Prokaryotic Transcriptrion FINAL
1/254
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
255 Terms
A gene is not just the protein-coding sequence- what two major parts does it include?
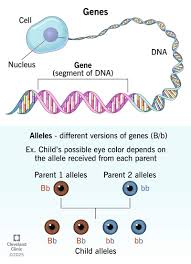
coding region
regulatory elements
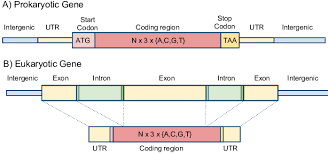
what is the coding region in a gene
the actual DNA sequence that will determine the amino acid sequence ( primary sequence) of a protein
what are some examples of regulatory elements within a gene
promoter
enhancers/silencers
transcription factor binding sites
A gene is used to create an RNA primary transcript. What is the mechanism for this?
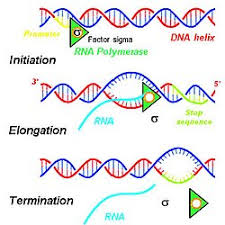
RNA polymerase binds the promoter
DNA strands separate
One strand (template strand) is used to build RNA
RNA is synthesized 5’ → 3’
Yielding a primary transcript
what is the primary transcript
the inital RNA copy
When a primary transcript is made for bacteria, the RNA that is first made is already usable by ribosomes, therefore there is no editing phase, unlike in eukaryotes that have pre-mRNA that needs to be edited. Why?
because they dont have a nucleus- so transcription and translation happen in the same space ( the cytoplasm) - so as soon as RNA is made ribosomes jump on it therefore making the primary transcript the functional mRNA
In eukaryotes primary transcript is NOT immediately translation ready - why is this?
because transcription happens in the nucleus, but translation happens in the cytoplasm
In eukaryotes primary transcript is NOT translation ready because transcription happens in the nucleus, but translation happens in the cytoplasm.
Therefiore the mRNA has to pass through the nuclear membrane via nuclear pores
What is the critical issue with this?
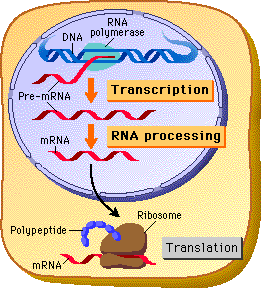
The nucleus does NOT allow incomplete or faulty RNA to leave. Therefore the processing steps are like a checklist to earn permission to exit the nucleus
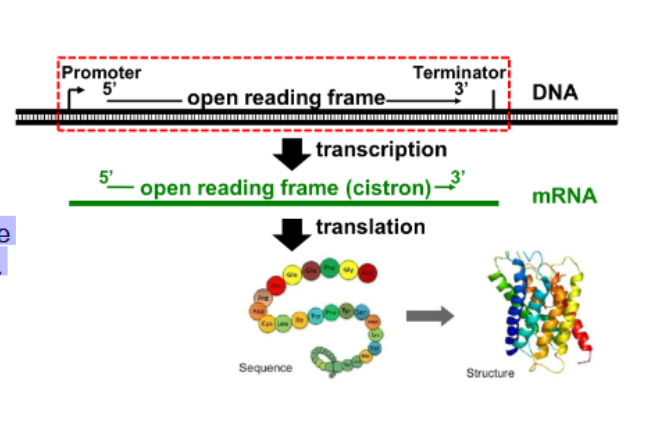
what is a continous sequence within the coding region of a gene from start codon to stop codon that is a potential protein coding stretch that has a start codon , ends at a stop codon , and has no stop codons in between
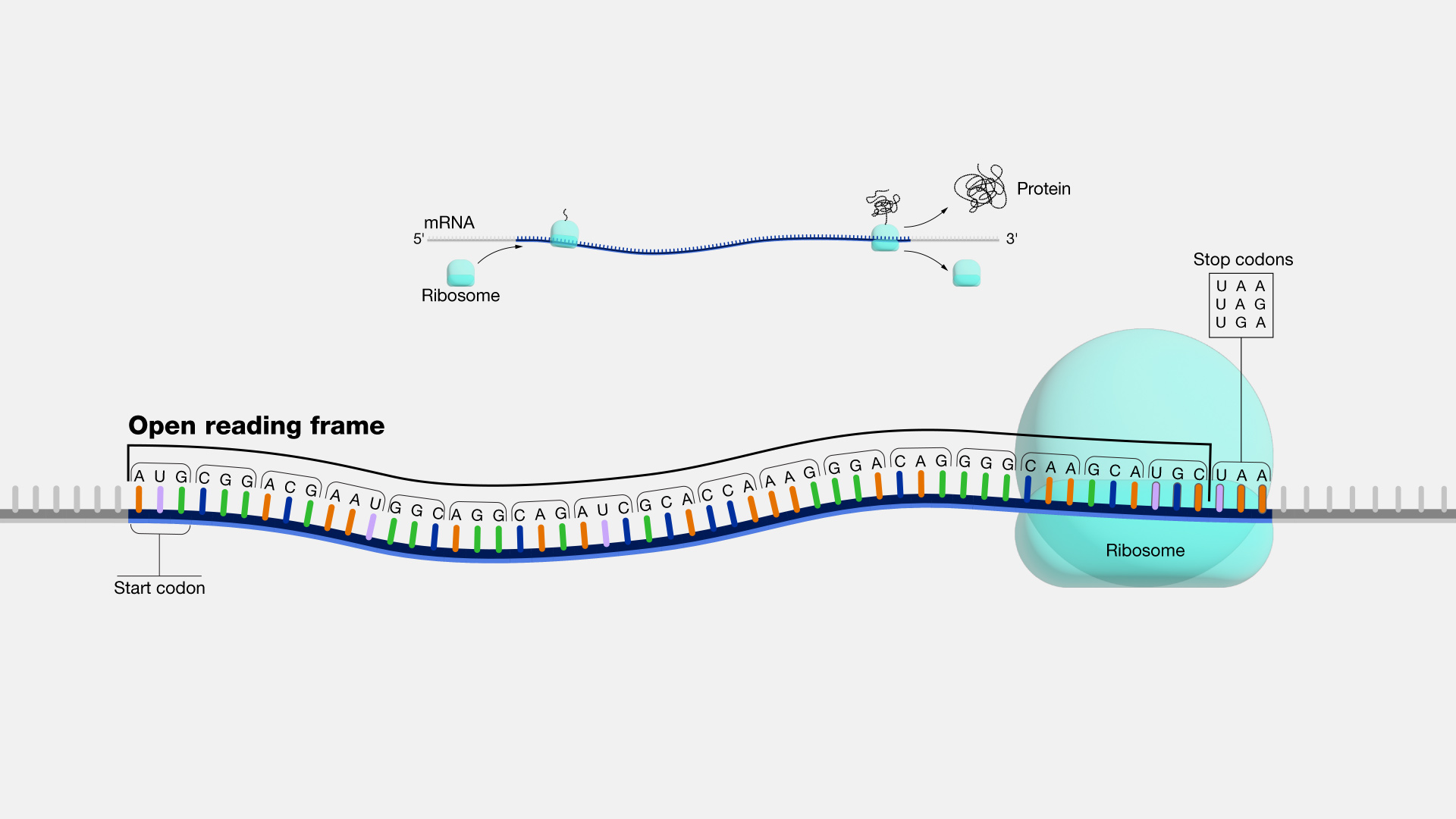
open reading frame
what is a cistron
a gene that encodes for one protein
where are cistrons found?
