BIOL 280 - Exam 4
1/456
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
457 Terms
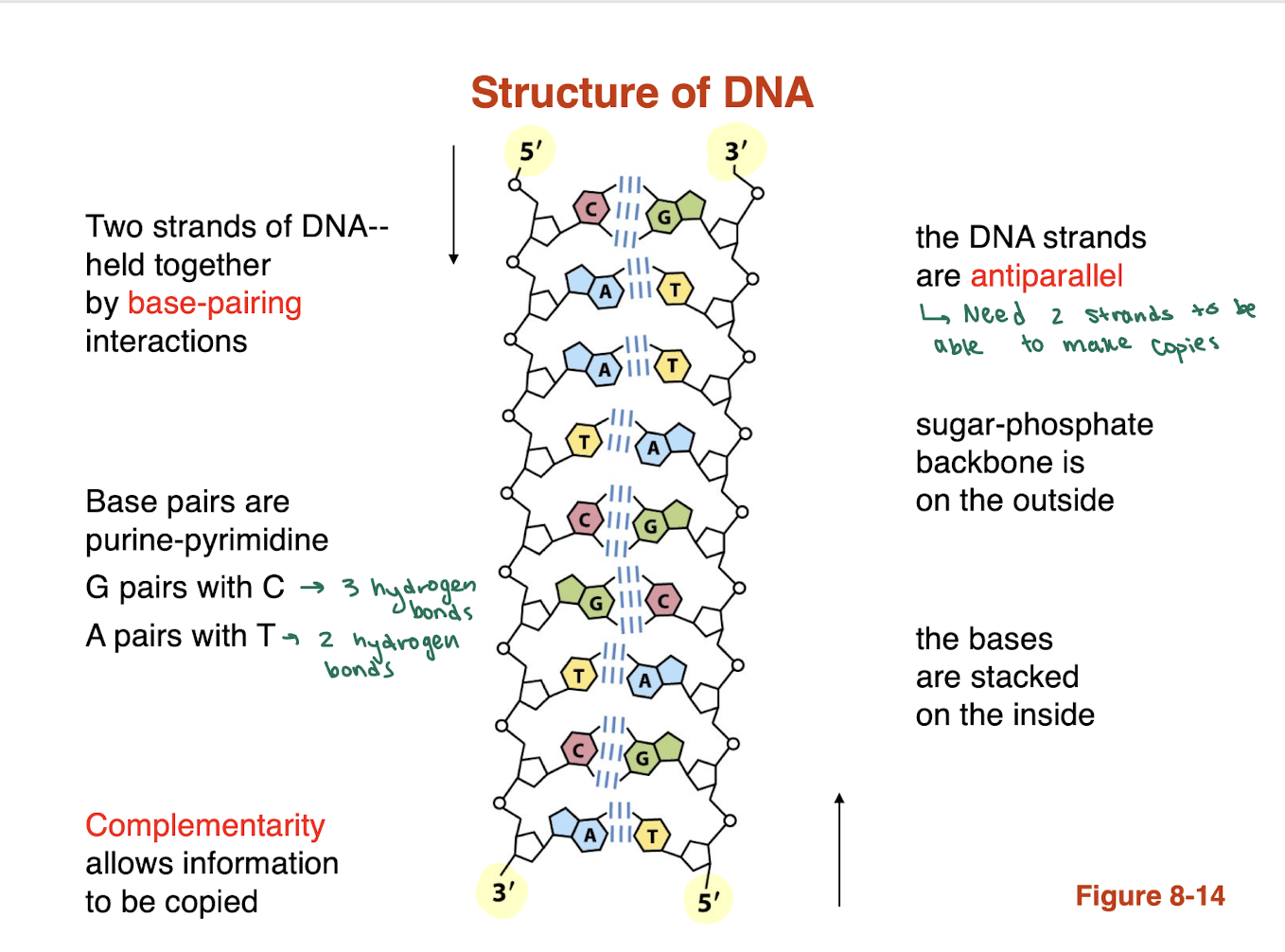
What is a phosphodiester bond and how does it form the DNA backbone?
A phosphodiester bond is a covalent linkage between the 3′ OH group of one nucleotide and the 5′ phosphate group of the next nucleotide.
This creates the sugar-phosphate backbone of DNA.
Repeats along the strand → forms a continuous chain
Leaves nitrogenous bases free for hydrogen bonding
What is DNA polarity and how is it represented?
DNA has directionality (polarity):
One end = 5′ end (free phosphate group)
Other end = 3′ end (free OH group)
Sequences are always written 5′ → 3′
Example: pACGTA
“p” = phosphate at 5′ end
“OH” (if shown) = 3′ end
This direction is critical for DNA replication and synthesis
nucleotides are linked together via?
phosphodiester bonds
What are hydrogen bonds in biological systems and what factors affect their strength?
Hydrogen bonds occur between a hydrogen donor (X–H, where X = O or N) and a hydrogen acceptor (O or N with lone pairs).
Strength depends on geometry:
Linear (straight) hydrogen bonds → stronger
Angled hydrogen bonds → weaker (common in proteins due to structural constraints)
Overall: Hydrogen bonds are weak individually but collectively stabilize biological structures.
hydrogen bonding helps with what kind of interactions in DNA?
base pairing
base pairs are purine and pyrimidine what are the two base pairings for DNA? and how many hydrogen bonds are associated with the pairings
G pairs with C —> 3 hydrogen bonds
A pairs with T —> 2 hydrogen bonds
BLANK allows for information to be copied
complementarity
in terms of the structure of DNA the sugar phosphate backbond is on the BLANK and the bases are stacked on the BLANK
sugar phosphate backbone is stacked on the outside
bases are stacked on the inside
DNA —> DNA is
replication
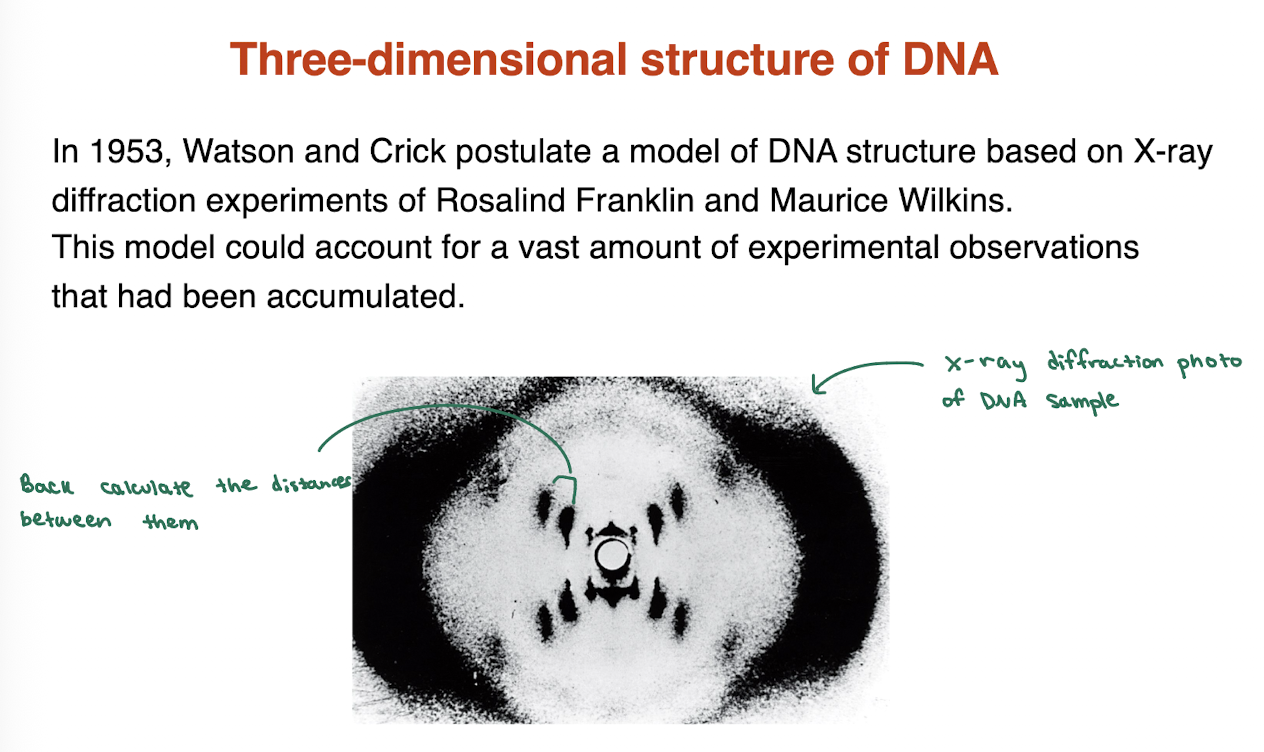
what experiments were utilized to get the three dimensional structure of DNA?
in 1953, Watson and Crick postulate a model of DNA structure based on X-ray diffraction experiments of Rosalind Franklin and Maurice Wilkins. This model could account for a vast amount of experimental observations that had been accumulated.
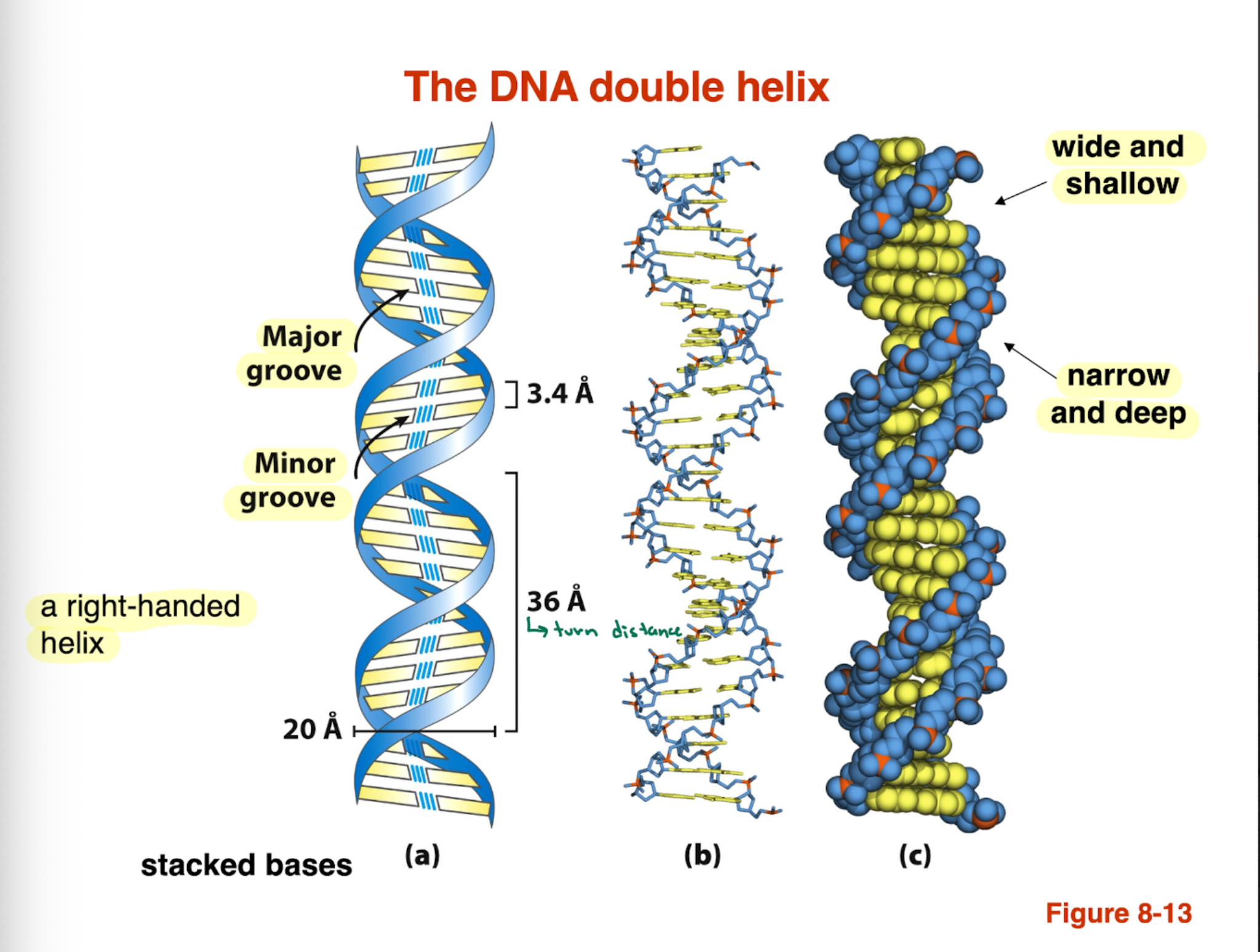
DNA forms a right or left handed helix?
What are the types of grooves and please describe them
DNA forms a right handed doubel helix
Major groove: wide and shallow —> important for protein binding
minor groove: narrow and deep
The DNA double helix is stabilized by
hydrogen bonding — between base pair
stacking interactions
what are the differences between the B form and A form of DNA?
B form DNA:
most stable
found in solution
10.5 bp per turn
right handed
A form DNA:
DNA-RNA, RNA-RNA helix
found in solution
11 bp per turn
right handed
A form DNA is more compacted
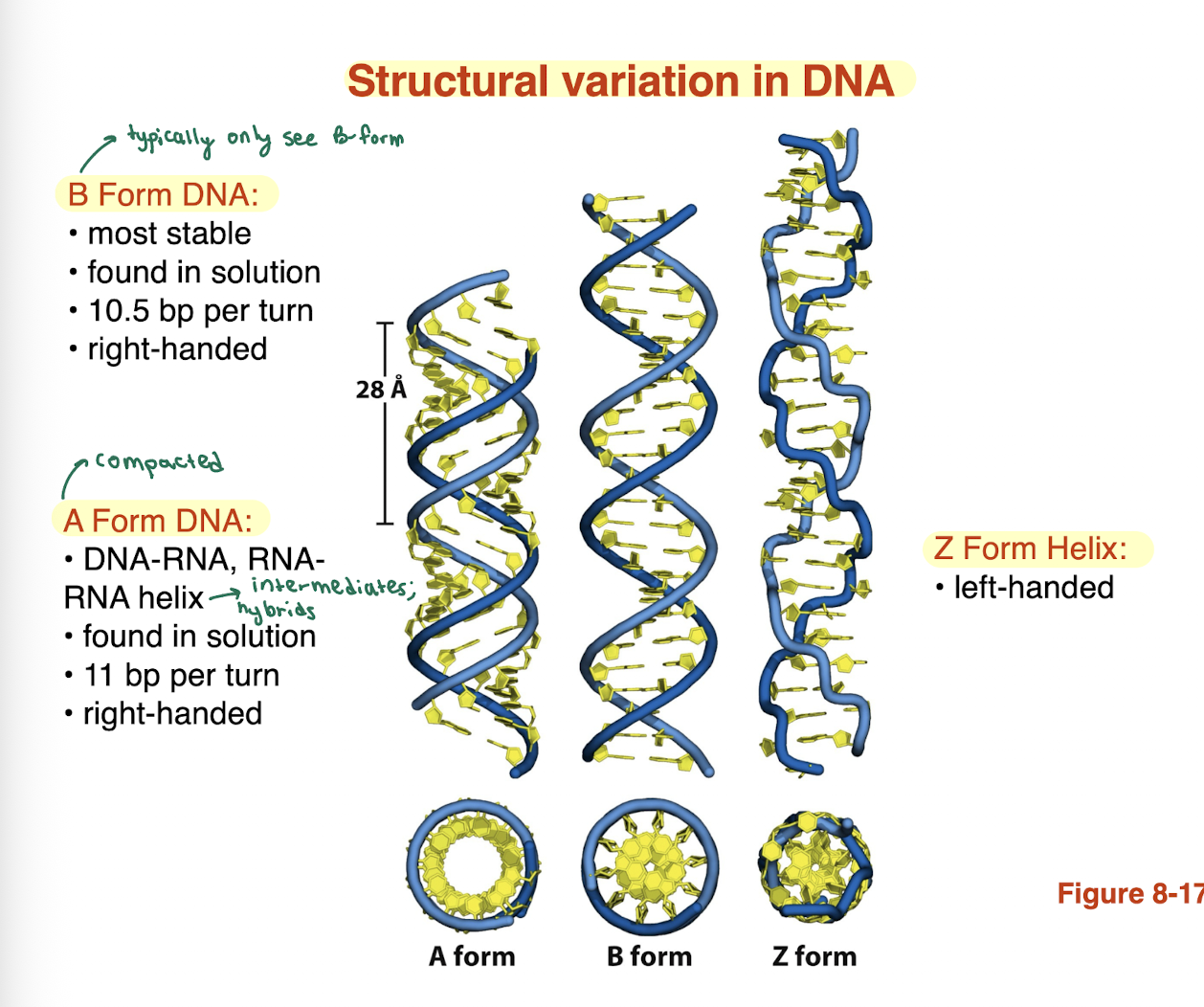
what is a Z form helix
left handed
what leads to the denaturation of DNA?
high temperature or pH
What is DNA denaturation and how is it measured?
