Week 3 - Eukaryotic DNA Replication
1/84
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
85 Terms
differences between eukaryotes and prokaryotes
The DNA of eukaryotes is enclosed in a nucleus
Prokaryotes lack a nucleus
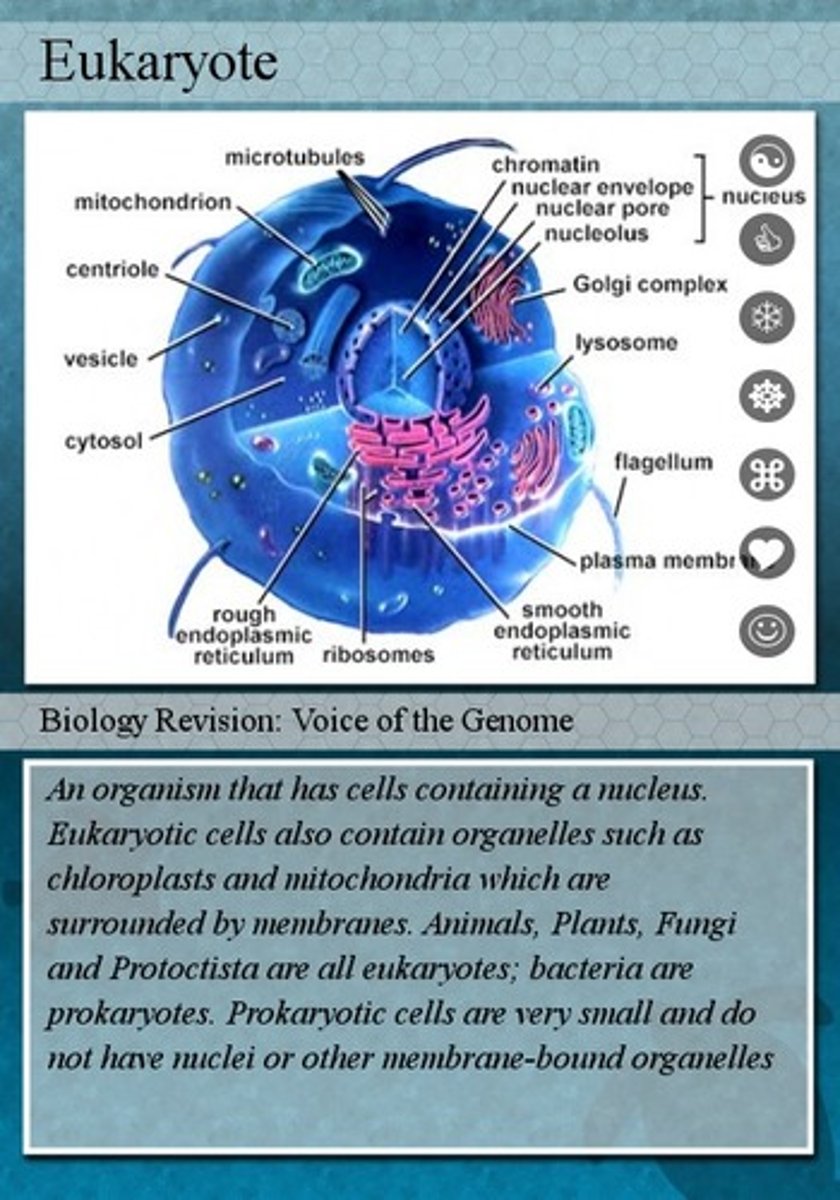
eukaryotes
consists of a true nucleus with a nuclear membrane and nucleoli; genetic material DNA is present inside the nucleus
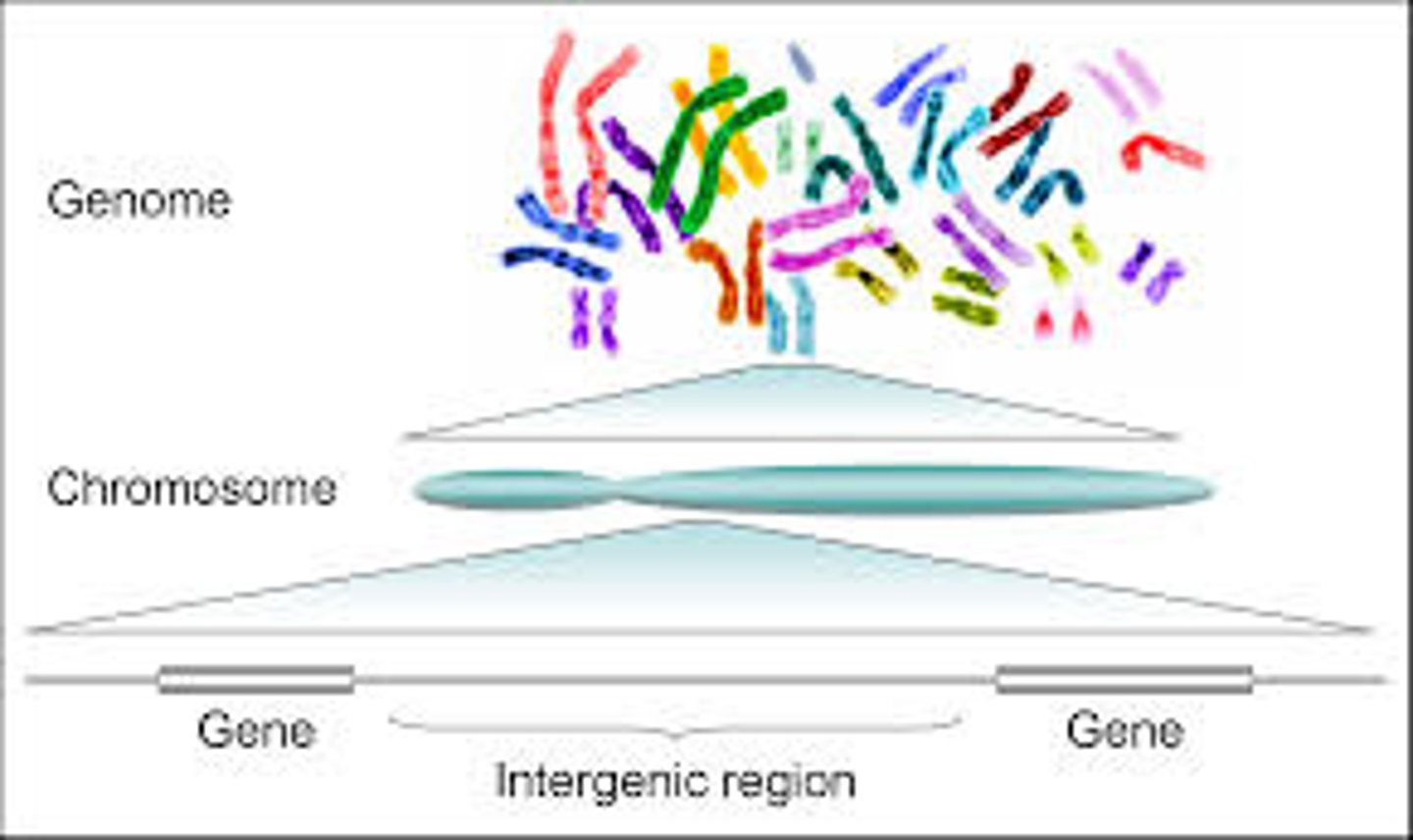
Genome
All the DNA of an organism is referred to as an organism
Prokaryotic DNA
Usually consists of a single circular chromosome
Found in the nucleoid region (no membrane-bound nucleus)
Typically has one origin of replication• Circular structure means no chromosome ends
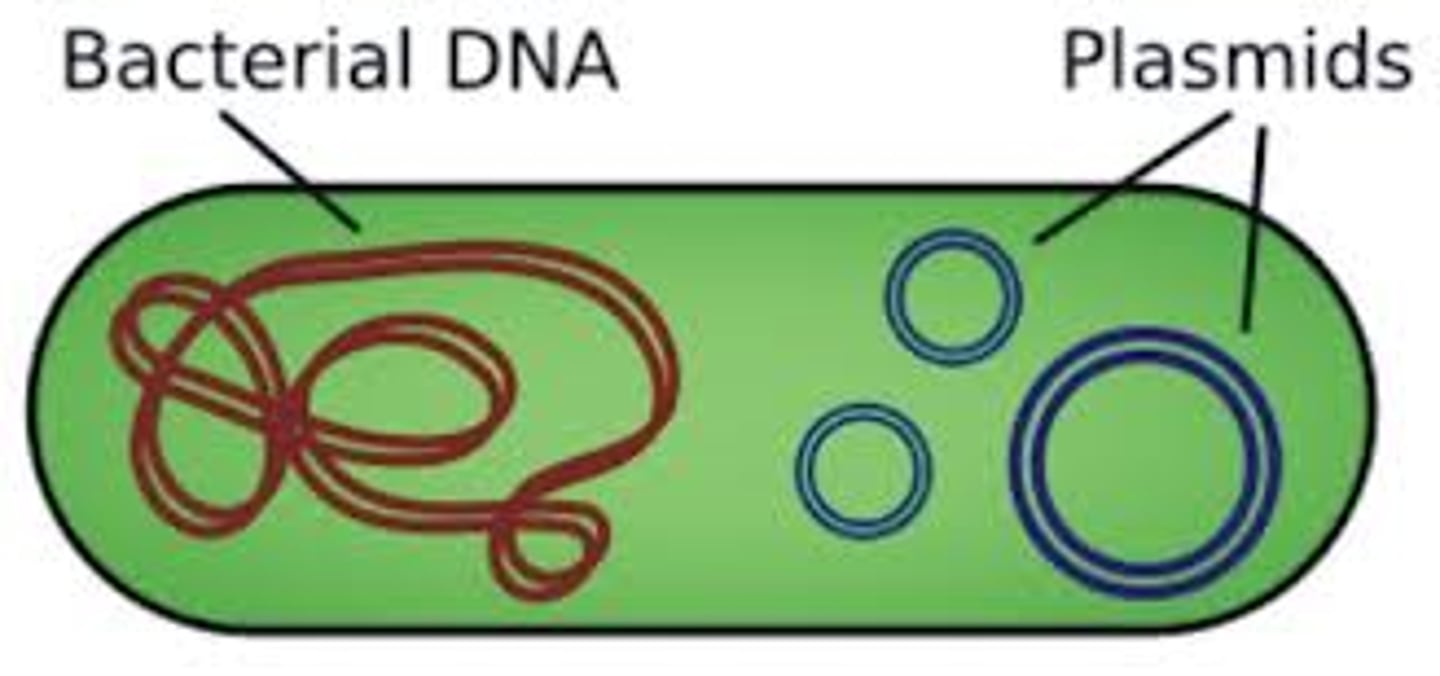
Eukaryotic DNA
Organized into multiple linear chromosomes
(Example human -46 chromosomes, dog -78 chromosomes)
Located inside a membrane-bound nucleus
Each chromosome contains multiple origins of replication
Linear structure creates chromosome ends (telomeres)
DNA in both the nucleus and mitochondria, but the nuclear DNA represents the genome
