DNA Structure, Topology, Recognition
1/40
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
DNA Structure
phosphodiester bonds
hydrogen bonding
double helix
Recognition of DNA by Proteins
DNA binding domains
hydrogen bonding
DNA Topology
linking number
supercoiling
topoisomerases
Higher Order Structure
histones
nucleosomes
chromatin
Information pathways - the central dogma
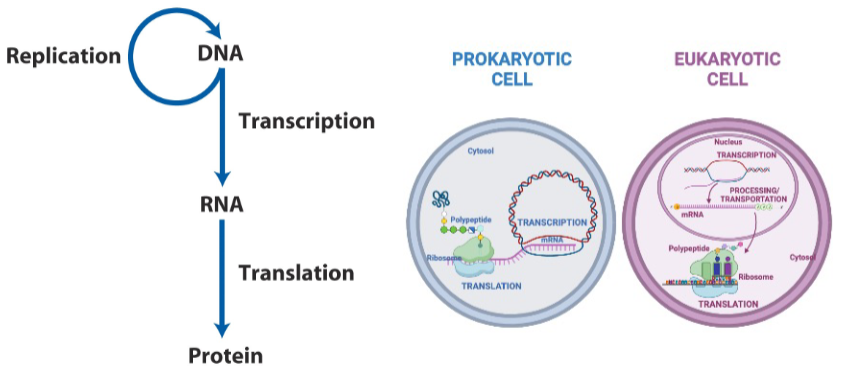
DNA is an informational molecule
within genes
outside of genes
polymer of _________________
DNA is an informational molecule
sequence of bases specifies genetic information
within genes
nucleotides specify the amino acid sequences of proteins
outside of genes
regulatory sequences direct:
DNA replication
mRNA synthesis
when, where, how much?
DNA is a polymer of deoxyribonucleotides
Deoxyribonucleotides
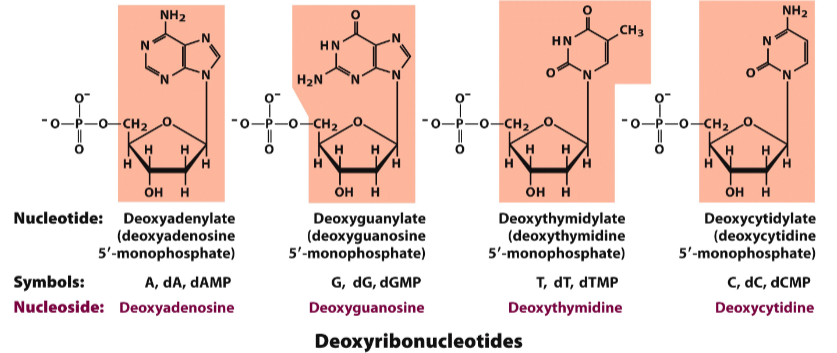
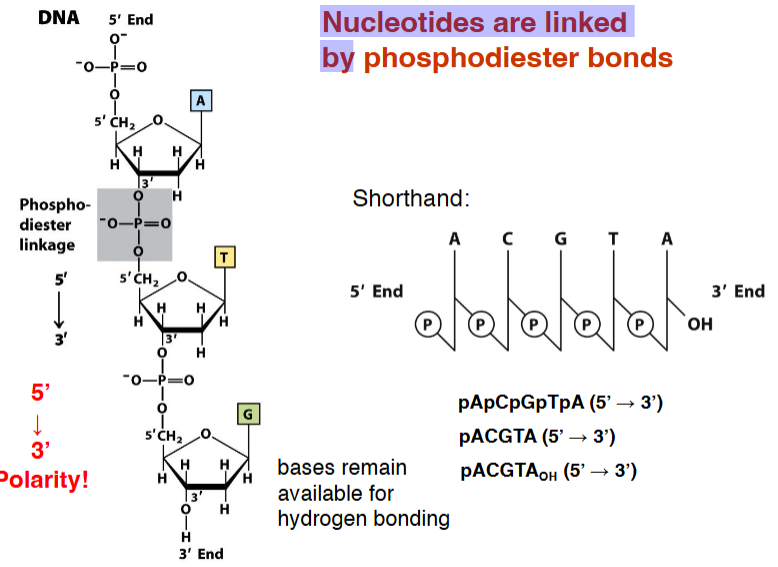
Nucleotides are linked by _____________ bonds
what direction polarity?
bases remain available for __________ ________
Common hydrogen bonds in biological systems
bond length?