Denaturation: DNA goes from double helix → random coil (single strands)
Measured by UV absorbance
Denaturation → ↑ UV absorbance
Due to loss of base stacking interactions
Called the hyperchromic effect
double helix has stacked bases so it would have LESS UV absorbance while single strand DNA will have more UV absorbance since it has unstacked bases and more exposure to light so it can absorb more
What is melting temperature (Tm) in regards to denaturation? it tells us what about DNA
Tm: temperature where 50% of DNA is denatured
Half helical, half random coil
Used to compare DNA stability
How does GC content affect DNA stability?
More G≡C → higher Tm → more stable DNA
Less G≡C → lower Tm → less stable DNA
On graph: curve with higher Tm = higher %GC
so higher amounts of GC require higher amounts of temperature to denature since the GC bp has 3 H bonds compared to 2. hence higher Tm compared to another can tell us that we have higher GC content than the other.
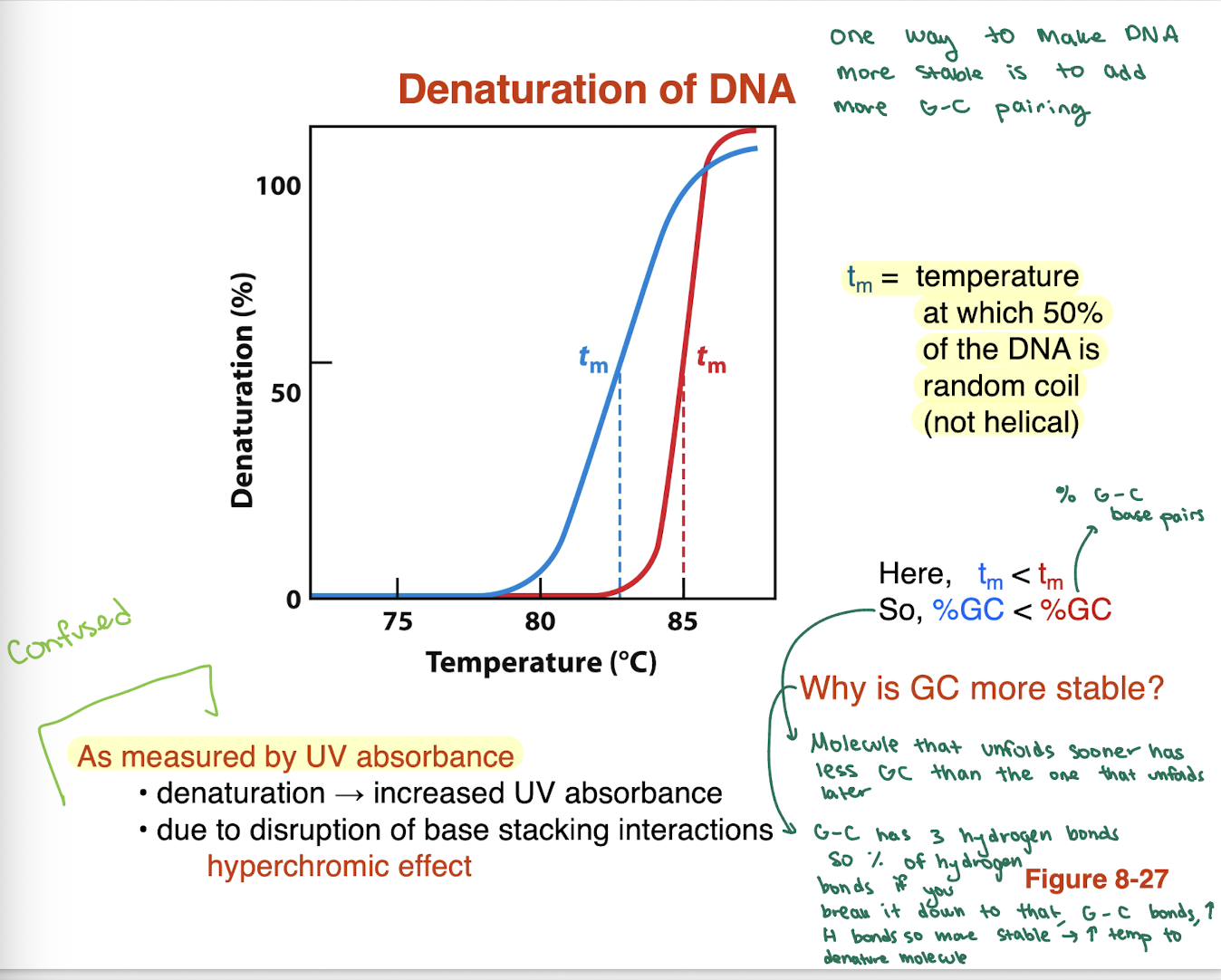
higher GC bp amount requires BLANK temperature to denature
higher
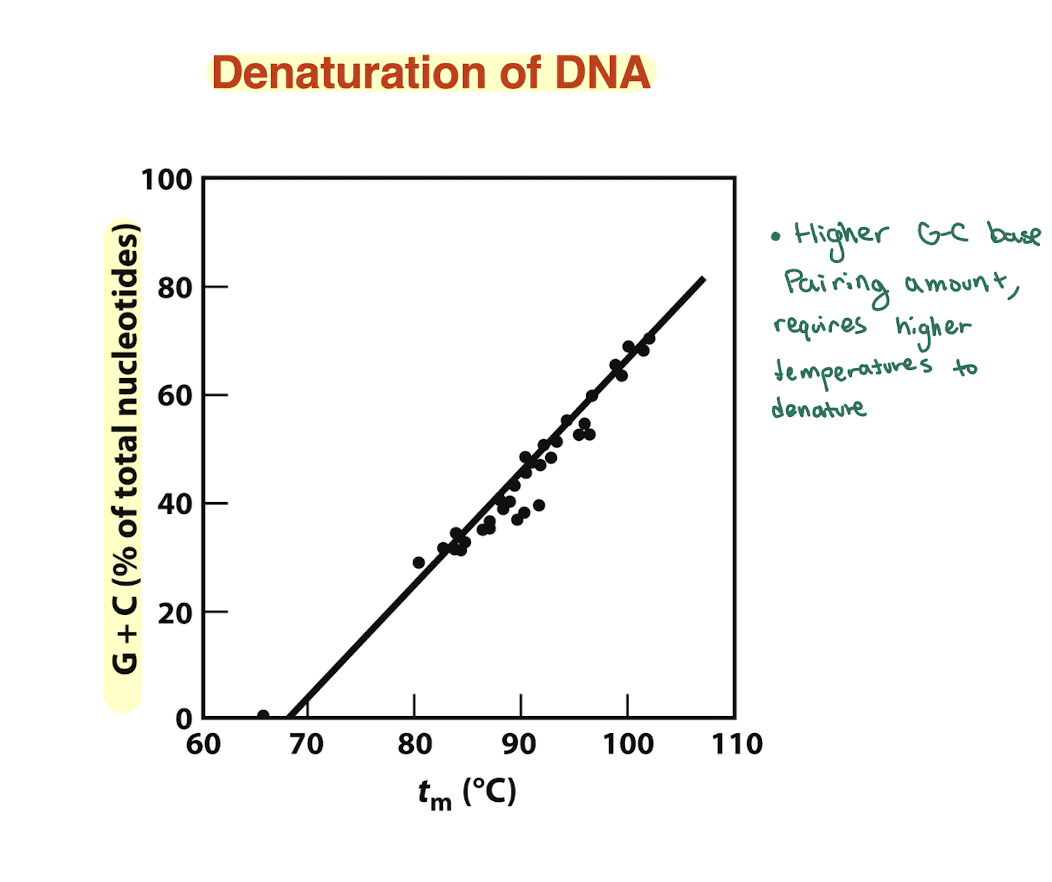
what is the overall structure of RNA?
single stranded molecule
forms a single stranded helix (not double helix like DNA)
can fold back on itself via base pairing
how does RNA form secondary structure?
intramolecular base pairing (within same strand)
creates double stranded regions
these regions adopt an A-form helix
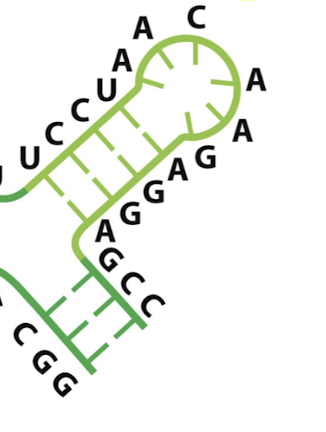
what secondary structure element is this?
hairpin
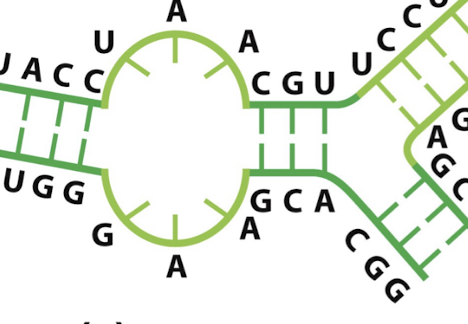
what secondary structure element is this?
internal loop
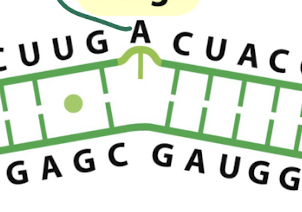
what secondary structure is this?
bulge: one or more unpaired bases on one side only
causes a distortion in the helix
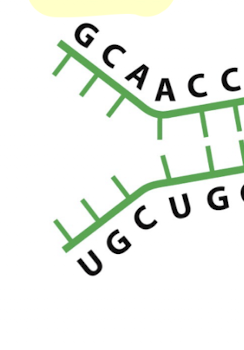
what secondary structure is this?
single stranded regions in RNA: regions with no base pairing
what type of helix do RNA double stranded regions form?
A form helix
more compact and rigid than B form DNA
occurs in RNA hairpins and stems
what base pairs occur in RNA?
canonical pairs: A-U, G-C
Non canonical pair : G-U (wobble)
G-U pairing helps RNA form complex secondary structures
what is RNase P?
an RNA enzyme (ribozyme) that cleaves RNA (tRNA processing)
how do DNA binding proteins recognize specific sequences?
Primarily through major groove interactions
Hydrogen bonding with exposed base edges
Major groove provides more chemical information than minor groove
→ Enables sequence-specific recognition without unwinding DNA
why is sequence specific DNA recognition important?
allows proteins to identify specific DNA sequences
essential for information transfer processes:
transcription
replication
DNA repaire
ensures correct genes are regulated or accessed
How does the major groove enable sequence-specific recognition?
Distinct donor/acceptor patterns for each base pair
Can distinguish:
AT vs TA
GC vs CG
Provides maximum chemical information
→ Main site for DNA-binding protein recognition
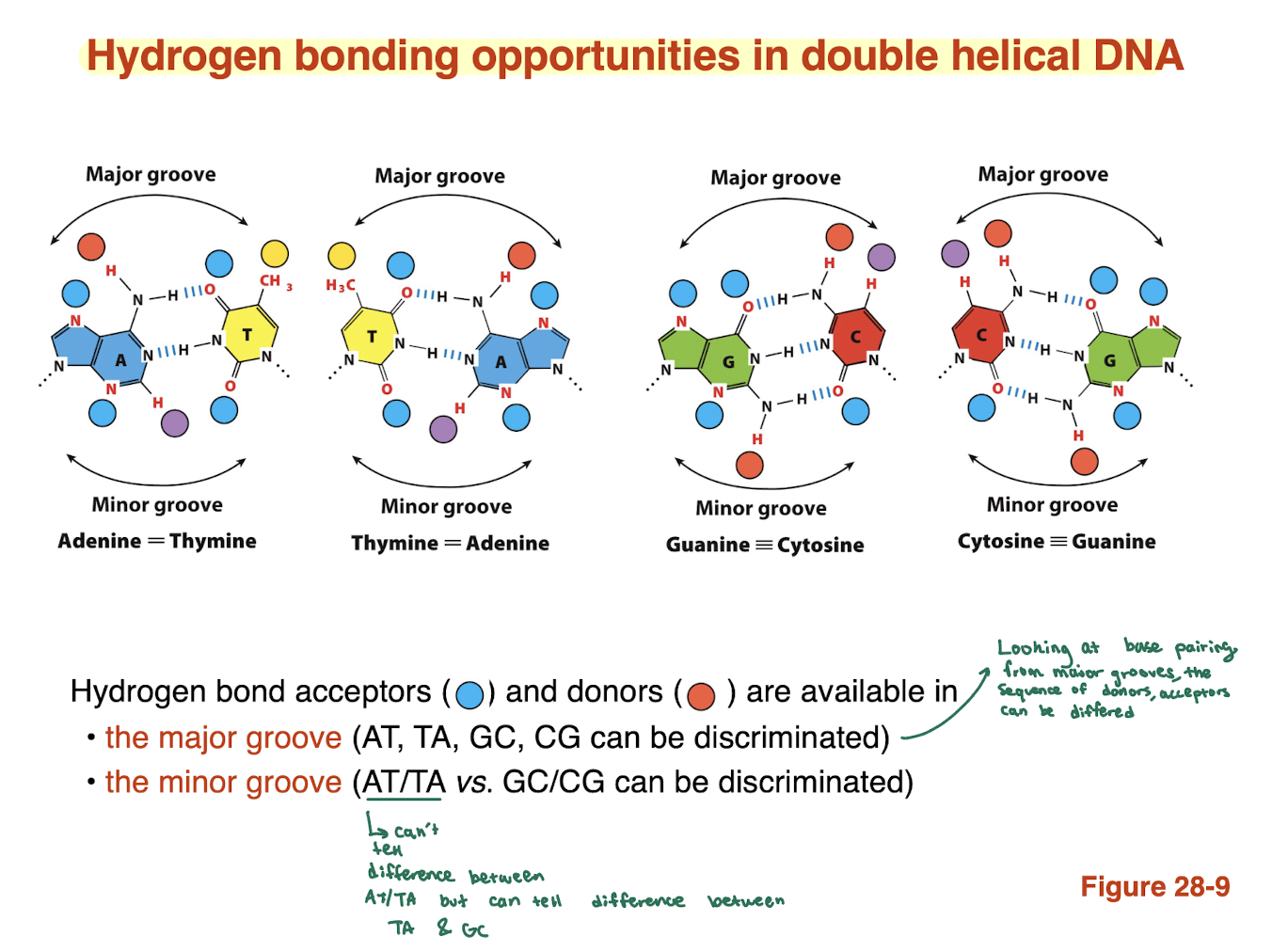
What information can be obtained from the minor groove?
Limited chemical information
Can distinguish:
AT/TA vs GC/CG
Cannot distinguish AT from TA
→ Less useful for precise sequence recognition
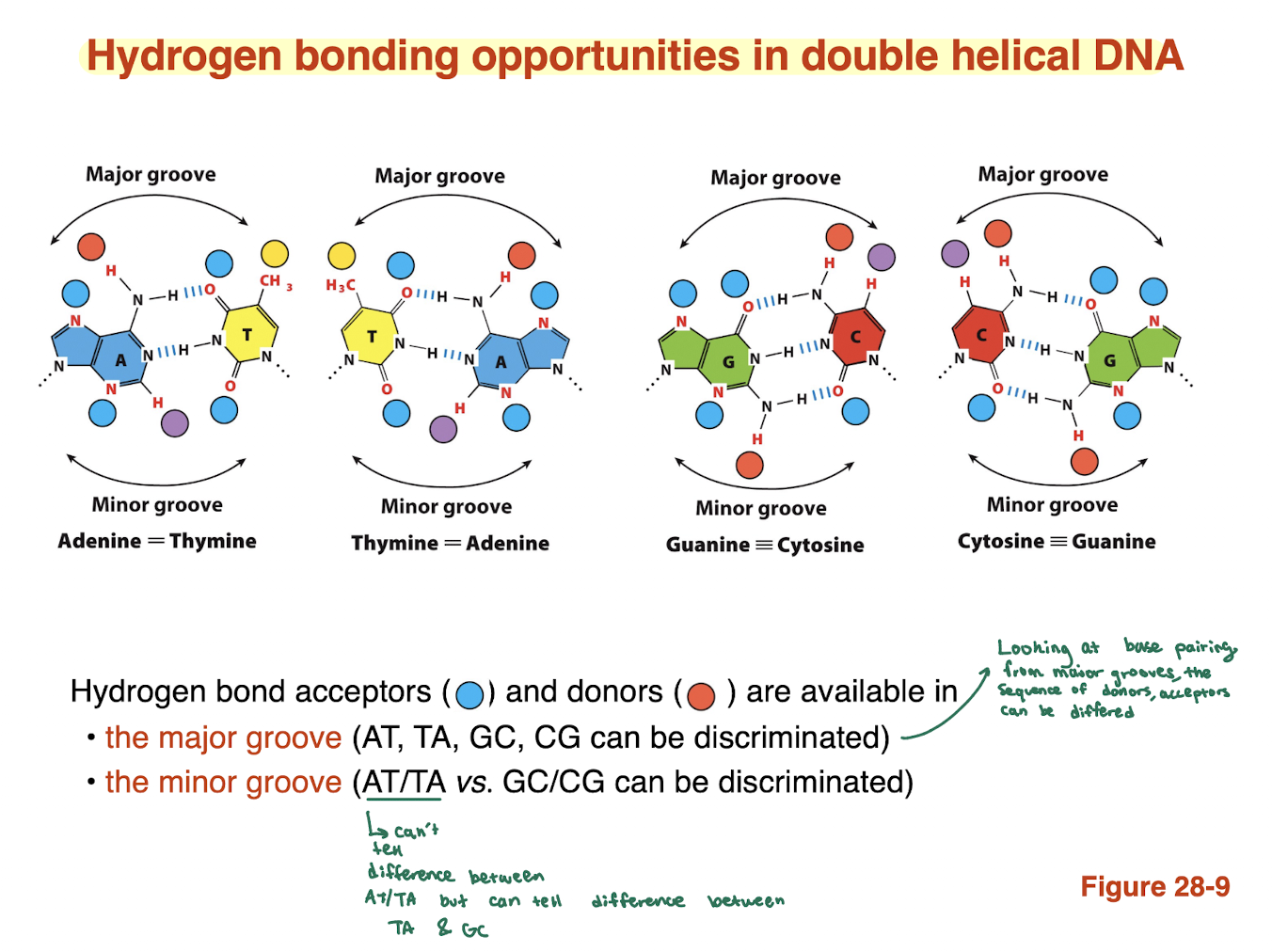
Why is the major groove more important than the minor groove?
