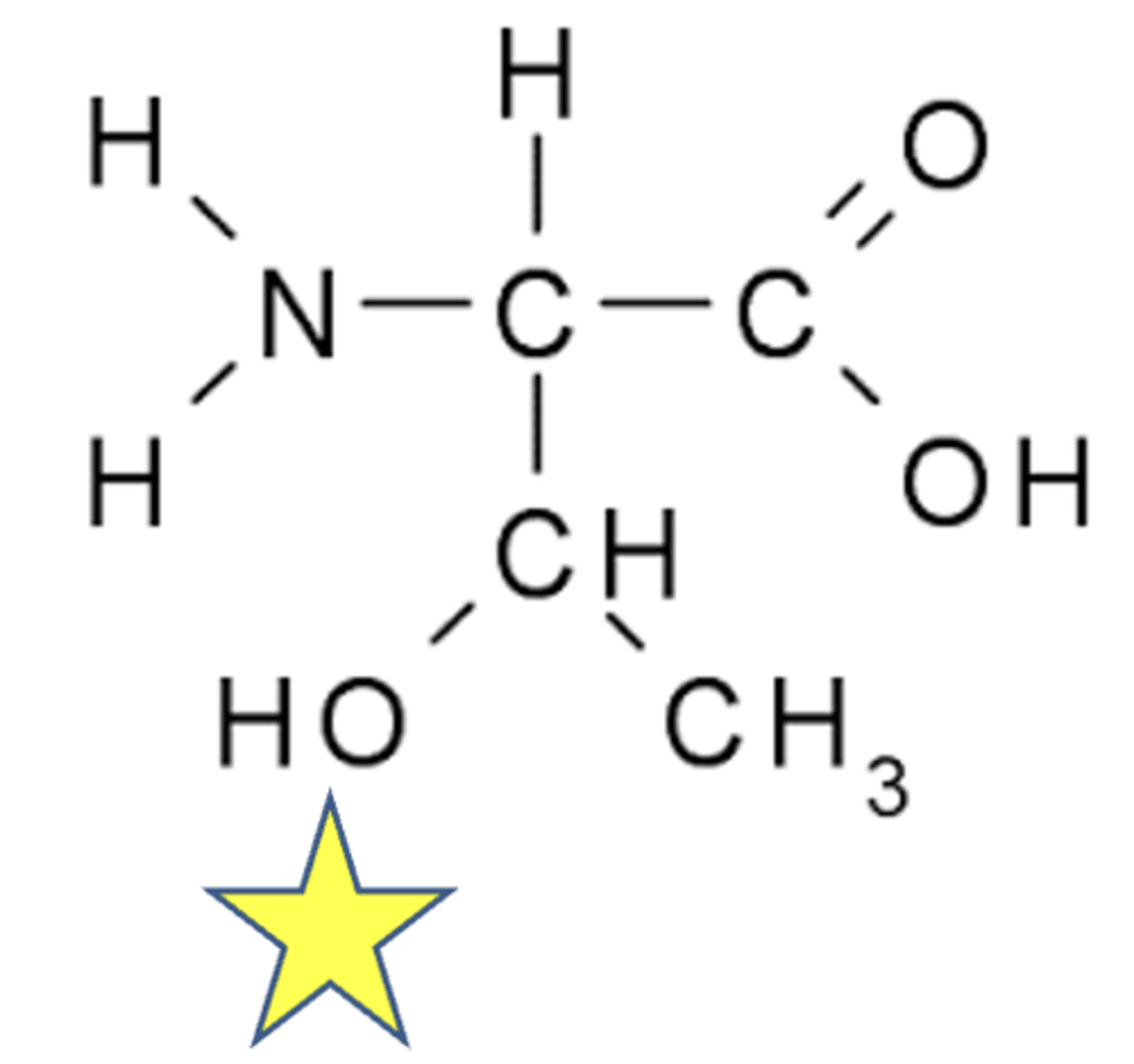
DNA damage: single base alteration (depurination)
It is a change in the DNA in which the purine (adenine or guanine) is removed from the deoxyribose sugar by hydrolysis of the b-N-glycosidic link between them.