in open reading frames which are found inside of the coding region of the gene
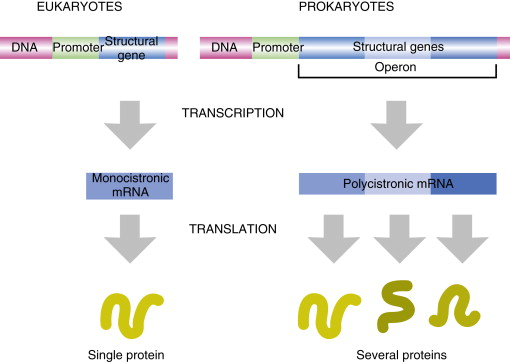
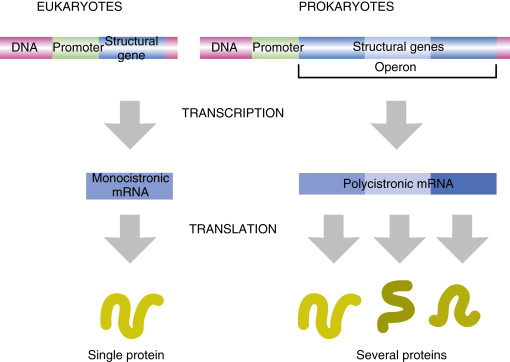
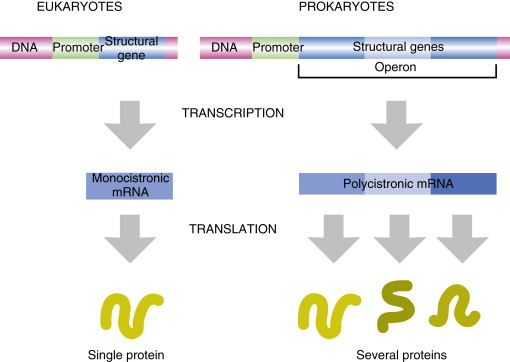
In bacteria, one mRNA can contain multiple cistrons. What does this mean?
one primary transcript yields multiple proteins
In bacteria, one mRNA can contain multiple cistrons therefore making it polycistronic. Why?
because genes involved in the same pathway are grouped together in operons
In eukaryotes one mRNA encodes for only one protein, what does that make it’s mRNA
monocistonic
Mainly in eukaryotes one mRNA yields one protein-making it monocistronic. Why is this and what does it allow?
because each gene has its own promoter and is transcribed independently therefore allowing much more precise regulation
True or False coding strand sequence is the actual blueprint for amino acids to make proteins
true
“ALL CDS (coding sequences) are ORFs but not all ORFs are CDS”
How are all CDSs ORFs
they have:
start codon
stop codon
and are continuous, fitting the definition of an ORF
“ALL CDS are ORFs but not all ORFs are CDS”
How are all ORFs not CDSs
the genome has many random ORFs but most are not actually used becasue they have no promoter, not transcribed, not translated, or have regulatory signlas missing
therefore ORFs have potential BUT CDS are confirmed, functional protein coding regions
What the big picture connection for genes?
A gene contains coding + regulatory info
It is transcribed → primary RNA
In:
Bacteria → ready immediately
Eukaryotes → must be processed
The RNA contains:
ORFs (possible proteins)
CDS (actual protein-coding regions)
Depending on organism:
Prokaryotes → multiple proteins per mRNA (polycistronic)
Eukaryotes → one protein per mRNA (monocistronic)
what is a group of genes controlled together with the key idea of multiple genes being on one contol system found in prokaryotes?
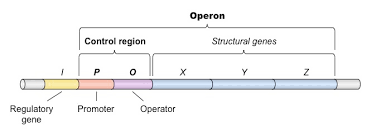
operons
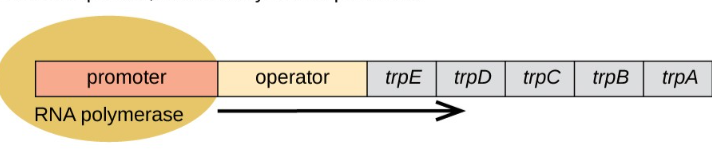
why are operons useful
If a cell needs a pathway (like making ATP), it needs all the proteins together, not one at a time therefore ensuring efficieny coordination
which direction are operons arranged in
5’ → 3’ all lined up one after another - being transcribed in one continuous pass
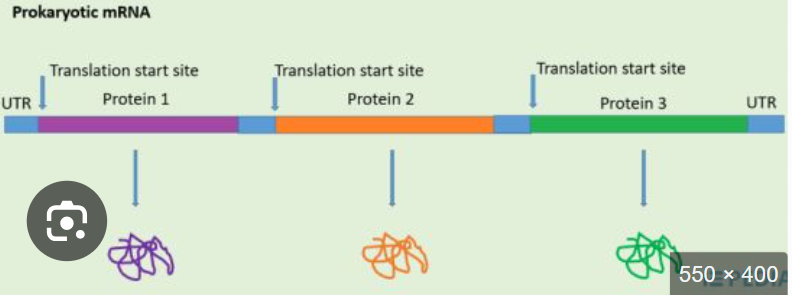
what would be the sequence of ORFs
Promoter → ORF1 → ORF2 → ORF3 → Terminator
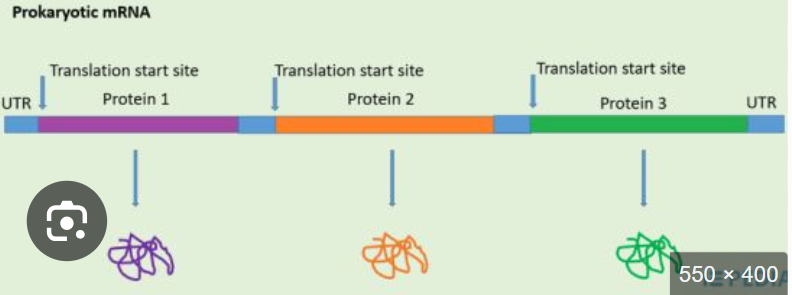
In an open reading frame, one promoter and one terminator yields a polycistronic RNA how does this happen?