Major groove exposes unique H-bonding patterns for each base pair
Minor groove patterns are more similar/redundant
→ Proteins bind major groove for accurate sequence reading
How do proteins recognize DNA sequences?
Amino acid side chains form hydrogen bonds with DNA bases
Interact with donor/acceptor groups in grooves (mainly major groove)
Recognition depends on matching H-bond patterns
→ Enables sequence-specific DNA binding
Which amino acids commonly interact with DNA bases? what is the mneomnic we can use?
Asn (asparagine)
Gln (glutamine)
Glu (glutamate)
Lys (lysine)
Arg (arginine)
These side chains can act as H-bond donors and/or acceptors
“Naughty Queens Grab Large Apples”
N → Asn (Asparagine)
Q → Gln (Glutamine)
G → Glu (Glutamate)
L → Lys (Lysine)
A → Arg (Arginine)
What determines sequence specificity in DNA-protein interactions?
Each base pair has a unique H-bonding pattern
Proteins use a specific arrangement of amino acids
Must match the pattern of bases in sequence order
→ “Reading” DNA = matching amino acid side chains to base patterns
DNA binding proteins typically bind where?
Generally bind DNA in the major groove — where it can recognize the sequence the beest
alpha helix fits nicely into the wide major groove
certain DNA binding motifs are common
helix turn helix
zinc finger
homeodomina
leucine zipper
basic helix loop helix (BHLH)
What is the helix–turn–helix DNA-binding motif and how does it recognize DNA?
Protein motif with two α-helices connected by a turn
One helix = recognition helix
Recognition helix fits into the major groove of DNA
Amino acid side chains interact with base pairs via H-bonding
→ Allows protein to read specific DNA sequences without unwinding DNA
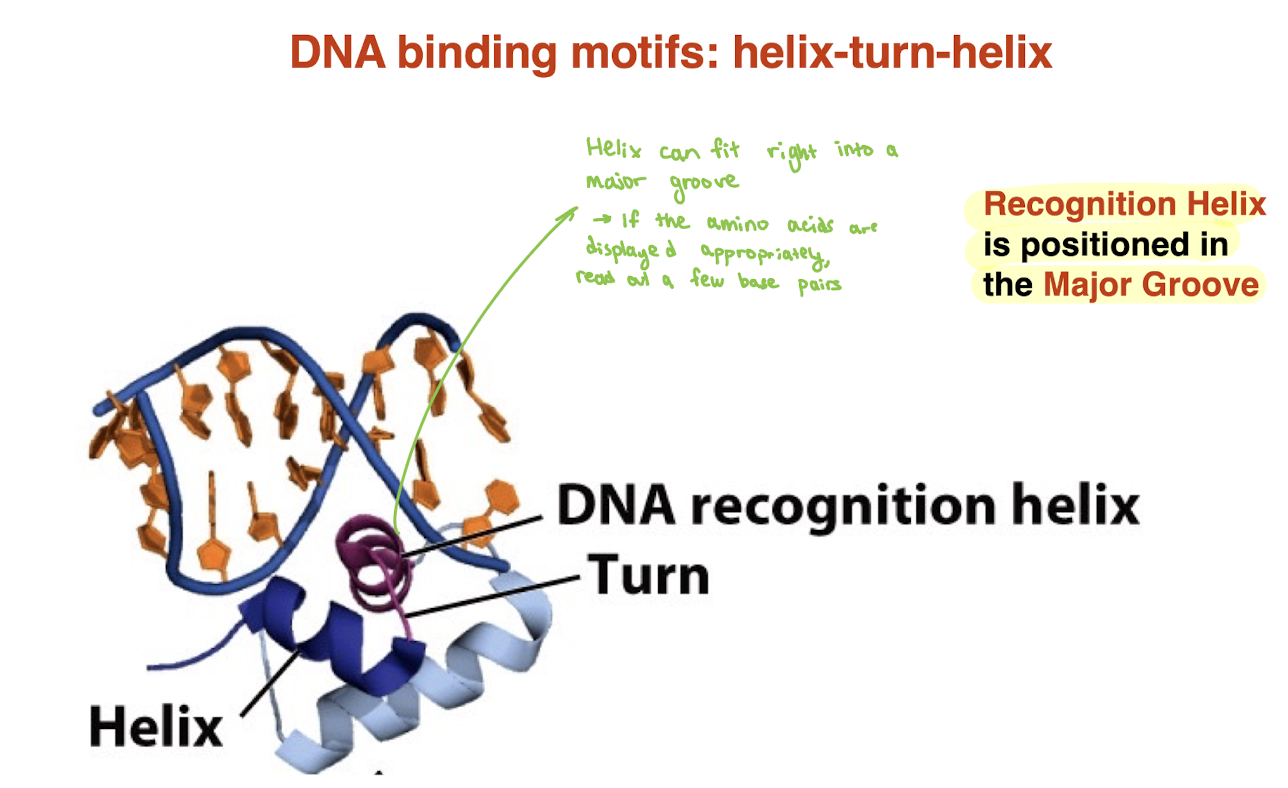
What is the zinc finger DNA-binding motif and how does it recognize DNA?
Protein motif stabilized by a coordinated Zn²⁺ ion
Zn²⁺ held by 4 Cys OR 2 Cys + 2 His residues
Contains a recognition helix that inserts into the major groove
Each “finger” recognizes a few base pairs
Multiple zinc fingers can be linked in tandem
→ Allows recognition of longer, specific DNA sequences
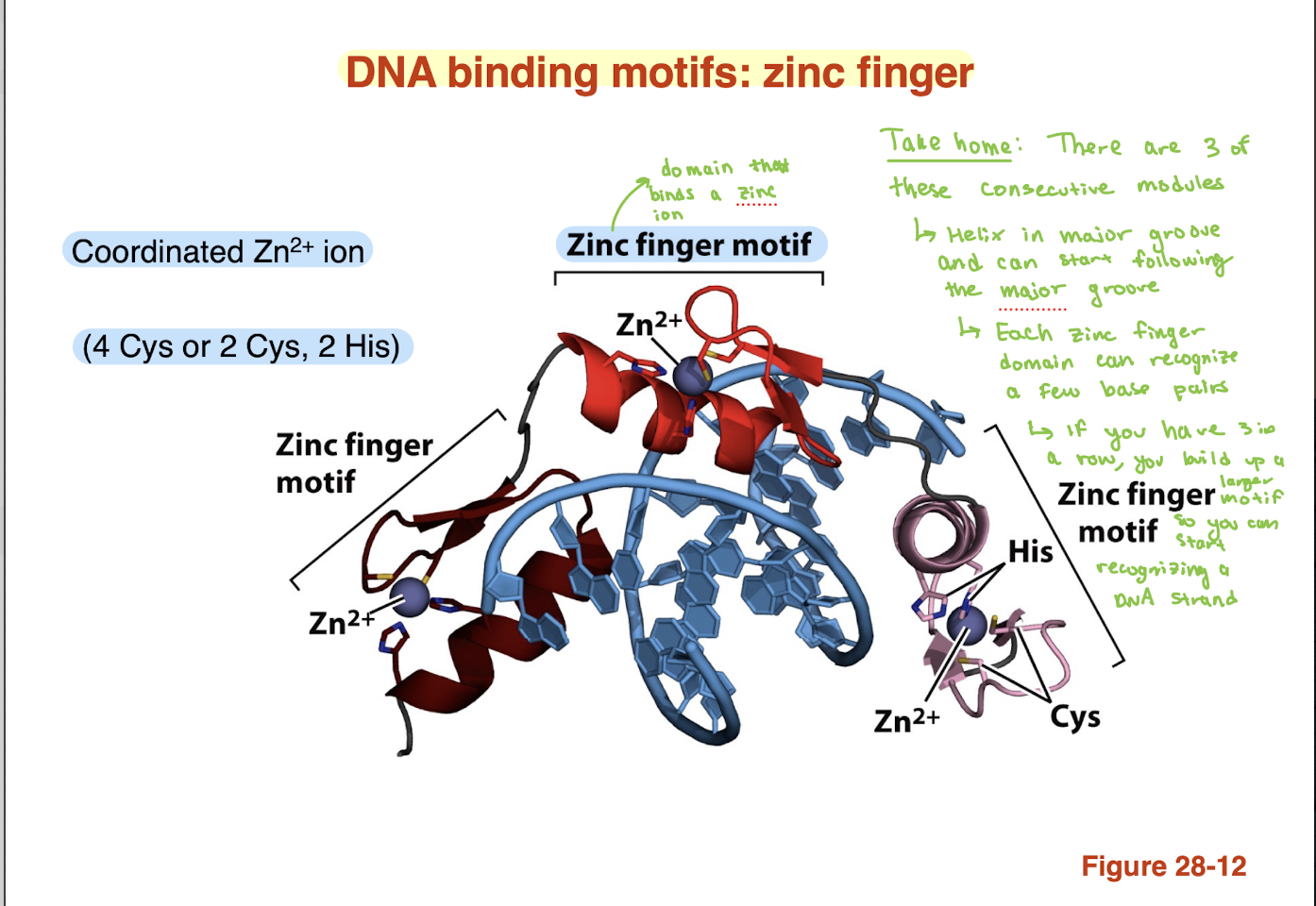
What is the homeodomain DNA-binding motif and how does it recognize DNA?
Homeodomain: conserved DNA-binding motif (~60 amino acids)
Contains helix–turn–helix structure
One helix = recognition helix
Recognition helix inserts into the major groove
Amino acid side chains form H-bonds with bases
→ Enables sequence-specific DNA binding (gene regulation
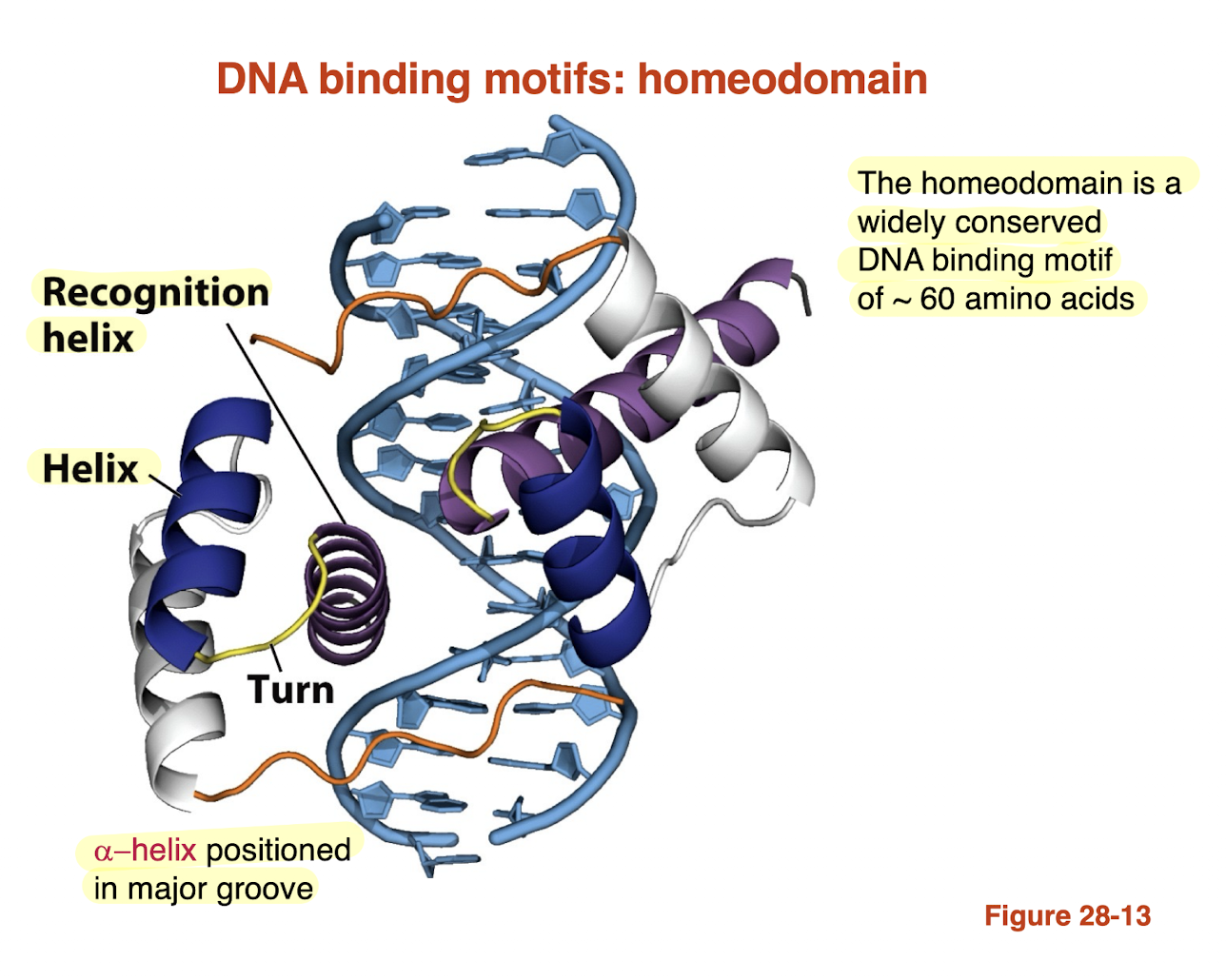
What is the leucine zipper DNA-binding motif and how does it bind DNA?
Leucine zipper: motif with repeating Leu residues that form a coiled-coil dimer (dimerization)
Two helices “zip” together → dimer formation
DNA-binding region contains basic residues (Lys, Arg)
These helices insert into the major groove
Protein binds DNA by clamping onto both sides
→ Combines dimerization (Leu) + DNA recognition (Lys/Arg in major groove)
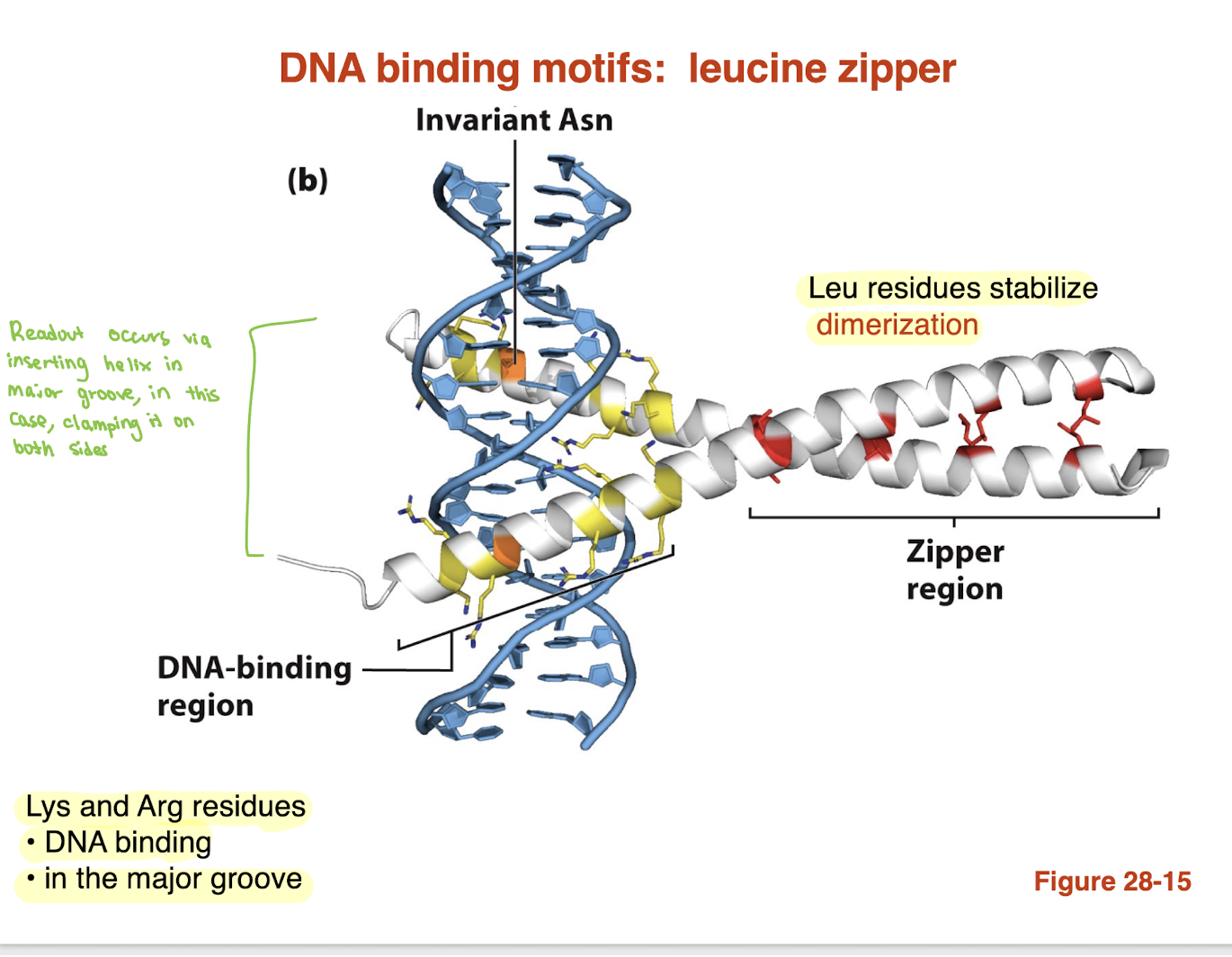
What is the helix–loop–helix (HLH) DNA-binding motif and how does it bind DNA?