- After depurination, an apurinic site is formed where the sugar phosphate backbone remains, and the sugar ring has a OH group in the place of purine.
- 5000 purines (a and g) can be lost per cell a day, but are efficiently repaired via BER pathway
single base alteration (depurination for cytosine and thymine)
Loss of pyrimidine base (cytosine and thymine) occurs by a similar mechanism, but at a substantially lower rate.
cytosine deamination
Deamination of amino group attached to C-4 in cytosine results in the formation of uracil
- Can spontaneously occur at rate of 100 bases per cell per day
- Uracil in DNA can not base pair with guanine, and this disrupts hydrogen bonding
- Cells can correct this spontaneous deamination by the removal of uracil and repair by Base Excision Repair mechanisms.
adenine deamination
Results in the formation of hypoxanthine
- Hypoxanthine can base pair with cytosine instead of thymine
- This results in a mutation where the original A-T base pair transforms into a G-C base pair.
oxidatatively damaged bases
Reactive oxygen species (superoxide ion O2, hydroxyl radicals and H2O2) generated in cellular metabolism (normal products), can cause single base alteration.
alkylation of base
- can cause apurnation (affects guanine)
- Occurs upon exposure to an alkylating agent. E.g., anticancer agents [nitrogen mustard (cyclophosphamide) used to treat lymphomas]
- They add ethyl or methyl groups to DNA. They are alkylating agents.
- Results in miss-pairing, strand breakage or depurination. Most susceptible site in DNA is N-7 of guanine which renders glycosidic bond susceptible to hydrolysis and loss of the base.
bifunctional alkylating agents
Alkylation of two bases by one alkylating agent (e.g., busulfan, anticancer drug used in CML=chronic myeloid leukemia).
- Results in interstrand or intrastrand cross-linkages.
UV induced DNA damage
UVA and UVB radiation that reaches the Earth’s surface contribute to the serious health effects
UV light induced dimers
UV induces covalent linkage between C-C of adjacent pyrimidine bases.
- T-T dimers are formed when cells are exposed to UV rays (as in sunlight) (when there are adjacent thymine bases); they are the most common type. They are the result of photochemical reactions.
- They interfere with base pairing during DNA replication.
Uv rays from the sun can cause melanoma
80,000 T-T dimers per cell within one hour of sunlight exposure.
Asymmetry
Borders (irregular)
Color (variegated)
Diameter (greater than 6 mm), about the size of a pencil eraser)
Evolving over time
miss match repair
Repairs miss-paired bases (well studied in bacteria)
base excision repair
Repairs Non-Helix distorting damage
E.g., Repairing Deaminated base or
Depurinated base.
nucleotide excision repair
Repairs bulky helix distorting damage.
E.g., repairing the T-T dimers.
In bacteria immediately after the replication, the parental strand is methylated whereas the daughter strand is not.
Methylation does not occur until sometime after replication is completed.
- Specific proteins (Mut proteins; Mut L, Mut S and Mut H) scan the newly synthesized DNA strand (not methylated strand) looking for mispaired bases`
- The recognition of the new strand is based on methylation status.
endonuclease
cuts the new strand at a site near the methylation point.
exonuclease
removes the portion of the DNA containing the defect
DNA polymerase
adds the correct nucleotides and DNA ligase seals the nicks in the back-bone.
base excision pair
Repairs damaged DNA throughout the cell cycle.
- It is responsible primarily for removing small, non-helix distorting damage (e.g., Deamination of Cytosine to uracil).
DNA glycosides (BER)
DNA glycosylases recognize the modified bases in the DNA
- Each DNA glycosylase is specific for an abnormal base. For example, in the diagram at right, cytosine is deaminated to give uracil.