Rna polymerase binds one promoter and transcribes straight through multiple ORFs, stopping at one terminator
even though it ones prokaryotic mRNA, it doesnt make one protein , what does it make
multiple indpendent translation units
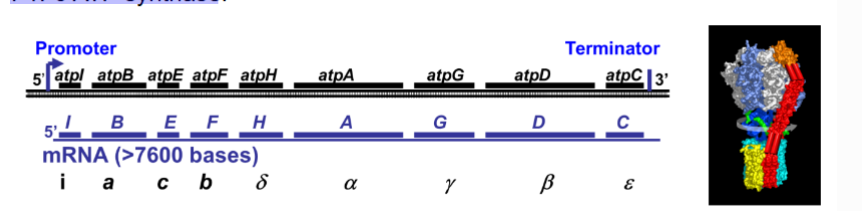
How is an atp operon in e.coli an example?
the atp operon contains 9 ORFS and encodes subunits of ATP synthase- they are all grouped becasued they all work together at the same time
Why do cis and trans-acting factors matter for transcription?
transcription only happens when cis elements are present- meaning there must be a promoter/operator DNA sequence and when trans factors bind those DNA sequences to activate or repress transcription
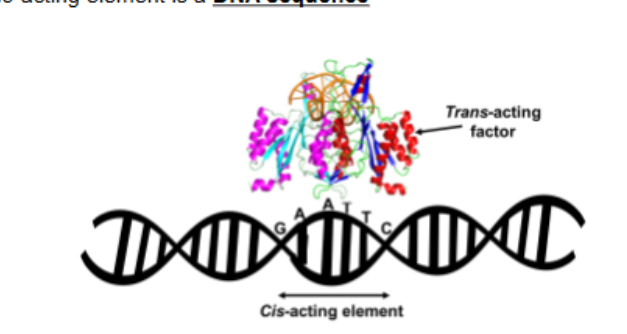
Transcription does not always require both cis elements and trans factors to start but rather that…
cis elements define where transcription can happen and trans factors decide whether it actually happens
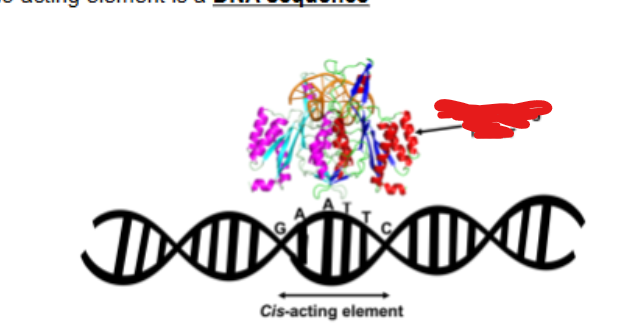
what are transacting factors usually
DNA- binding proteins
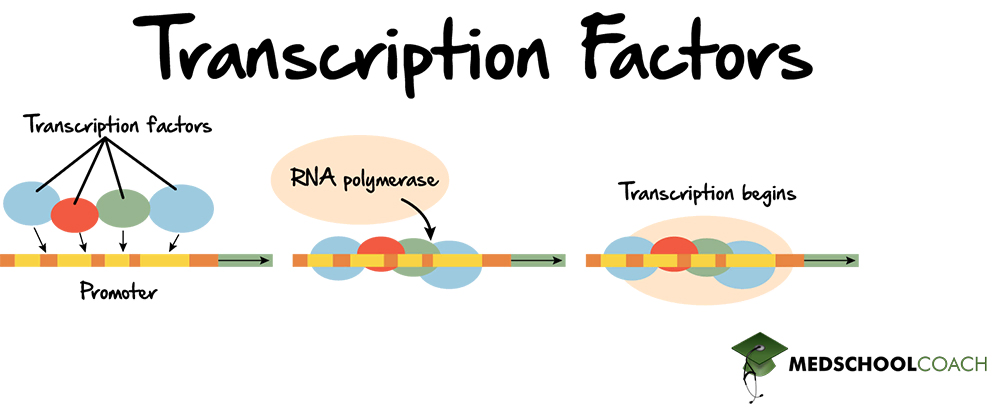