Consists of two α-helices connected by a loop
One helix = recognition helix (binds DNA)
Other helix = dimerization helix
Amphipathic helices allow dimer formation
Basic residues (Lys, Arg) mediate DNA binding
Recognition helix inserts into the major groove
→ Combines dimerization + sequence-specific DNA binding
DNA molecules can be very long hence DNA must be BLANK to fit into cells
compacted
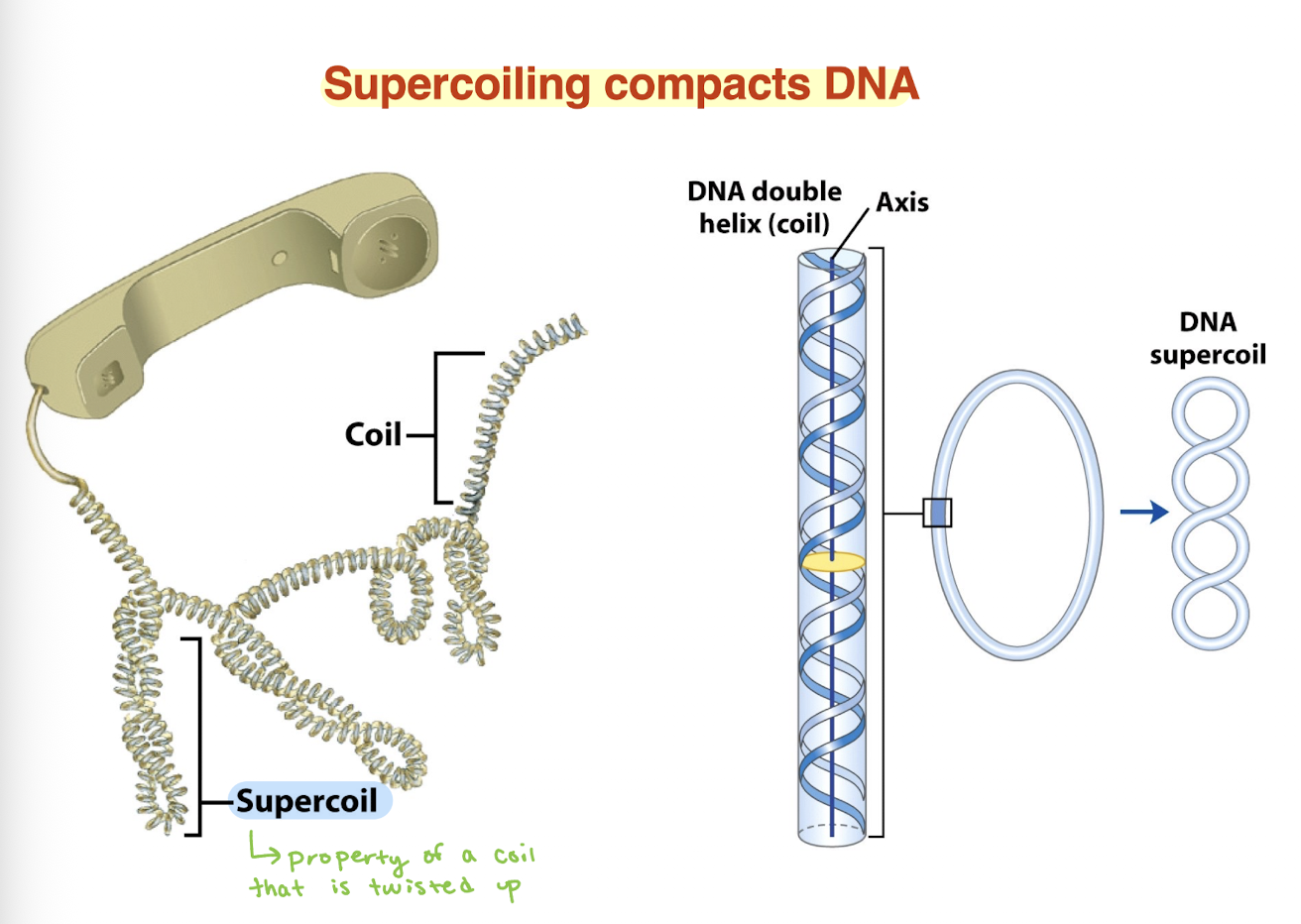
how does supercoiling relieve torsional strain in DNA?
Separating DNA strands creates torsional strain (overwinding)
DNA cannot freely rotate → strain builds up
DNA relieves this by forming supercoils (writhe)
Supercoiling converts twist → coiling in space
→ Lowers overall energy and relieves strain
AKA DNA supercoils to release twisting stress
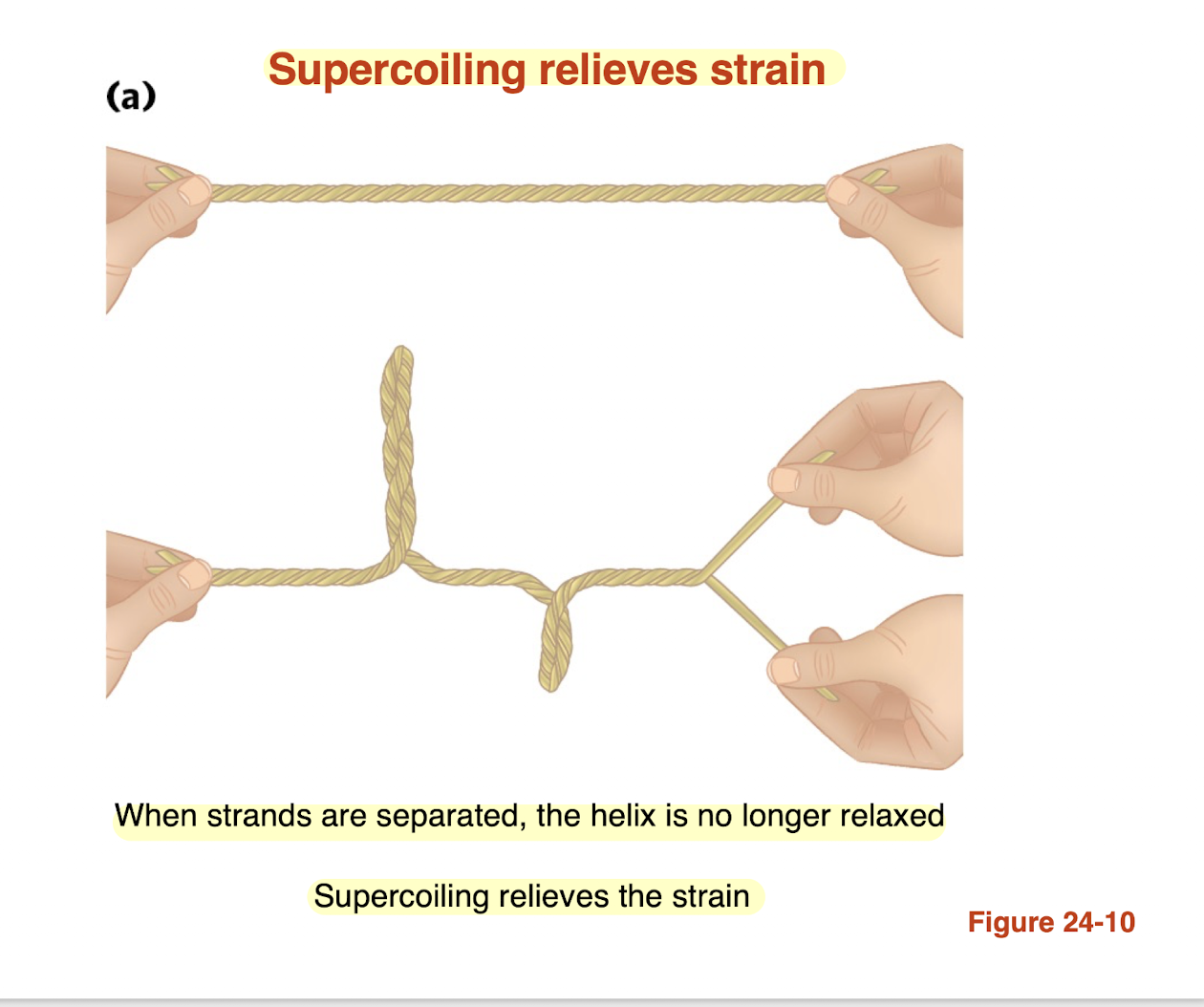
polymerase does separates the strands during replication and introduces BLANK
Strain
Supercoiling is an intrinsic property of DNA’s BLANK structure
supercoiling is an intrinsic property of DNA’s tertiary structure
it occurs in all cellular DNAs and is highly regulated by each cell
what does it mean for DNA to be underwound?
fewer turns than relaxed DNA which creates torsional strain or less tightly twisted
Relaxed DNA: ~10.5 bp/turn
Underwound DNA: more bp/turn (e.g., ~12 bp/turn)
how does supercoiling stabilize underwound DNA?
Underwound DNA is energetically strained
DNA forms supercoils to compensate
Converts underwinding (twist) → writhe (coils)
→ Stabilizes the molecule by lowering energy
Why is underwound DNA biologically useful?
Facilitates strand separation
Makes DNA easier to open
Important for:
Replication
Transcription
What is the linking number (Lk) in DNA?
Lk = number of times one DNA strand wraps around the other
Applies to closed (circular) DNA
Represents total helical turns
→ Topological property (cannot change without breaking DNA)
How are Lk, ΔLk, and superhelical density (σ) related in DNA?
Lk = # base pairs ÷ bp per turn
Example: 2100 bp ÷ 10.5 = Lk₀ = 200 (relaxed DNA)
ΔLk = Lk − Lk₀
Underwound example: ΔLk = −2 → Lk = 198
σ (superhelical density) = ΔLk / Lk₀
Example: σ = −2 / 200 = −0.01
Key points:
Nick (strand break) → Lk undefined
Negative ΔLk = underwound DNA
underwound DNA is what kind of supercoil?
negative
overwound DNA is what kind of supercoil?
positive
what are topoisomers?
two forms of a circular DNA that differ only in a topological property such as linking number are referred to as topoisomers
What does topoisomerase I do?
changes Lk in increments of 1
cleaves one DNA strand
relaxes positive and negative supercoils
does NOT require ATP
Example: Topoisomerase I
what does Topoisomerase II do?
changes Lk in increments of 2
cleaves both DNA strands
relaxes positive and negative supercoils
can introduce negative supercoils (prokaryotes)
requires ATP (hydrolyzes ATP)
examples: DNA gyrase, Topoisomerase II
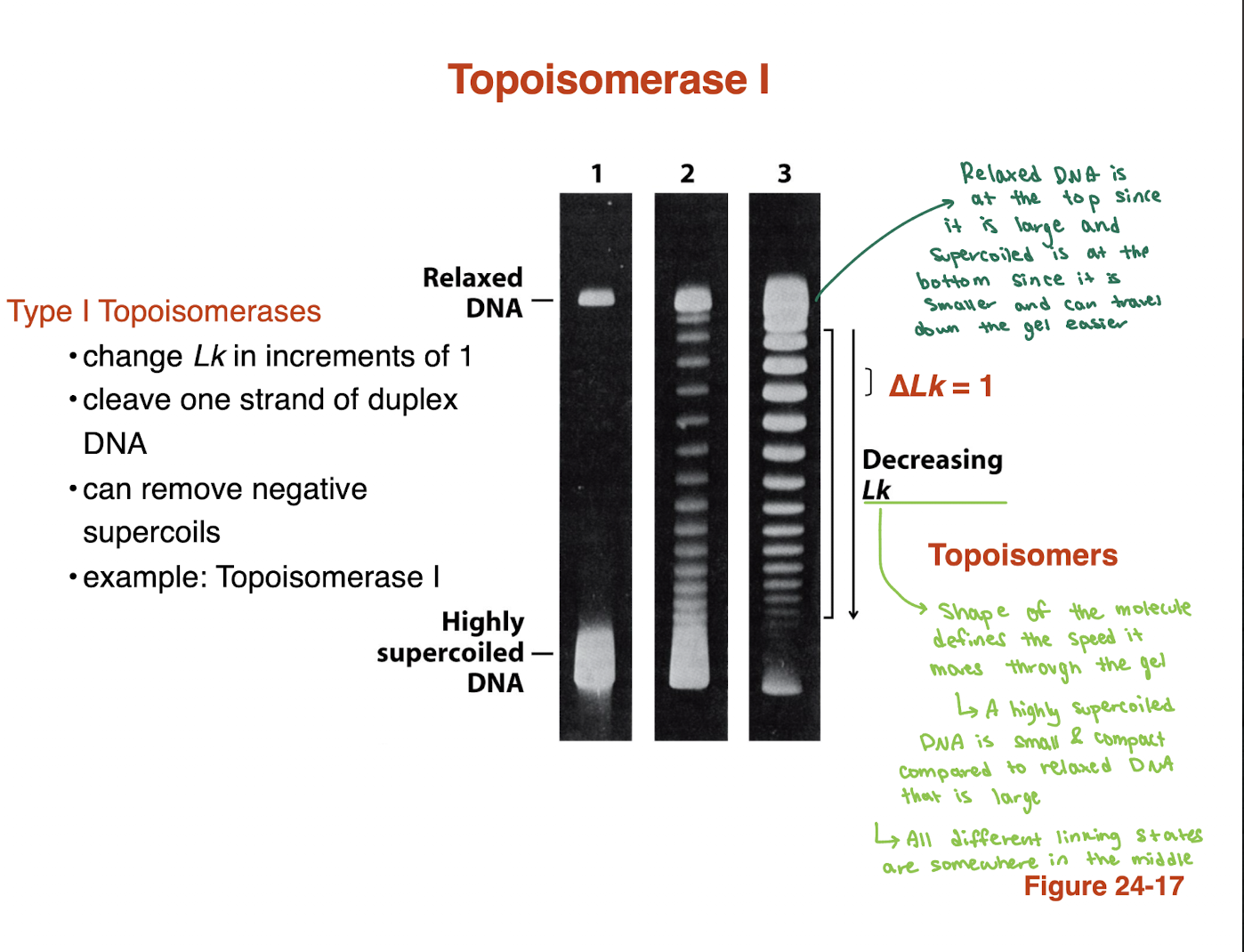
How does DNA supercoiling affect migration in gel electrophoresis?