Angled hydrogen bonds are weaker why?
2.6-3Å bond length
Angled hydrogen bonds (due to constraints in protein structures, for example, are weaker.
Hydrogen bonding mediates _____________________- in DNA
Hydrogen bonding mediates base-pairing interactions in DNA
structure of dna
6 points regarding strand interactions, location of backbone vs bases, base pairing, how information is copied
Two strands of DNA– held together by base-pairing interactions
the DNA strands are antiparallel
Sugar-phosphate backbone is on the outside
the bases are stacked on the inside
Base pairs are purine-pyrimidine
G pairs with C
A pairs with T
Complementarity allows information to be copied
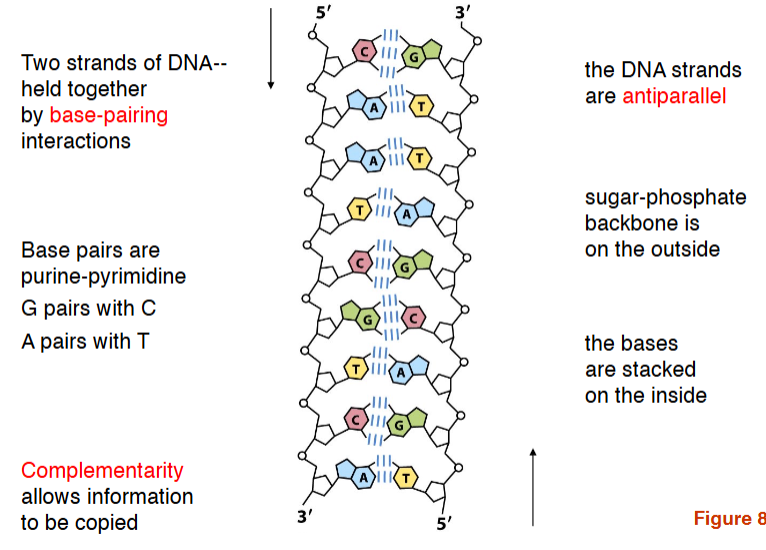
dna double helix
how is it stabilized?
hydrogen bonding + stacking interactions
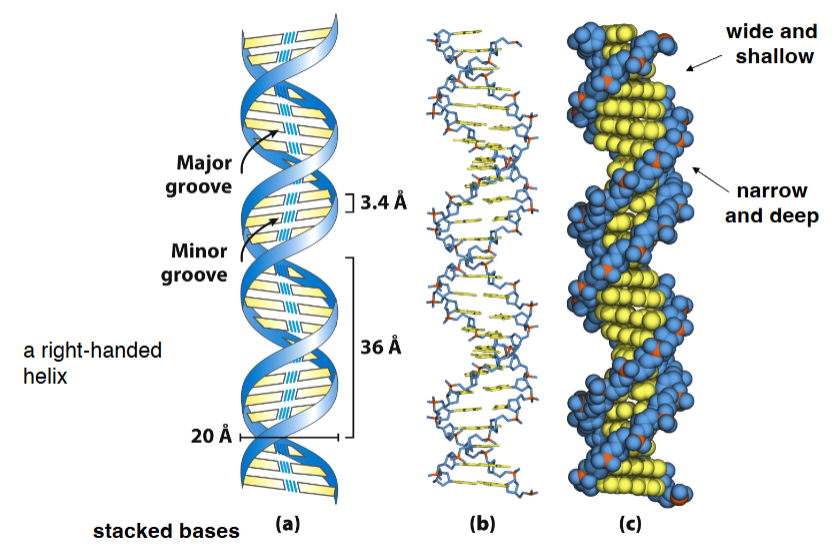
Structural variation in DNA
A Form DNA: | B Form DNA: | Z Form Helix: |