- uracil DNA glycosylase recognizes the abnormal base and removes it.
gene expression
Conversion of the information in a gene into a protein product
The edge of each base pair is exposed at the surface of the double helix, presenting a distinctive pattern of
- hydrogen bond donors
- hydrogen bond acceptors
- hydrophobic patches
- for proteins to recognize in both the major and minor groove.
In most cases, the protein makes a series of contacts with the DNA, involving
- hydrogen bonds
- ionic bonds,
- hydrophobic interactions
gene control region
The entire section of the DNA involved in regulating and initiating gene transcription
- This includes promoter as well as regulatory sequences.
In animals and plants, it is not unusual to have the gene regulatory region spread over
50,000 base pairs of DNA, much of these DNA serve as spacers
- In humans, average gene length is 3000 base pairs.
It is the gene regulatory proteins that enable the genes of an organism to be turned on or off.
enhancers
DNA sequences that increase transcription.
repressors
proteins that prevent transcription.
translational control
Mechanism 1: Specific proteins interfere with binding to ribosomes.
Mechanism 2: Regulating the activity of translation factors via phosphorylation.
Eukaryotic cells decrease their overall rate of protein synthesis in response to a variety of stressful situations, including
- deprivation of growth factors
- nutrients
- infection by viruses,
- sudden increase in temperature.
This decrease is caused by phosphorylation of the translation eukaryotic initiation factor (eIF2) by specific protein kinases.
Suppression of Gene Expression
Through RNA Interference (RNAi)
Recently discovered pathways (1998) that cause inhibition of translation and degradation of mRNAs
Small Interfering RNAs (siRNAs)
very similar to miRNAs in terms of functions. However, the way they are formed are different
miRNA is coded by the cellular genome.
siRNA are synthesized and introduced by transfection
or virally coded RNAs introduced into the cells by viral infection.
The RNAi technique
introduces into a cell or organism a double stranded RNA (dsRNA) molecule whose nucleotide sequence matches that of part of the gene to be inactivated.
The ds RNA is processed by
Dicer enzyme ultimately to a 21 nts long double stranded RNA.
The Argonaute associates with ds siRNA and converts it to
single stranded siRNA and forms the RISC.
RISC directs the siRNA to hybridize with
mRNAs and causes its degradation
external factors of cancer
tobacco, infection (HPV), chemicals and radiation.
sunlight, diet, alcohol, carcinogens…..
internal factors of cancer
inherited mutations (e.g., BRCA1 and BRCA2), hormones, immune conditions and mutations that occur from metabolism, obesity, immunosuppression, chronic inflammation, age…..
Cancers are Genetic Diseases
Loss of Control over Cell Cycle!
It is a collection of diseases that arise from the loss of control over cell division.
Because cell cycle is controlled by proteins, and proteins are encoded by genes, cancers are genetic diseases.
In cancer, cells proliferate abnormally and invade tissue locally or at locations distant from their site of origin.
Cancers arise due to mutations in
Proto-oncogenes and Tumor Suppressor genes
EGF-R, HER2, Src kinase (Proto-oncogenes)
P53, Rb, BRCA-1 and BRCA-2 (Tumor
suppressor genes)
CYCLINS and
CYCLIN DEPENDENT
KINASES
Regulate Cell Cycle
tumor / neoplasm
An abnormal cell that grows and proliferates out of control
benign
neoplastic cells do not become invasive, the tumor
cancerous tumor
its cells have acquired the ability to invade surrounding tissue.
two heritable properties of cancer cells
reproduce in defiance of the normal restraints on cell growth and division
They invade and colonize territories normally reserved for other cells
adenoma
benign epithelial tumor with a glandular organization
adenocarcinoma
corresponding type of malignant tumor
- vast majority of the colorectal cancer are this type because colon has numerous glands within its tissue.