transacting factors are typically DNA binding proteins which are usually known as
transcription factors
the process of DNA template-dependent RNA synthesis and is catalyzed by RNA polymerase
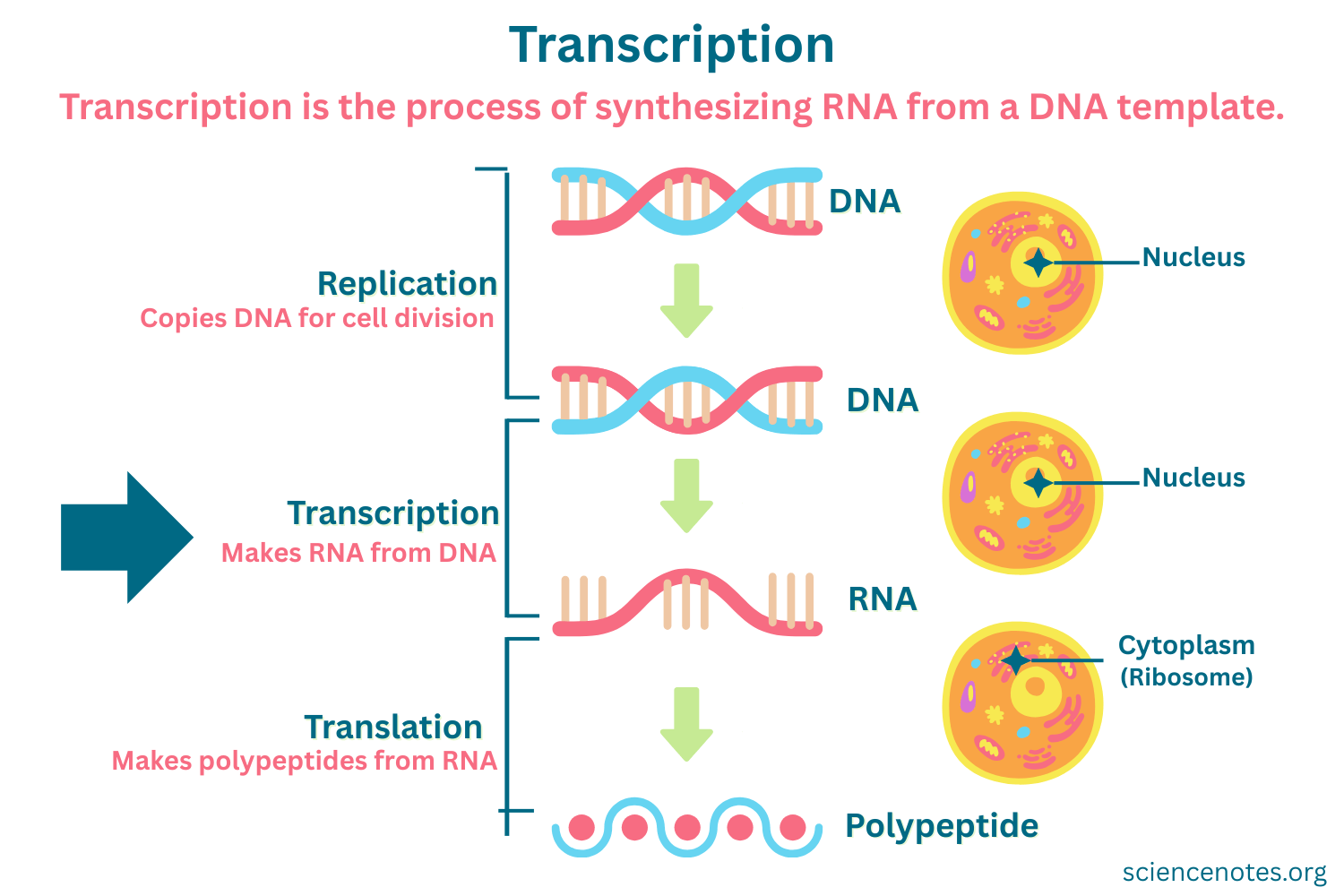
Transcription
While DNA replication replicates the entire chromosome/genome… what is the difference with RNA synthesis
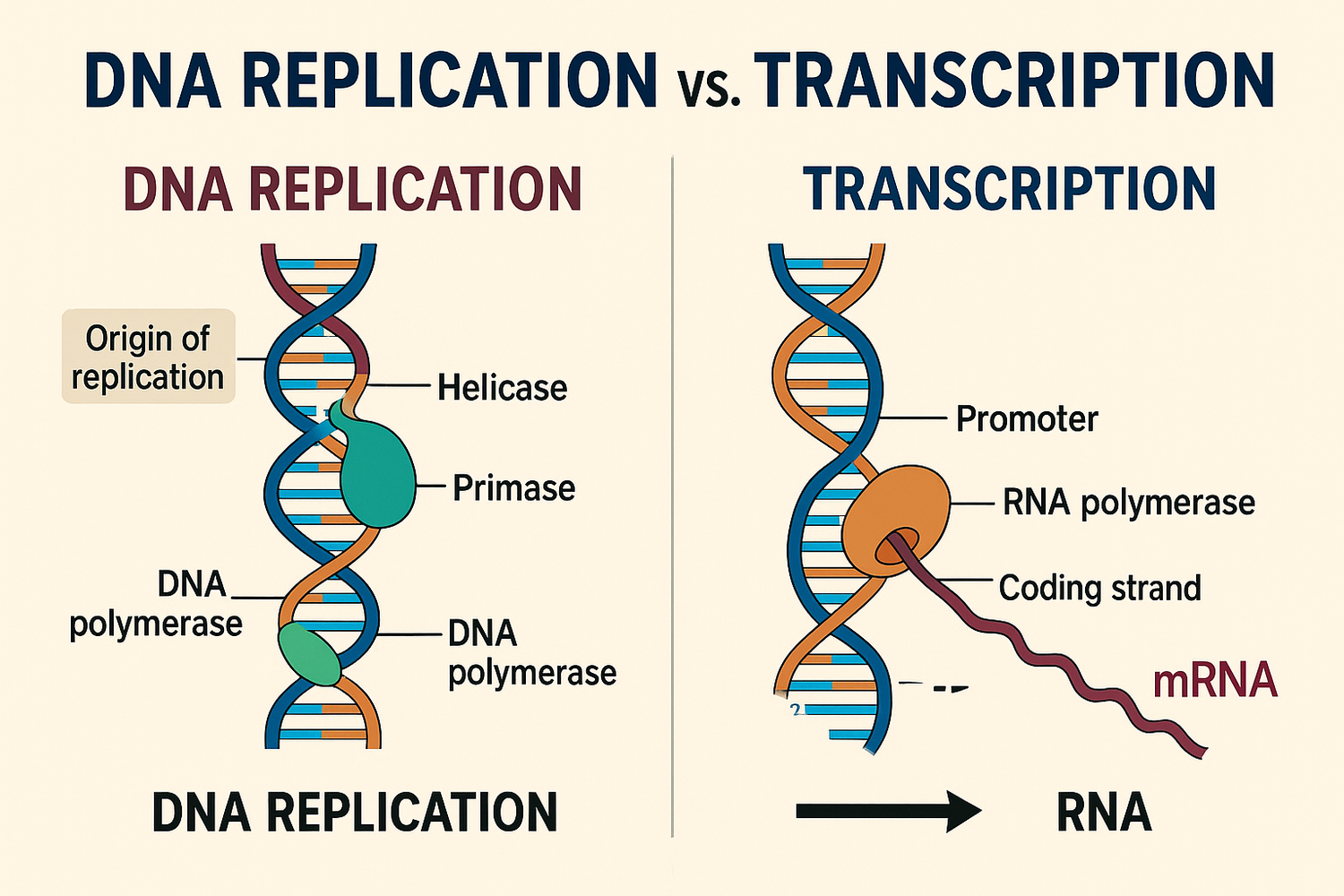
RNA synthesis much more selective
When does transcription start and end
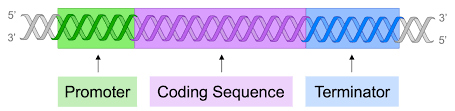
it starts at a promoter and ends at a terminator
what is the finished RNA molecule after transcription
the primary transcript
what is the transcription start site defined as
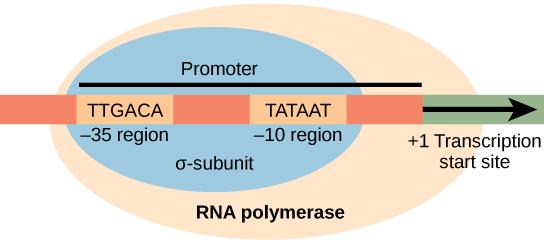
the exact first nucleotide where RNA synthesis begins
where is the transcription start site always located
+1
what is important to know about +1
it is NOT the promoter, it is the first base that gets copied into RNA
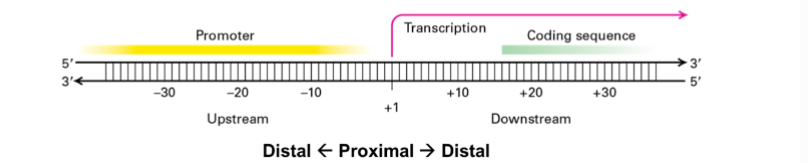
how is the start of transcription always at +1 if transcription starts at the promoter?