• DNA-RNA, RNA- RNA helix • found in solution • 11 bp per turn • right-handed | • most stable • found in solution • 10.5 bp per turn • right-handed | • left-handed |
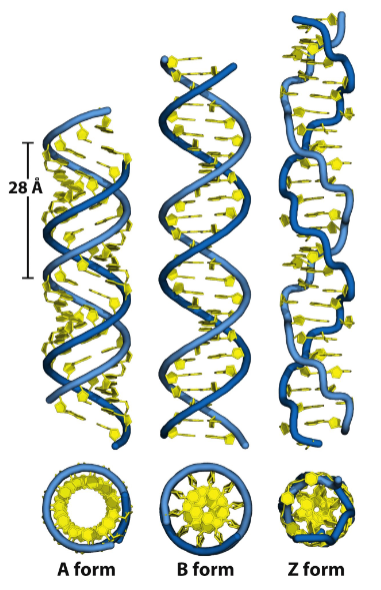
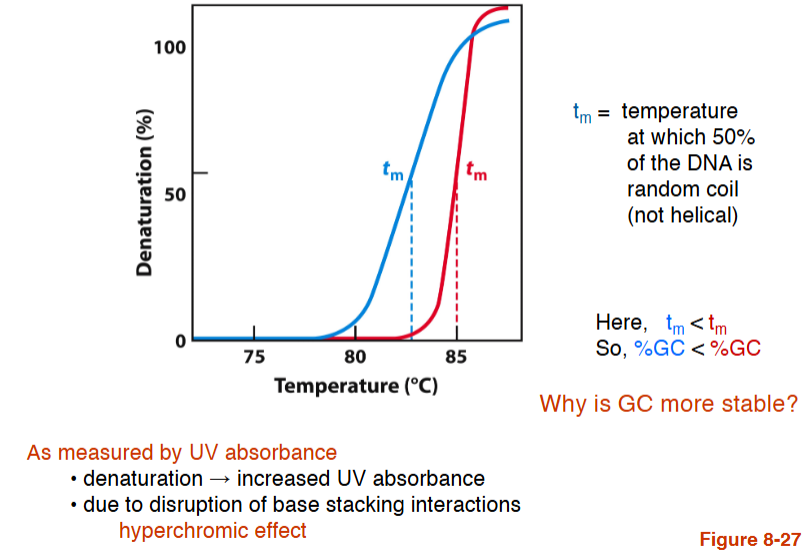
Denaturation of DNA
tm = temperature at which 50% of the DNA is random coil (not helical)
Here, tm < ™
So, %GC < %GC
Why is GC more stable?
As measured by UV absorbance, denaturation → __________?
due to _____________ = what effect?
High temperature or pH
tm increases linearly with number of GC bonds
RNA Structure
single strand helix, hairpin double helix (secondary structure)
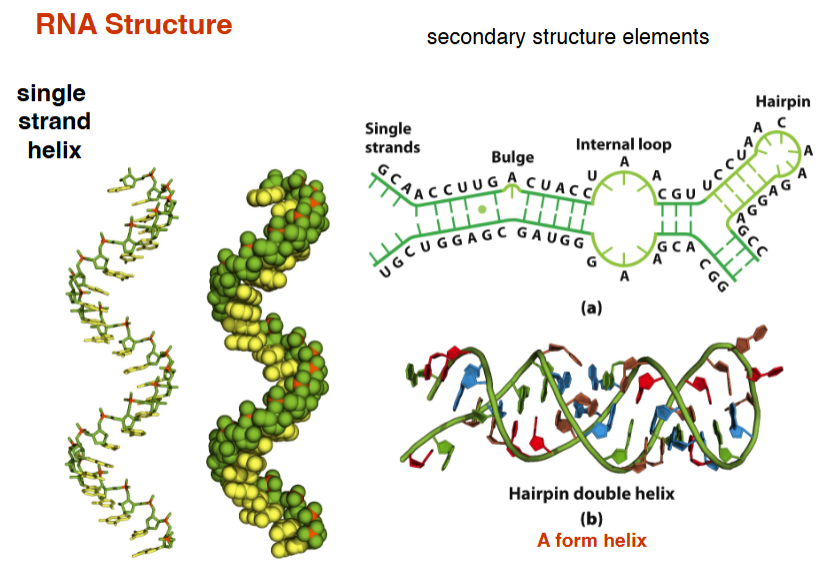
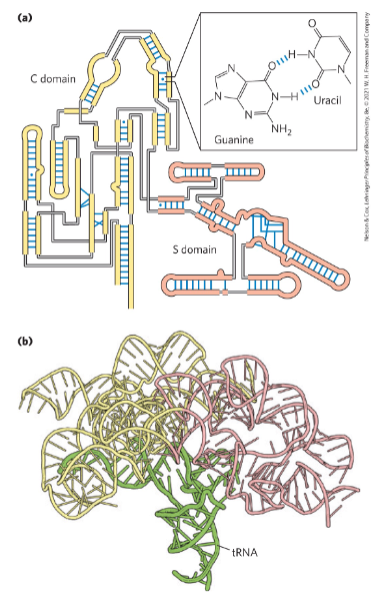
An example of RNA structure: RNase P
Base pairs in RNA:
A-U, G-C
AND
G-U
DNA sequences are recognized by DNA binding proteins
Sequence-specific DNA recognition is key to what?
How are specific DNA sequences recognized?
Sequence-specific DNA recognition is key to carrying out the steps in information transfer