Highly supercoiled DNA → more compact → travels faster (lower on gel)
Relaxed DNA → more open/large → travels slower (higher on gel)
Different bands = topoisomers (different Lk values)
Topoisomerase I produces a ladder with ΔLk = 1 between bands
👉 Key idea: Shape (compact vs relaxed), not length, determines migration speed
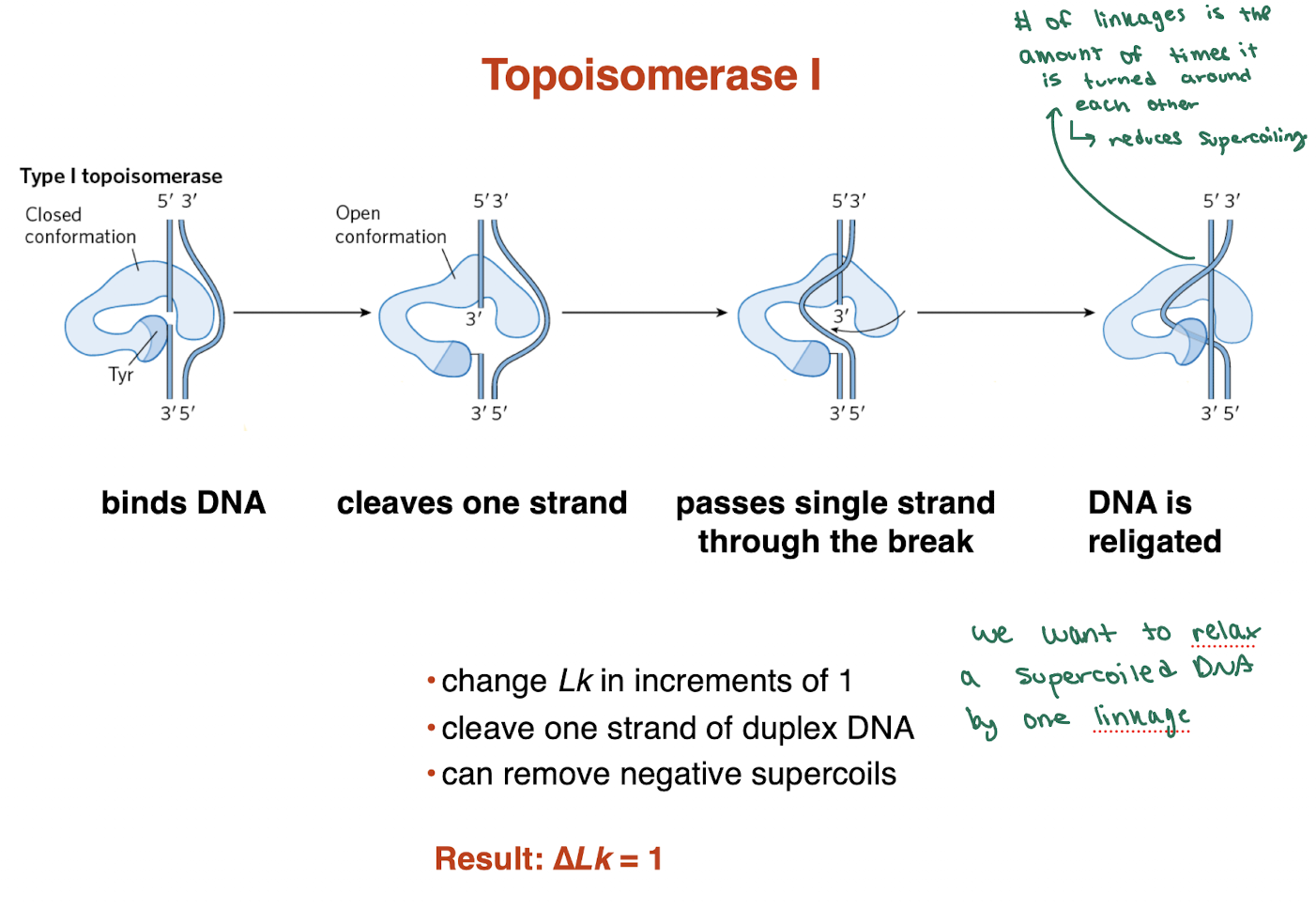
What is the mechanism of Topoisomerase I?
Binds DNA
Cleaves one strand (via Tyr residue)
Passes the uncut strand through the break
Religates DNA
→ Relieves supercoiling without fully breaking the helix
How does Topoisomerase I change DNA topology?
remove supercoils (especially negative supercoils)
reduces torsional strain
helps maintain DNA stability during replication/transcription
What is the mechanism of Topoisomerase I?
Binds DNA
Cleaves one strand (via Tyr residue)
Passes the uncut strand through the break
Religates DNA
→ Relieves supercoiling without fully breaking the helix
How does Topoisomerase I change DNA topology?
Changes Lk in increments of 1
ΔLk = ±1 per cycle
Gradually relaxes supercoiled DNA
→ One “turn” removed at a timecan remove negative supercoils
What is the function of Topoisomerase I?
Removes supercoils (especially negative supercoils)
Reduces torsional strain
Helps maintain DNA stability during replication/transcription
TOPOISOMERASE I ALWAYS MOVES DNA TOWARD LK=0
draw topoisomerase I reaction
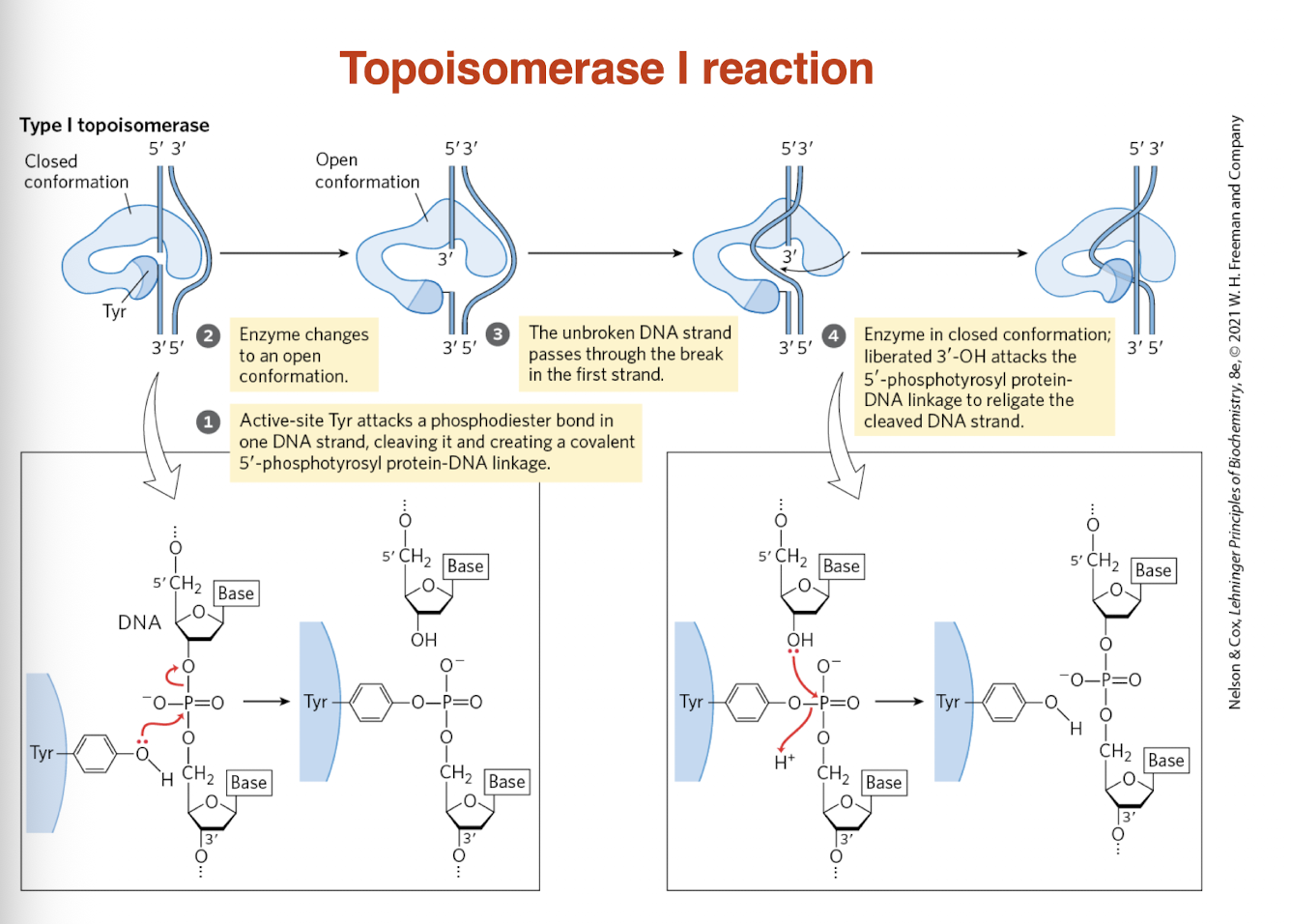
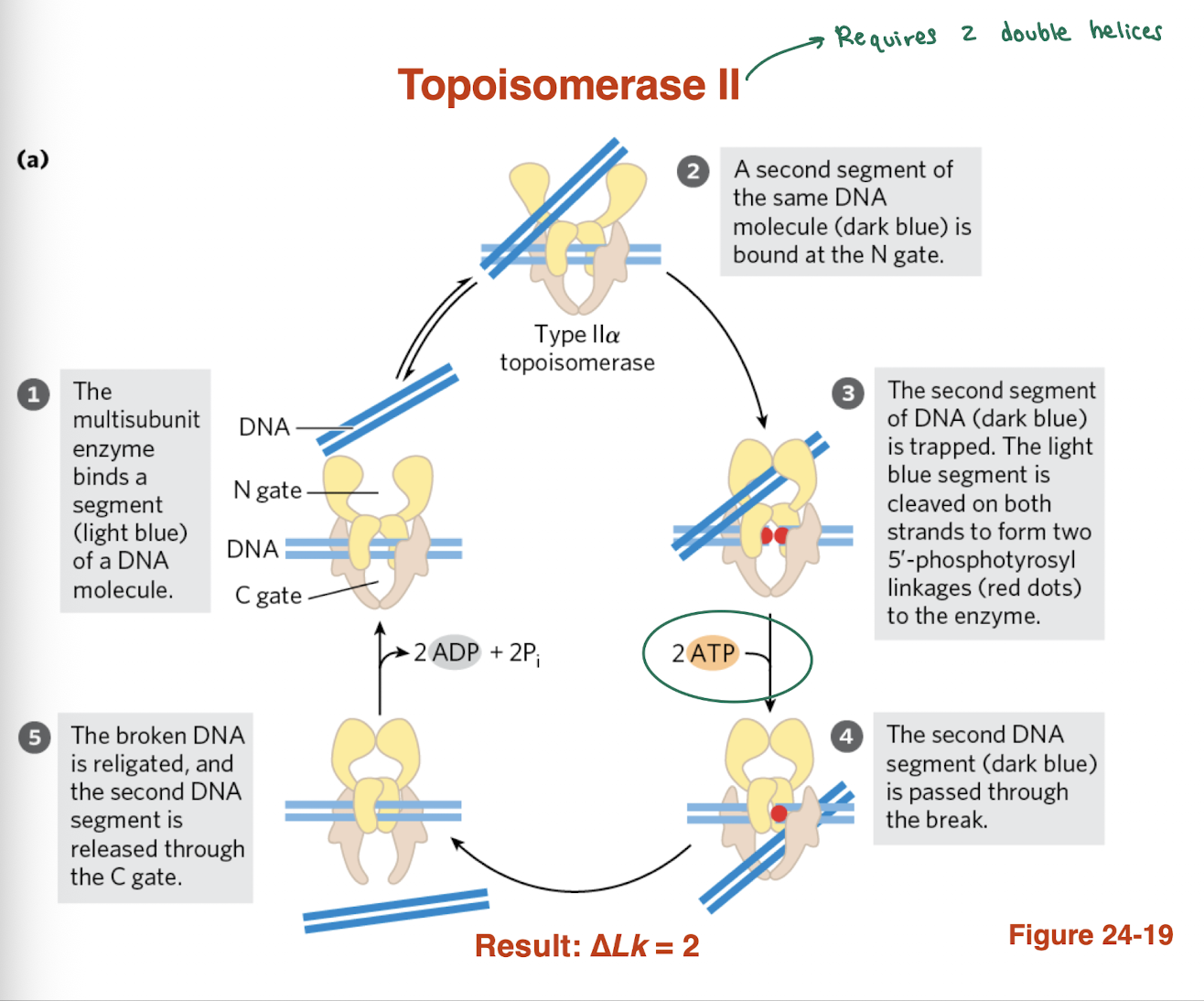
what is the mechanism of Topoisomerase II? what does it require?
Binds two DNA double helices (segments)
Cleaves both strands of one DNA segment
Uses ATP (2 ATP hydrolyzed)
Passes second DNA segment through the break
Religates the cut DNA and releases strand
→ Changes Lk in increments of 2 (ΔLk = ±2)
→ Can relieve supercoils and introduce negative supercoils (prokaryotes)
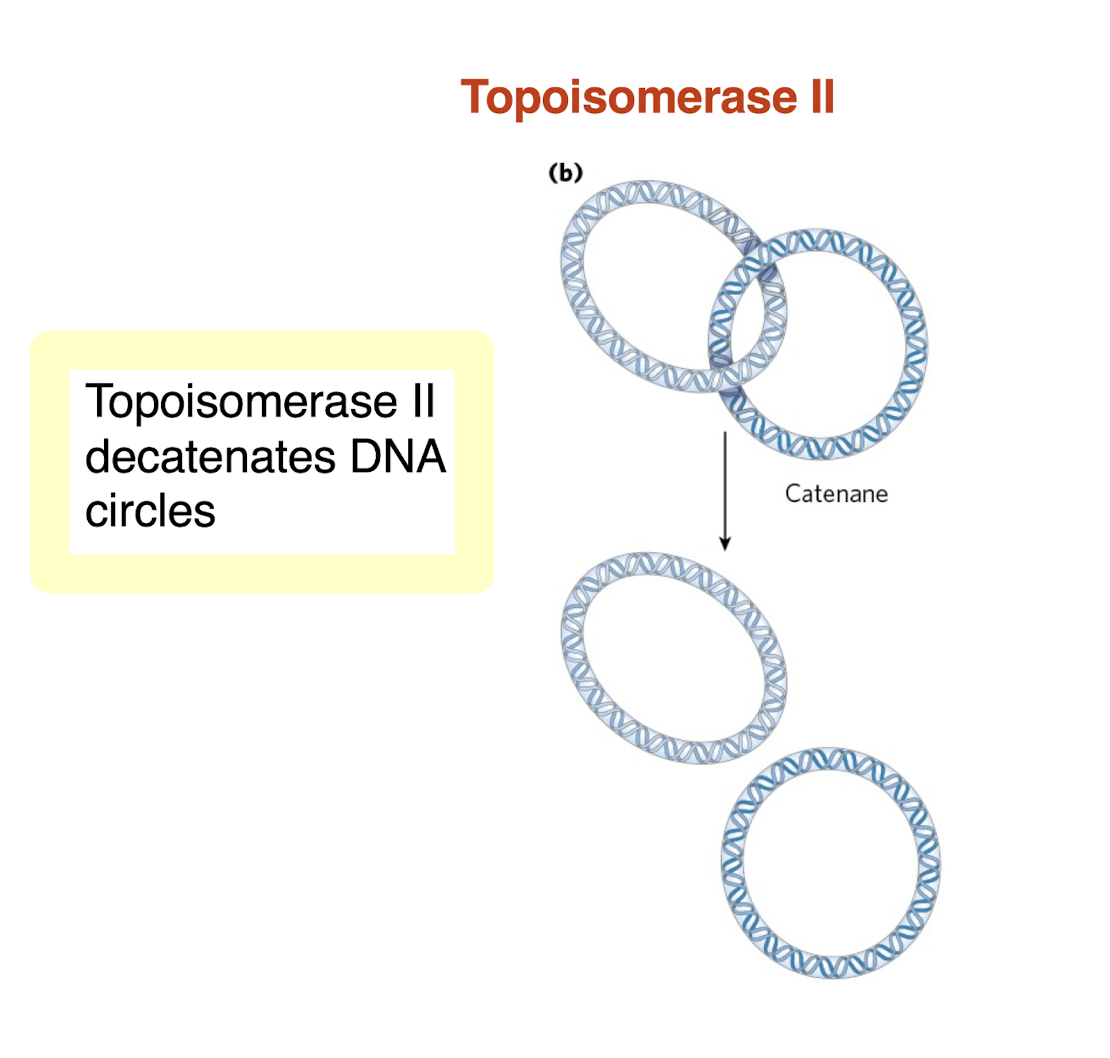
what does topoisomerase III do in DNA decatenation?
decatenates or separates interlinked circular DNA molecules
What is plectonemic supercoiling and what is its limitation?