A benign glandular tumor (an adenoma) remains inside
basal lamina
- marks the boundary of the normal structure whereas a malignant glandular tumor destroys duct integrity
cancer incidences
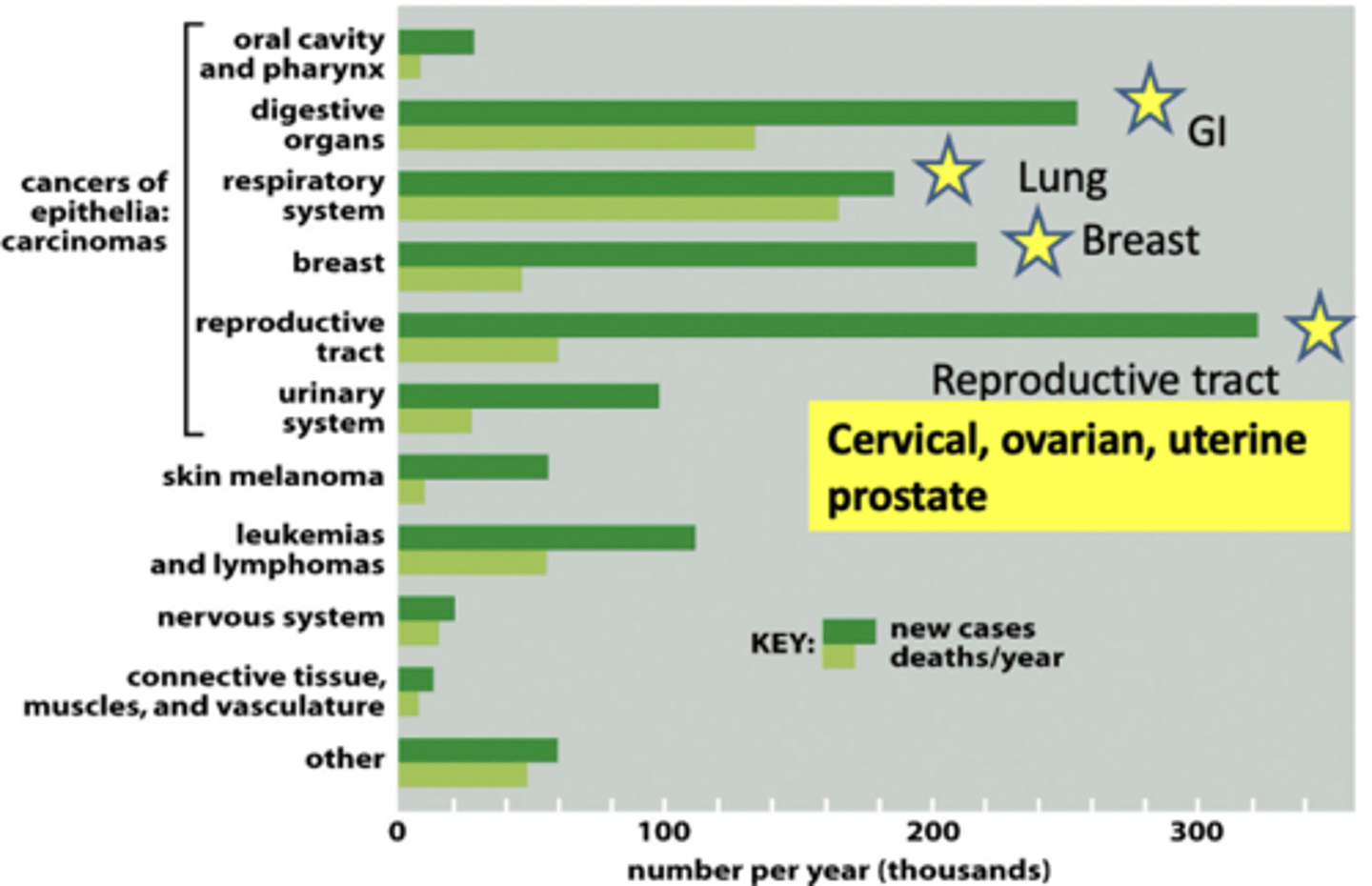
proto oncogene
a gene that is present in the cell that when mutated or expressed at very high levels can transform a normal cell to a cancer cell
c-Src (cellular Src in contrast to the viral Src); type of protooncogene
Regulated expression of proto-oncogenes is important for normal cellular functions such as growth and differentiation.