The promoter recruits RNA polymerase, but the +1 site is where RNA synthesis begins.
RNA polymerase binds the promoter, but transcription starts at +1.
There are directional terms based on transcription movement.
What is upstream?
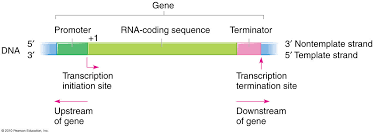
where polymerase came from
regulatory control region
usually noted by (-)
There are directional terms based on transcription movement.
What is downstream?
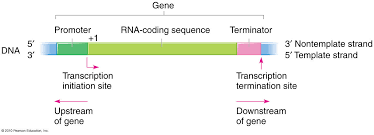
where polymerase is going
coding region of a gene
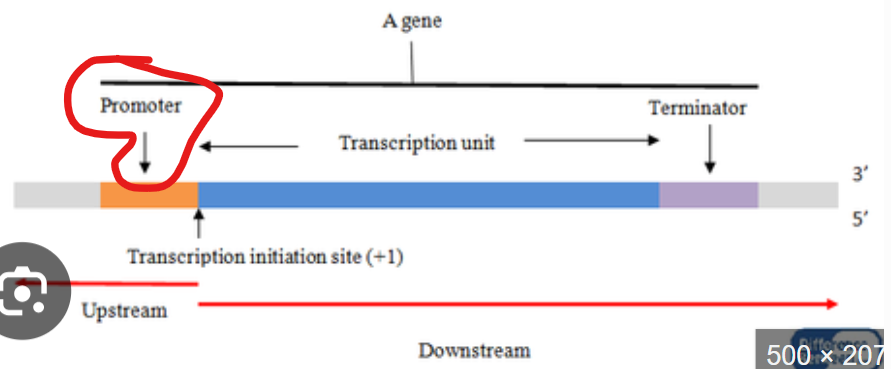
what does it mean when something is proximal?
close to the gene
often near promoter
what does it mean when something is distal?
Far away from gene
Can still regulate gene (especially in eukaryotes via enhancers)
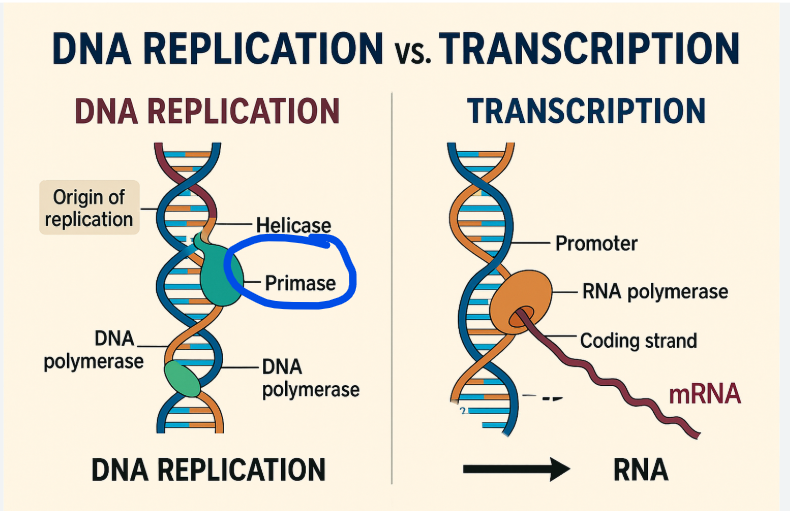
What is something RNA polymerase can do that DNA polymerase cannot. DNA polymerase needs this for replication and RNA polymerase does not
Rna polymerase can start synthesizing RNA de novo— without a primer
why does it matter that RNA polymerase can start synthesizing de novo
because it can start the first bond on its own and initiate RNA synthesis directly at +1 which is why transcription can begin immediately at a gene
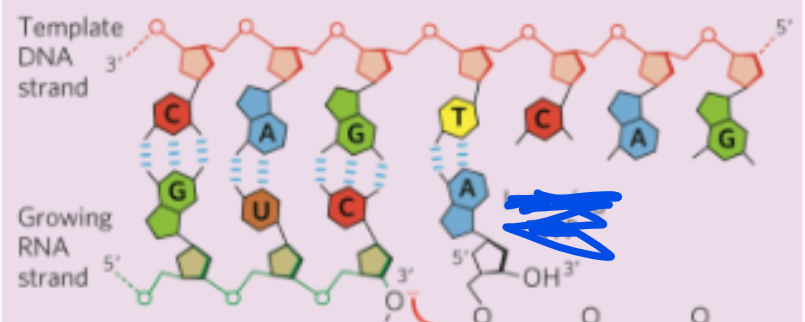
what are the substrates of RNA polymerase
NTPs (ribonucleoside triphosphates) such as ATP, GTP, CTP, UTP
the substrates of RNA polymerase are NTPs what does each one contain
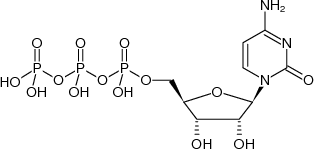
a sugar
a base
three phosphate groups
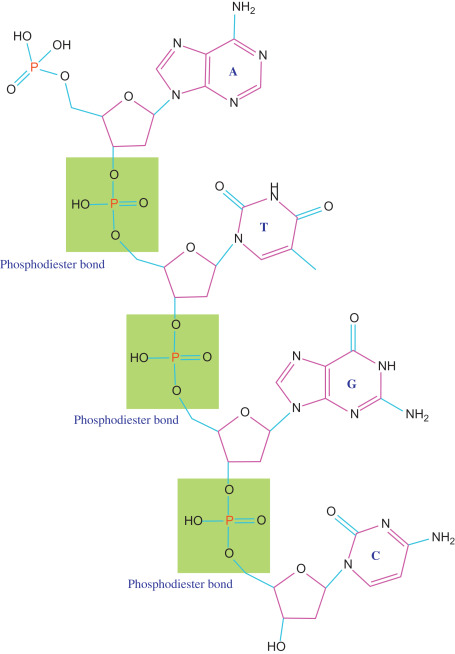
RNA polymerase links nucleotides together by forming what kind of bonds
phosphodiester bonds
How do the phosphodiester bonds form?