How are specific DNA sequences recognized?
• hydrogen bonding
• major groove interactions
Hydrogen bonding opportunities in double helical DNA
Hydrogen bond acceptors (🔵) and donors (🔴) are available in
• the major groove (AT, TA, GC, CG can be discriminated)
• the minor groove (AT/TA vs. GC/CG can be discriminated
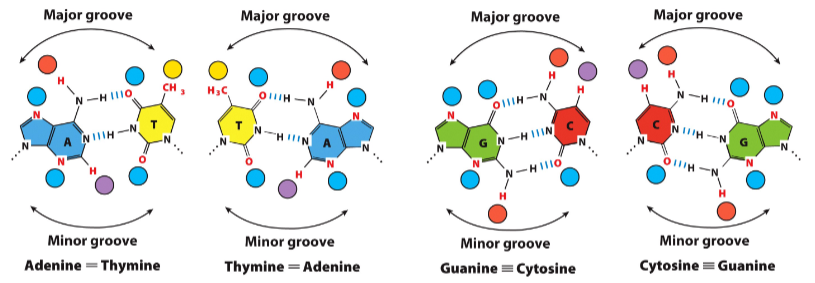
Amino acid side chains form hydrogen bonds with bases in double helical DNA
Sequence-specific DNA recognition is key to carrying out the steps in information transfer
Asn
• Gln
• Glu
• Lys
• Arg
DNA binding proteins
• Generally bind DNA in the Major Groove
• a-helix fits nicely into the wide major groove
Certain DNA Binding Motifs are common (5)
Helix-Turn-Helix (HTH)
Zinc Finger
Homeodomain
Leucine Zipper
Basic Helix-Loop-Helix (BHLH)
DNA Binding Motifs: Leucine Zipper
Lys and Arg residues ➡DNA binding + in the major groove
