L15. Antibiotic resistance in the environment
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
What is the concept of ‘One health’?
An interdisciplinary approach that recognizes the interconnectedness of human, animal, and environmental health, advocating for collaborative efforts in managing health issues to improve outcomes across all three domains.
i.e. Treatment effectiveness across human, animal, and environmental health, microbes are the biological link
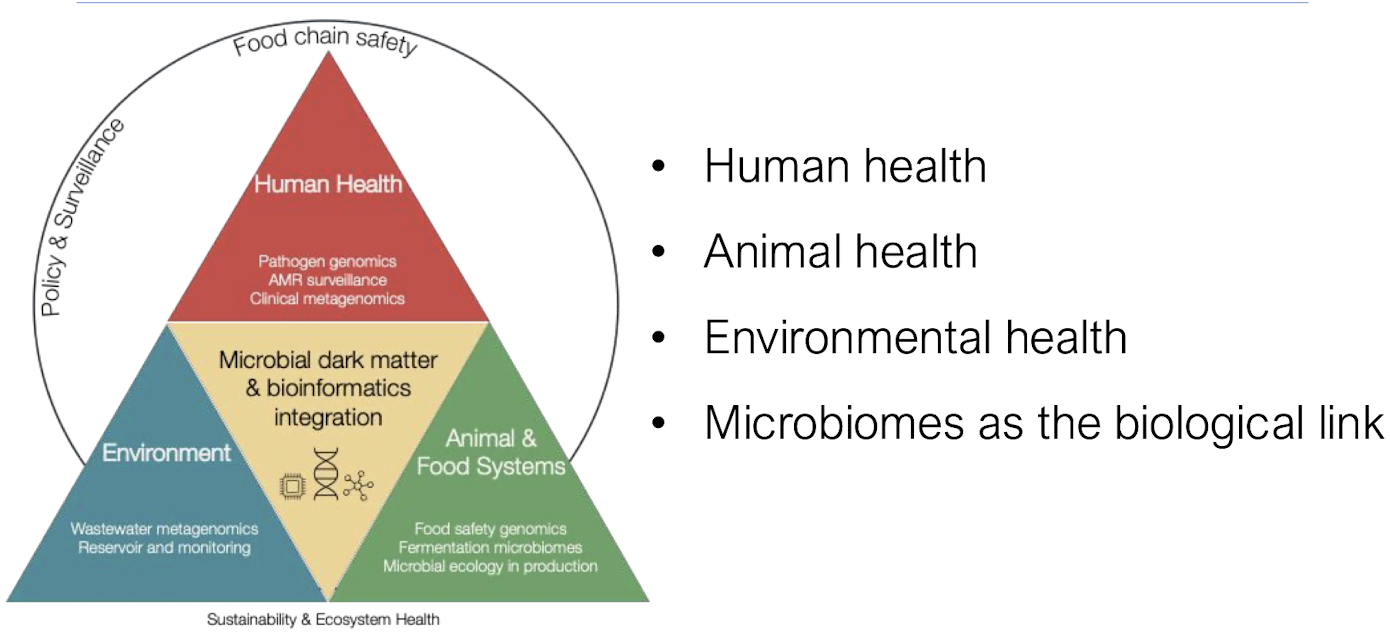
What is the scale of antibiotic use?