Plectonemic supercoiling: DNA coils around itself forming interwound helices
Creates supercoil axis and branch points
Common form of DNA supercoiling in cells
Limitation:
Does NOT compact DNA enough for full cellular packaging
→ Additional structures (e.g., chromatin) are needed for higher-order compaction
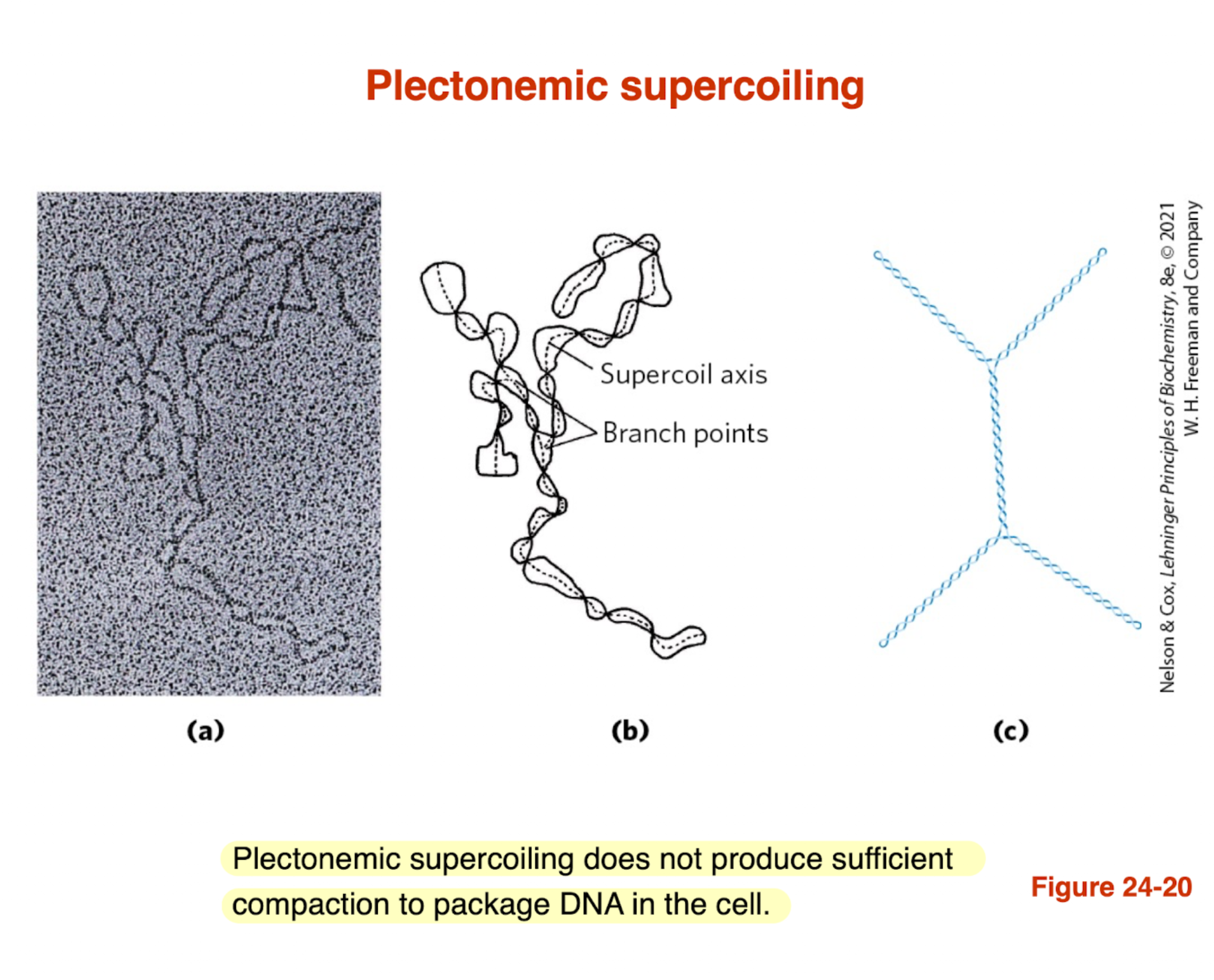
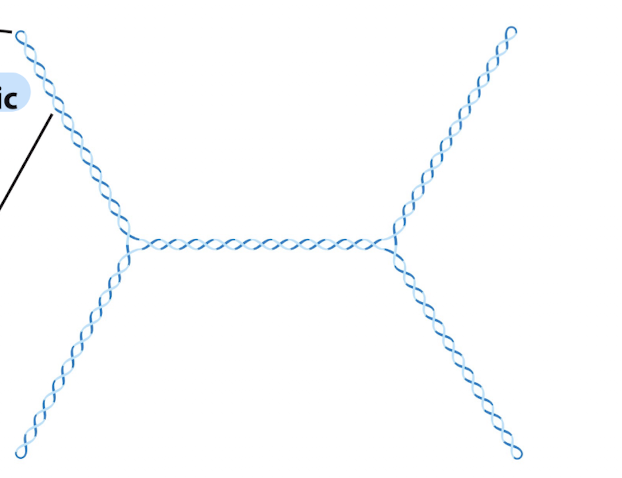
this is an example of what kind of supercoiling and it is MORE or LESS compact?
less compact and plectonemic
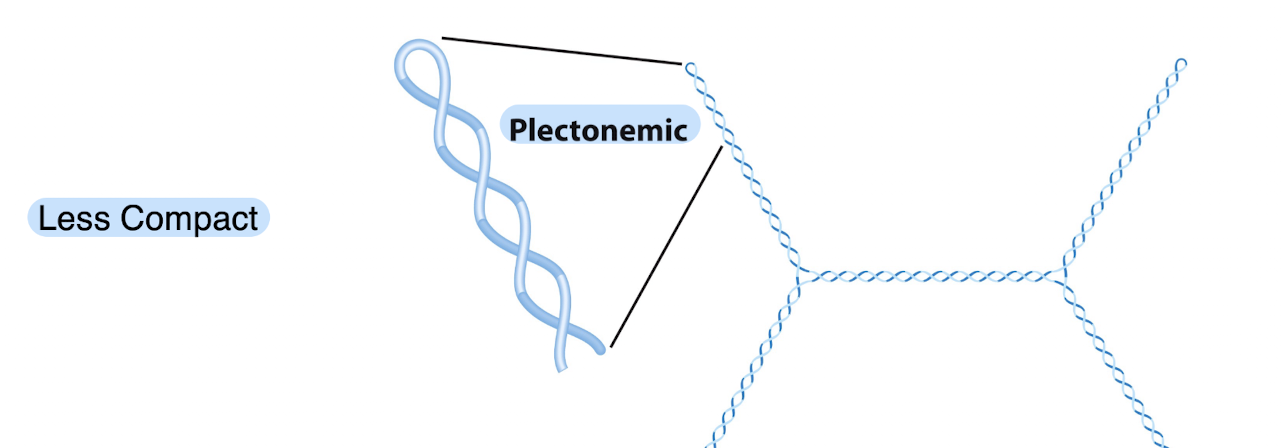

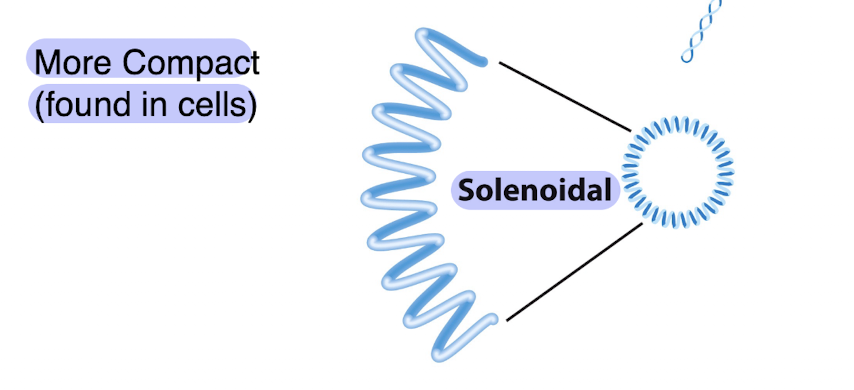
what kind of supercoiling is this?
solenoidal and more compact — found in cells
What is a nucleosome and how does it package DNA?
Nucleosome = DNA wrapped around a histone core
Forms “beads on a string” structure
→ Basic unit of DNA packaging in eukaryotes
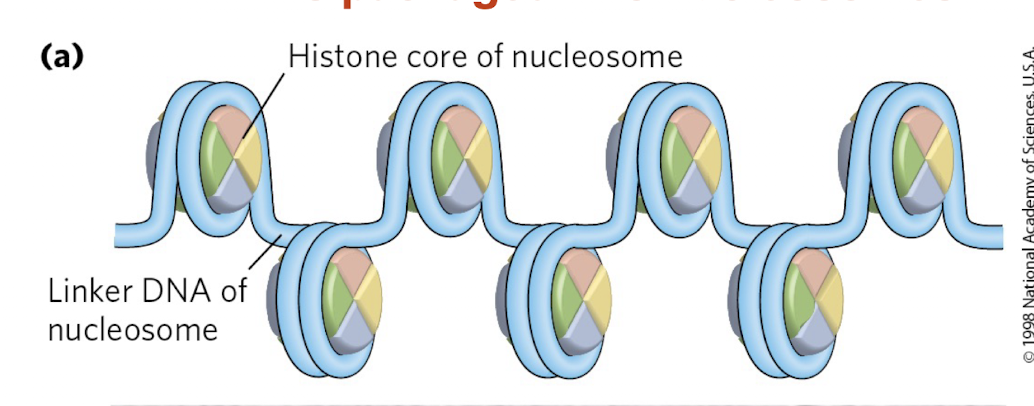
What is the role of histones and linker DNA in chromatin?
Histones: proteins that compact DNA
Linker DNA: stretches between nucleosomes
Allows DNA to be organized and further compacted
👉 Key idea: Histones enable DNA to fit inside the cell while maintaining structure
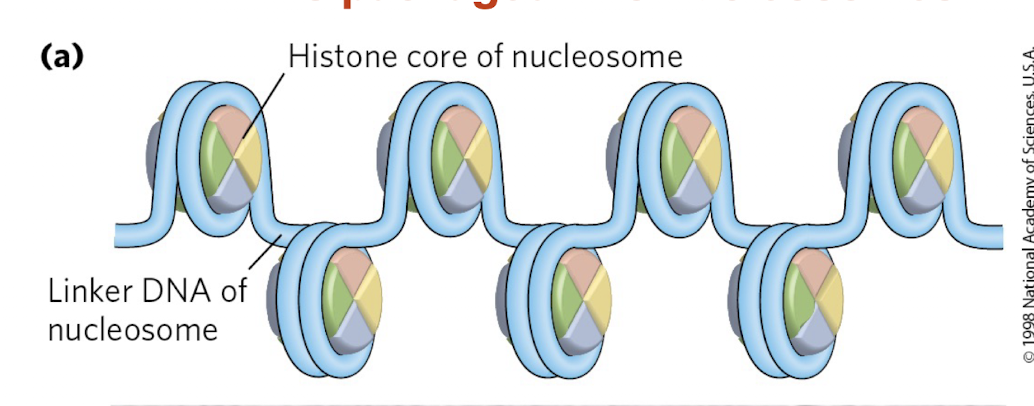
histones are found in BLANK of all eukaryotic cells?
chromatin
what is histones main purpose as a protein?
main purpose of histones is to allow DNA to wrap around it so it needs to have a surface that DNA likes
What is the nucleosome core particle? — not sure if high yield but just konw
Core made of 8 histone proteins (histone octamer):
2 × H2A, 2 × H2B, 2 × H3, 2 × H4
~146 bp of DNA wrapped around the histone core
DNA wraps ~1.7 turns
146 bp of DNA wrapped around histone care
→ Fundamental unit of chromatin structure and DNA packaging
essentially nucleosome core particle is just DNA or 146 bp wrapped around a histone protein
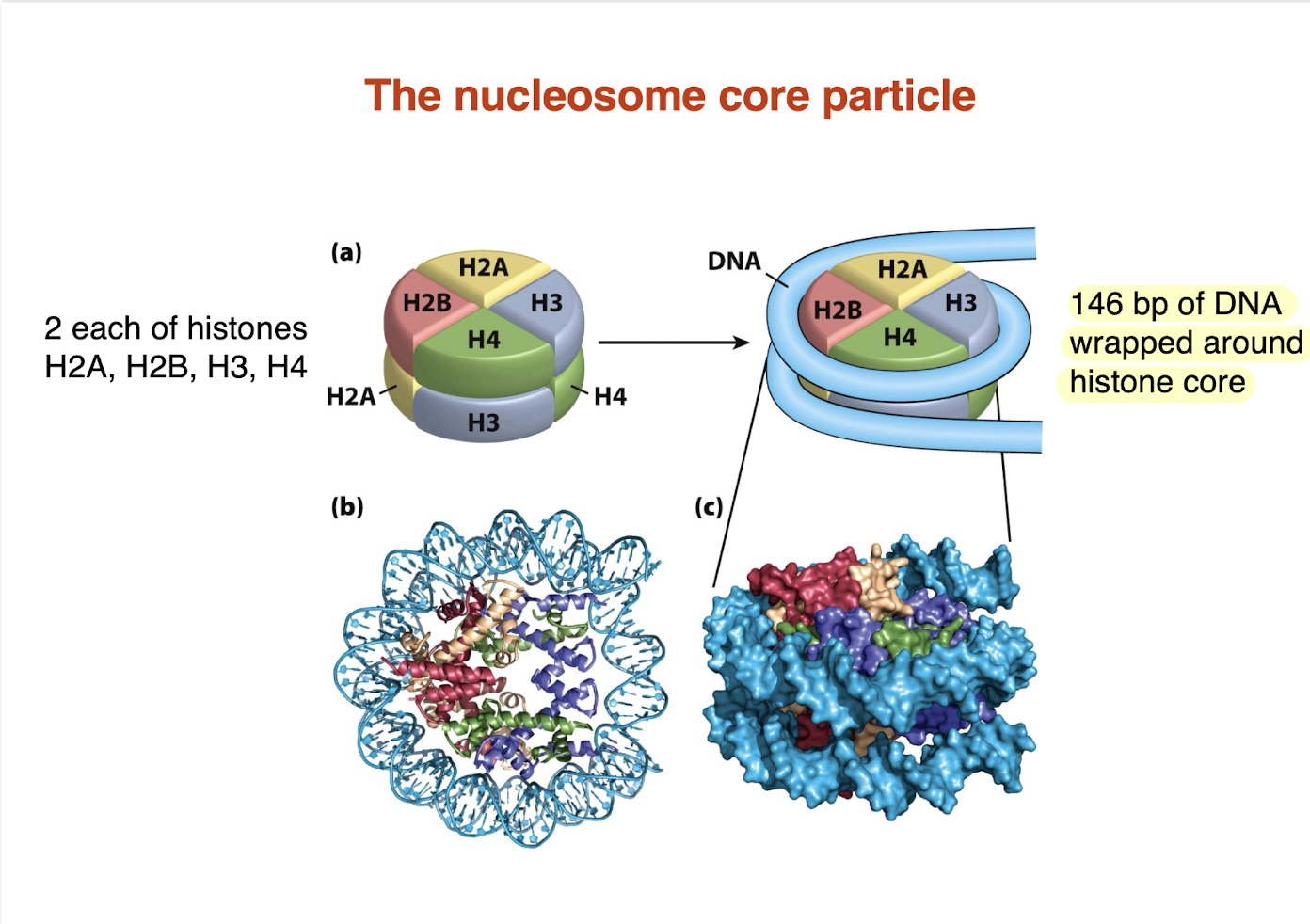
What are histone tails?
Amino-terminal (N-terminal) extensions of histone proteins
Protrude outward from the nucleosome core
Flexible and accessible outside the DNA-histone structure
“amino terminal tails of histone proteins that produce from the core particles. these tails are extensively post translationally modified and also paritpcien in DNA packaging“
What is the function of histone tails?
Extensively post-translationally modified (e.g., acetylation, methylation)
Help regulate DNA packaging (tight vs loose chromatin)
use tails for information, to define where we are, and change information on tails
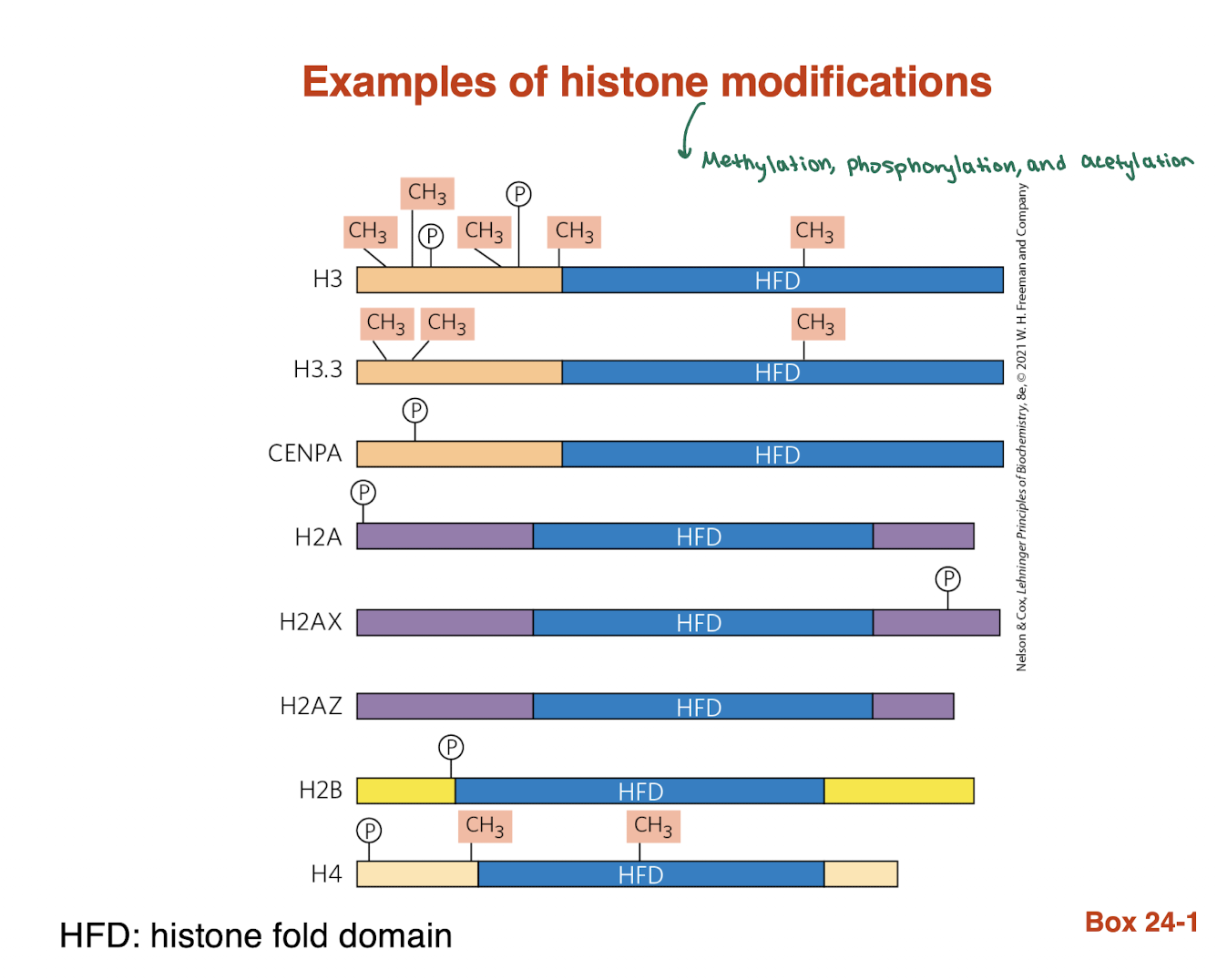
what are examples of histone modifications?