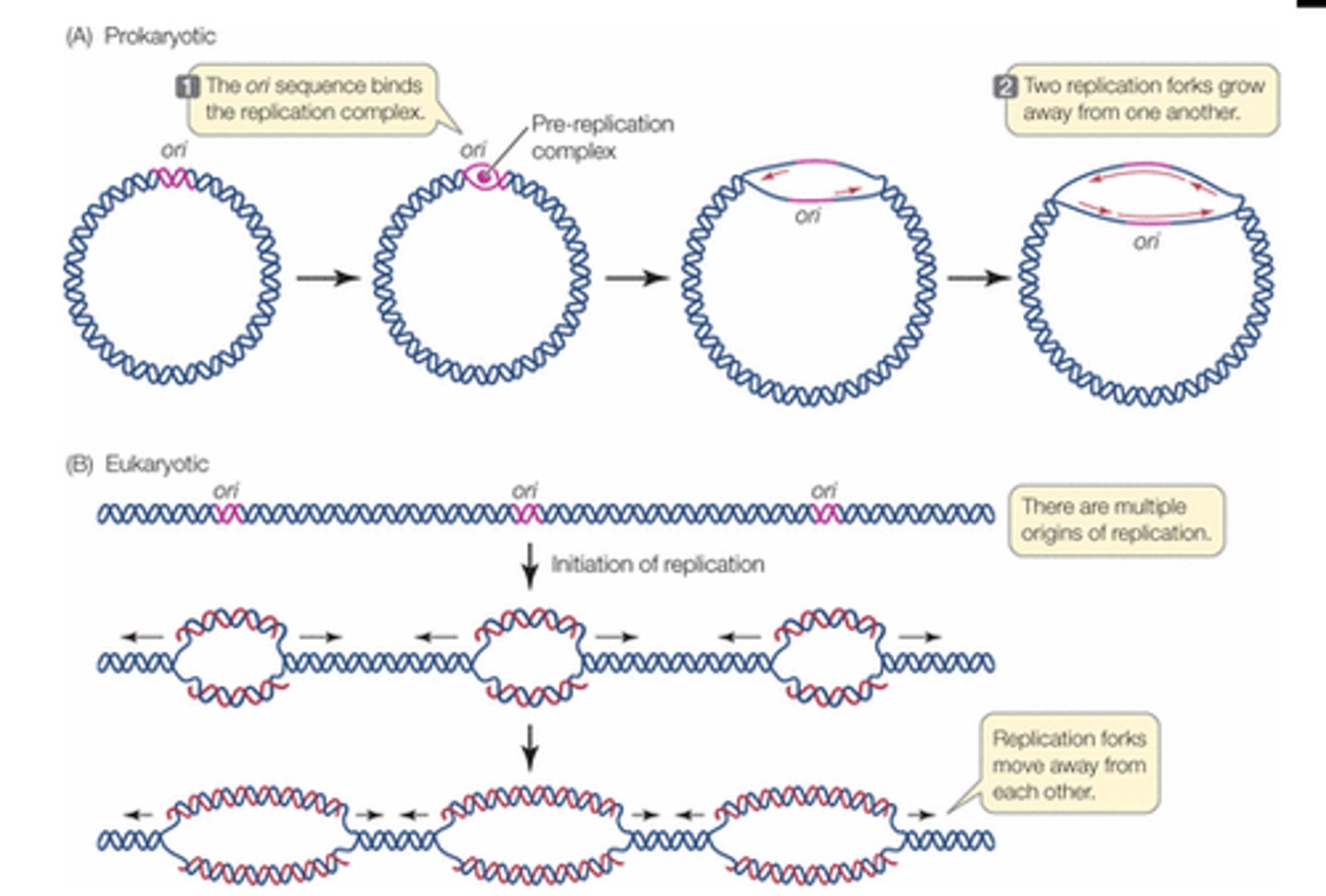
Replication of the genome occurs in the nucleus
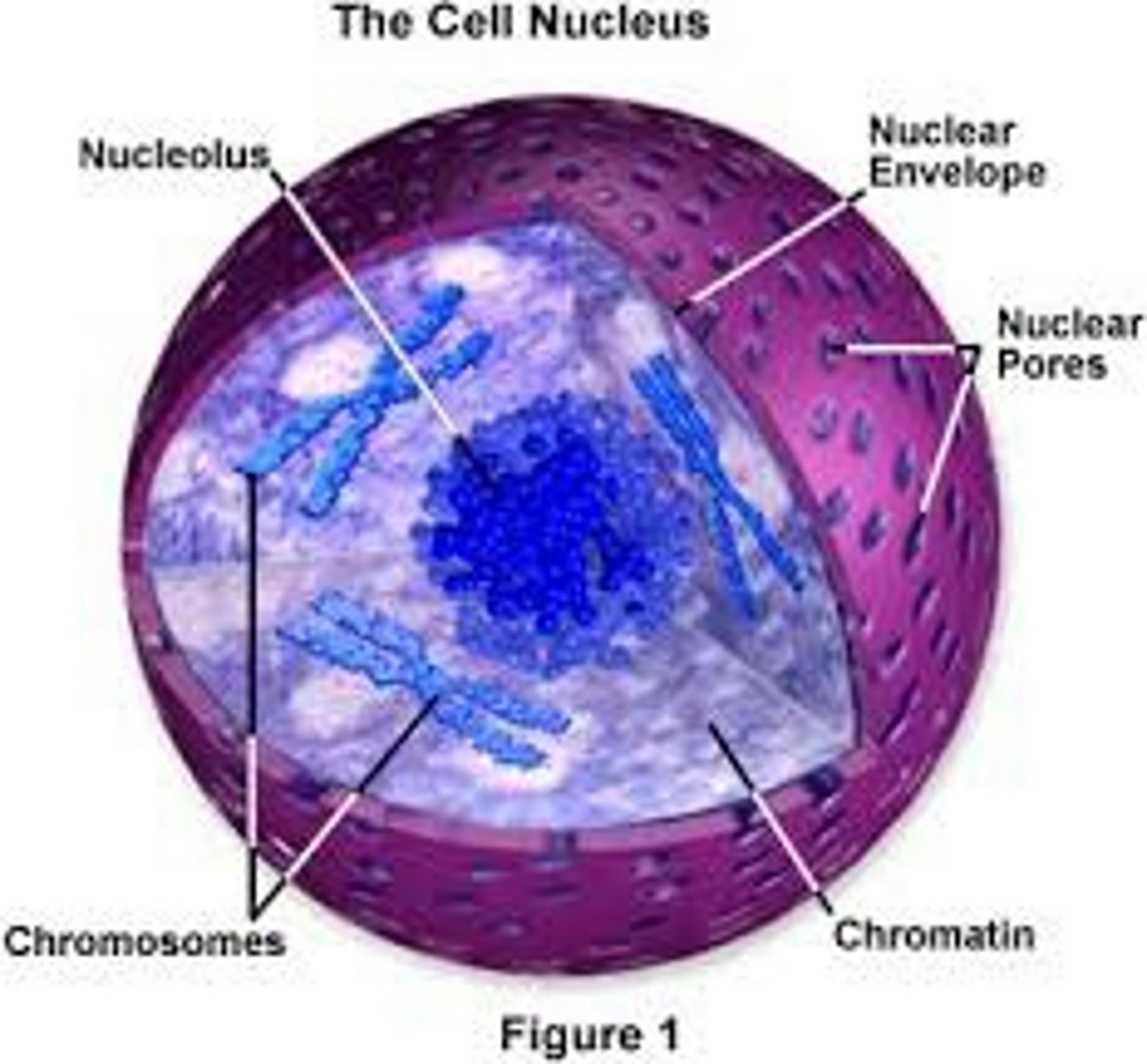
nucleolus
Found inside the nucleus and produces ribosomes
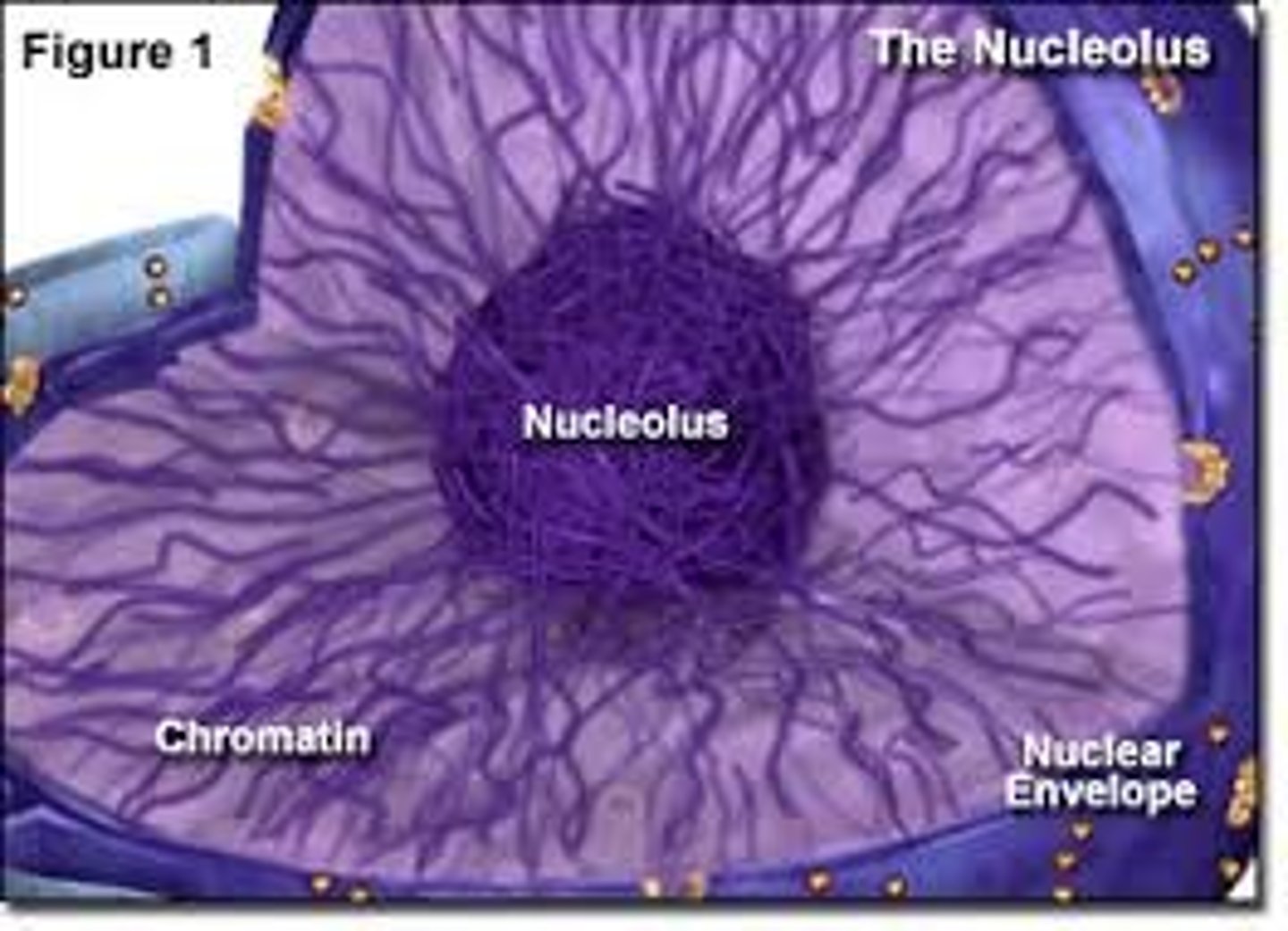
chromatin
Substance found in eukaryotic chromosomes that consists of DNA tightly coiled around histones
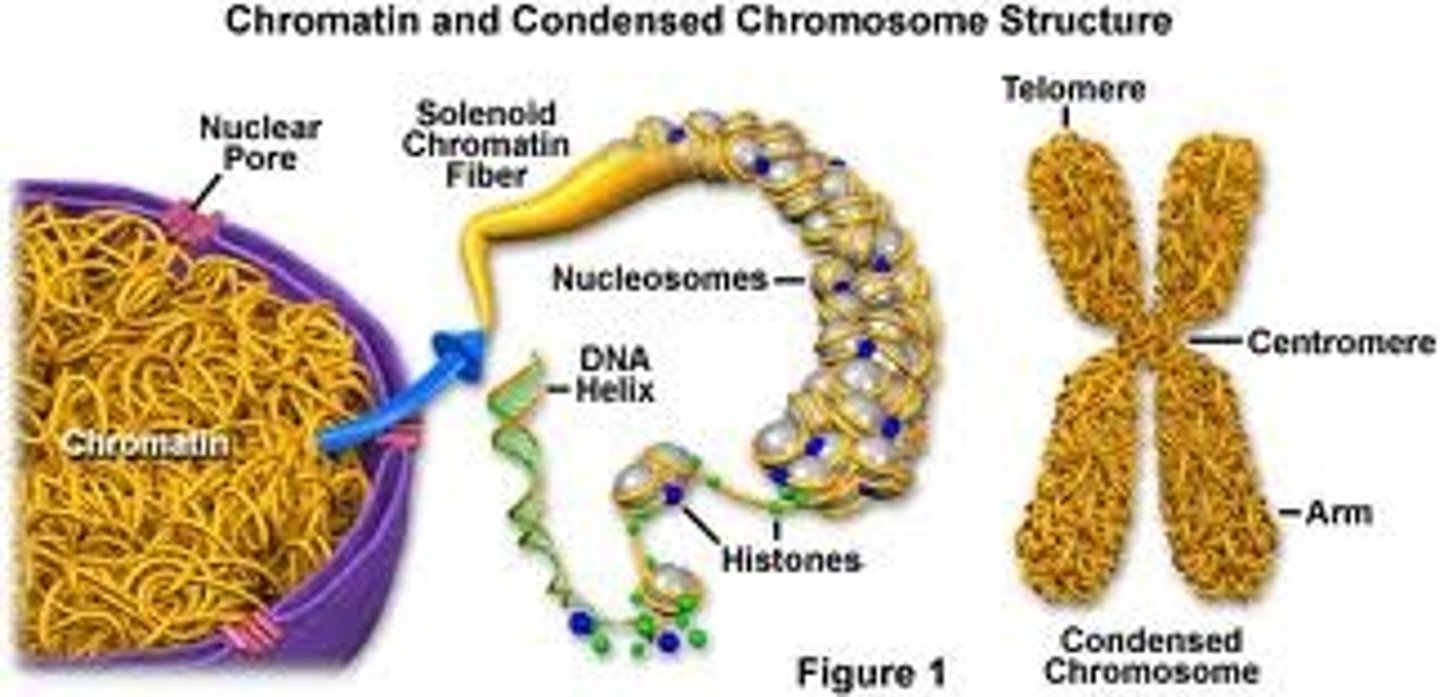
nuclear pore
a protein-lined channel in the nuclear envelope that regulates the transportation of molecules between the nucleus and the cytoplasm
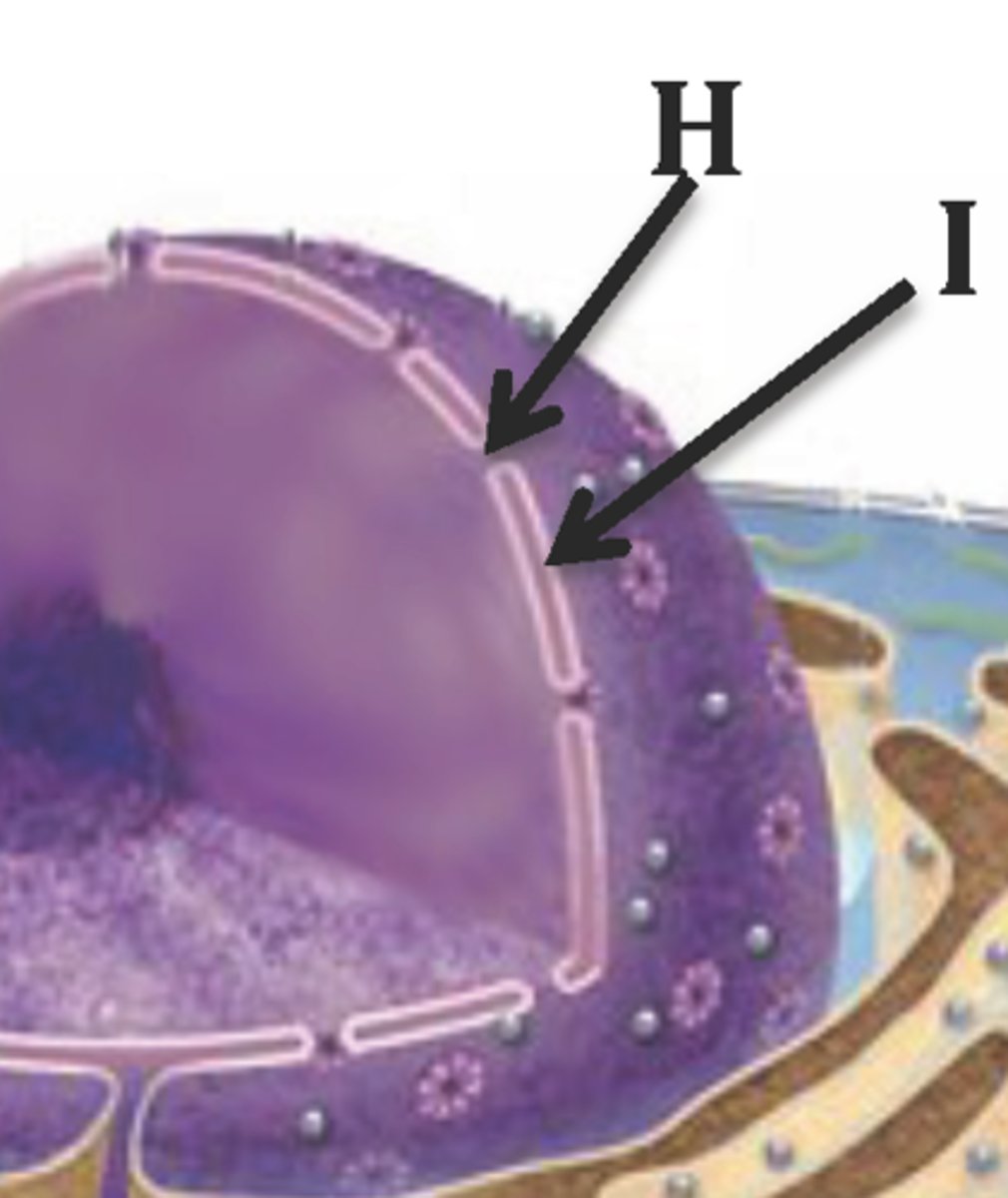
nucleoplasm
semi-solid fluid inside the nucleus that contains the chromatin and nucleolus
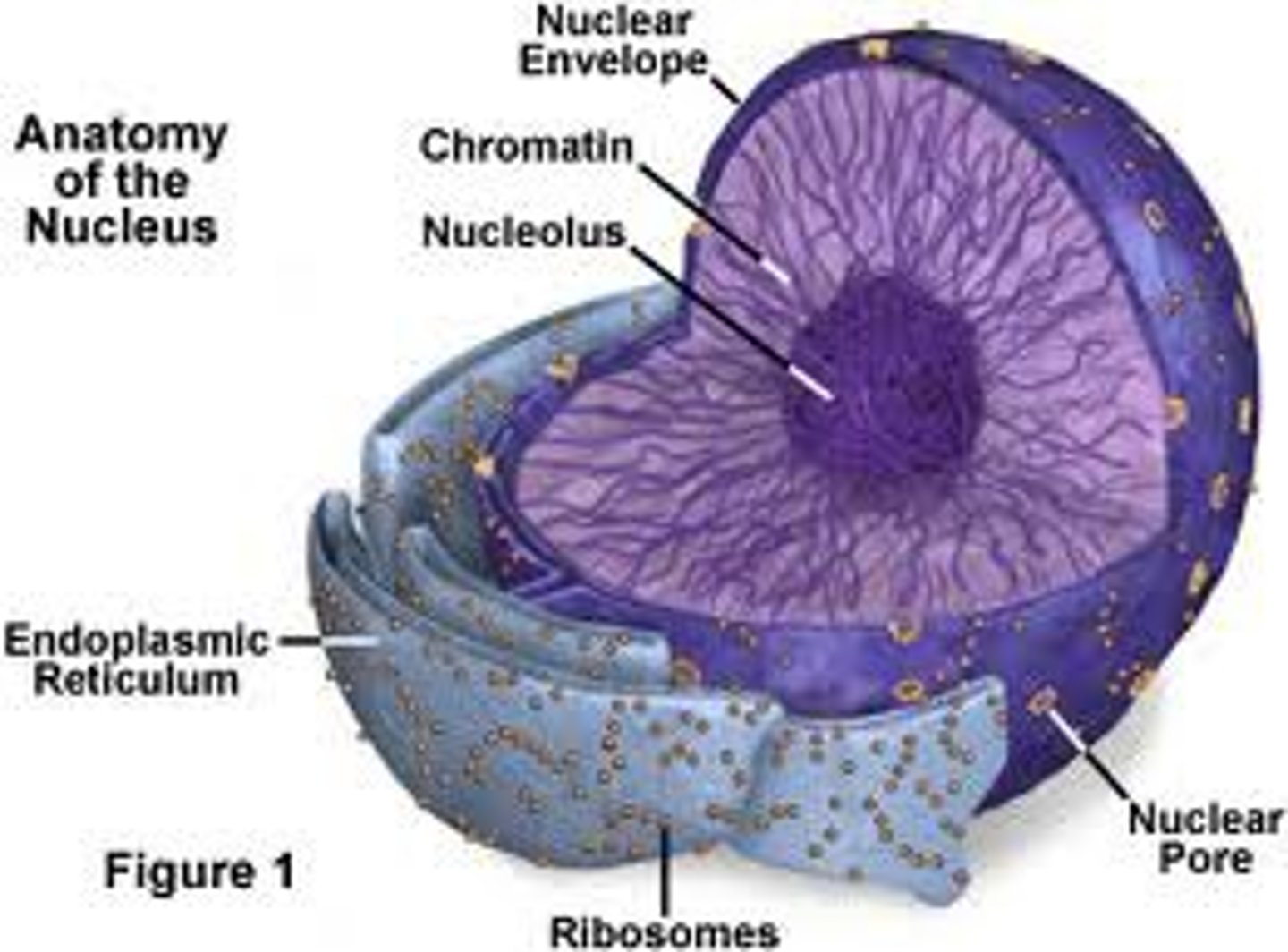
nuclear envelope
layer of two membranes that surrounds the nucleus of a cell
cell cycle
series of events that cells go through as they grow and divide
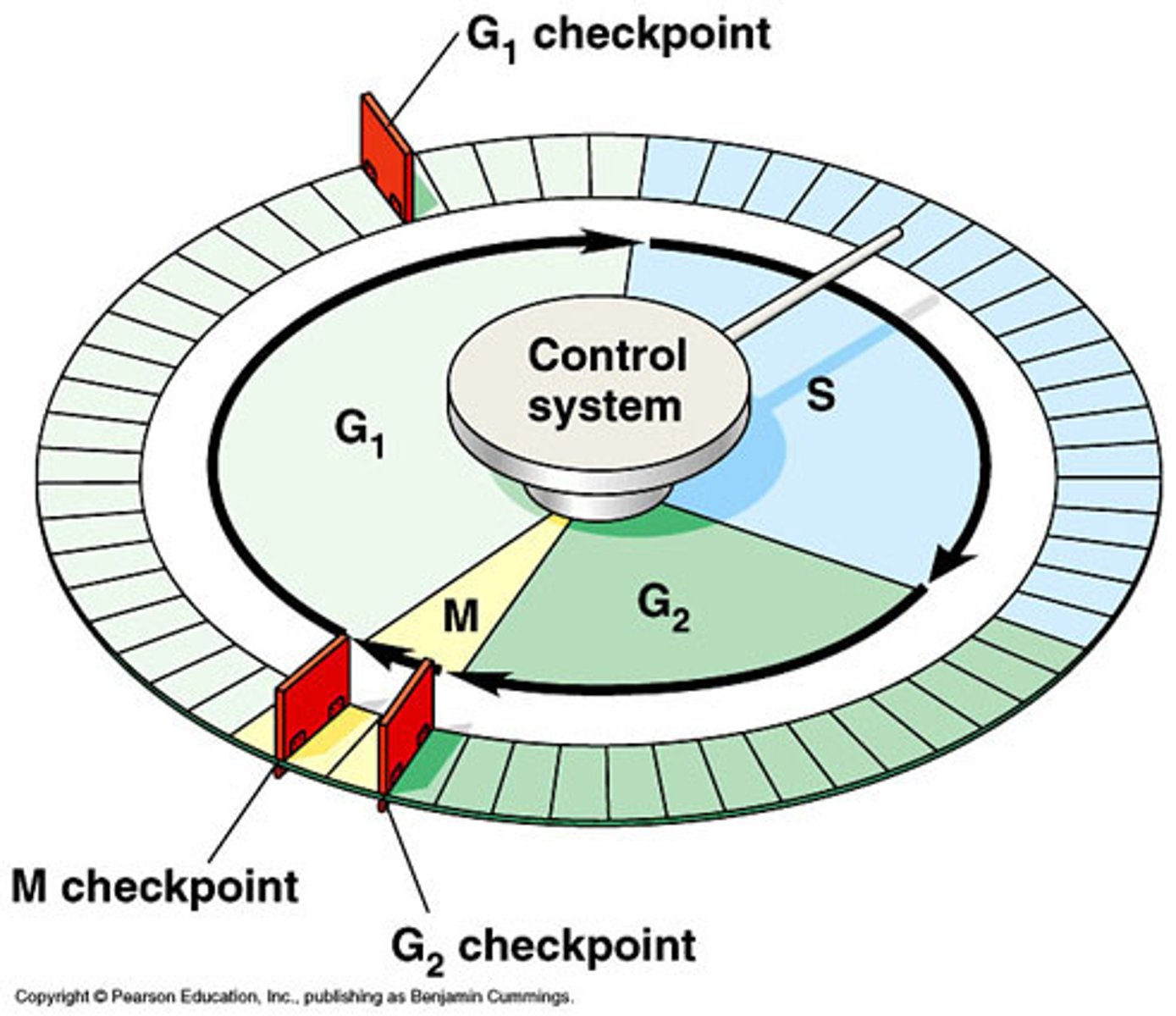
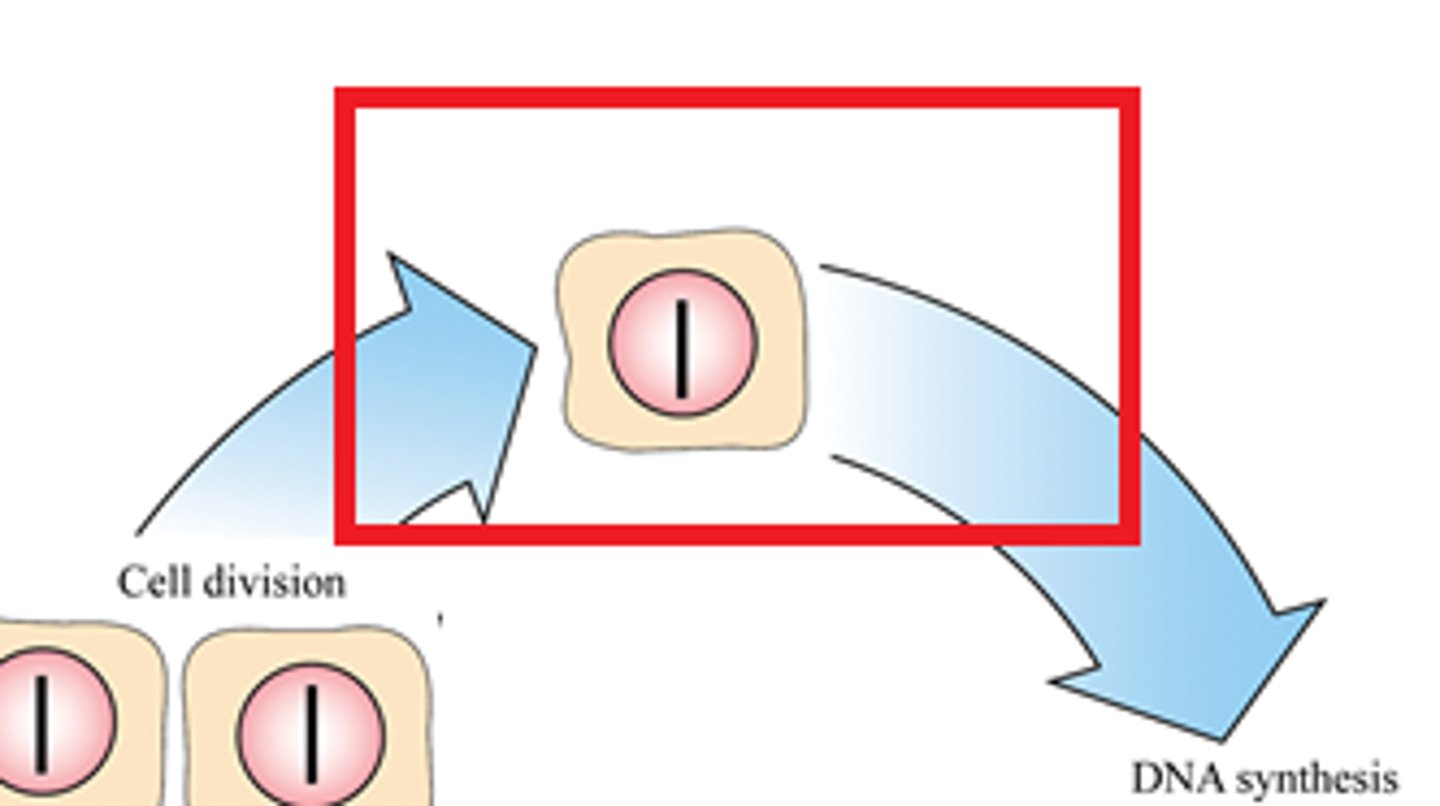
G1 phase
The first gap, or growth phase, of the cell cycle, consisting of the portion of interphase before DNA synthesis begins.