the 3’ OH of the growing RNA chain attacks the incoming NTP and two phosphates are released (pyrophosphate)
what type of base pairing does RNA polymerase use
watson-crick base pairing
what is the big difference between DNA polymerase and RNA polymerase
there is no proofreading in RNA polymerase
Since theres no proofreading in RNA polymerase what is the error ratew
much higher
what is the main flow of transcription
RNA polymerase binds promoter
↓
positions at +1
↓
uses DNA template strand
↓
selects NTPs via base pairing
↓
forms phosphodiester bonds
↓
RNA elongates 5’ → 3’
↓
no proofreading → occasional errors
What does the RNA transcript resemble
the coding strand
How does RNA temporraily base pair with DNA
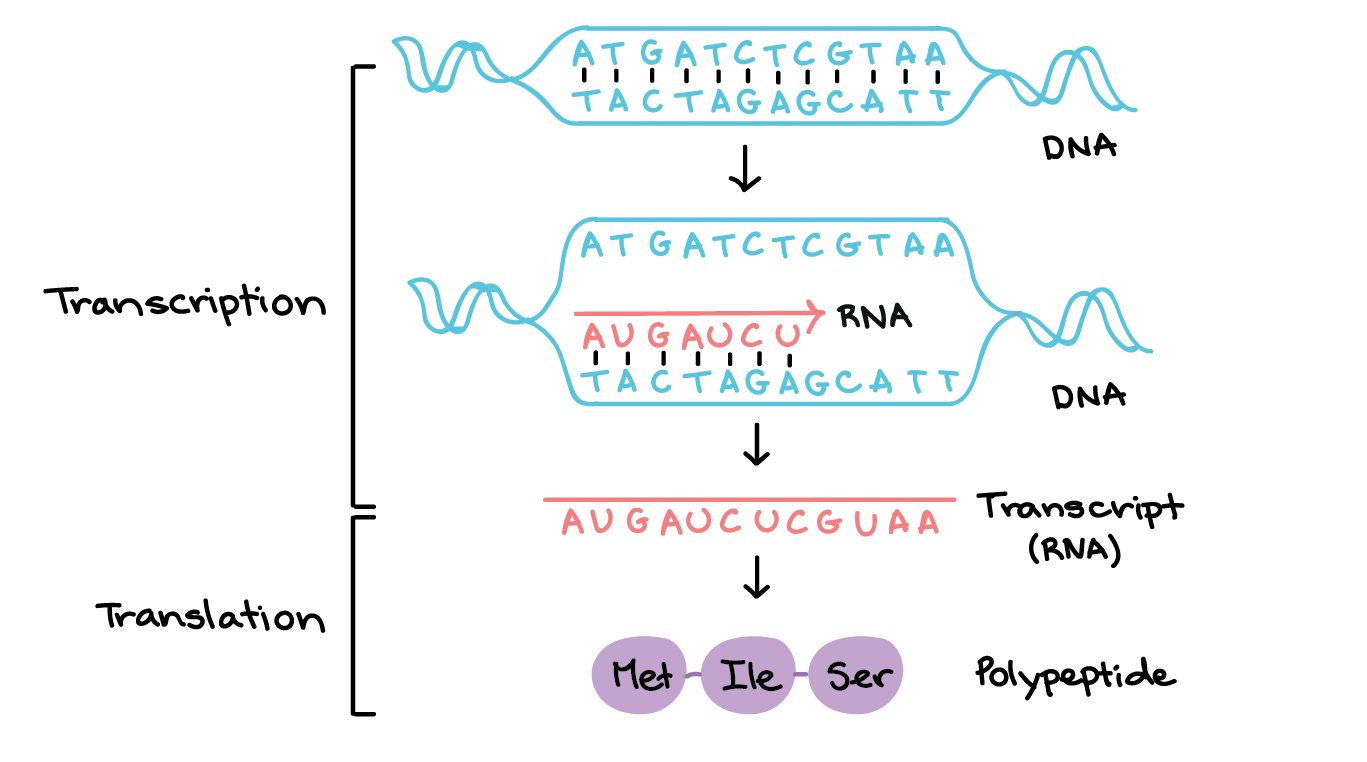
during transcription RNA polymerase opens a small region of DNA and the new RNA being made briefly pairs with the DNA template
During transcription:
RNA polymerase opens a small region of DNA
The new RNA being made briefly pairs with the DNA template
What does this create?
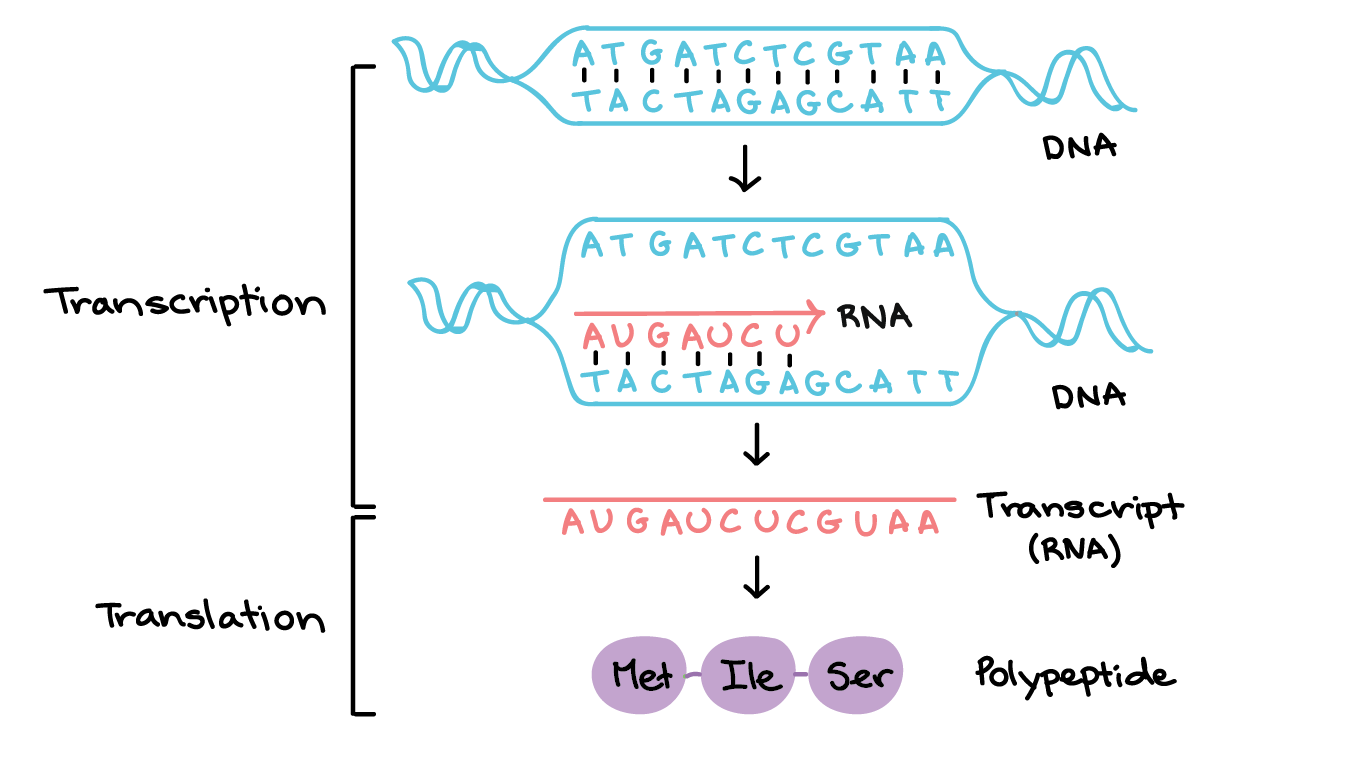
a temporary RNA-DNA hybrid helix until RNA peels away and the DNA re-closes
what is the template strand?