• Large fraction used in agriculture, even more so than human use
• Growth promotion and prophylaxis - used just in case / merely for improving crop yields
• Environmental exposure as molecules remain in the environment via manure, runoff, wastewater
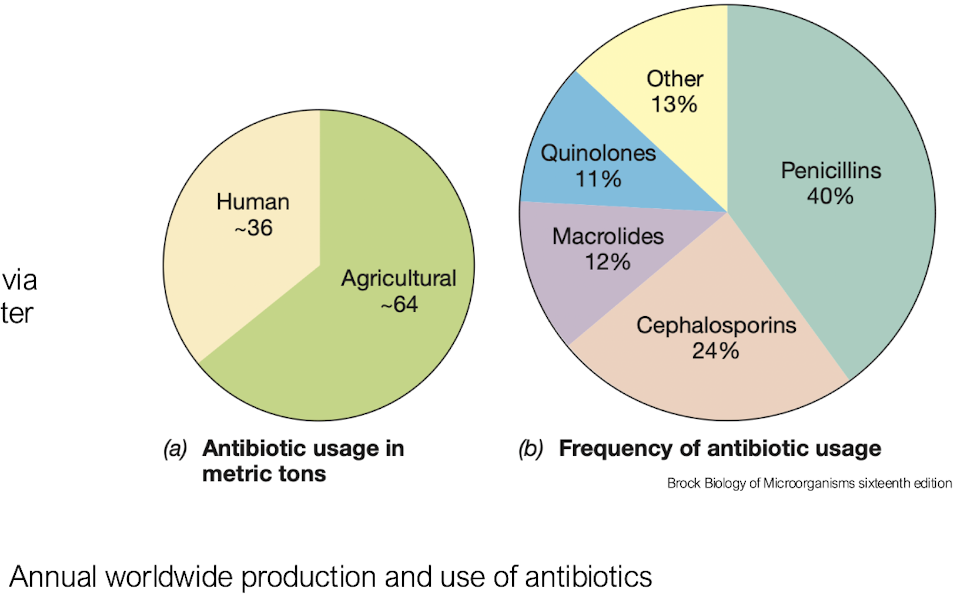
What are the main modes of action of antibacterial agents?
• Cell wall inhibitors that are different from the host to not damage host cells e.g. penicillin, vancomycin
• Protein synthesis inhibitors e.g. erythromycin (macrolides)
• DNA/RNA synthesis inhibitors e.g. puromycin
• Metabolism inhibitors e.g. sulfonamides and trimethoprim (folic acid)
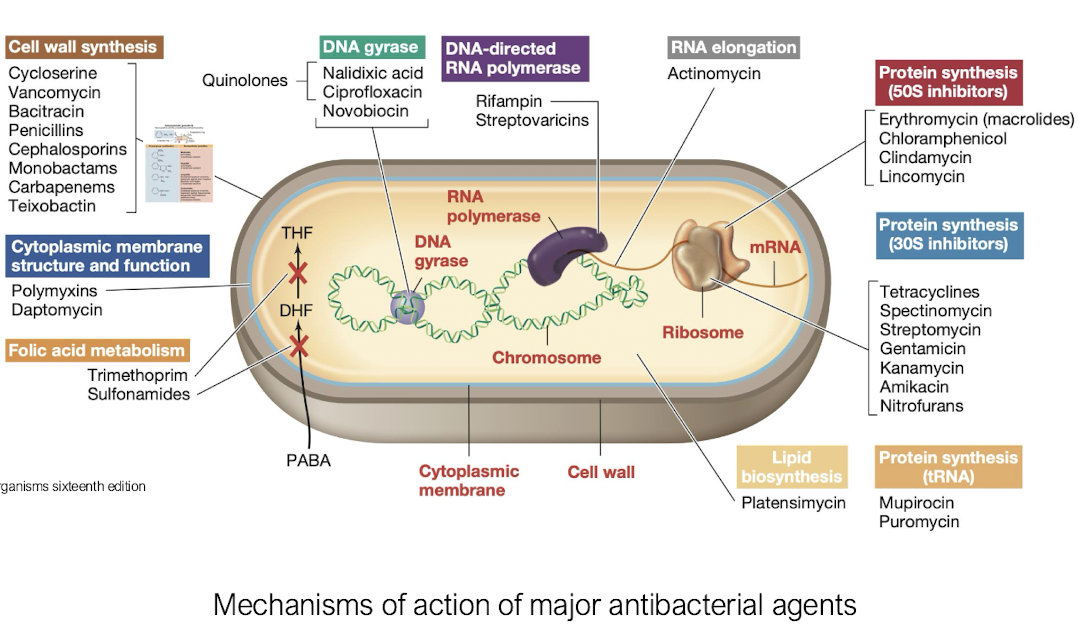
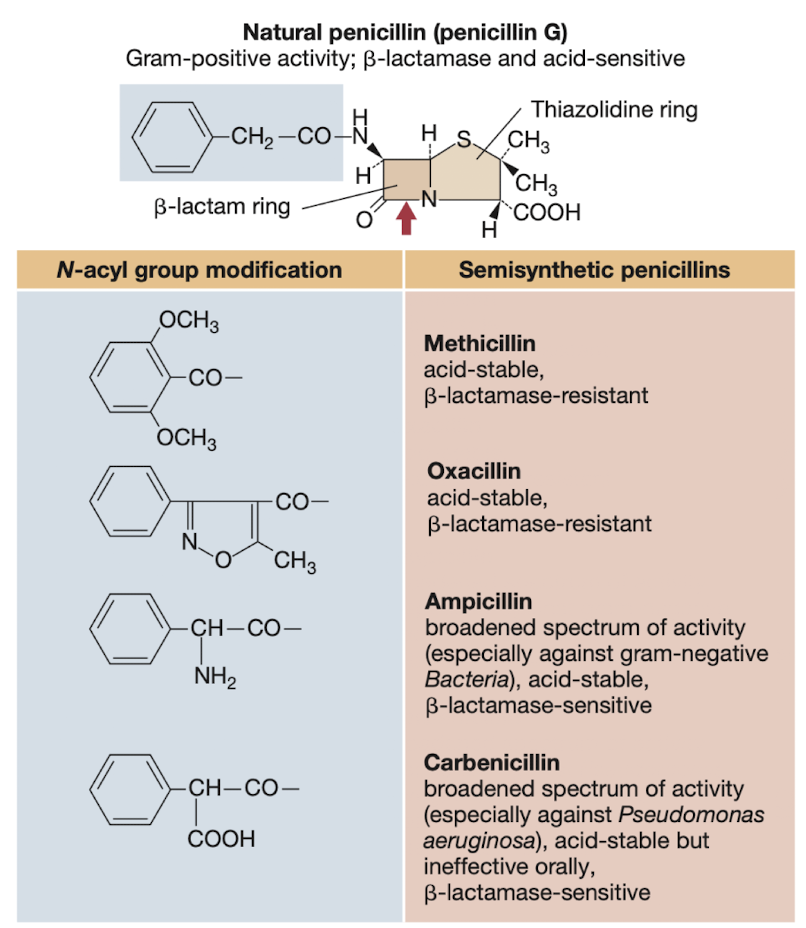