When c-Src is mutated,
its expression is abnormal (very high) and it can cause cancer.
Mutated c-Src is an oncogene as it has the potential to cause cancer.
examples of protooncogenes
Human Epidermal Growth Factor Receptor 1 (HER1) and 2 (HER2)-these are Receptor tyrosine kinases.
B-RAF- a cytoplasmic serine/threonine kinase.
C-Src- a cytoplasmic tyrosine kinase.
c-Myc- a transcription factor
In normal epithelial cells, expression of -------- is tightly regulated
EGF-R (epidermal growth factor receptor)
In normal cells, when the EGF receptor is not phosphorylated, it is
inactive
In normal cells, binding of the EGF to EGFR causes
dimerization of the receptor and tyrosine phosphorylation of several residues
- This activates signal transduction and causes cell proliferation.
- Receptor dephosphorylation and or internalization turns off signal transduction pathways.
reason proto ongoene becomes an oncogene (point mutation / deletion)
translates into a changed amino acid that has different biochemical properties than the amino acid the DNA sequence originally encoded for
This change in amino acid can mimic phosphorylation, or inhibit binding of negative regulatory molecules, resulting in constitutively active protein.
reason proto ongoene becomes an oncogene (gene aplification)
normal protein produced in much higher amounts
reason proto ongoene becomes an oncogene (chromosome rearrangement
- placement of strong enhancer causes overproduction of normal protein
- fusion to another actively transcribed gene results in either increased levels of fusion or fusion protein is hyperactive
How Point Mutations or Deletions Can Cause Cancer
There are 4 members in Epidermal Growth Factor Receptor (EGFR) family. They are HER1, HER2, HER3 and HER4.
- They are receptor tyrosine kinases localized to the plasma membrane. They are important for cell growth, survival, adhesion, migration and differentiation.
- Mutations and deletions in HER1 can lead to colon and lung cancer
HER-1
Mutations or NH2 terminal deletion in HER1 gene could lead to its constitutive activation and results in cancer. Observed in Lung and colon cancers.
- Genzyme and Qiagen make kits to detect EGFR mutations in cancer tissue.
drugs that target HER-1 (EGFR1)
- Monoclonal antibodies (Intravenously given):
- Cetuximab (Erbitux) (Imclone Systems)
- Panitumumab (Amgen)
- Matuzumab (Merck Serono in collaboration with Takeda pharmaceuticals)
Tyrosine kinase Inhibitors
- Gefitinib also known as Iressa -1st selective tyrosine kinase inhibitor, marketed by AstraZeneka.
- Erlotinib (Genentech, OSI pharmaceuticals, Roche)
MOA of cetiximab
It is a human monoclonal antibody.
Used to treat metastatic colorectal cancer and head and neck cancers.
Binds to the extracellular domain of the EGFR and inhibits signaling.
MMOA of gifitinib
It is the first selective inhibitor that targets mutant EGFR
- Binds to the ATP binding pocket in EGFR1 cytoplasmic domain.
Used to treat lung cancers.
HER-2
It is a member of Epidermal Growth Factor Receptor (EGFR) family.
Approximately 12-40% of breast cancers have an amplification of the HER2 gene.
This leads to overstimulation of the receptor and abnormal cell proliferation.
and colon cancer
interactions of Herceptin with HER2
Binding to HER2 interferes with the dimerization of the receptor.
Receptors are down- regulated and disappear from the membrane.
herceptin
monoclonal antibody
- It was the 1st pharmacogenomics based targeted treatment for breast cancer.