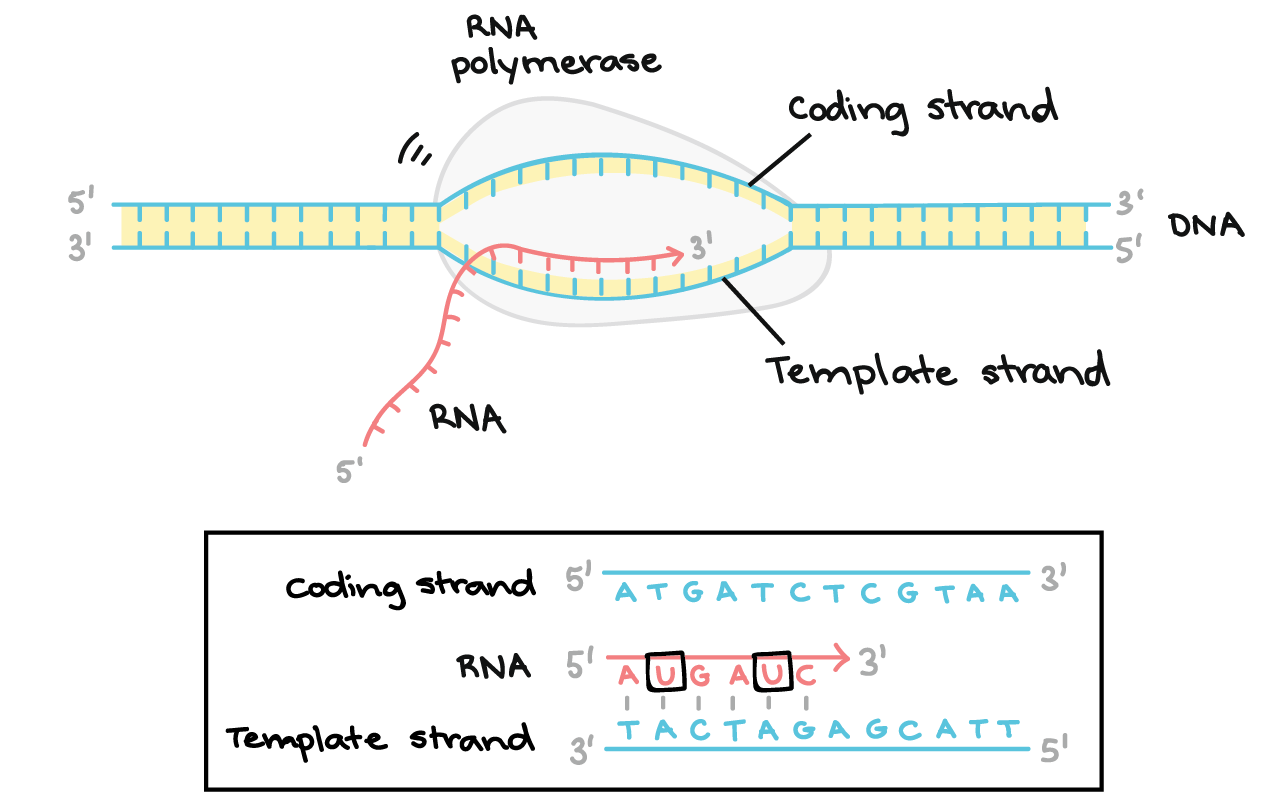
the dna strand RNA polymerase reads
what does RNA polymerase use the template strand for?
to decide which NTP to add

what is the noncoding strand?
it is not used for copying, but matches the RNA sequence
the nontemplate (noncoding) strand is NOT used for copying, but matches the RNA sequence - why does it match RNA?
because RNA is complementary to the template strand and the coding strand is already complementary to the template- the only difference is that RNA uses U instead of T
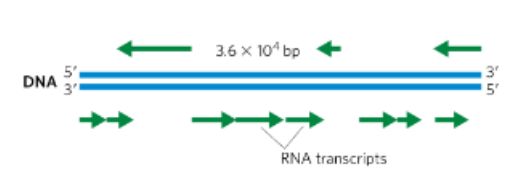
what does it mean when the coding strand can be in either DNA strand?
DNA is double stranded so for one gene the top strand might be coding and for another gene the bottom strand might be coding - therefore there is NO universal coding strand
what does rifampin target
it inhibits bacterial transcription by binding RNA polymerase
what does rifampin specifically bind to in RNA Polymerase
the β subunit which is essential for RNA synthesis
RNA polymerase binds the promoter, starts transcription at +1 and builds RNA from NTPs. This is essential for making mRNA, making proteins, and keeping the cell alive.
What does rifampin do to this process?
rifampin binds RNA polymerase before or during early transcription and blocks initiation- this prevents RNA polymerase from properly extending the RNA chain after starting

what is the result of rifampin
RNA cant be made and proteins cannot be made and bacteria die
what is the key idea behind rifampin
no transcription means no gene expression which means no survival of bacteria
how does rifampin work clinically
the bacteria relies heavily on transcription to survive in host cells and respond to stress—if you were to block (rifampin) transcription it would create a powerful antibiotic effect
How does resistance to rifampin happen
when theres a mutation in the rpoB gene
what is rpoB
a gene that encodes the β subunit of RNA polymerase
what happens when β subunit of RNA polymerase mutates?
The structure of the β subunit changes slightly and rifampin can no longer bind effectively allowing RNA polymerase to still work normally for bacteria
How does the use of Rifampin tie everything together?