What is a common feature of molecules in antibiotics
N-acyl group modification often enhances their antibacterial activity and stability.
How do antibiotics contribute to environmental contamination?
Antibiotics are only partially metabolised in humans and animals -> leave residues → wastewater
• Wastewater treatment plants do not fully remove antibiotics or ARGs
• Agricultural manure contains antibiotics, resistant bacteria, and resistance genes -> Runoff transports contaminants into soils, rivers, and groundwater
• Aquaculture releases antibiotics directly into aquatic environments
• Low concentration but biologically active - Low-dose, chronic exposure creates continuous selection pressure
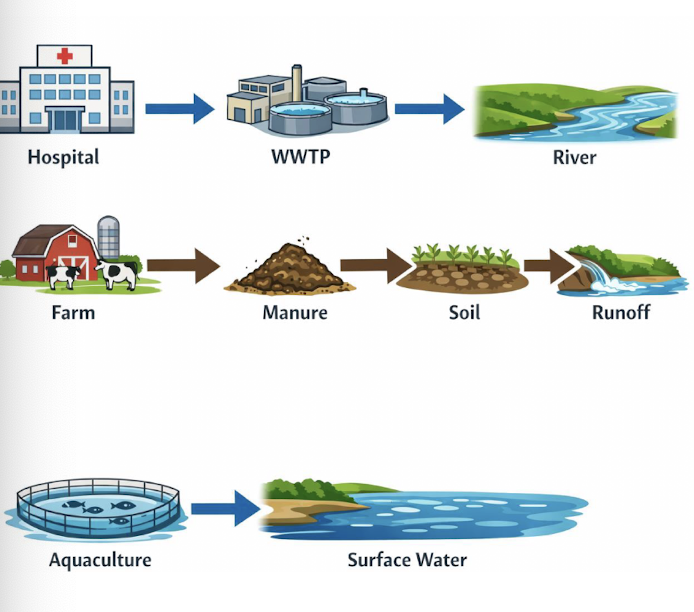
How does resistance arise?