- It interferes with HER2 receptor and blocks cell signaling
- Monoclonal antibodies (written as mAb) are identical antibodies produced by one type of B cells and are obtained from clones of a single cell
chronic myelogenous leukemia
hematologic cancer resulting from malignant transformation of pluripotent stem cells
- A specific chromosomal abnormality called Philadelphia chromosome (Ph) is observed in ~95 % of patients
philidelphia chromosome
- This chromosomal abnormality results from the translocation of parts of chromosome 9 and 22
A proto-oncogene called Abelson or c-abl located on chromosome 9 is translocated to chromosome 22 near a gene called Bcr (break point cluster region protein), and a part of chromosome 22 breaks off and moves to chromosome 9.
Abl is a tyrosine kinase that requires cytokine to be activated. In its monomeric form, Abl is inactive. When cytokines are present, Abl proteins are dimerized, phosphorylated and activated.
treatment of CML
Gleevec (Imatinib mesylate)
- It is a p210 tyrosine kinase (BCR/Abl) inhibitor.
- It inhibits proliferation and induces apoptosis in bcr-abl positive cells.
Dasatinib- from Bristol- Meyers Squibb
- for treatment of adults with ALL (Acute Lymphoblastic Leukemia) and it is also used to treat CML patients. (Dasatinib is a Abl-Bcr kinase inhibitor, also inhibits Src kinase).
tumor supressor genes