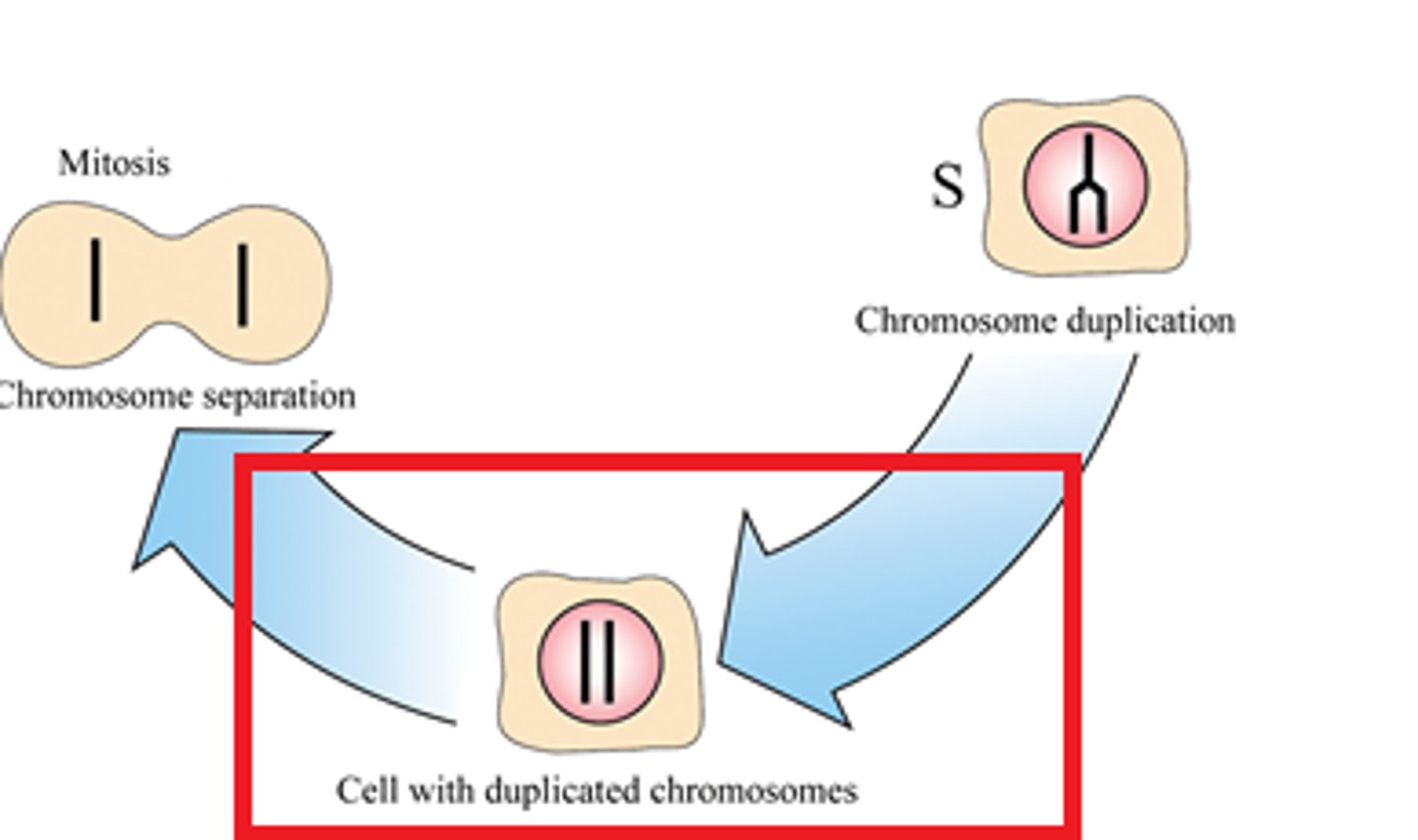
S phase
Replication is restricted to S phase to ensure that the genome (total DNA) is duplicated once and only once per cell cycle before cell division, preventing genome instability
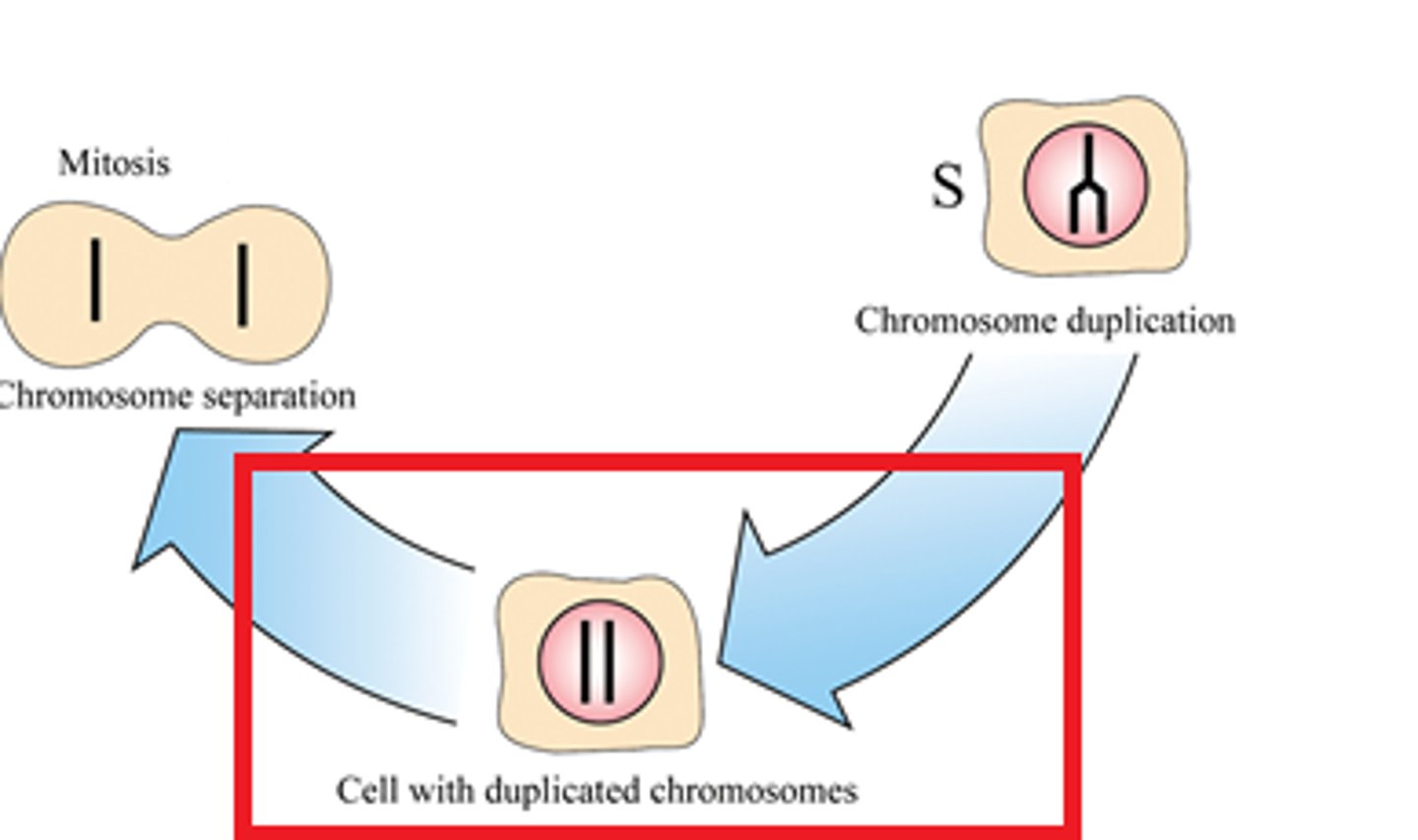
G2 phase
The second growth phase of the cell cycle, consisting of the portion of interphase after DNA synthesis occurs.
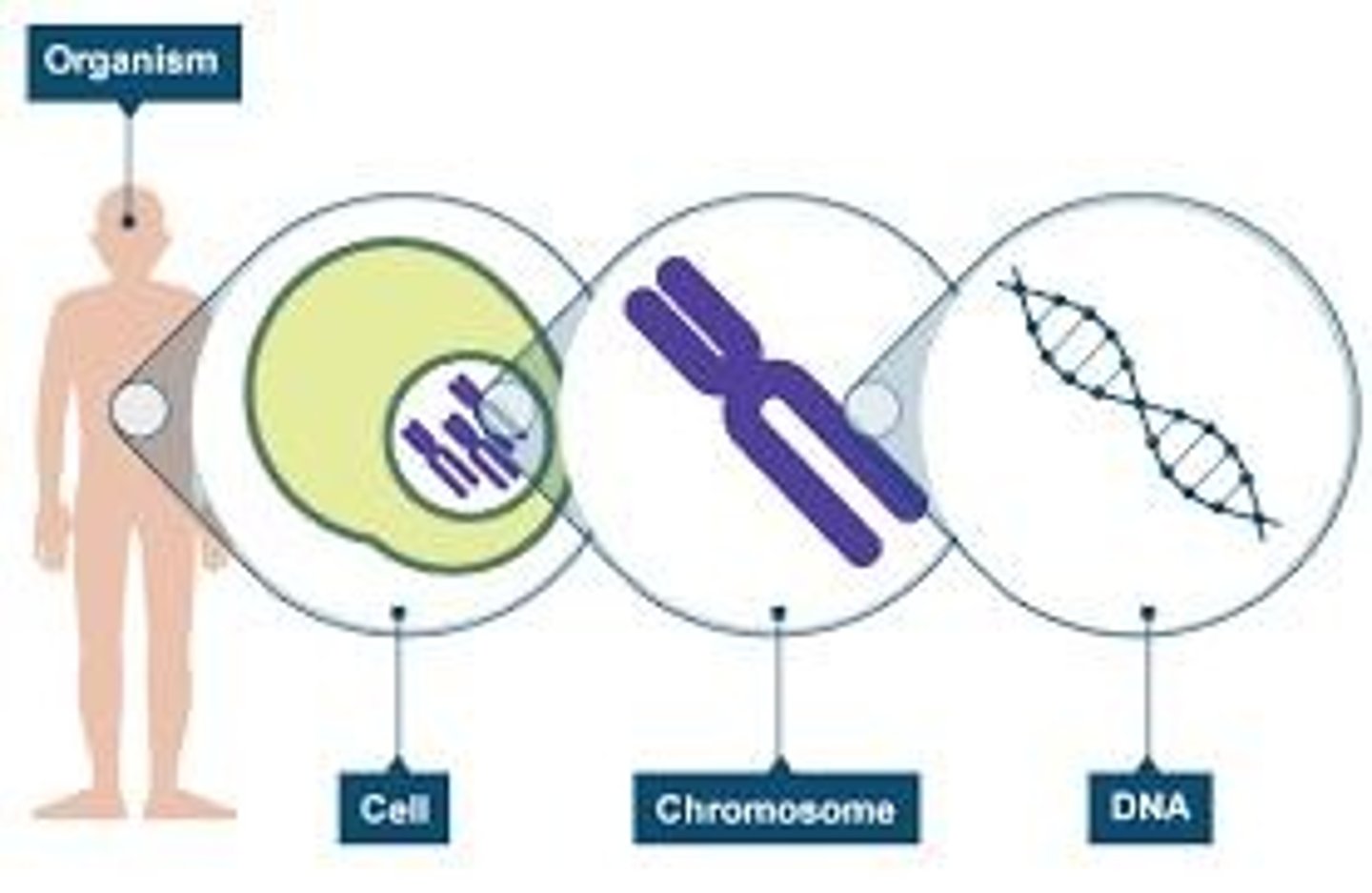
Humans are diploid
Each somatic cell contains two copies of the chromosome
One from each parent
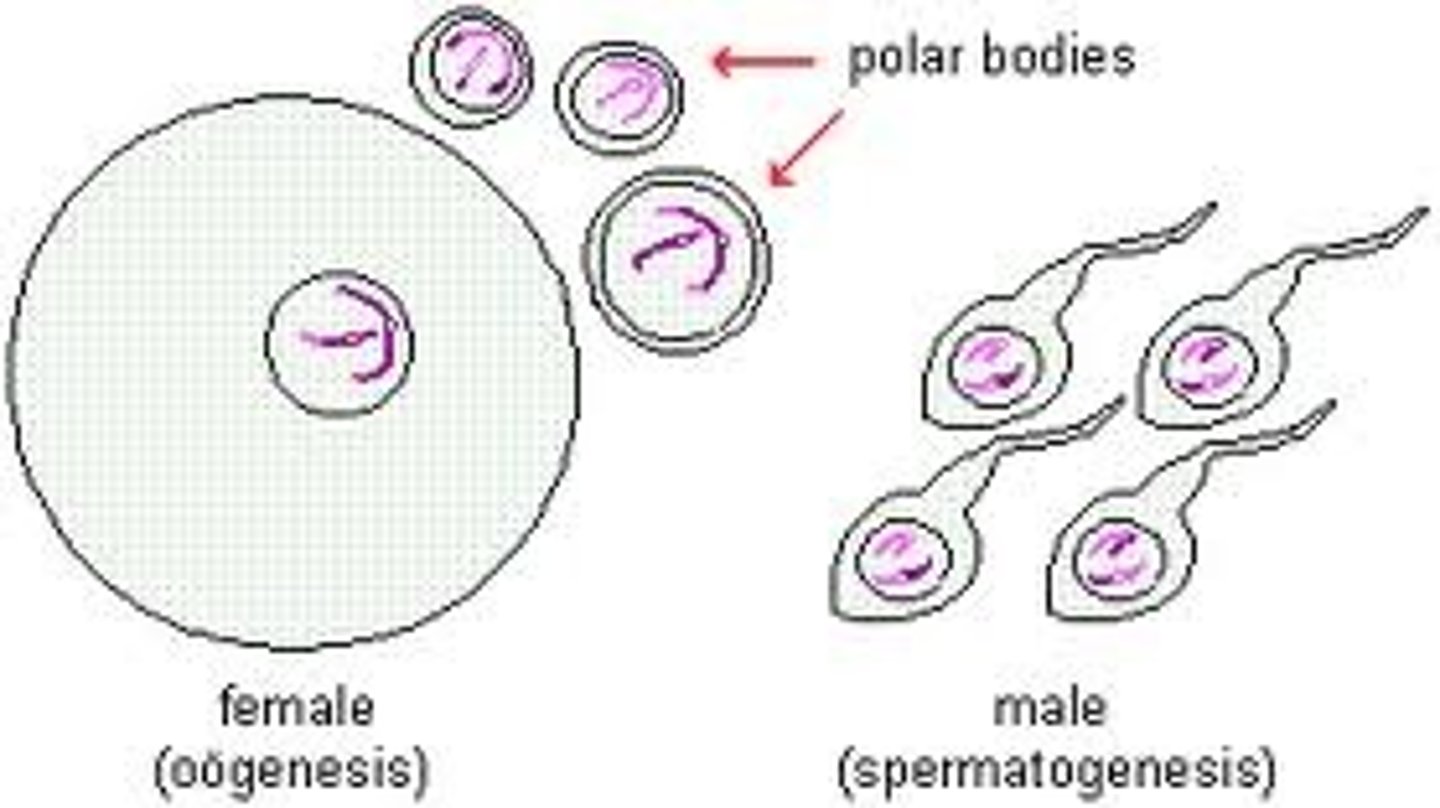
gametes
Eggs and sperm are haploid cells, meaning they contain half the number of chromosomes of the organism. In humans, each egg and each sperm contains23 chromosomes
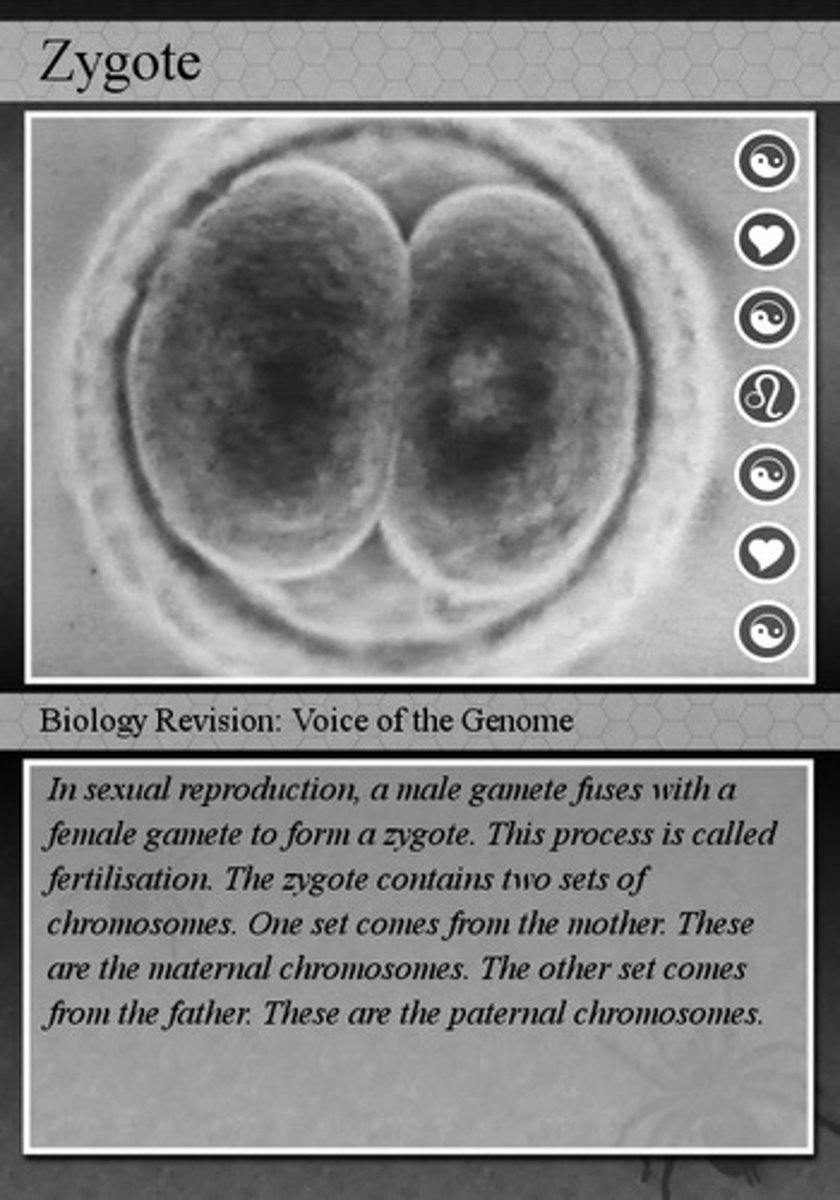
zygote