RNA polymerase builds DNA from DNA so it need to initiate transcription at promoters. Rifampin blocks initiation at promoters. This shuts down RNA synthesis
Why doesnt rifamplin kill human cells
there are structural differences with bacterial rna polymerase and eukaryotic rna polymerase. Rifamplin fits bacterial enzymes and does NOT effectively bind human enzymes
Bacterial RnA has 2 modes- what are they
core enzyme
holoenzyme
what does the core enzyme do?
build RNA (elongation)
what does the holoenzyme do?
finds where to start (initiation)
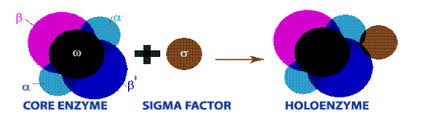
what makes up the holoenzyme
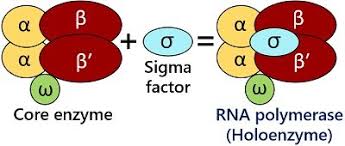
the core and the σ
Unlike eukaryotes (which have multiple RNA polymerases) for transcription, what does bacteria like e.coli use for transcription?
one enzyme for all transcription
Unlike eukaryotes (which have multiple RNA polymerases), bacteria like E. coli use one enzyme for all transcription. So what does it have to do?
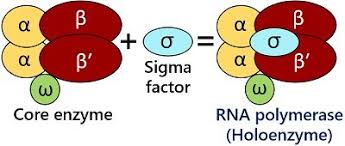
find promoters, start transciption, and elongate RNA- thats why it uses different subunits to specialize tasks
the core enzyme is the RNA building machine- what makes up the core enzyme
α₂ β β′ ω
what does the core enzme do specifically?
Elongation
what does elongation do?
add nucleotides to RNA
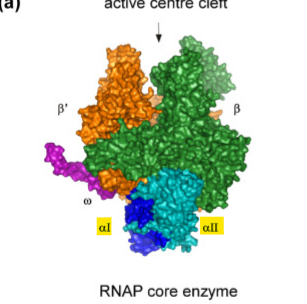
what do the 2 copies of α do as the core enzyme subunit?
they help assemble the enzyme and interact with activator proteins for regulation
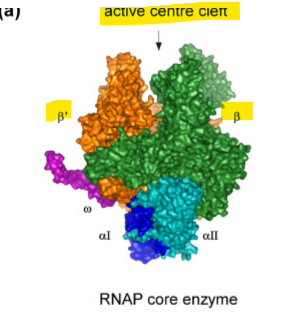
what does the β and β′ (beta and beta prime) do as the subunit in the core enzyme?
form the catalytic center and actaully link NTPs together- forming the phosphodiester bonds and growing the RNA chain
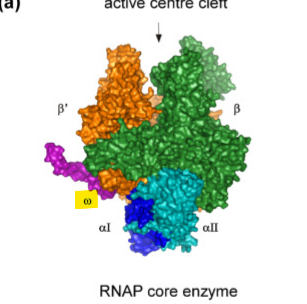
what does ω do in the RNA subunit?
stablizes the structure
what is the key idea for the core enzyme
it can MAKE RNA, but it doesnt know where to start
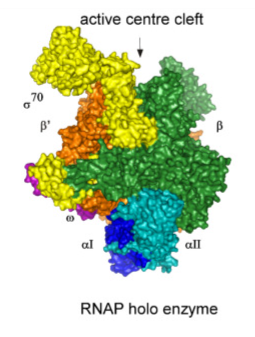
what does the holoenzyme do?
it starts the transcription correctly
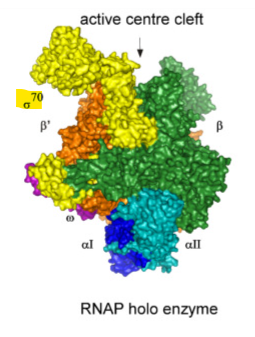
what does the σ do in the holoenzyme
σ recognizes promoters
Why is it so important that σ recognizes promoters
it tells RNA polymerase where a gene begins and where to bind DNA
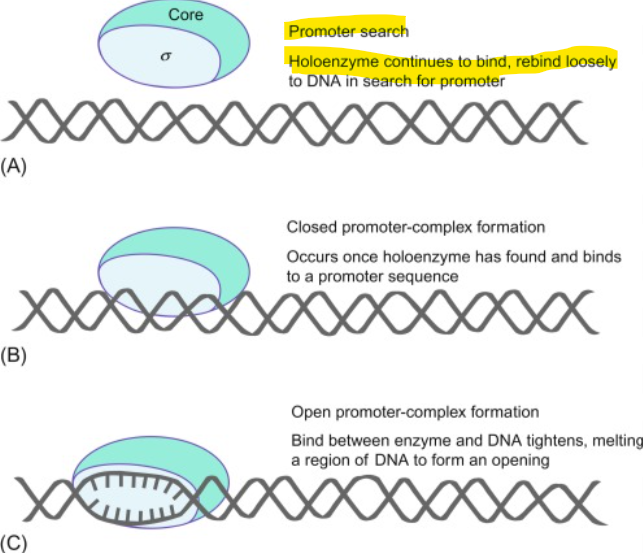
What is the mechanism of the holoenzyme
σ binds to specific DNA sequences (promoter)
RNA polymerase is positioned correctly
Transcription begins at +1
What happens if we dont have a σ
RNA polymerase would bind DNA randomly with no proper gene expression
what happens after initiation
the first 10 nucleotides are made with the sigma often falling off and the core enzyme continuing elongation
What is the flow if initiation?
Holoenzyme (with σ) → finds promoter → starts RNA
↓
σ leaves
↓
Core enzyme → continues elongation
different σ factors mean different genes which means not all promoters are the same. So what does this mean for bacteria?
Bacteria have multiple σ subunits and each recognizes different promoter sequences
Different σ factors mean different genes which means not all promoters are the same. This means that bacteria has multiple σ subunits and each recognizes different promiter sequences. What is a primary example of this
σ⁷⁰
what is significant about this σ⁷⁰
it is a default promoter which is a housekeeping gene
what do the other σ factor do?
they respond to stress and heat shock
why is it important to have different σ
it allows bacteria to rapidly switch gene expression and respond to the environment. By changing σ, you change which genes are transcribed
due to promoters being within the DNAsequence what can you classify promoters as
cis elements- DNA sequence
since having a σ factor determines if transcription properly occurs what can you classify σ factors as
trans factor - proteins that recognize it
The core RNA polymerase ____________, while the σ subunit_______________________________
synthesizes RNA; allows the holoenzyme to recognize promoters and initiate transcription at the correct site
what is a prokaryotic promoter
a DNA region (on the coding strand) that tells RNA polymerase where to bind and start transcription.