inhibit multiple steps in signal transduction pathways or modulate transcription; thus, contribute to inhibition of cell proliferation.
They regulate and inhibit inappropriate cell growth and cell proliferation.
examples of tumor supressor proteins
are RB-1 (retinoblastoma gene) and p53.
RB-1 MOA
Progression from G1 to S requires E2F transcription factors. RB-1 can bind to E2F and inhibit its activity.
RB-1 Retinoblastoma Tumor Suppressor Gene
is inactivated by
HPV encoded protein
belongs to the pocket protein family, whose members have a pocket for the functional binding of other proteins
E6 protein can inactivate another tumor suppressor protein p53 and can lead to
cervical cancers
- Vaccine called Gardasil (Merck) and Cervarix (Glaxo). The vaccine contains virus like particle but no genome, so it is non-infectious
- Vaccination can prevent cervical HPV-induced cancers
Main Functions of p53the Guardian of the Genome
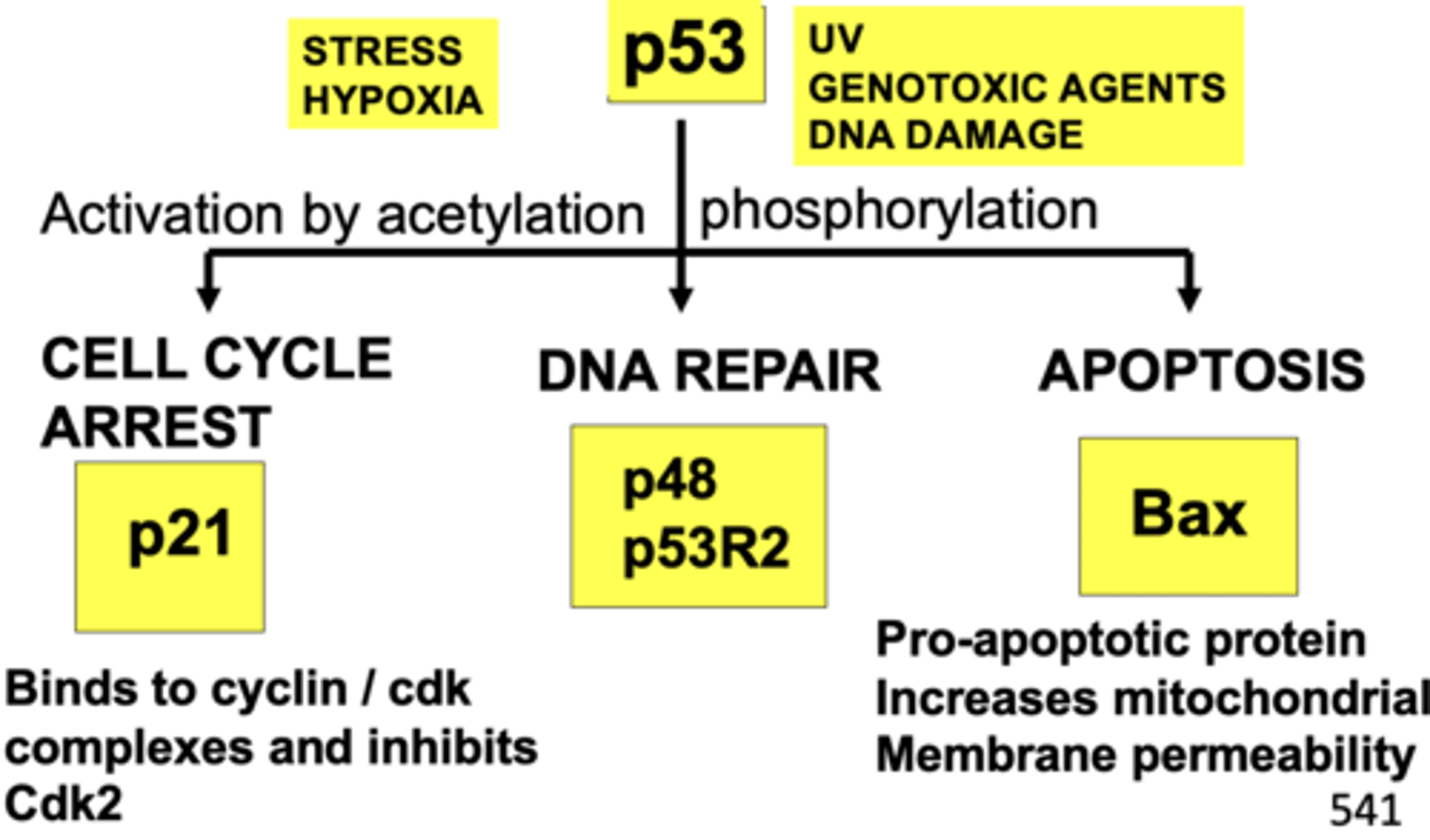
apoptosis
programmed cell death
- In multicellular organisms, cells that are no longer needed or are a threat to the organism are destroyed by a tightly regulated cell suicide process known as programmed cell death
- This happens naturally during embryonic development
also occurs in cells that contain extensive DNA damage. So, it is a normal process of eliminating the unwanted cells
Many anti-cancer drugs kill cancer cells by causing