When an egg and a sperm fuse during fertilization, the resulting cell becomes diploid, containing the full number of chromosomes for the organism —has 46 chromosomes and is diploid
Human Haploid genome
(one set (23) of chromosomes): ≈ 3.2 billion nucleotides in the23 chromosomes
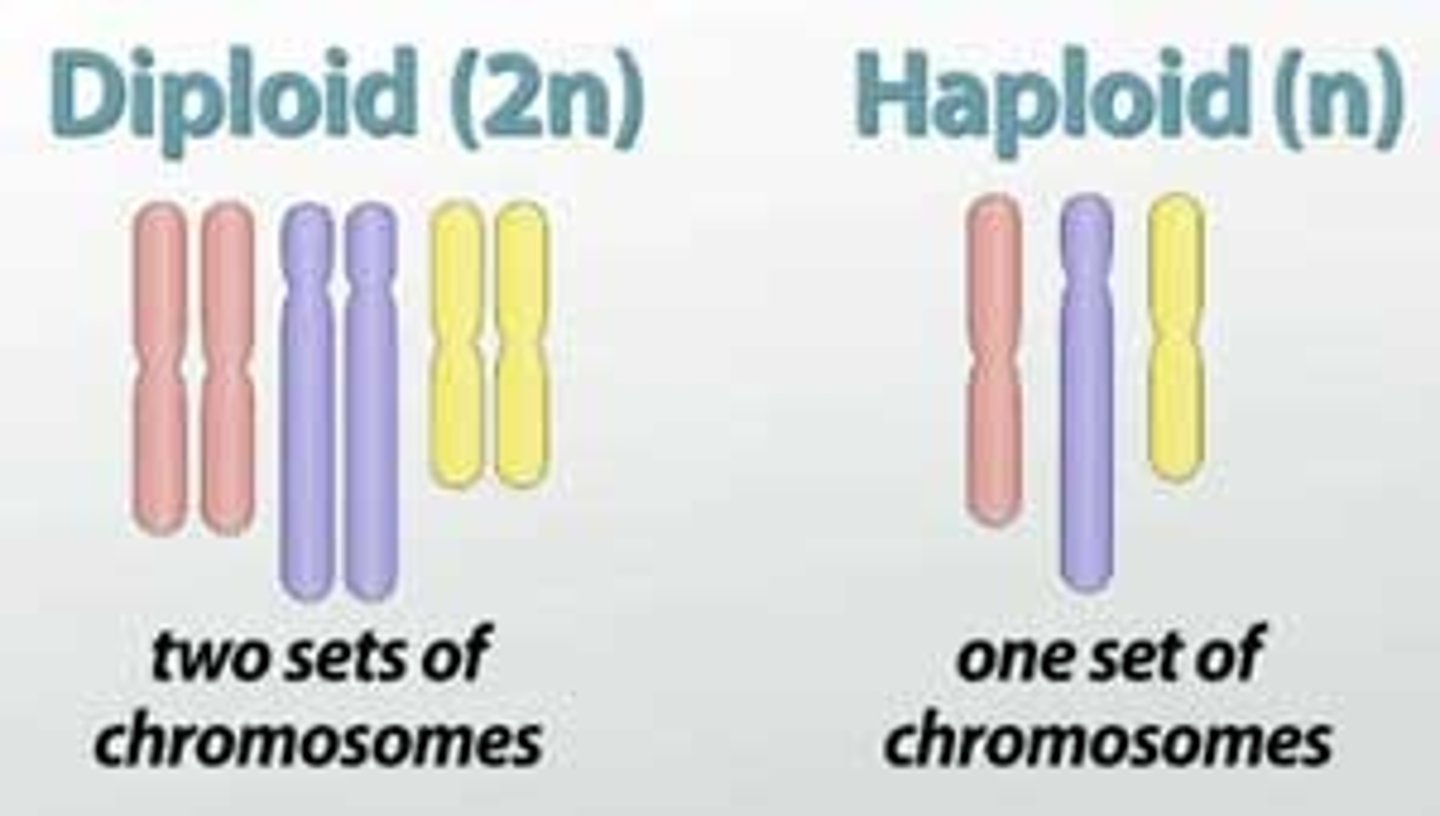
Human Diploid somatic cell
(two sets (46) chromosomes): ≈ 6.4 billion nucleotides in the46 chromosomes
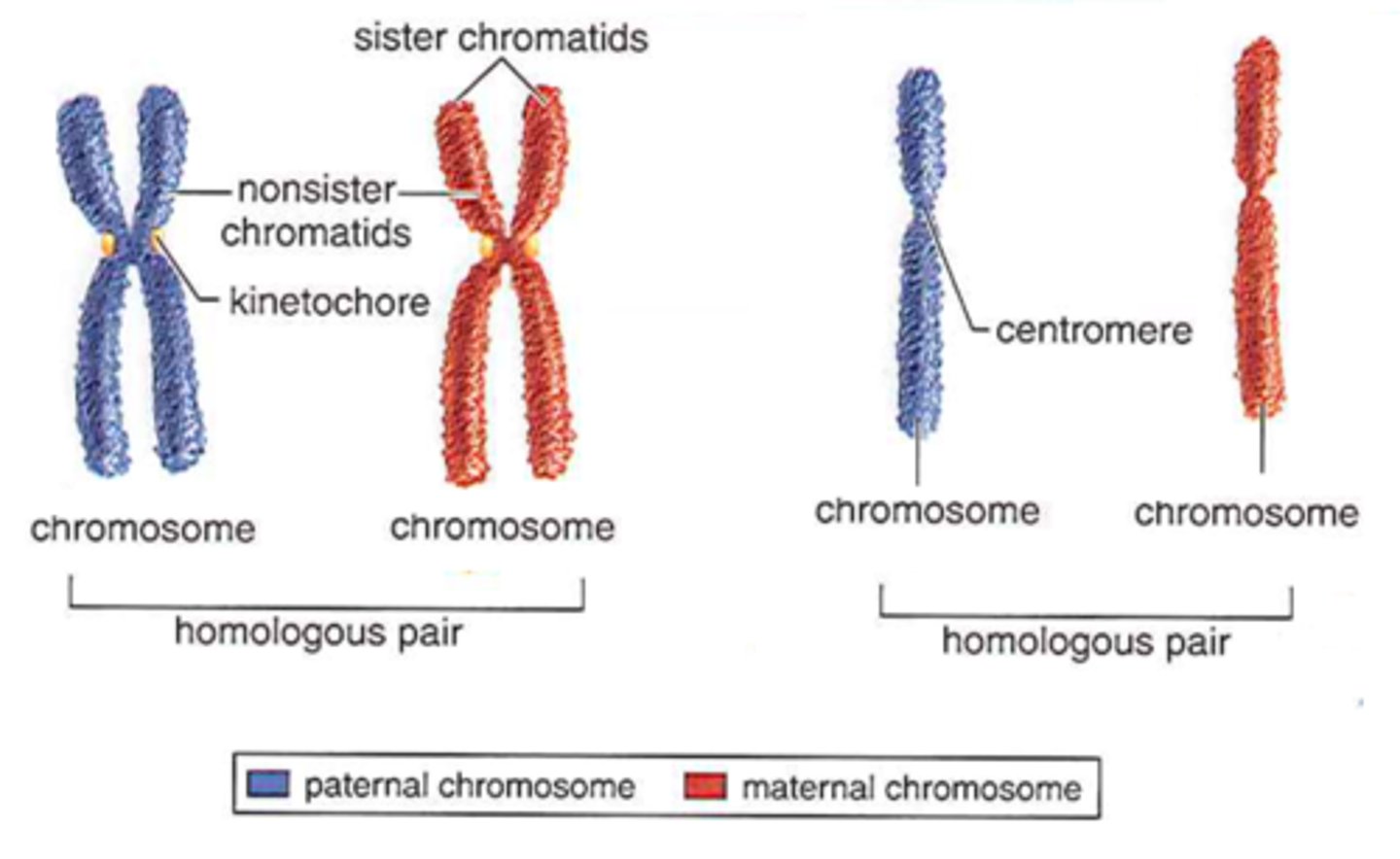
21st chromosome
Only chromosome can be duplicated (mutation) and still produce viable human life (other than sex chromosomes)
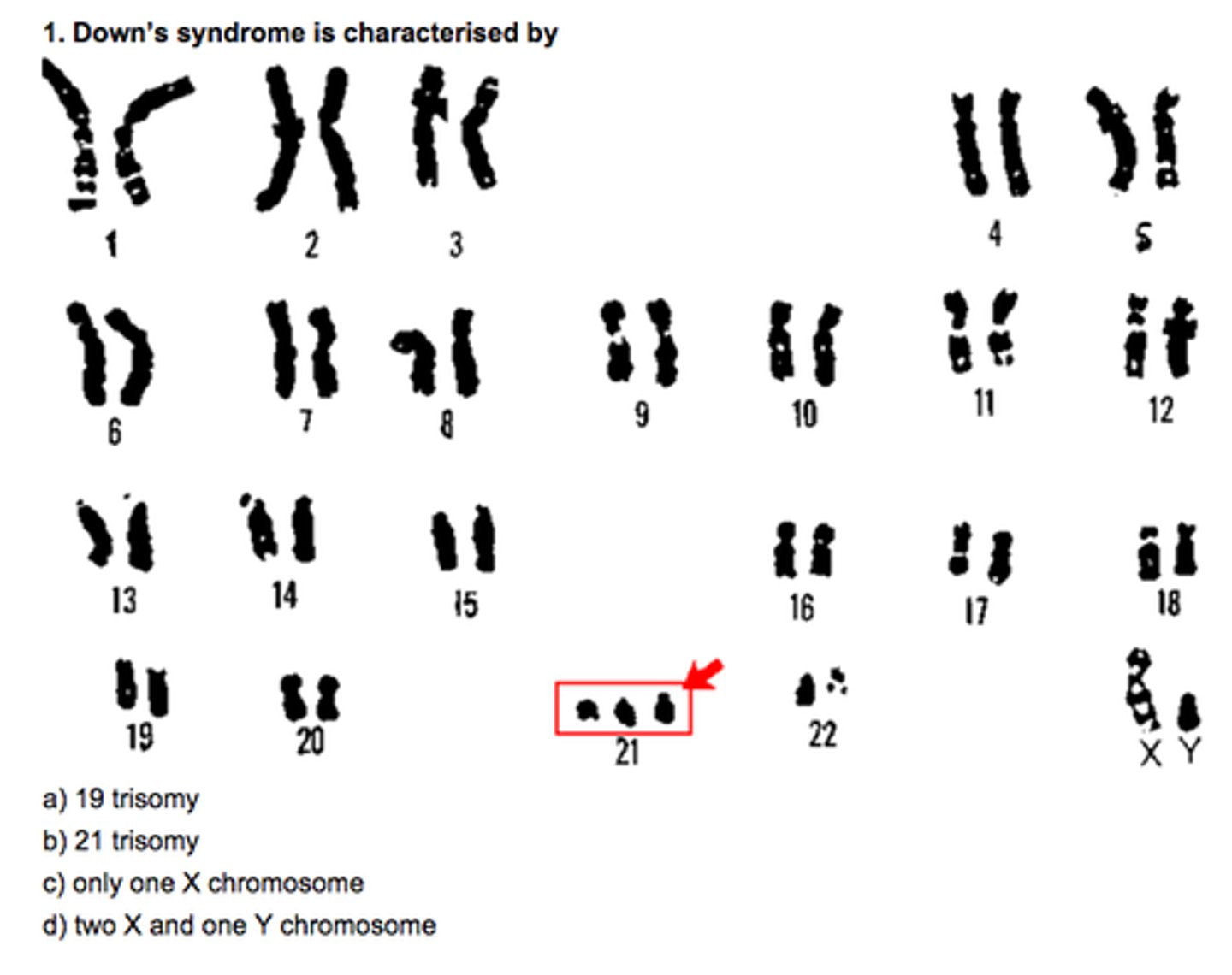
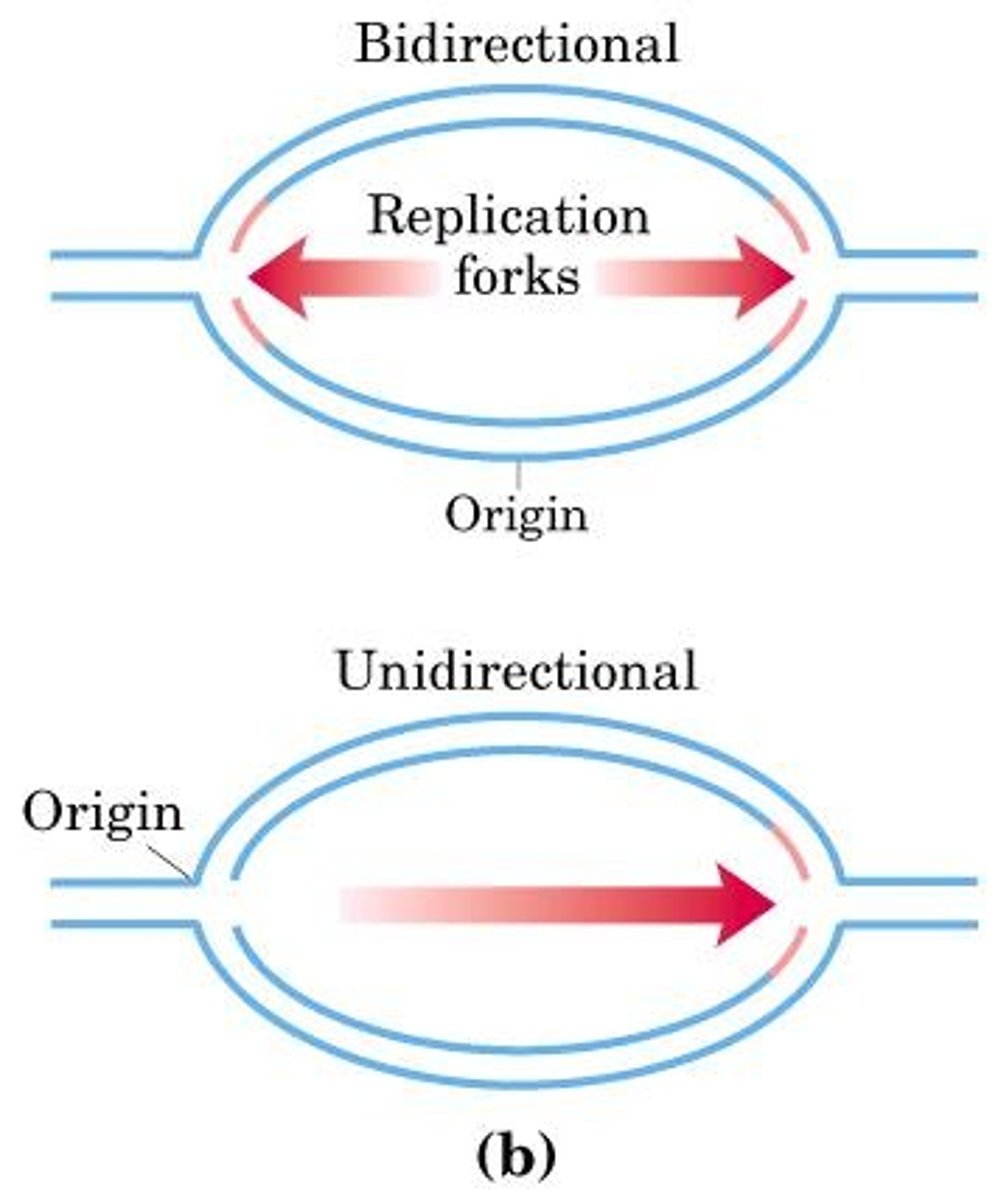
Origin of replication (ORI)
Each mammalian chromosome is composed of many replication sites
Multiple origins allow eukaryotes to replicate their larger DNA in a relatively short time
Replication must be fast and accurate --> cell cycle