Leu residues stabilize dimerization
DNA binding motifs: homeodomain
The homeodomain is a widely conserved DNA binding motif of ~ 60 amino acids
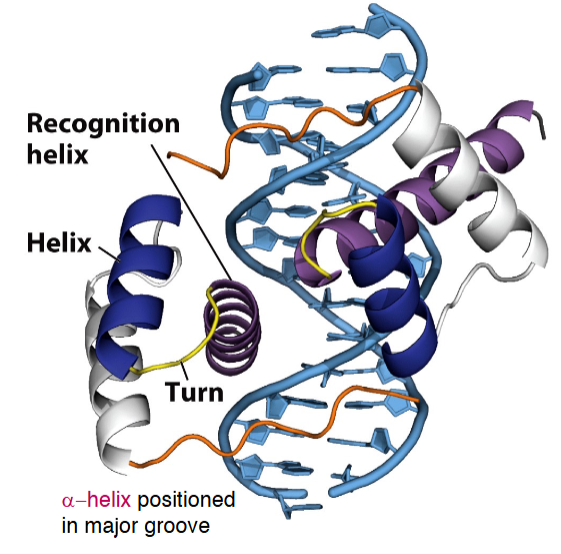
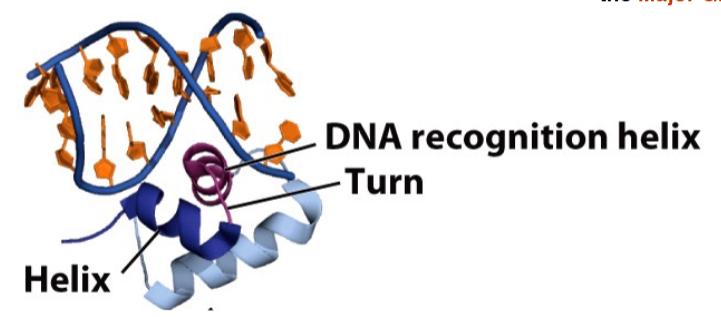
DNA binding motifs: helix-turn-helix
Recognition Helix is positioned in the Major Groove
Helix-turn-helix domains in the lac repressor protein
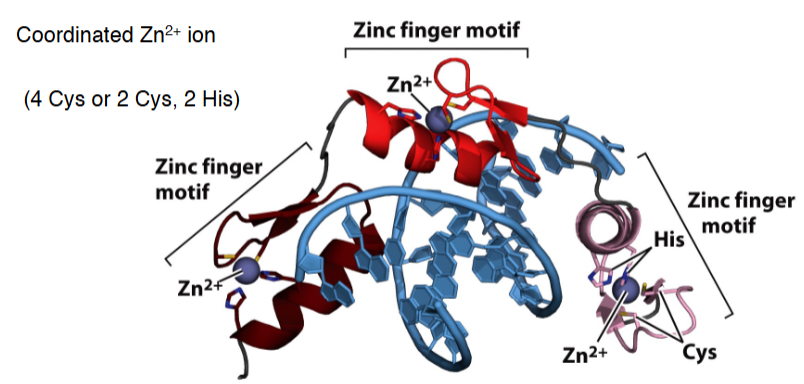
DNA binding motifs: zinc finger
Coordinated Zn2+ ion (4 Cys or 2 Cys, 2 His)
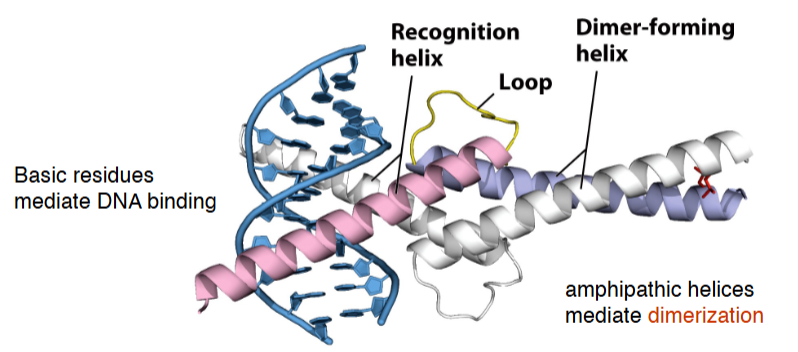
DNA binding motifs: helix-loop-helix
amphipathic helices mediate dimerization
Basic residues mediate DNA binding
DNA must be compacted to fit in cells
length details?
The length of T4 DNA is several hundred times the size of the viral particle.
If you add up the length of the DNA in all cells of an average human
(~1014 cells) the total DNA length is ~ 2 x 1011 km.
The distance between the earth and the sun is 1.5 x 108 km.