• Resistance arises through random genetic mutation
• Antibiotics do not create resistance; they select for it
• Sub-inhibitory concentrations can still select for resistance
• Compensatory mutations can reduce fitness costs
• Resistance can persist even after antibiotic removal
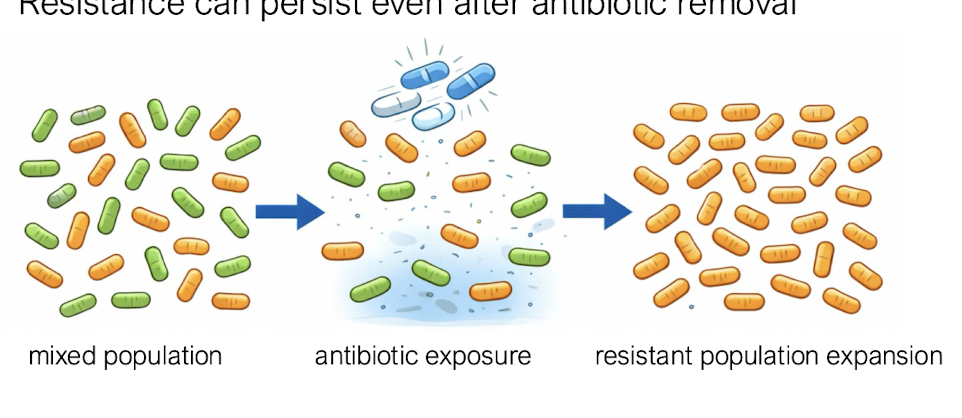
How is enzymatic inactivation & target modification a mechanism of resistance?
• Beta-lactamase production degrade beta-lactam antibiotics, evolved overtime after exposure to the antibiotic
• Modifying enzymes alter drug structure → loss of function
• Mutation of antibiotic binding sites
• Ribosomal protection proteins
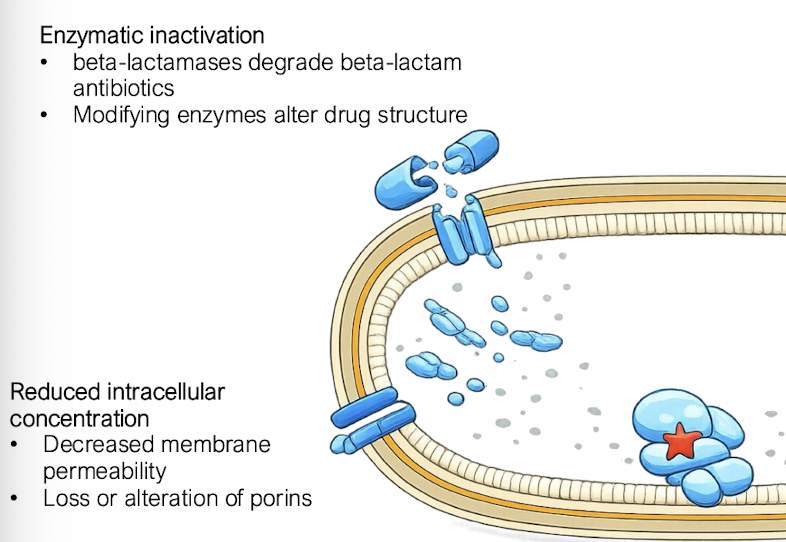
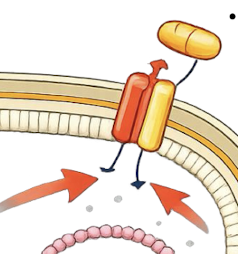
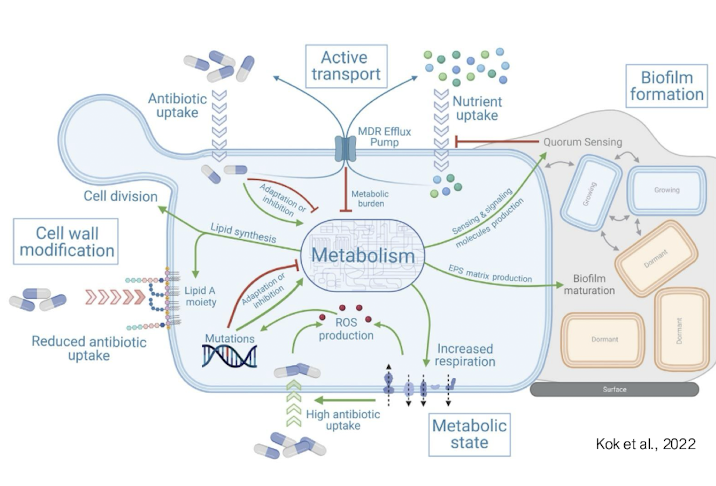
How is active efflux a mechanism of resistance?