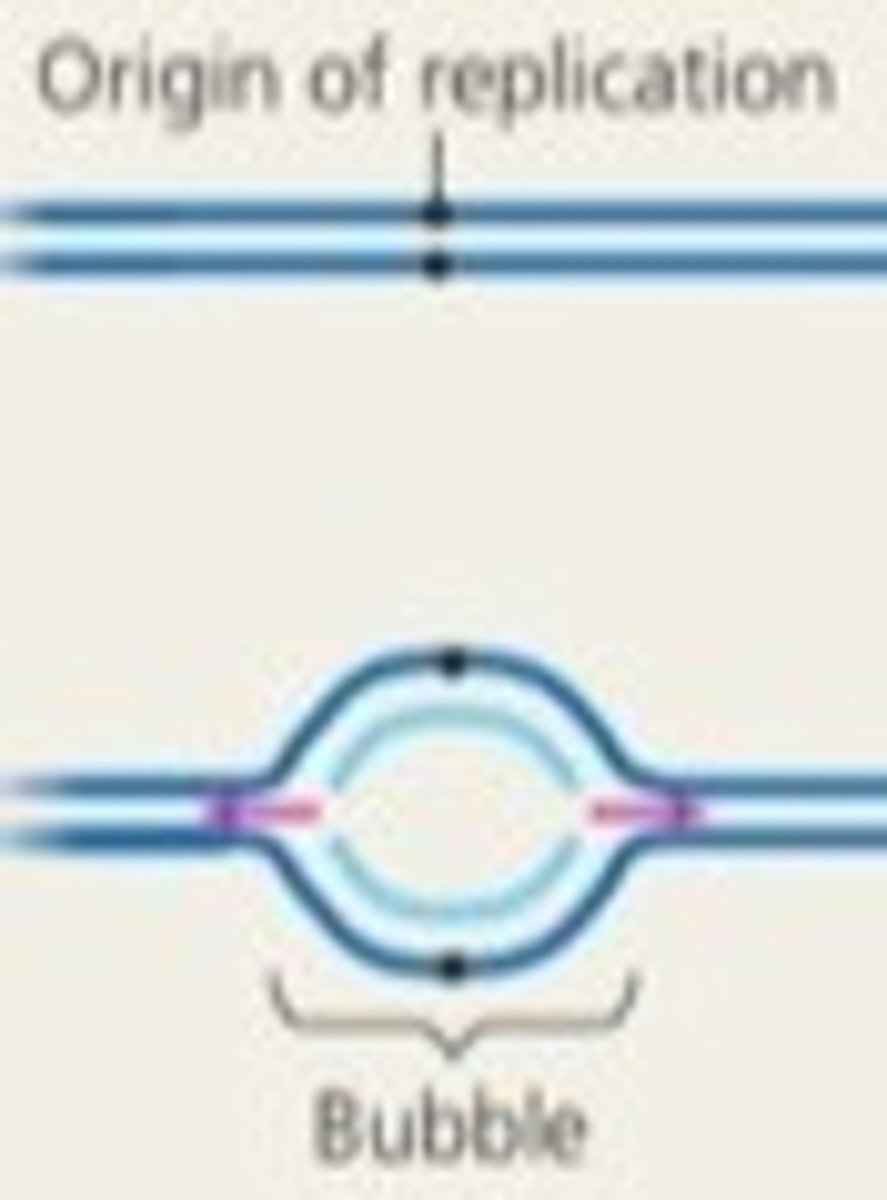
telomeres
Repeated DNA sequences at the ends of eukaryotic chromosomes
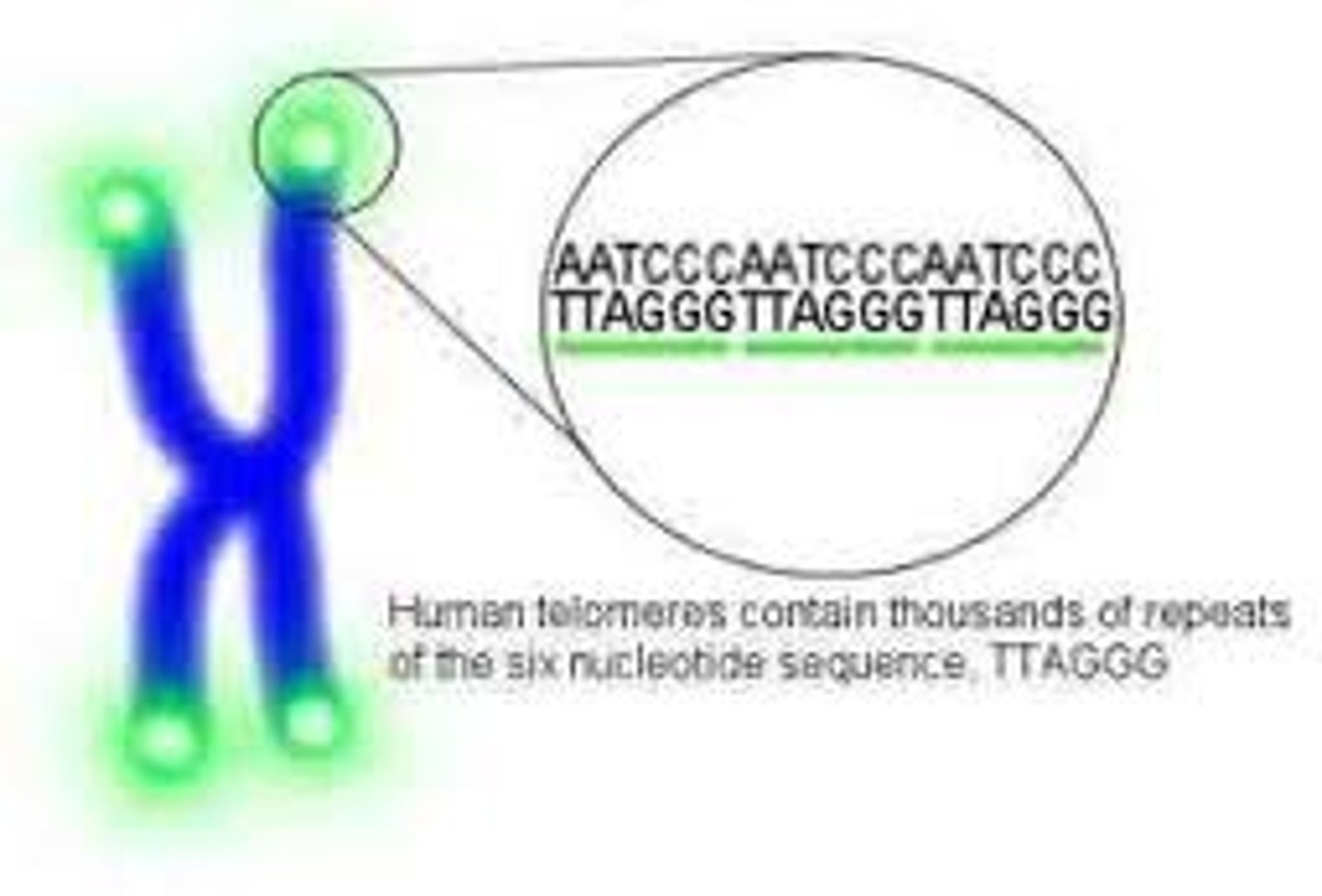
DNA polymerase
The enzymes which synthesize or polymerize the new DNA strand during DNA replication
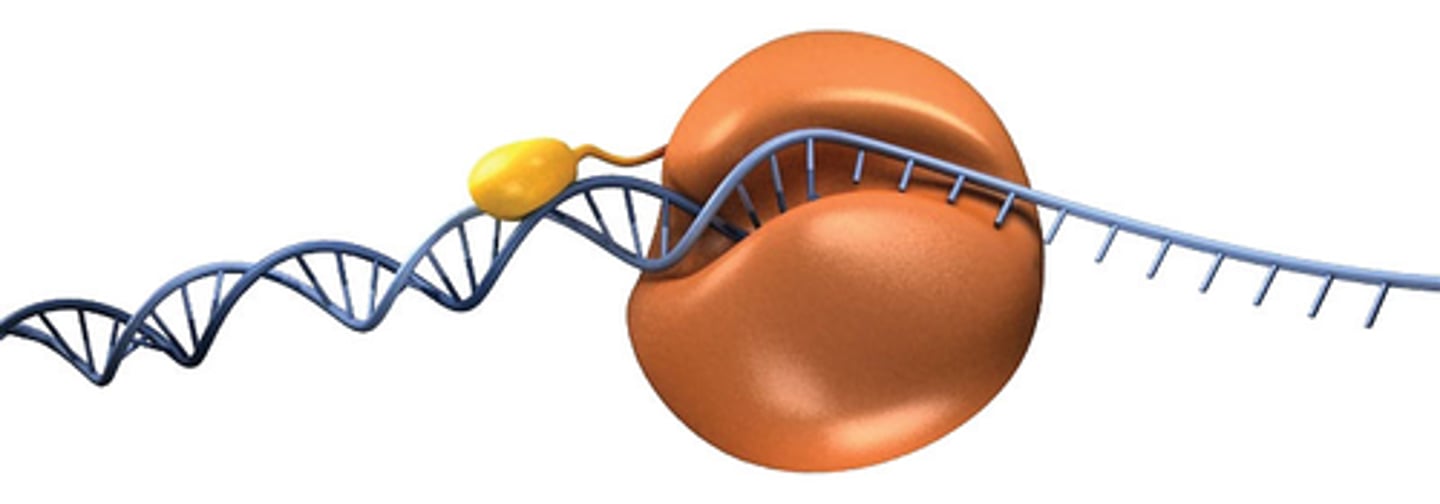
How DNA polymerase works
DNA polymerase uses energy stored in triphosphate of the incoming nucleotide to form the phosphodiester bone and extend DNA strand
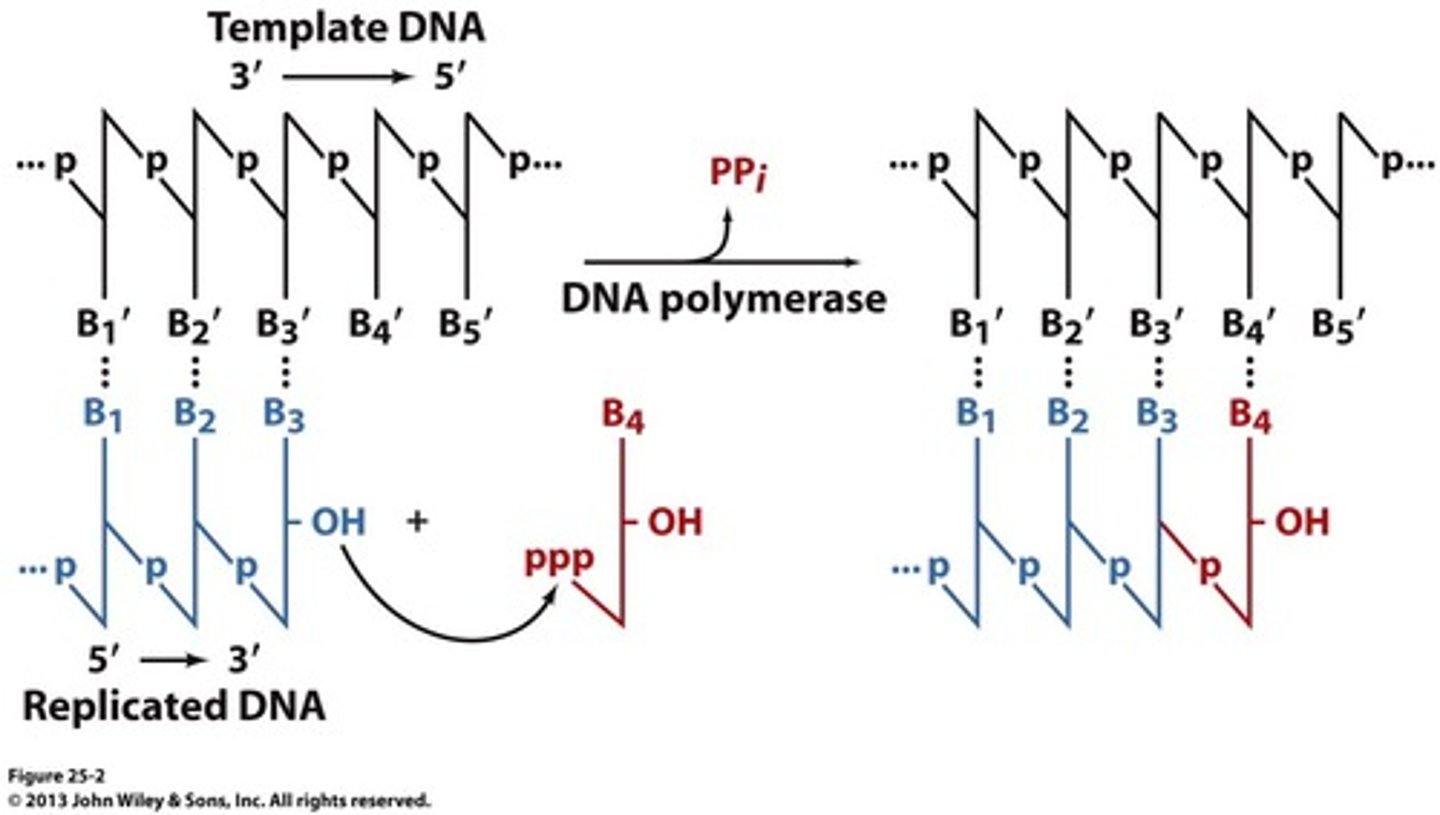
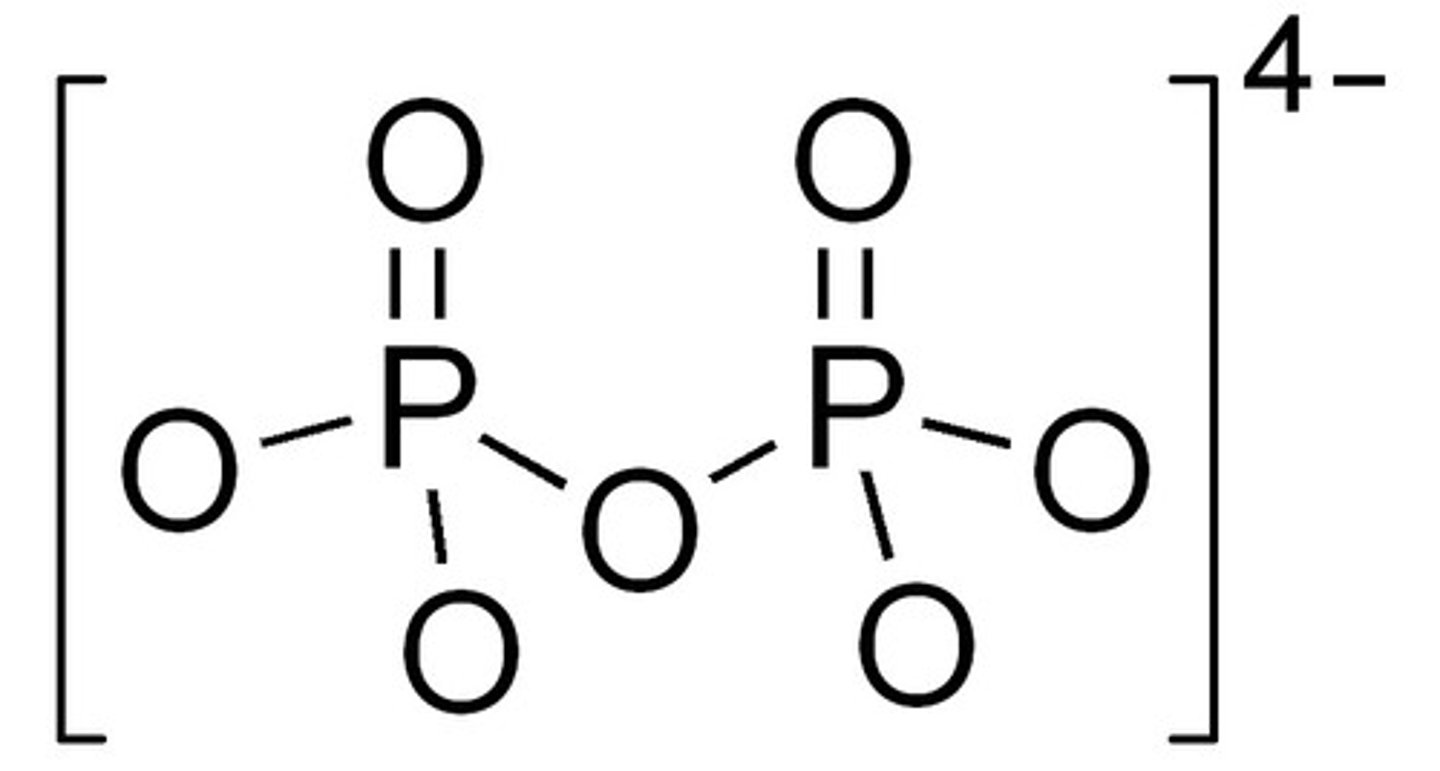
pyrophosphate
byproduct of DNA synthesis which hydrolyze to form 2 Pi; makes the reaction irreversible
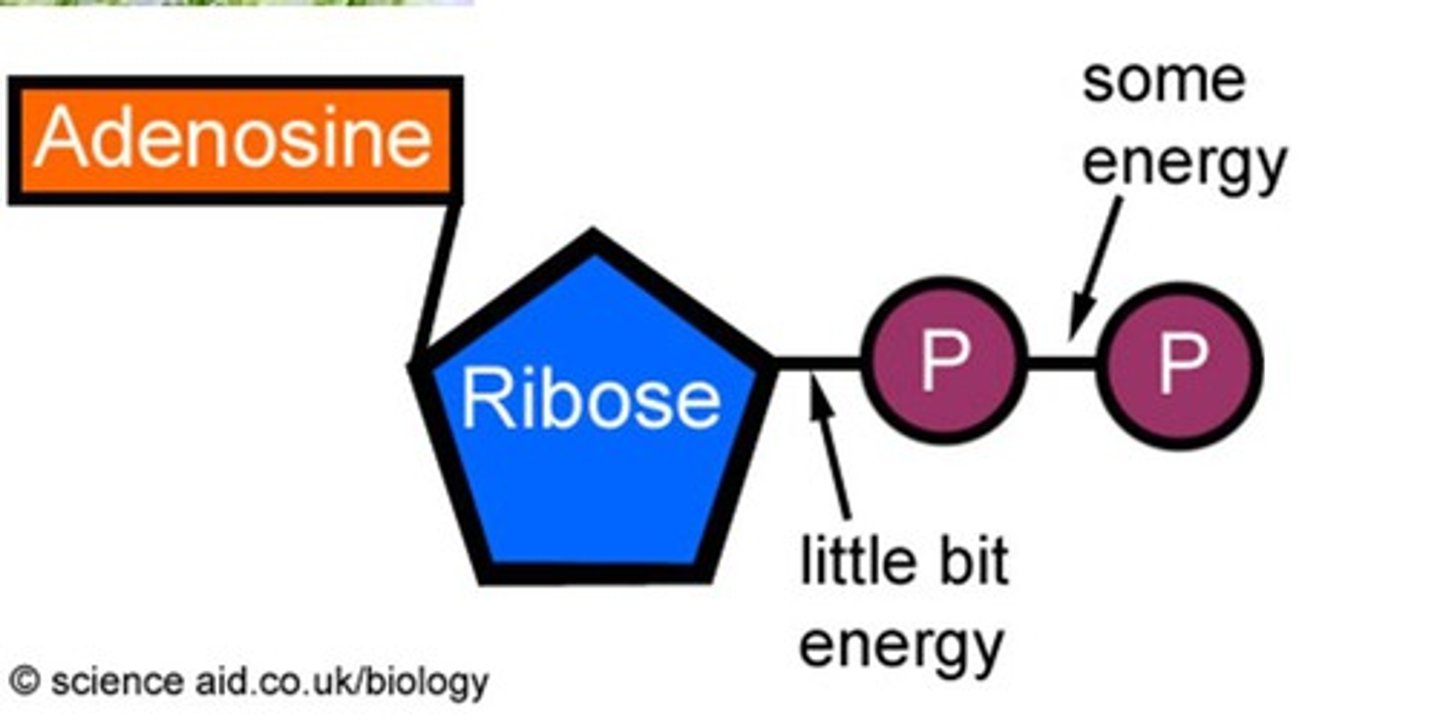
What if a nucleoside used a diphosphate
the reaction would be closer to equilibrium and reversible
DNA Exonucleases
Enzymes which remove nucleotides DNA