methylation, phosphorylation, and acetylation
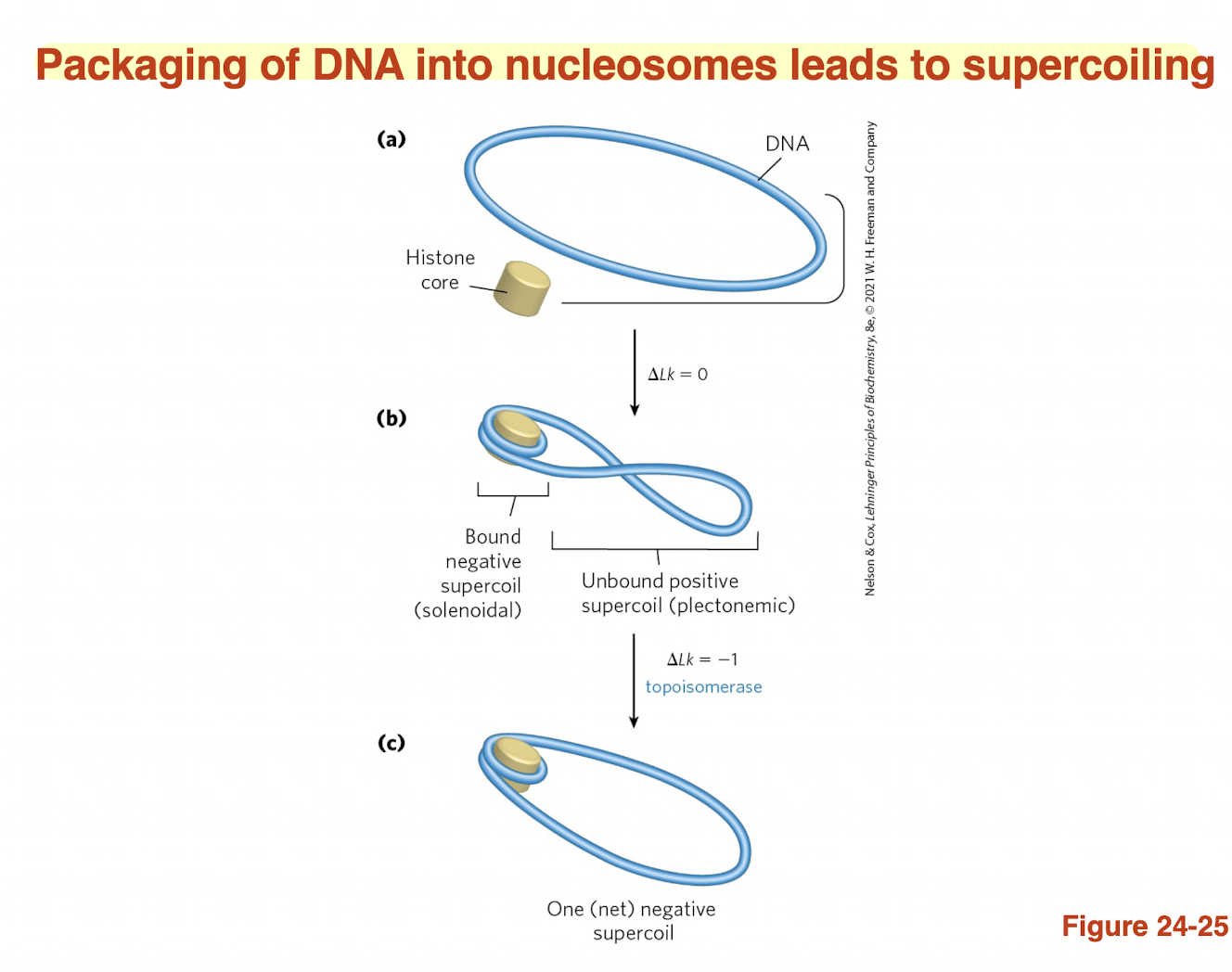
packaging of DNA into nucleosomes leads to BLANK?
supercoiling
How is DNA organized at a higher-order level beyond nucleosomes?
DNA is organized into looped domains
Loops are anchored to a protein scaffold (chromosomal scaffold)
This scaffold becomes visible when histones are removed
Helps compact DNA and organize chromosomes efficiently
Key idea: DNA is not random → it is structured into loops for packing + regulation
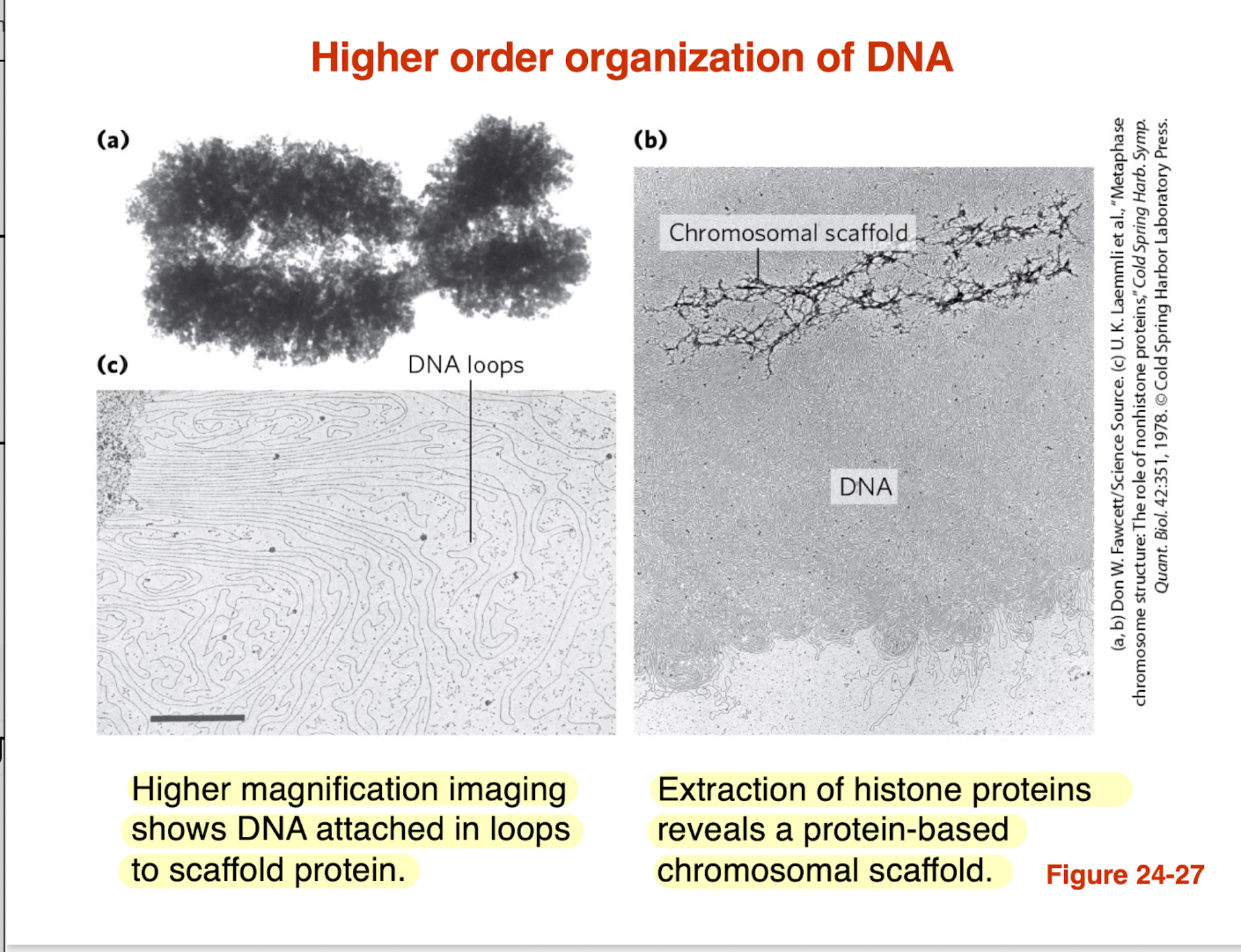
how is DNA organized at a higher order level beyond nucleosomes?
DNA is divided into active (euchromatin) and inactive (heterochromatin) compartments
CTCF (binding protein) helps organize DNA into loops called TADs (Topologically Associated Domains)
Active regions = loosely packed, transcriptionally active
Inactive regions = tightly packed, transcriptionally silent (heterochromatin)
Key idea: 3D DNA organization controls gene expression
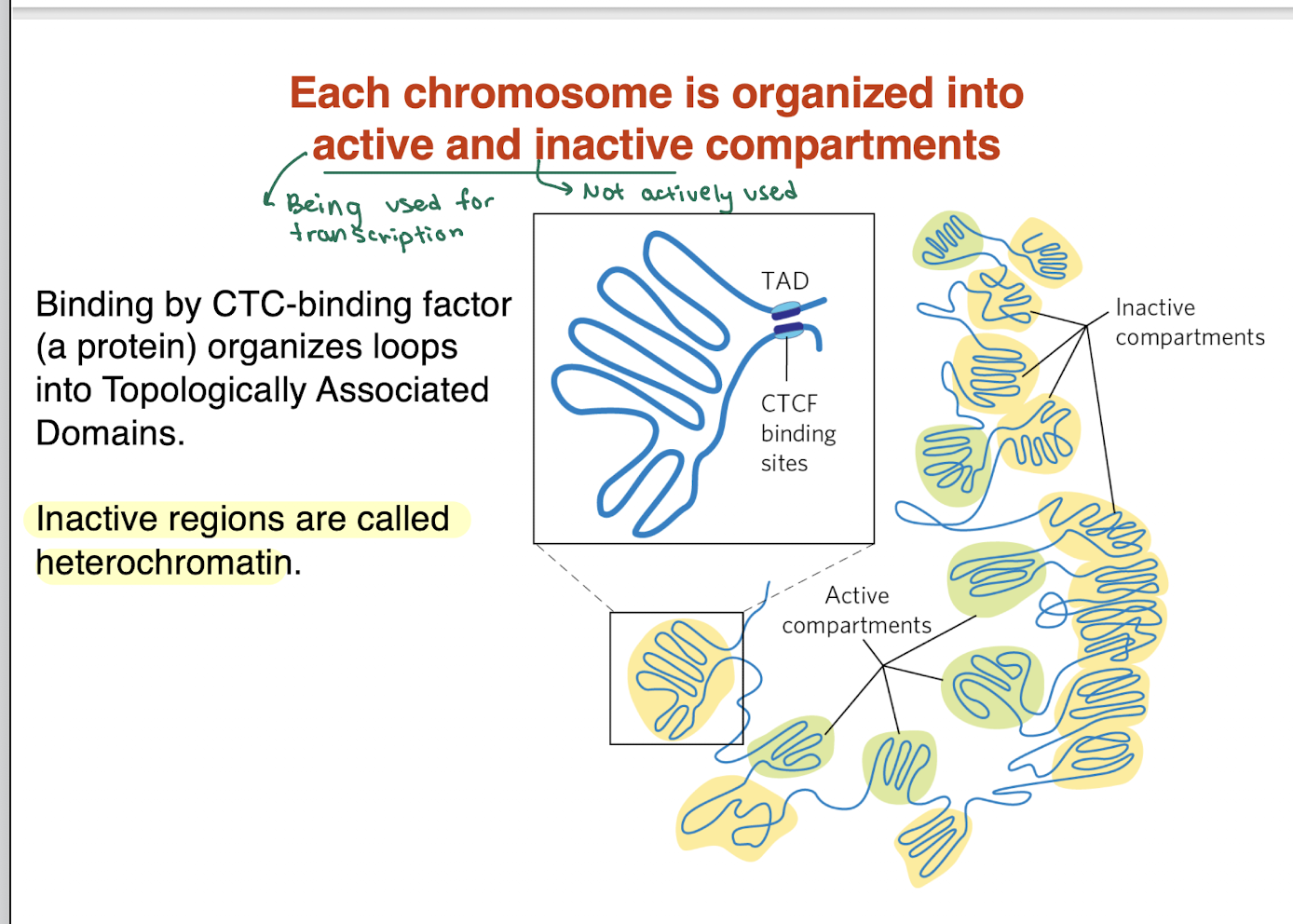
long non coding RNAs and associated proteins also organize DNA in BLANK
chromosomes
BLANK AND BLANK organize DNA in the eukaryotic cell cycle
Cohesins and condensins
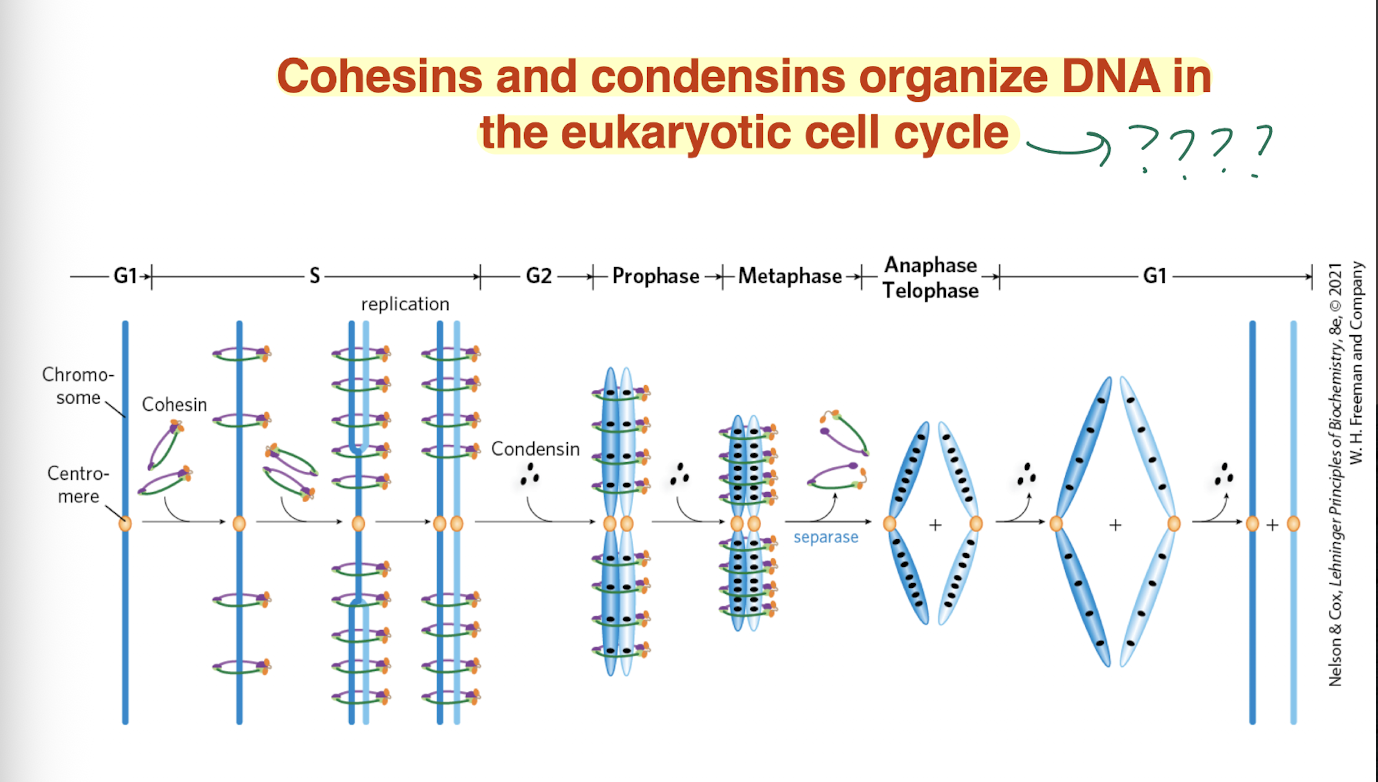
what is the difference between a semiconservative models vs a conservative model?