• Efflux pumps export antibiotics from the cell
• Often confer multidrug resistance
Expelling liquid with the antibiotics out out the cell, drug not initially intended for bacteria to have these systems
How is reduced intracellular concentration a mechanism of resistance?
• Decreased membrane permeability
• Loss or alteration of porins
How do plasmids confer antibiotic resistance?
Plasmids are small, circular DNA molecules that can carry antibiotic resistance genes and be transferred between bacteria through horizontal gene transfer.
Chromosomal mutations
Plasmid-borne resistance genes
Allows for the rapid spread of resistance traits within bacterial populations.
Resistome =
= comprises ALL the antibiotic resistance genes (ARGs)
Includes resistance elements found in both pathogenic bacteria and antibiotic-producing bacteria - expressed as genes
Cryptic resistance genes (which are not necessarily expressed) that are present in bacterial chromosomes.
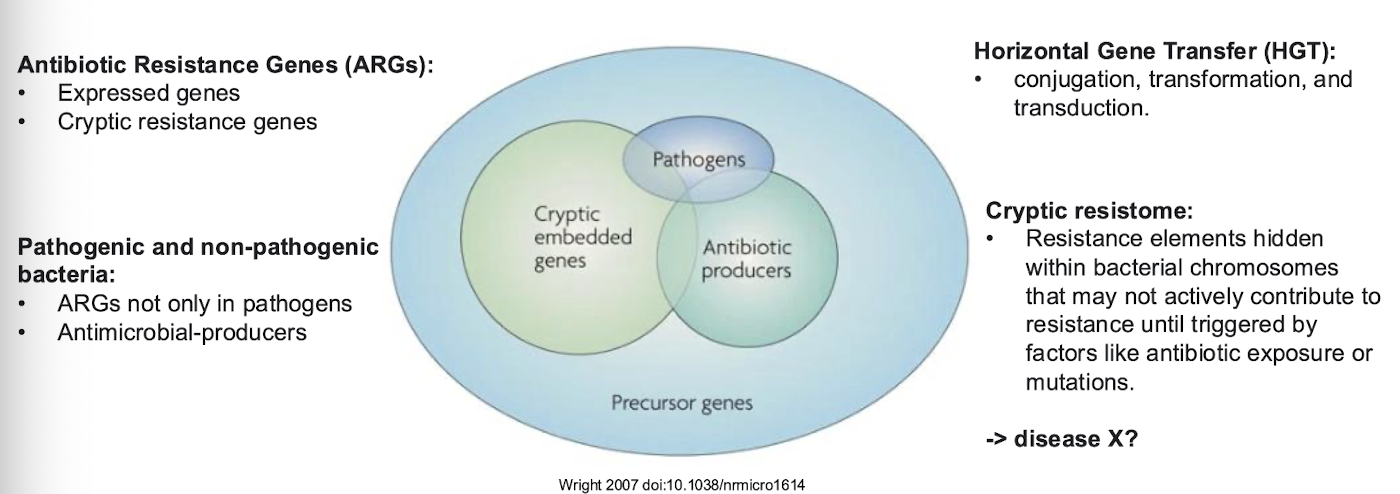
Cryptic resistome
Resistance elements hidden within bacterial chromosomes that may not actively contribute to resistance until triggered by factors like antibiotic exposure or mutations
Review of HGT
Horizontal Gene Transfer (HGT) refers to the process by which bacteria exchange genetic material, including antibiotic resistance genes, through mechanisms such as transformation, transduction, and conjugation:
• Conjugation: plasmids + conjugation pilli ‘sex pili’
• Transformation: free DNA
• Transduction: bacteriophages (virus) transfer genes, often stress genes
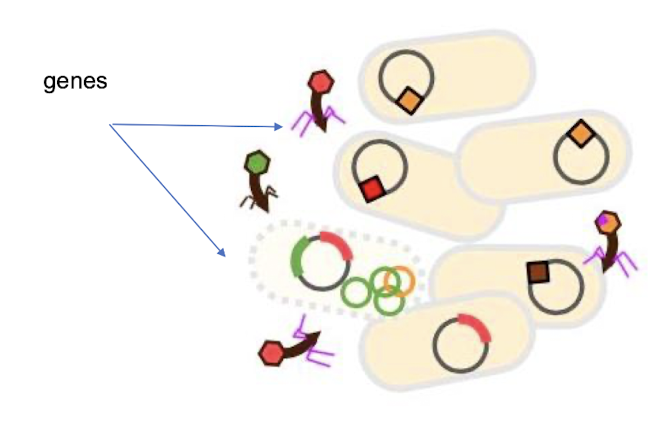
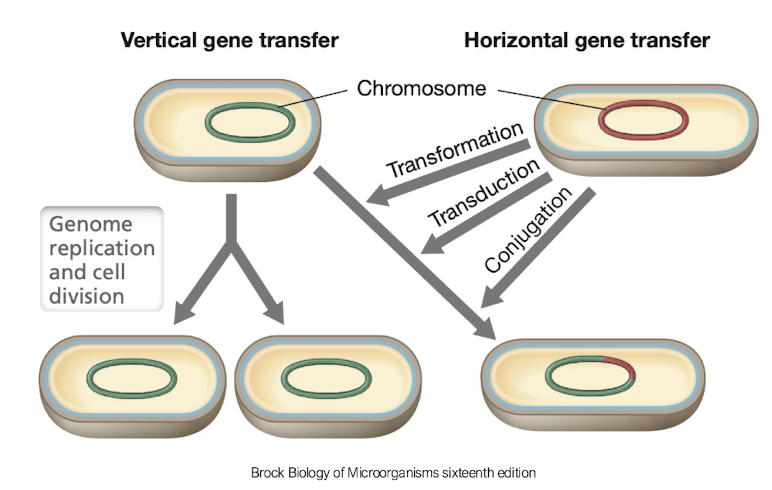
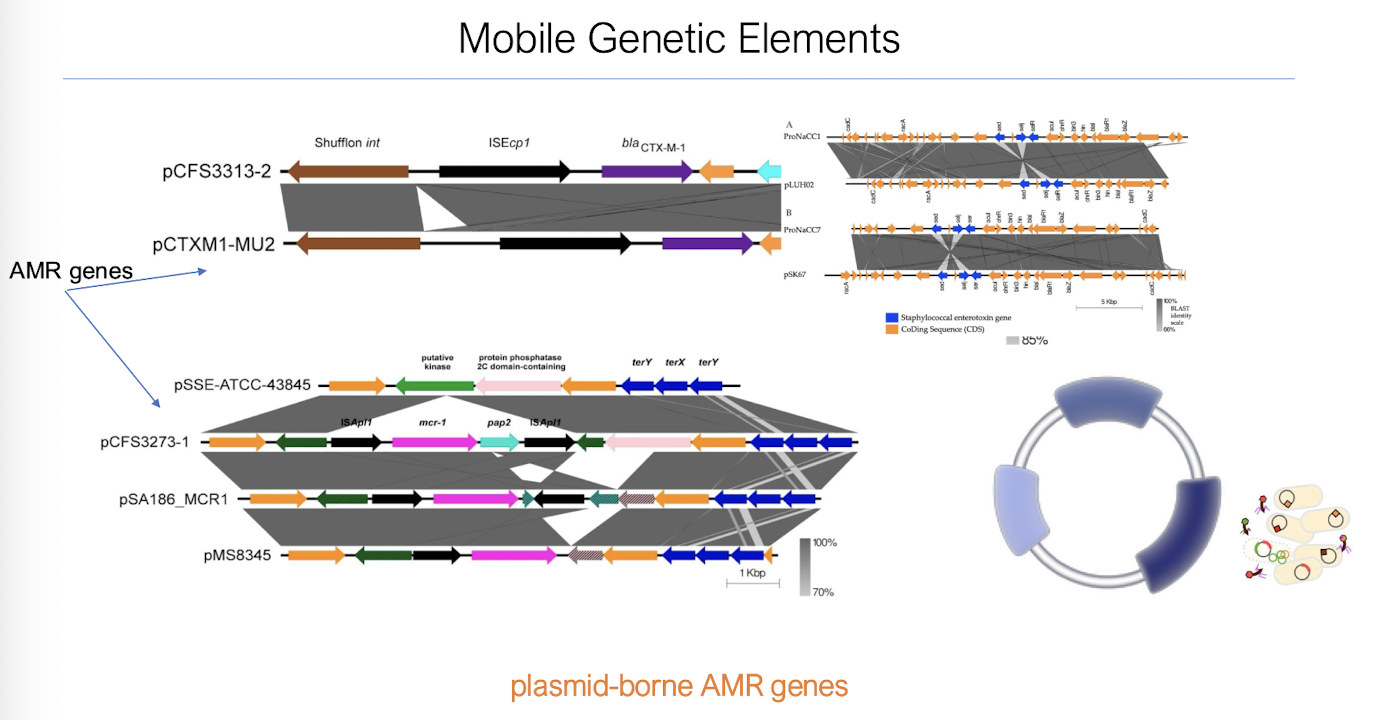
Where are AMR genes found in the bacterial genome?