specific exonucleases can remove nucleotides from the 3' end or 5' end of DNA
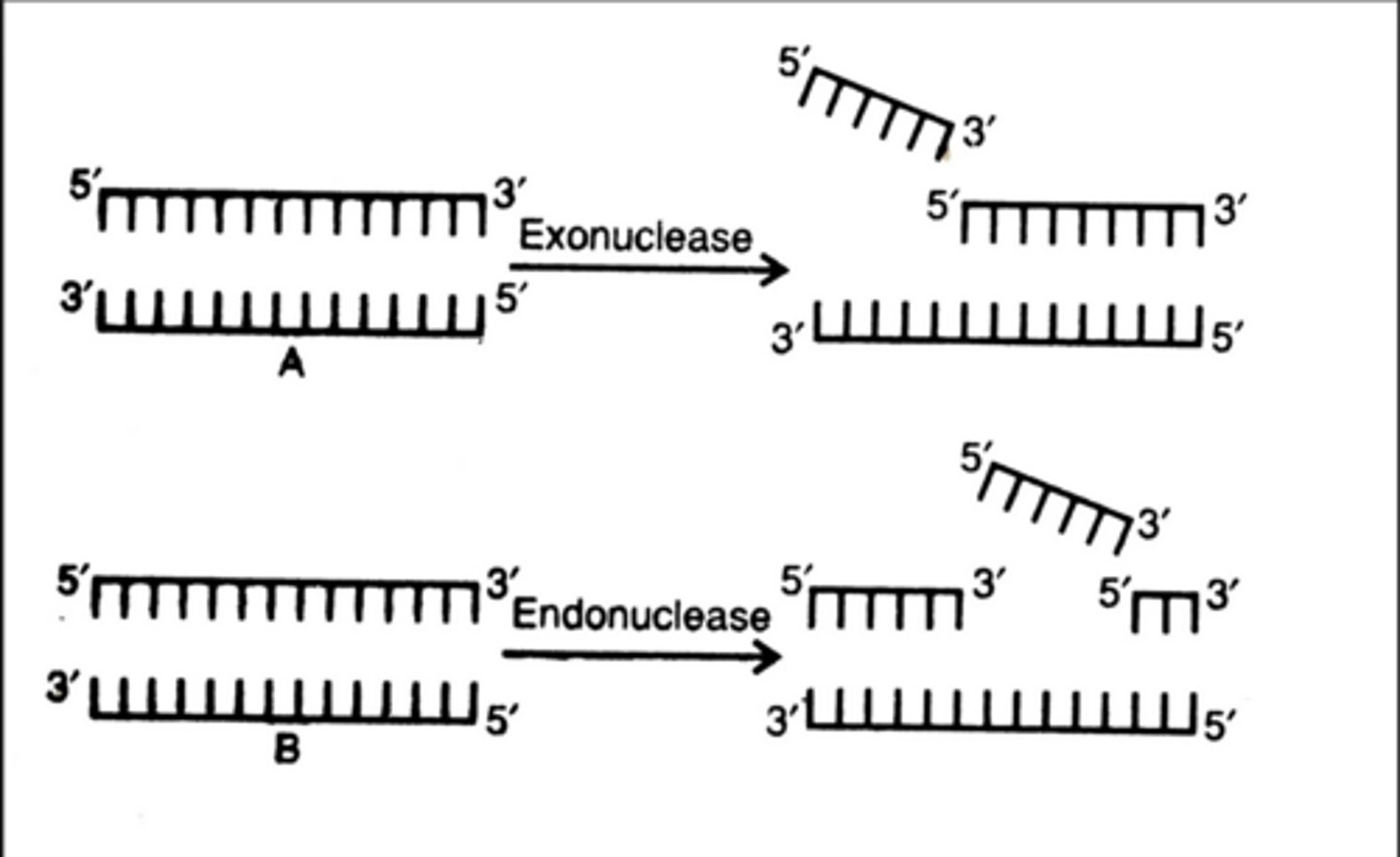
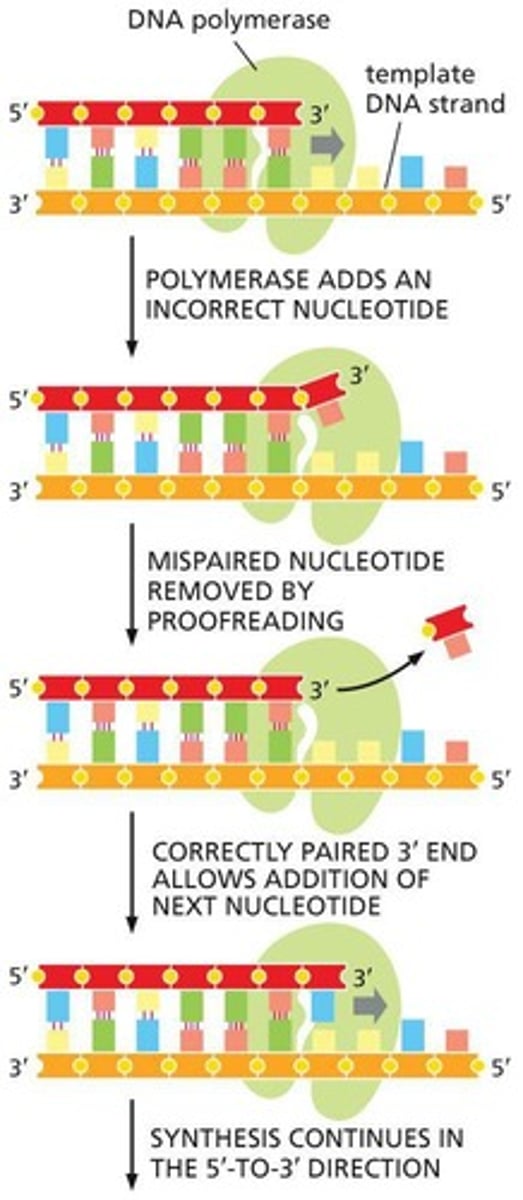
3' to 5' exonuclease
The repair mechanism in DNA Polymerase that cuts out DNA to repair it when there is a mismatched basepair
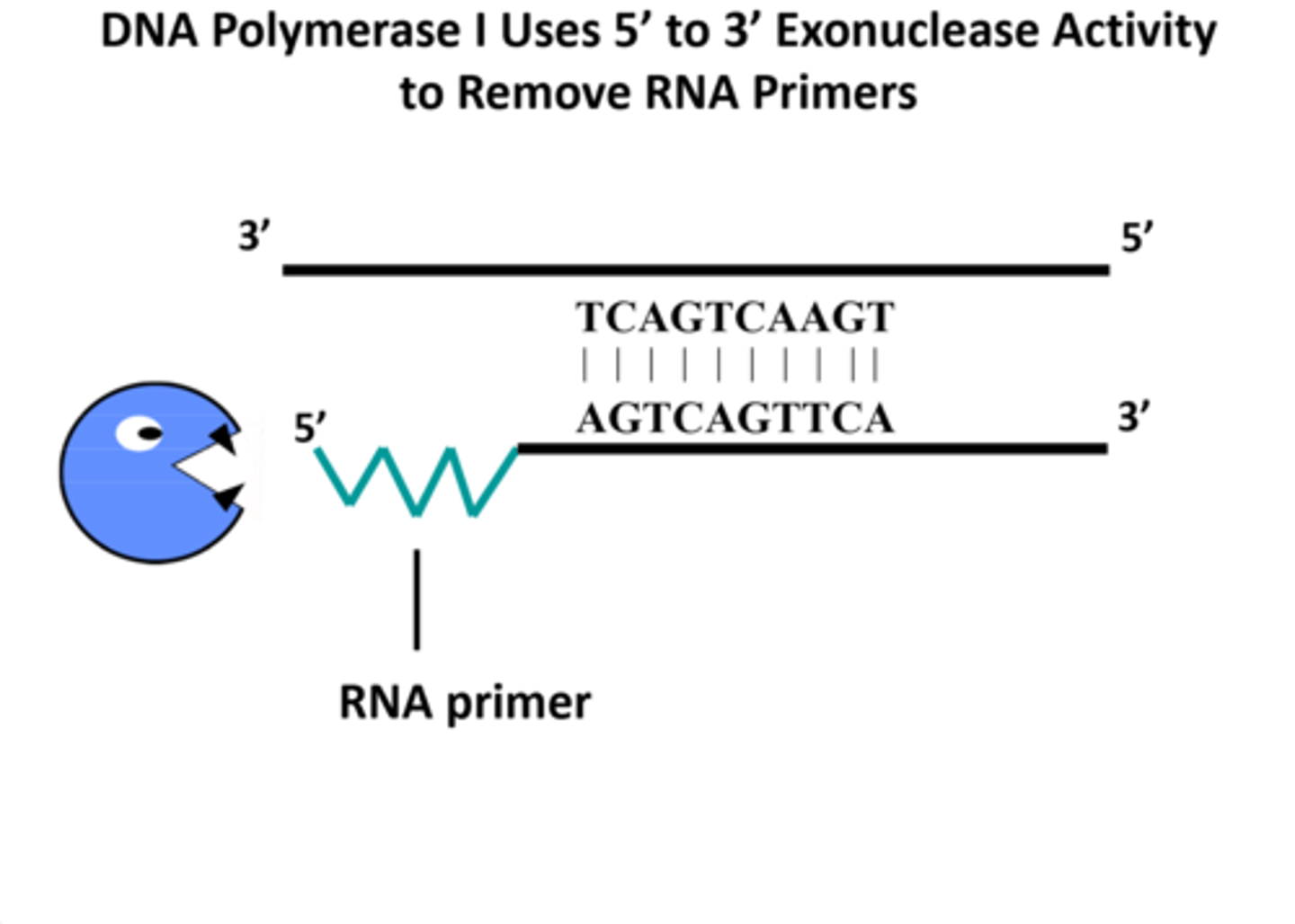
5' to 3' exonuclease
removes RNA primer
POLYMERIZING DNA polymerases
an only polymerize (add nucleotides to the DNA strand) in one direction, the 5′ to 3′ direction
Alpha polymerase
only one that can do de novo synthesis
RNA-DNA hybrid primer synthesis on both leading and lagging DNA strands
POL A/primase complex initiates DNA synthesis in eukaryotic replication
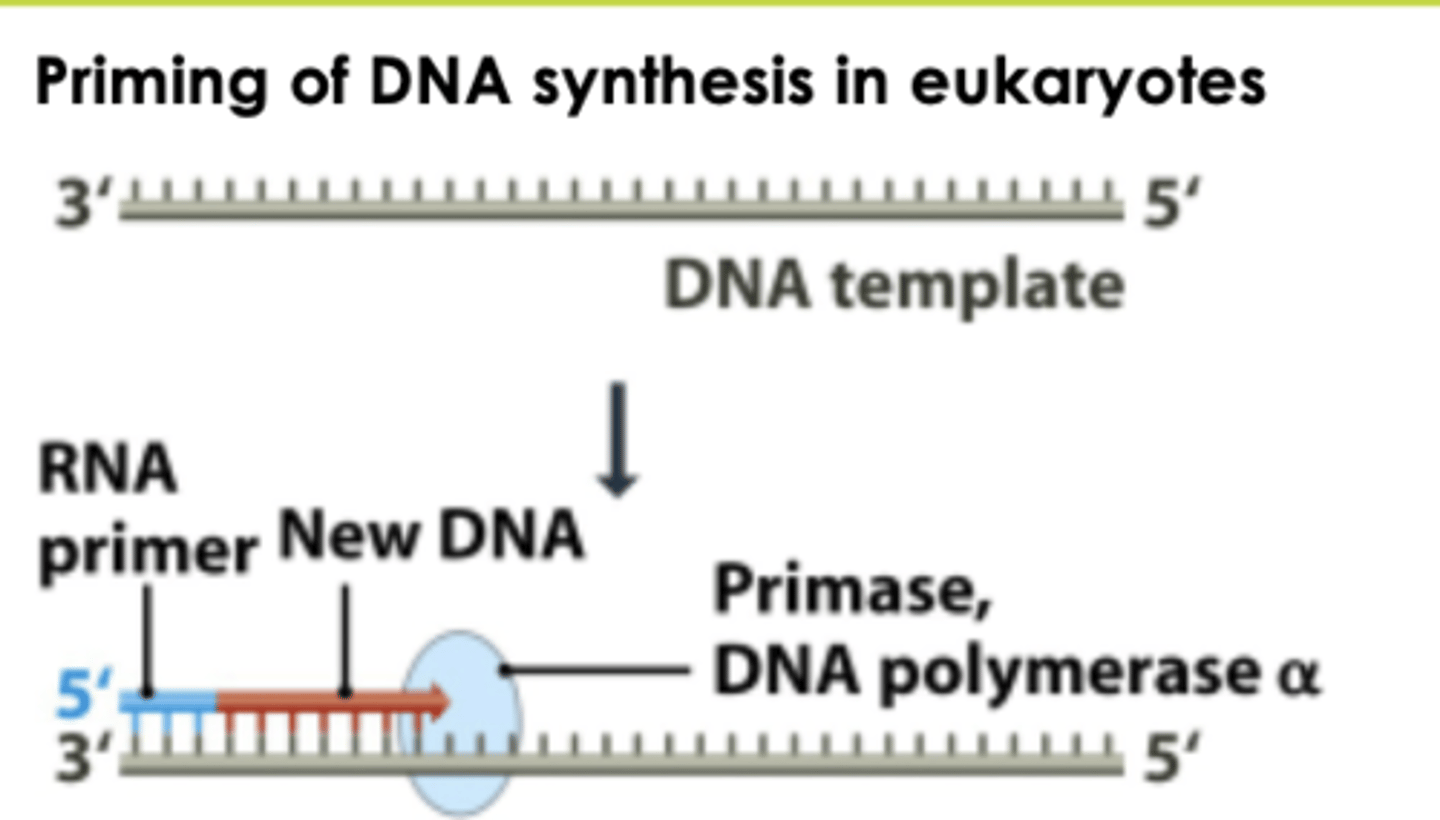
Beta polymerase
DNA repair
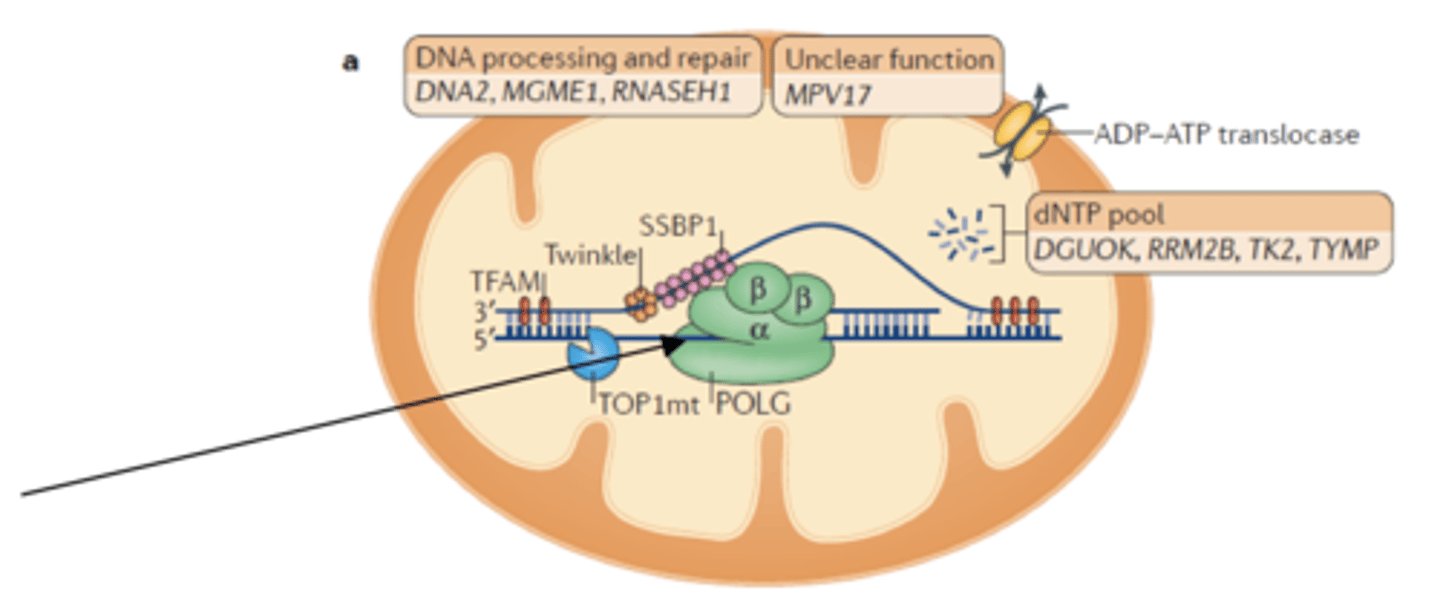
Gamma polymerase
replicate mitochondria DNA
- 3' to 5' exonuclease (proofreads)