semiconservative model: hybrid duplex of old and new strand —> RIGHT
conservative model: duplex of only old or only newly synthesized DNA
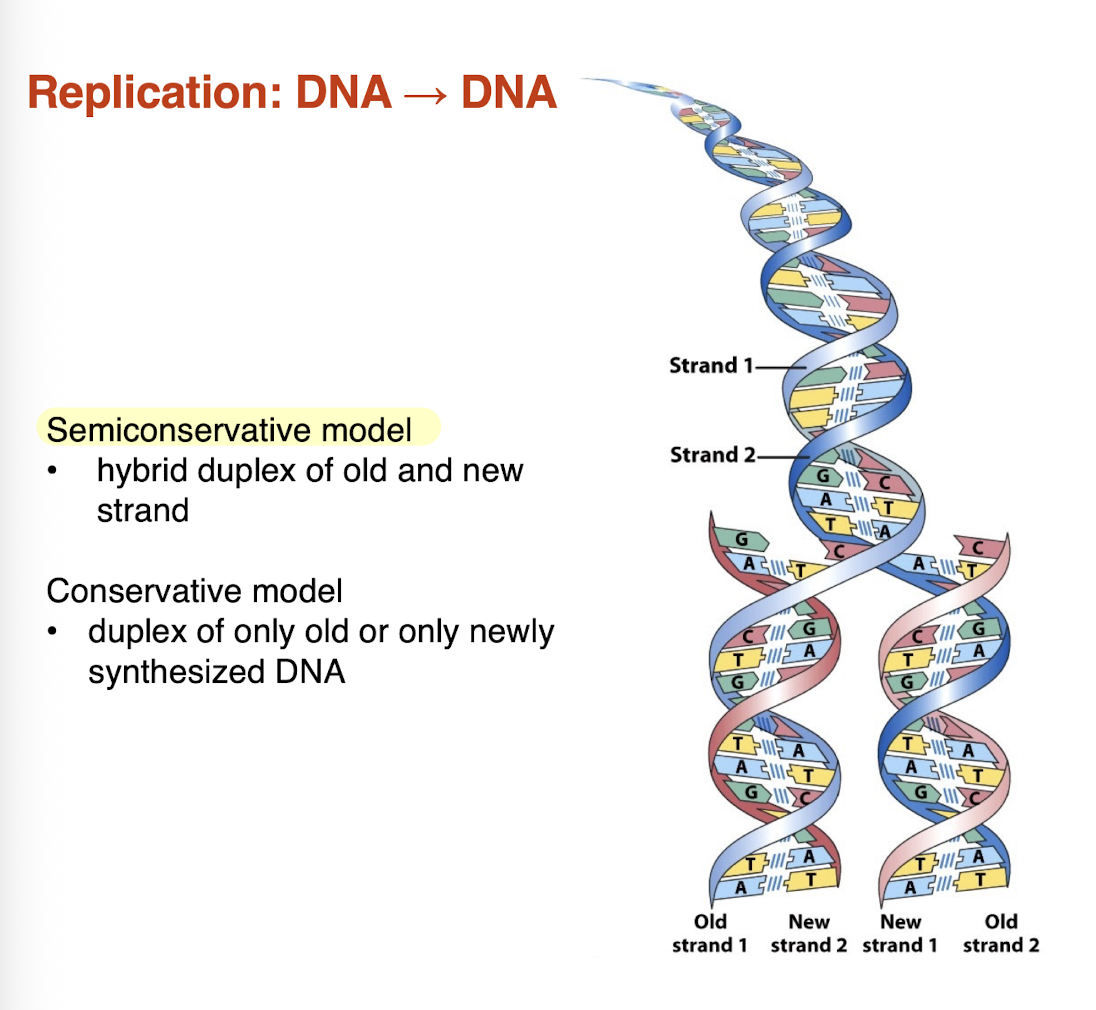
DNA synthesis is performed by what enzyme?
DNA polymerases
DNA synthesis requires what and catalyzes what?
require:
template strand to copy
primer strand with 3’ OH
dNTP substrates
catalyze:
nucleophilic attack by 3’ OH
phosphodiester bond formation
5’ —> 3’ synthesis
DNA synthesis is always X’ —> Y’
DNA synthesis is always 5’—> 3’
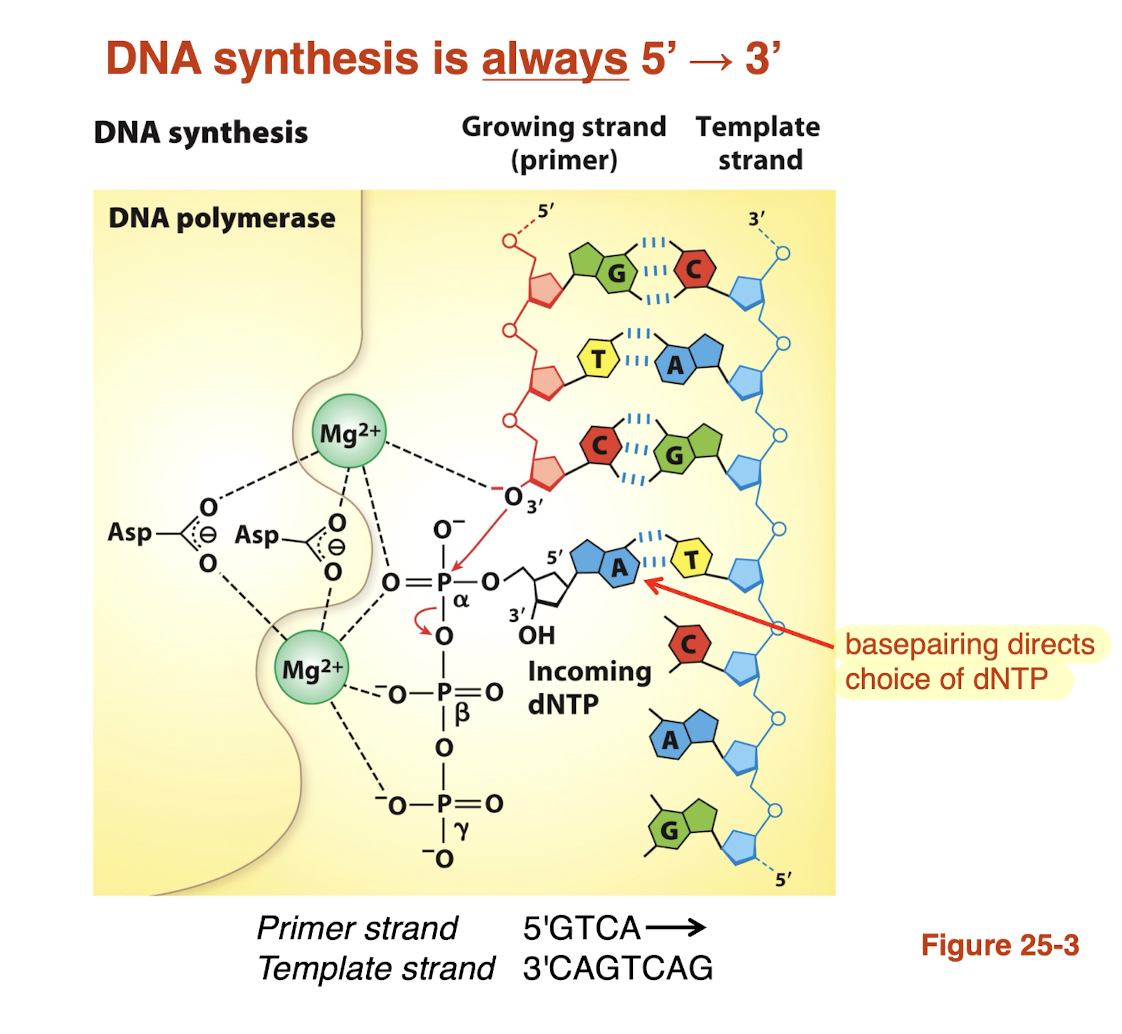
Why is base pair geometry important for DNA replication fidelity?
DNA polymerase active site is shaped to fit correct pairs (A-T, G-C)
correct pairs have proper geometry —> fit perfectly
incorrect pairs have distorted shape —> don’t fit well
this allows polymerase to select the right nucleotide
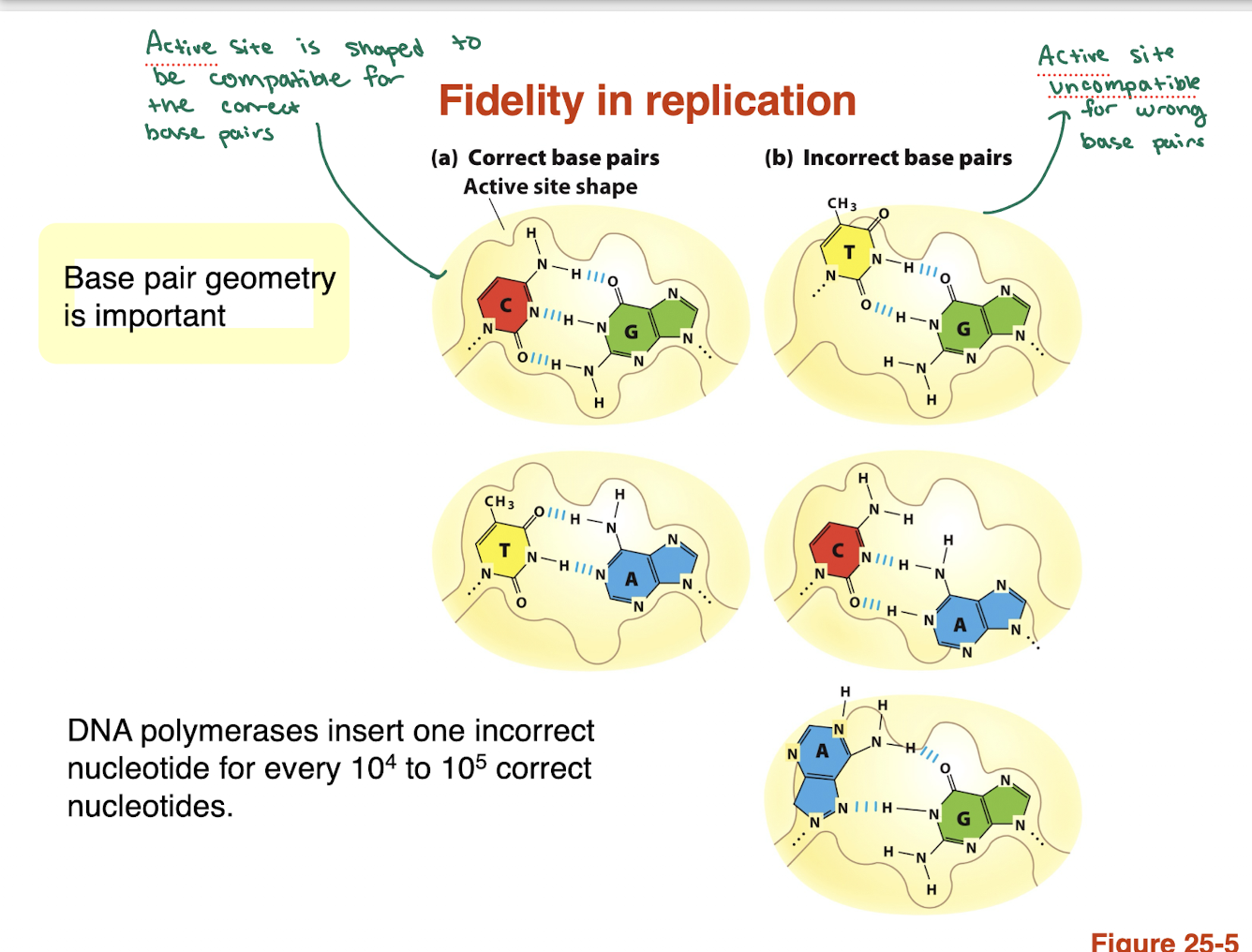
how accurate is DNA polymerase and why?
Error rate: ~1 mistake per 10⁴–10⁵ nucleotides
High fidelity comes from:
Base pair geometry recognition (shape-based selection)
Incorrect bases are rejected due to poor fit in active site
Key idea: shape matters more than just hydrogen bonding
What is the role of 3’ → 5’ exonuclease activity in DNA replication?
Provides proofreading function
Removes incorrectly added nucleotides
Works in the 3’ → 5’ direction
Increases replication accuracy significantly
How does DNA polymerase correct a mistake?
Incorrect base is added
DNA is shifted to exonuclease site
Wrong nucleotide is cleaved off
DNA returns to polymerase site
Correct nucleotide is added
DNA replication has three major stages what are they?
initiation
elongation
termination
What is DUE? it typically has a bp region that is heavily concentrated with?
DNA unwinding element
A-T rich region —> easier to separate strands (fewer H bonds)
site where DNA first unwinds
replication starts at specific sequences, where proteins bind and AT rich DNA unwinds first
what does the DnaA protein do?
able to recognize the origin sequence
recognizes the oriC sequence; opens duplex at specific sites in origin
what is the DnaB protein or helicase do?
unwinds the DNA
what does the DnaC protein do?
required for DnaB binding at the origin
How does DNA replication initiate at the origin (oriC) in bacteria?
DnaA binds origin (oriC):
DnaA-AAA+ binds specific sequences → wraps & bends DNA
DNA unwinding (DUE):
AT-rich DUE region melts → forms an open complex (bubble)
DNA bending proteins help:
Proteins like IHF assist in DNA deformation → promotes opening
Helicase loading:
DnaC loads DnaB helicase onto single-stranded DNA
Replication begins:
DnaB helicase further unwinds DNA → replication machinery assembles
what does initiation involve for DNA replication?
Initiation = protein binding → DNA bending → AT-rich unwinding → helicase loading → replication starts
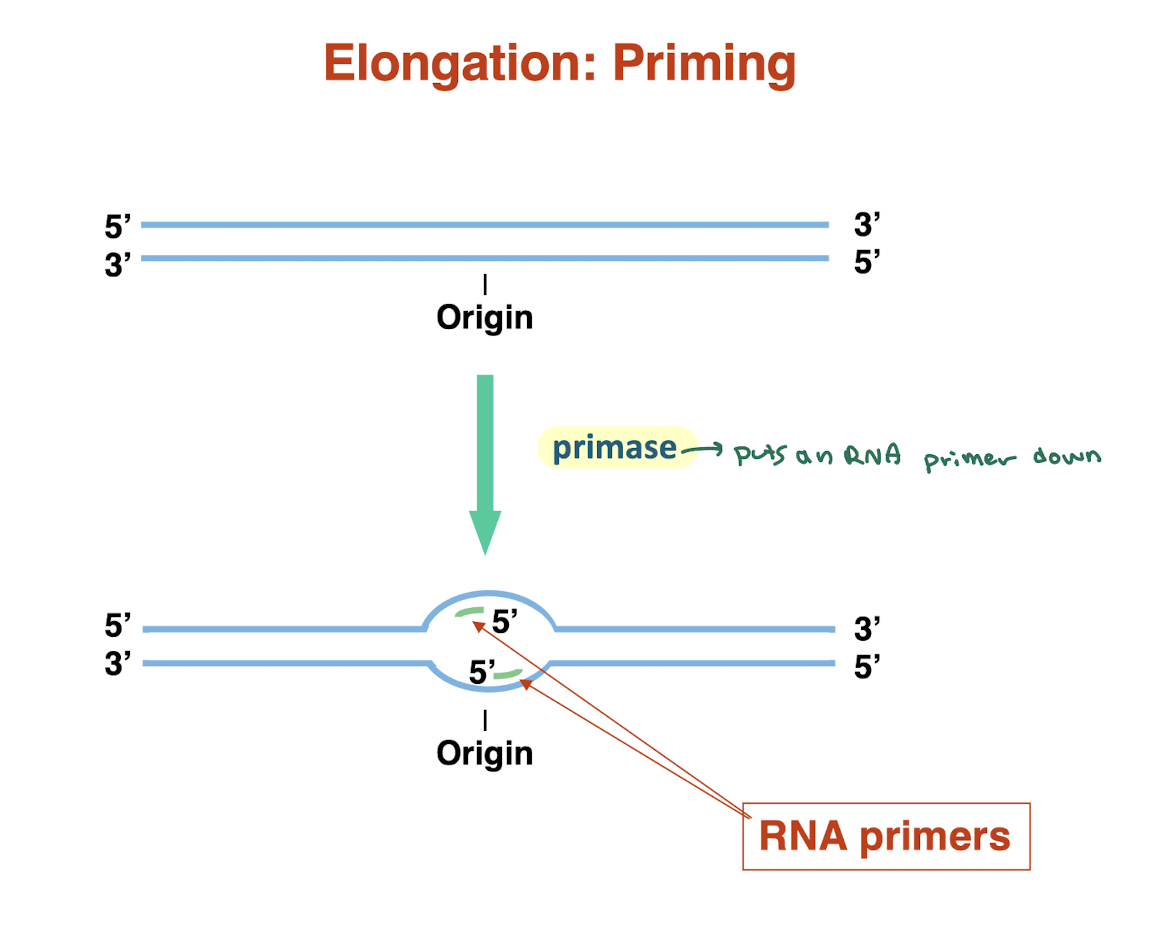
what is priming in DNA replication and why is it necessary? what enzyme does it use?
Primase synthesizes short RNA primers at the origin
Provides a free 3′-OH group for DNA polymerase to start synthesis
DNA polymerase cannot start de novo → must extend from a primer
Primers are laid down on both strands as replication begins
How does DNA polymerization occur during replication, and in what direction is DNA synthesized? what enzyme does it utilize? DNA replication reads the leading strand from what?
DNA polymerase III extends from RNA primers
Synthesizes DNA only in the 5′ → 3′ direction
Moves along the template strand reading 3′ → 5′
Replication proceeds bidirectionally from the origin
Both strands are copied simultaneously, but differently (leading vs lagging)