Supercoiling does 2 things:
Supercoiling compacts DNA
Supercoiling relieves strain
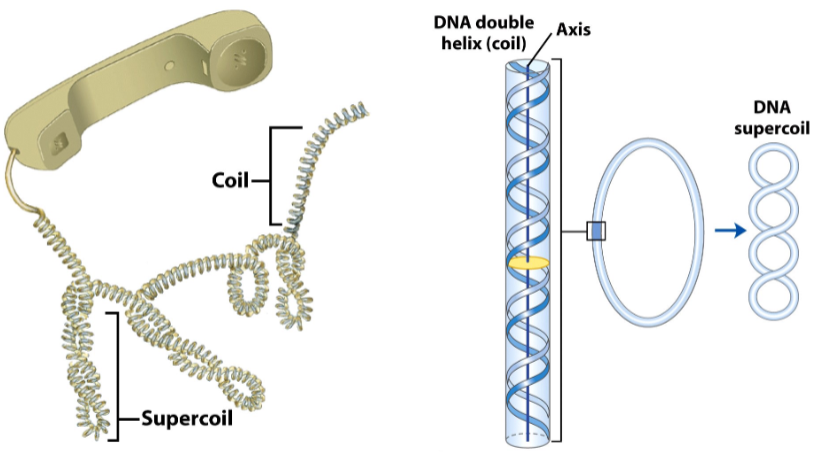
Polymerases separate the strands and introduce strain
Supercoiling is an intrinsic property of DNA tertiary structure.
• It occurs in all cellular DNAs and is highly regulated by each cell.
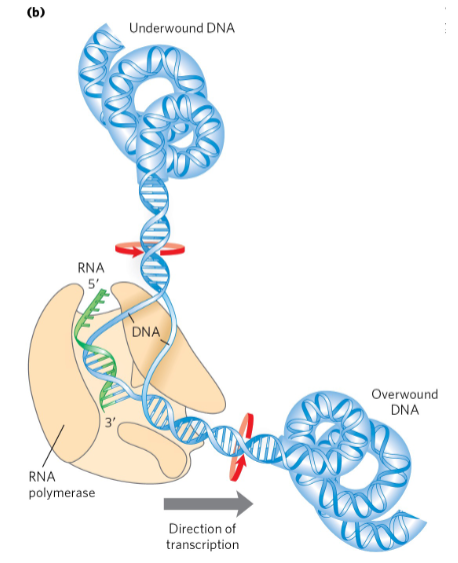
Most cellular DNA is underwound: Supercoiling _________ __________
Underwinding facilitates strand separation
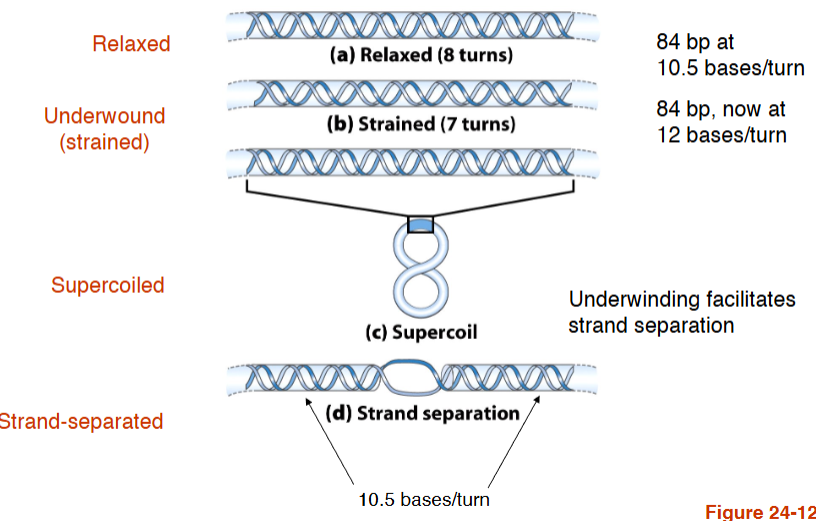
DNA underwinding is defined by the topological linking number, Lk
Separation of two strands of a double-stranded circular DNA.