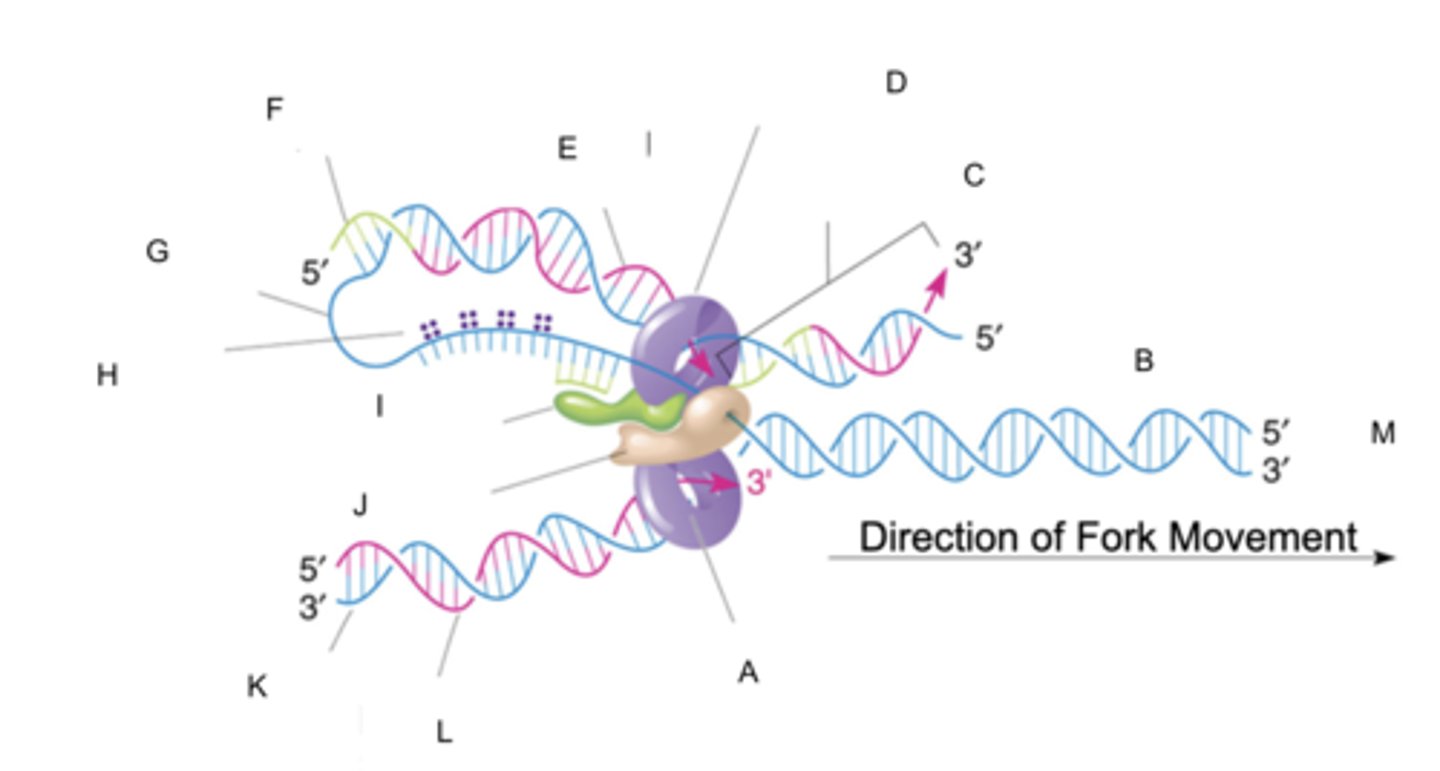
Delta polymerase
polymerize (replicate) DNA on lagging strands
fill DNA gaps when primer is removed
- 3' to 5' exonuclease (proofreads)
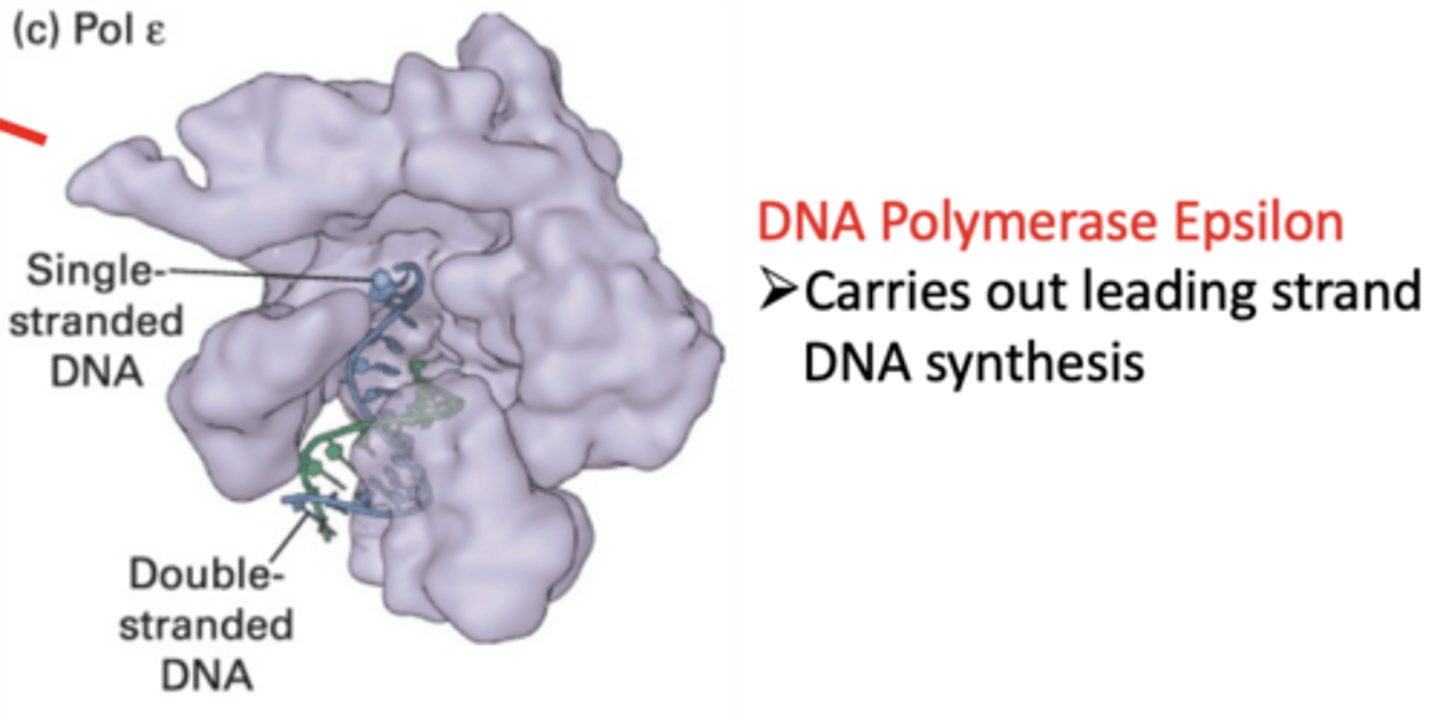
Epsilon polymerase
polymerize (replicate) DNA on leading strand
- 3' to 5' exonuclease (proofreads)
POLYMERIZING DNA polymerases can only polymerize nucleotides in one direction
the 5′ to 3′ direction
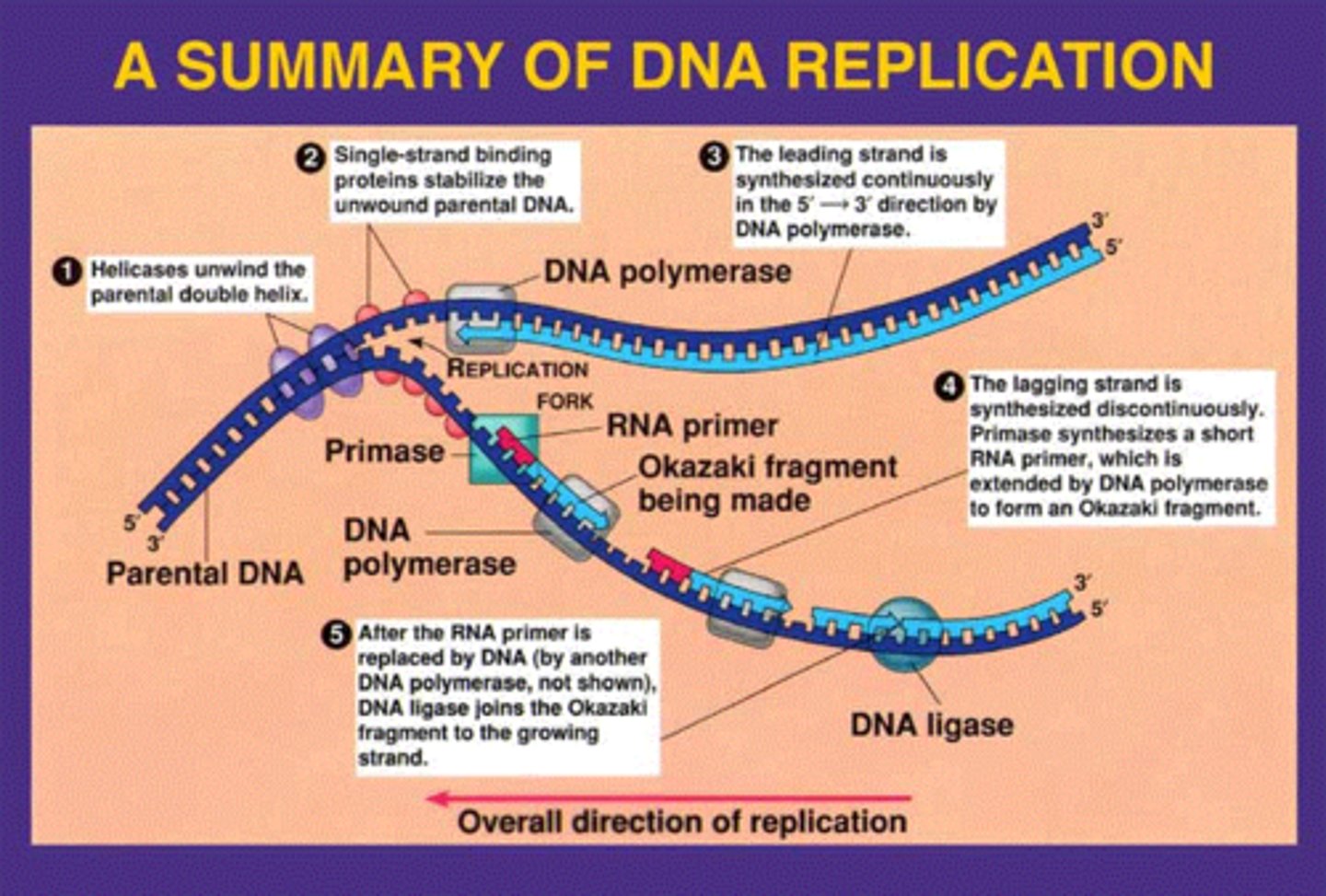
DNA Chain Elongation by
occurs in 5' to3' direction by adding one nucleotide at a time to 3'-OH of the nucleotide
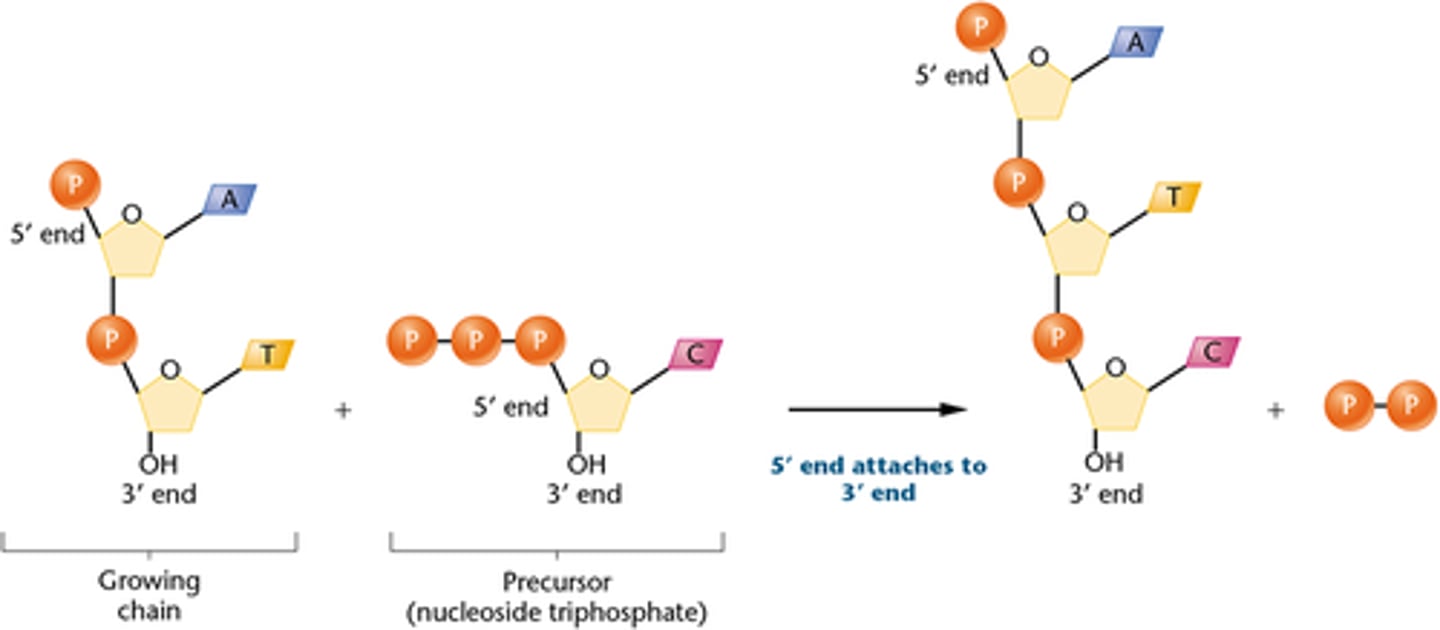
when a nucleotide is added
two terminal phosphates cleaved off:
provides newly exposed 3'- OH
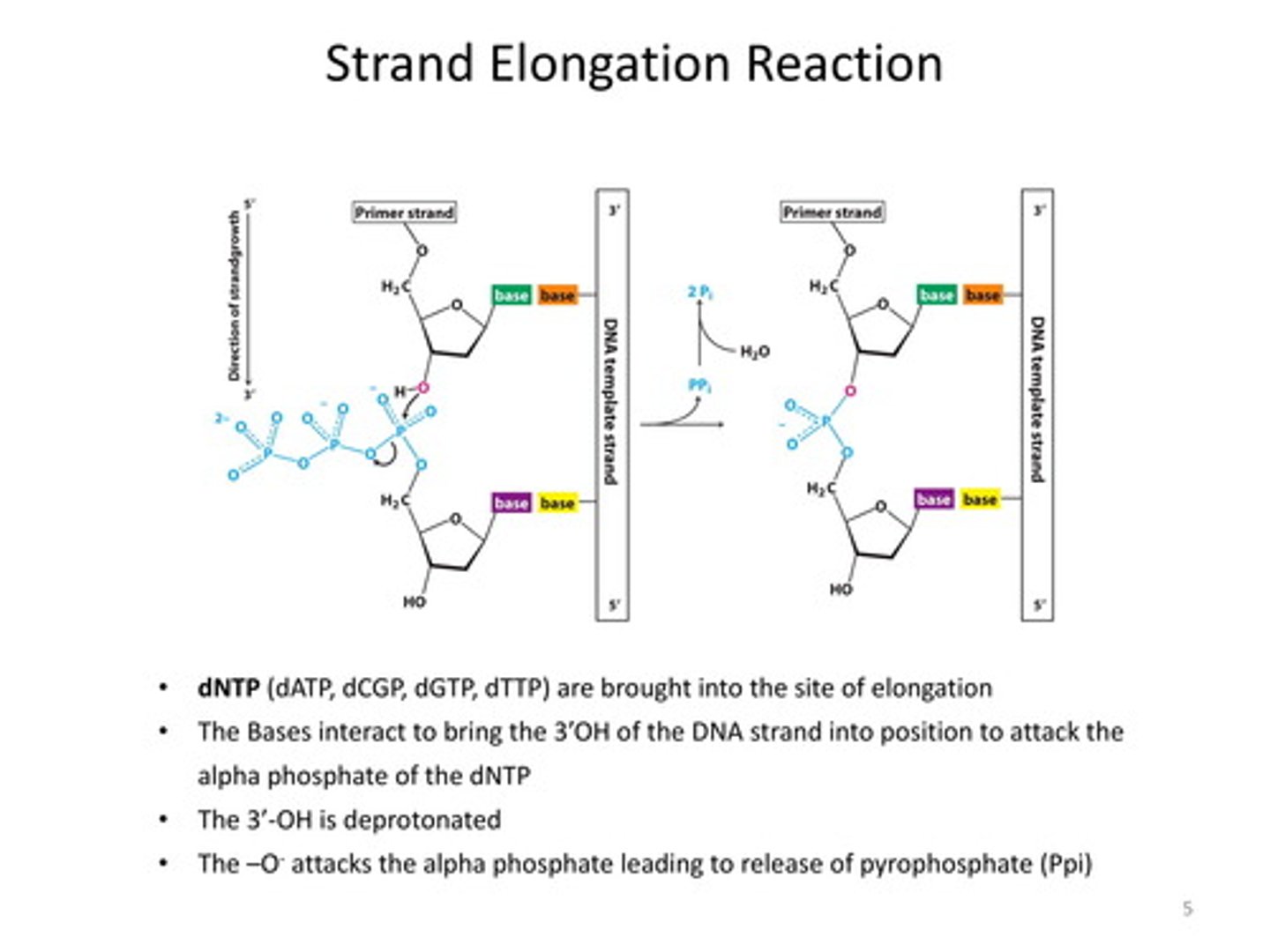
3'-OH end
an participate in addition of another nucleotide as DNA synthesis proceeds.