Antimicrobial resistance (AMR) genes are primarily found in the bacterial genome within plasmids, chromosomes, and transposons, enabling their transfer and expression.
S. aureus can encode for enterotoxins within their plasmids, and AMR genes.
How do food systems act as interfaces for AMR transmission?
• Livestock exposure to therapeutic and prophylactic antibiotics
• Animal gut microbiomes act as reservoirs
• Manure application spreads resistant bacteria into soil
• Crops can be contaminated via soil and irrigation water
• Raw and minimally processed foods can carry ARGs
• Long food chains and spread
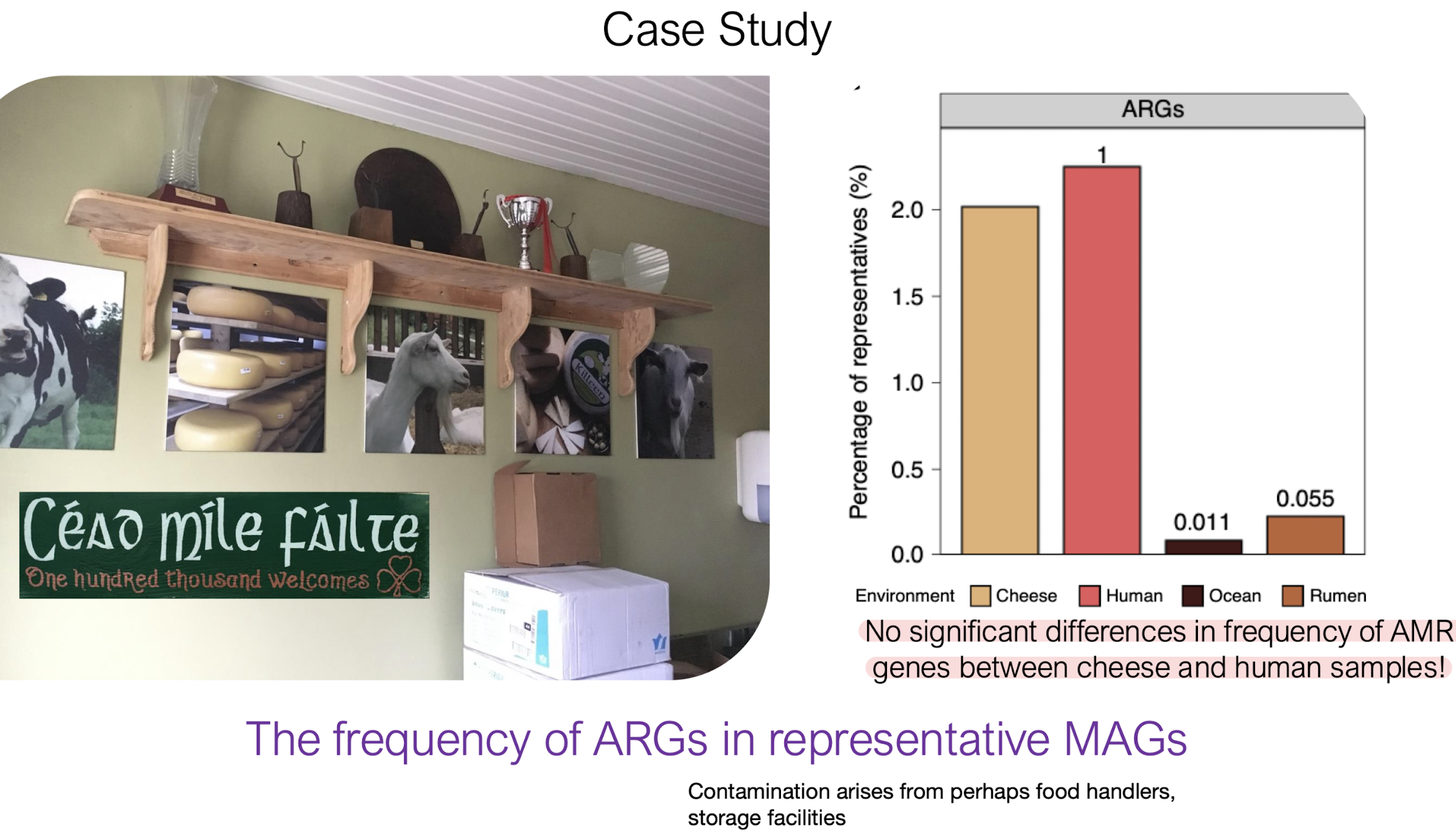
What did the case study in the powerpoint reveal?