In DNA, Lk specifies the number of helical turns in a closed circular DNA
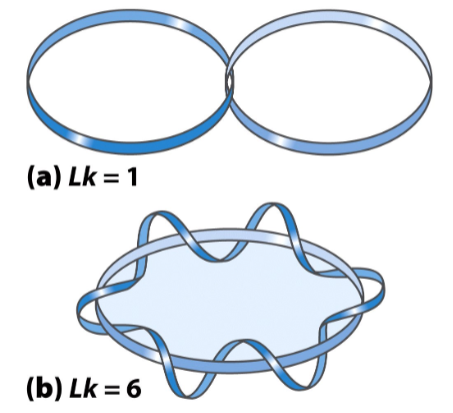
Lk = #bps ÷ bps per turn
Relaxed DNA:
2100 bp DNA / (10.5 bp/turn)
Lk = 2100 ÷ 10.5
Lk = 200
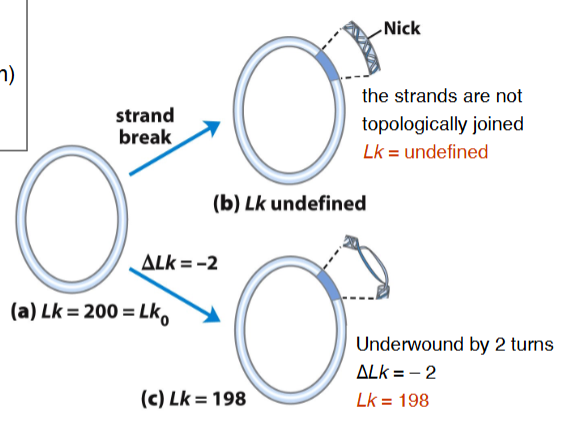
Superhelical Density
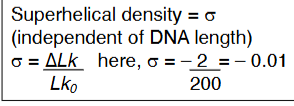
Negative and positive supercoils
Two forms of a circular DNA that differ only in a topological property such as linking number are referred to as _________
Topoisomers.
Underwind by 2 turns ➡∆Lk = –2
So, Lk = 200 – 2 = 198
Overwind by 2 turns ➡∆Lk = +2
So, Lk = 200 + 2 = 202
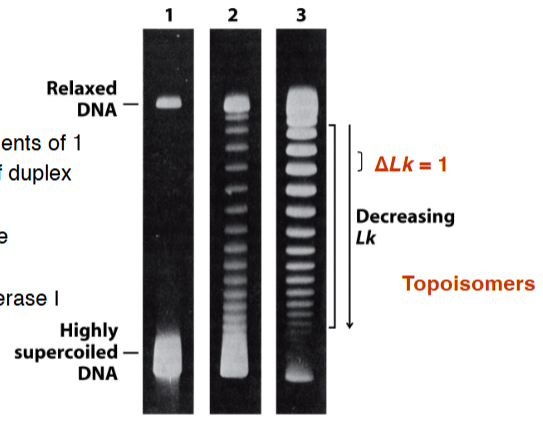
topoisomerase
Type I • changes Lk in increments of 1 • cleaves one strand of duplex DNA • can relax positive and negative supercoils • example: Topoisomerase I | Type II • changes Lk in increments of 2 • cleaves both strands of DNA • can relax positive and negative supercoils • can introduce negative supercoils (prokaryotes only) • hydrolyzes ATP • examples: DNA gyrase, Topoisomerase II |
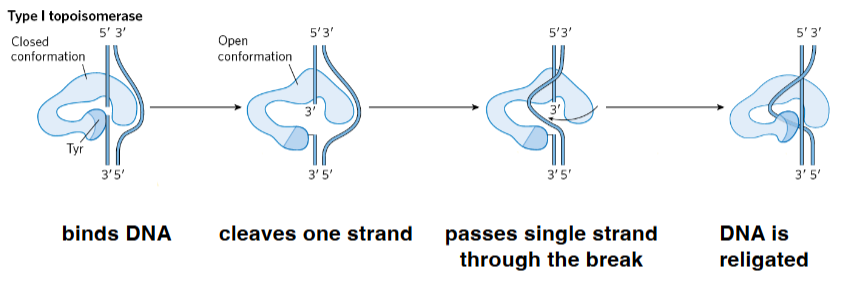
Type I Topoisomerases
• change Lk in increments of 1
• cleave one strand of duplex DNA
• can remove negative supercoils
Topoisomerase I steps
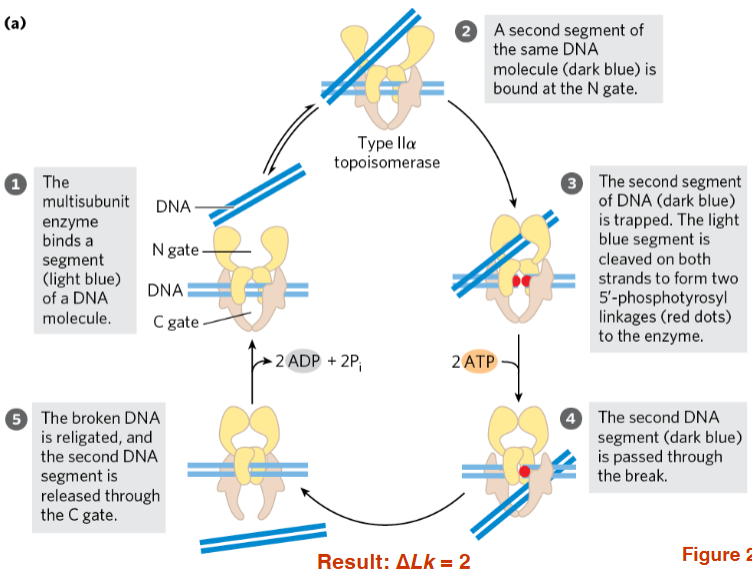
Topoisomerase II reactions
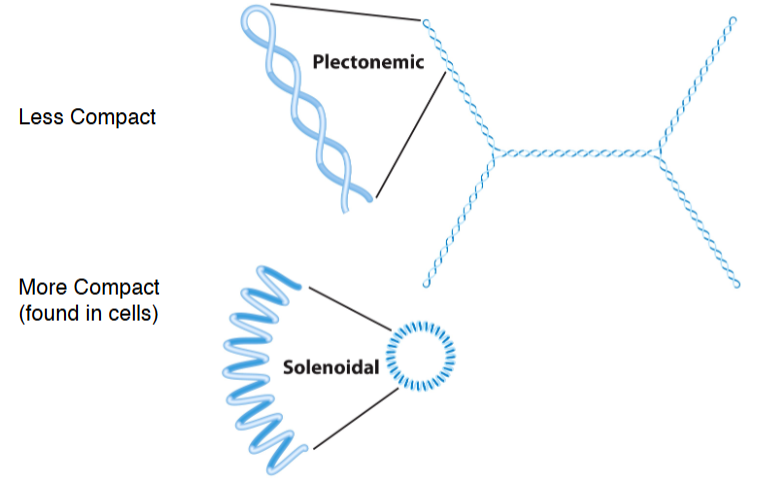
Plectonemic and solenoidal supercoiling
Plectonemic supercoiling does not produce sufficient compaction to package DNA in the cell.
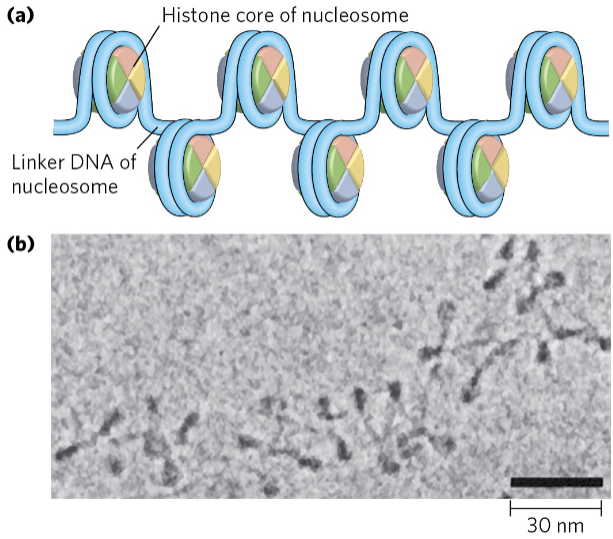
DNA is packaged into nucleosomes
Histone proteins act to package the DNA