DNA damage and through induction of apoptosis
- Most of the cytotoxic anticancer drugs in current use have been shown to induce apoptosis in susceptible cells
- Topoisomerase I and II inhibitors, alkylating agents etc.,
Bax
pro-apoptotic
Increases the permeability
Bcl2
anti-apoptotic
Decreases the permeability
In normal mitochondria Bcl2 binds to
bax
- Quenches the action of Bax
In DNA damaged cells p53 accumulates in the
cytoplasm
- it interacts with Bcl2 and activates Bax
apoptotic caspade cascade
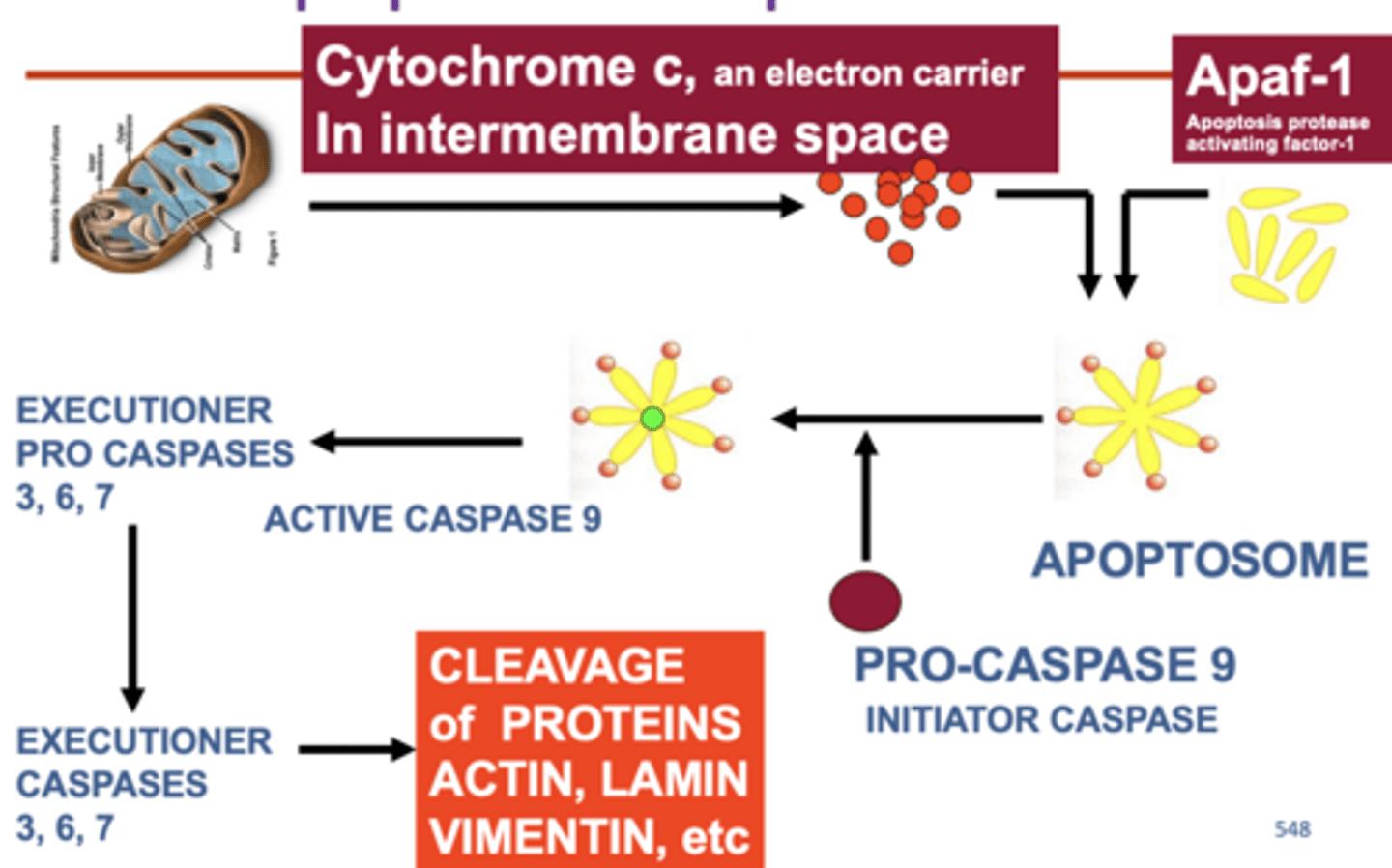
p53 functions as a
tetrameter
People who inherit only one functional copy of the P53 gene will most likely develop
tumors in early adulthood
present in 15 of the 16 tetramers. Mutant p53 exerts dominant negative effect. This may cause loss of p53 function.
BRCA 1 and 2
Breast Cancer type 1 and 2 Susceptibility Proteins
- belongs to the tumor suppressor gene family
- involved in the repair of DNA double strand breaks
where does BRCA 1 and 2 bind to
a protein called RAD51 to repair the breaks in the DNA
These breaks occur during:
Naturally during:
- chromatid exchange
- Medical radiation
- Environmental exposure
BRCA1 and BRCA2 helps to repair double stranded DNA breaks during
S and G2 phase of cell cycle.
In cancer, protooncogenes are
activated
- EGFR2 (HER2), c-Src, c-Myc, B-Raf
in cancer, tumor suppressor genes are
deactivated
- P53, RB-1, BRCA1 and 2