No significant differences in frequency of AMR genes between cheese and human samples!
How can long distance transport allow for mobile genetic elements to spread between species?
Via:
• Global food trade
• Migratory birds carry resistant bacteria across continents
• International travel
• River systems transport downstream
• Dust and aerosols
How can heavy metals e.g. copper, zinc, co-select for ARGs?
• Heavy metals (e.g. copper, zinc) can co-select for ARGs
• Biocides and disinfectants exert selective pressure, worrying trend
• Resistance genes are often linked on the same plasmid
• Selection for metal resistance can maintain antibiotic resistance
• Sub-lethal stress can enhance HGT
How can we detect ARG’s in the environment?
Using metagenomics:
Detecting mobile genetic elements e.g. plasmids and other genetic markers associated with antibiotic resistance.
How can we use metatranscriptomics to study antibiotic resistance?
Metatranscriptomics allows for the analysis of mRNA in environmental samples.
Captures RNA transcripts to reveal which genes are actively expressed in response to antibiotics.
It helps understand how these genes are expressed in various conditions and their roles in microbial communities.
How can we use metabolomics to study antibiotic resistance?
• Identification of metabolic pathways that may enable resistance or resilience under antimicrobial pressure.
• Complements gene-level data by linking metabolic function with resistance phenotypes
Which sector accounts for the majority of global antibiotic use according to the lecture?
C. Agriculture
Which of the following contributes most directly to environmental antibiotic contamination?
C. Manure runoff containing antibiotics and ARGs
Low‑dose, chronic environmental antibiotic exposure is problematic because:
B. It creates continuous selective pressure
Which antibiotic class targets DNA gyrase?
B. Quinolones
Which antibiotic class inhibits folic acid metabolism?
B. Sulfonamides and trimethoprim
Which semisynthetic penicillin is β‑lactamase‑resistant?
C. Methicillin
Which statement best describes how antibiotic resistance evolves?
B. Antibiotics select for pre‑existing resistant mutants
Which mechanism reduces intracellular antibiotic concentration?
C. Loss or alteration of porins
Efflux pumps contribute to resistance by:
B. Pumping antibiotics out of the cell
The resistome includes:
C. All antibiotic resistance genes, including cryptic ones
Which HGT mechanism involves bacteriophages?
C. Transduction
Cryptic resistance genes are characterised by:
C. Being silent until triggered by stress or mutation
Which statement about AMR transmission in food systems is correct?
B. Raw foods may carry ARGs
In the Irish cheese study, which phylum had the highest number of MAGs?
B. Firmicutes
What was the key finding regarding ARG frequency in cheese vs human samples?
C. Cheese and human samples showed no significant difference
Which non‑antibiotic factor can co‑select for ARGs?
B. Heavy metals
Which process enhances AMR spread across continents?
B. Migratory birds
Metagenomics is particularly useful because it:
B. Identifies mobile genetic elements such as plasmids
Metatranscriptomics provides insight into:
B. Which genes are actively expressed
Metabolomics contributes to AMR surveillance by:
B. Identifying metabolic pathways linked to resistance