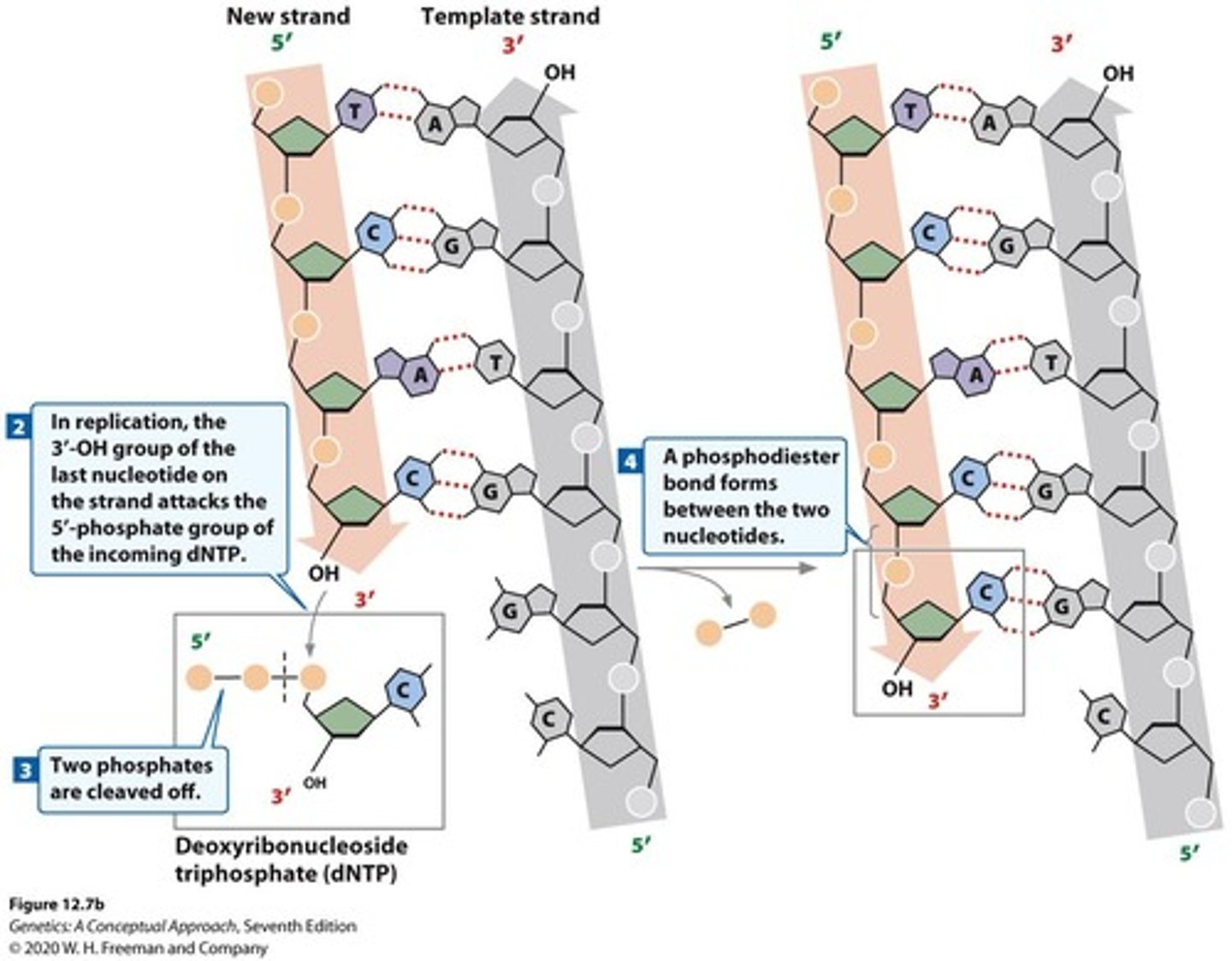
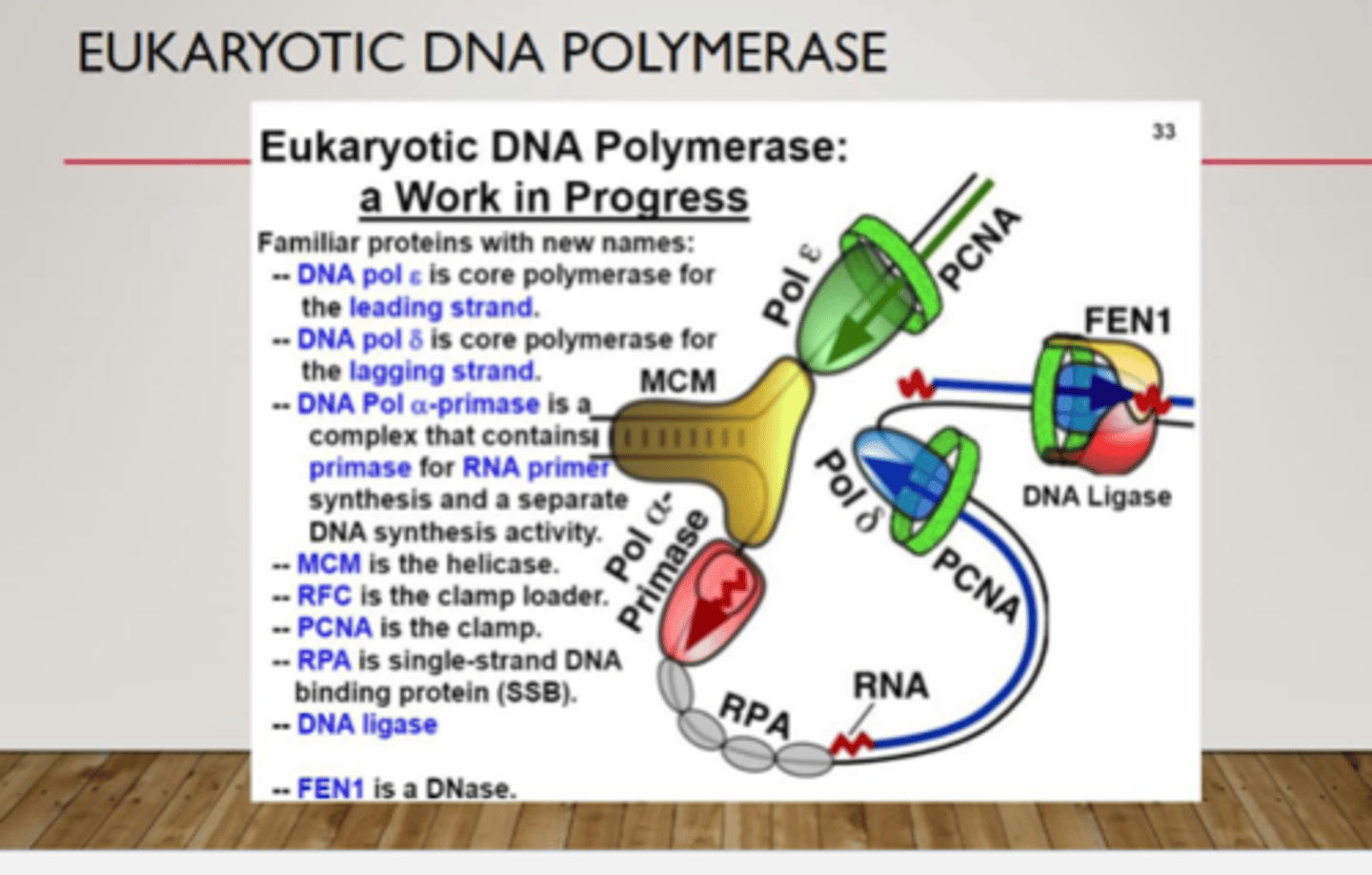
DNA polymerase delta and epsilon
contain exonuclease activity 3'-5' for proofreading their polymerase activity
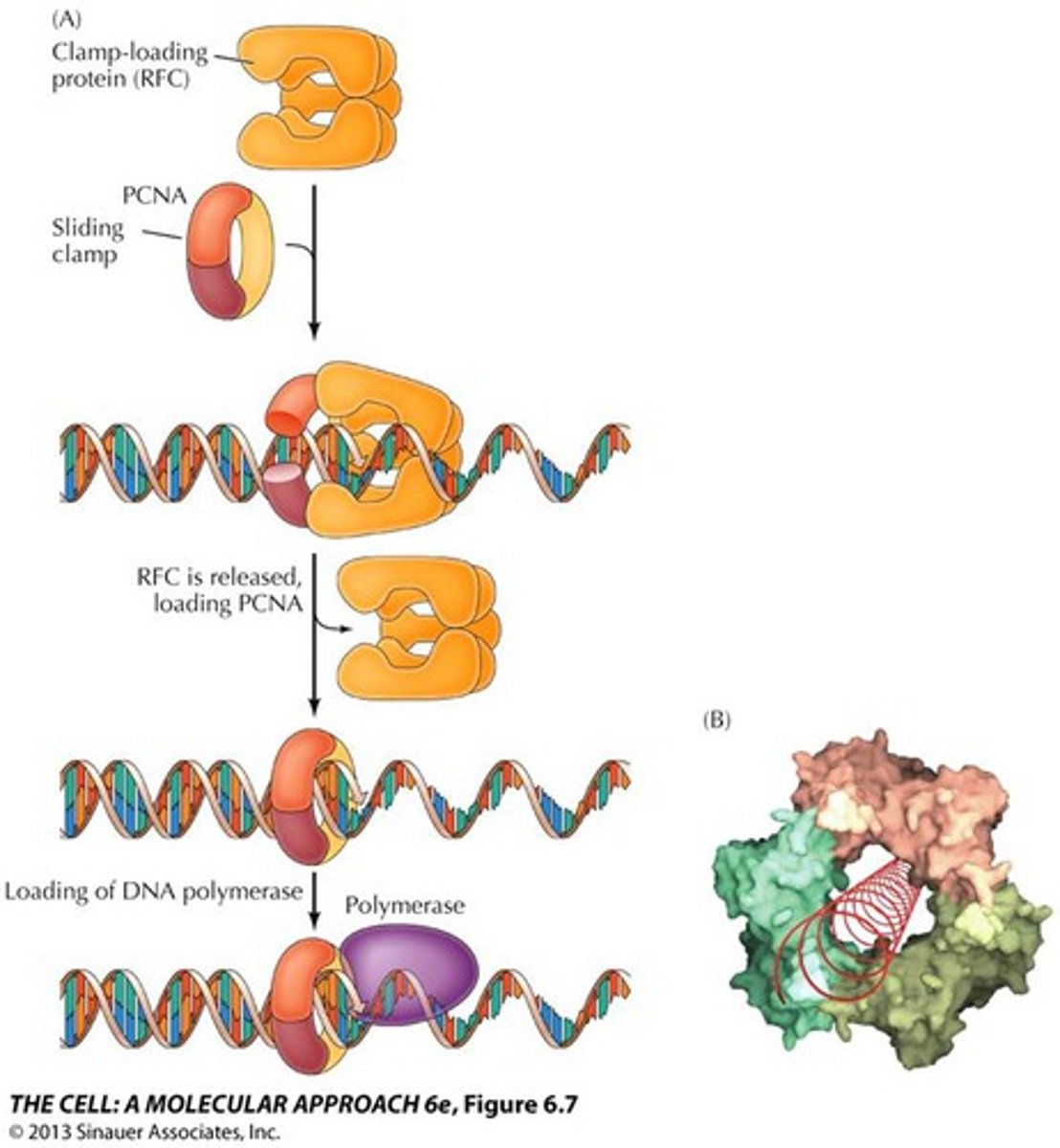
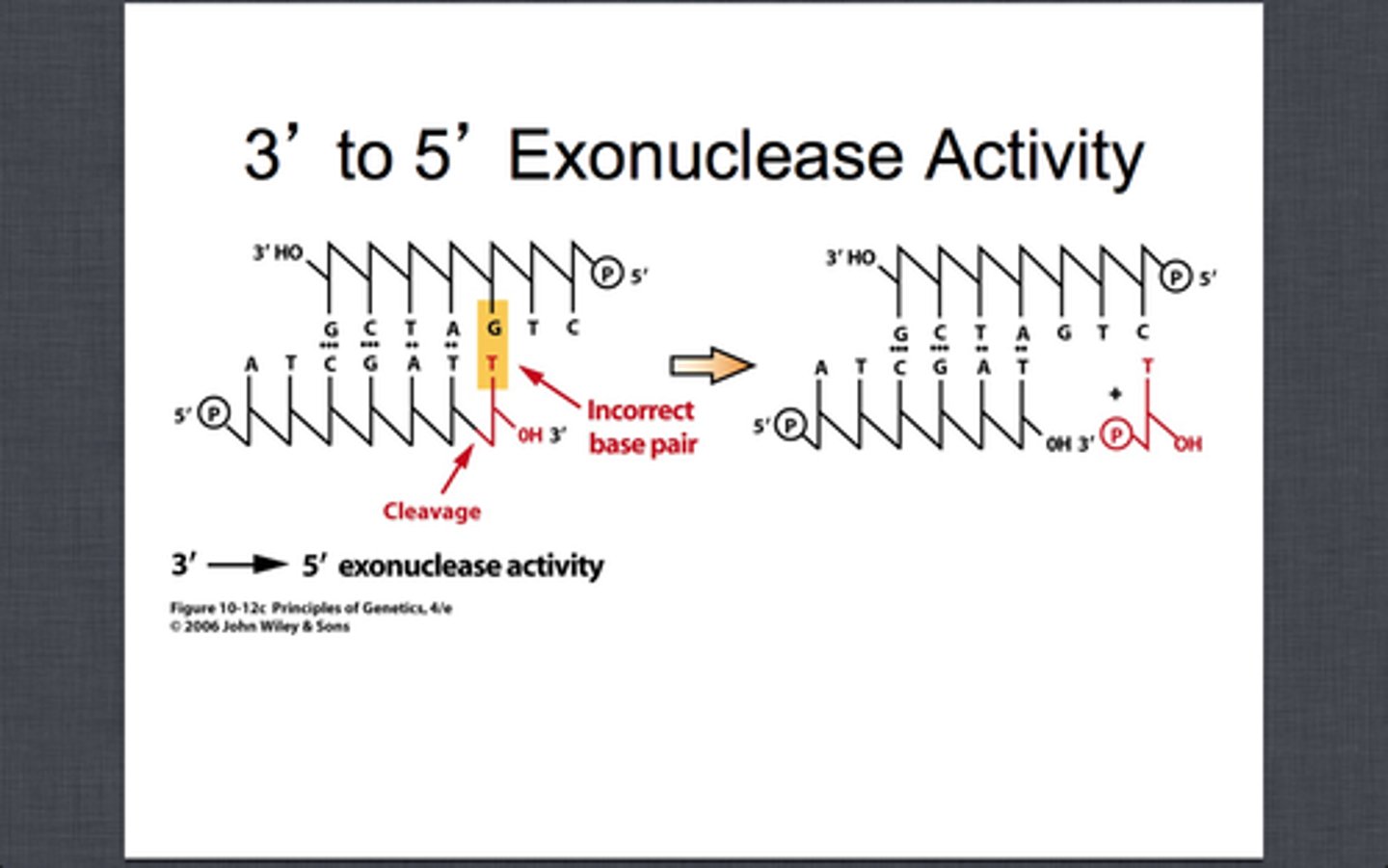
Exonuclease activity 3'-5'
proofread newly synthesized DNA, remove/replace incorrect nucleotides
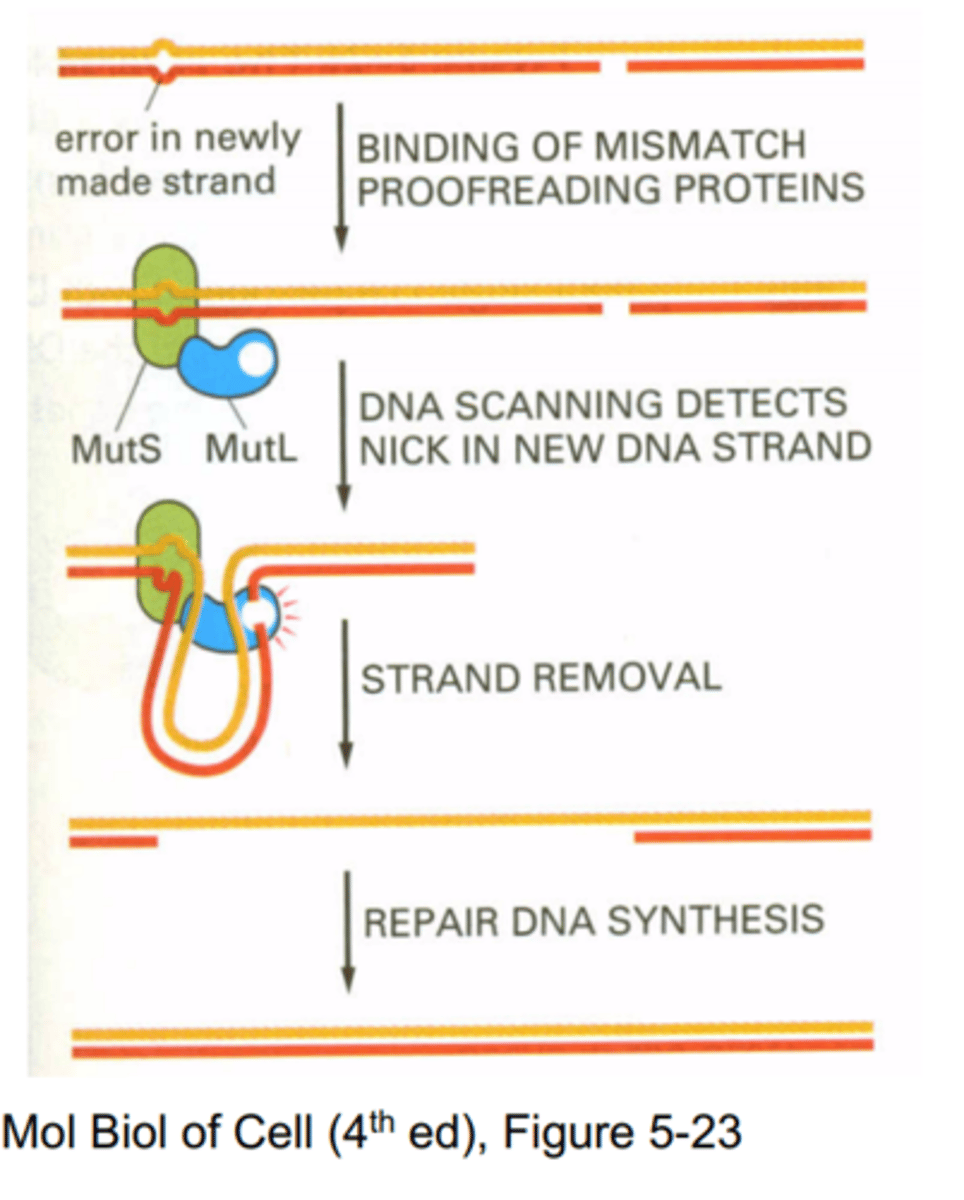
DNA mismatch repair
Mechanism for recognizing and correcting incorrectly paired nucleotides—those that are noncomplementary.
Consensus sequence
a sequence of DNA having similar structure and function in different organisms
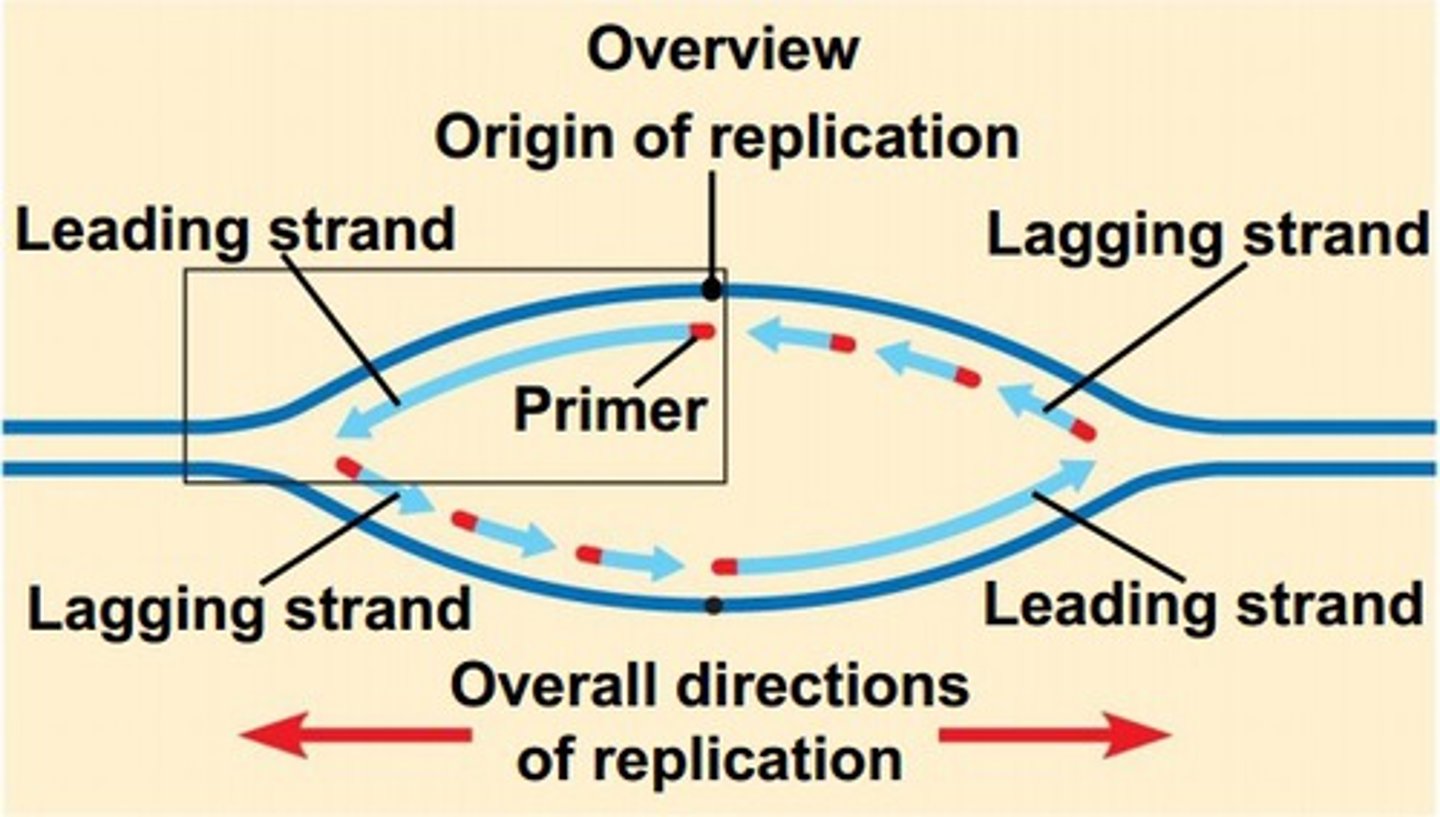
origin of replication
Site where the replication of a DNA molecule begins; consensus sequence; eukaryotic cells have multiple
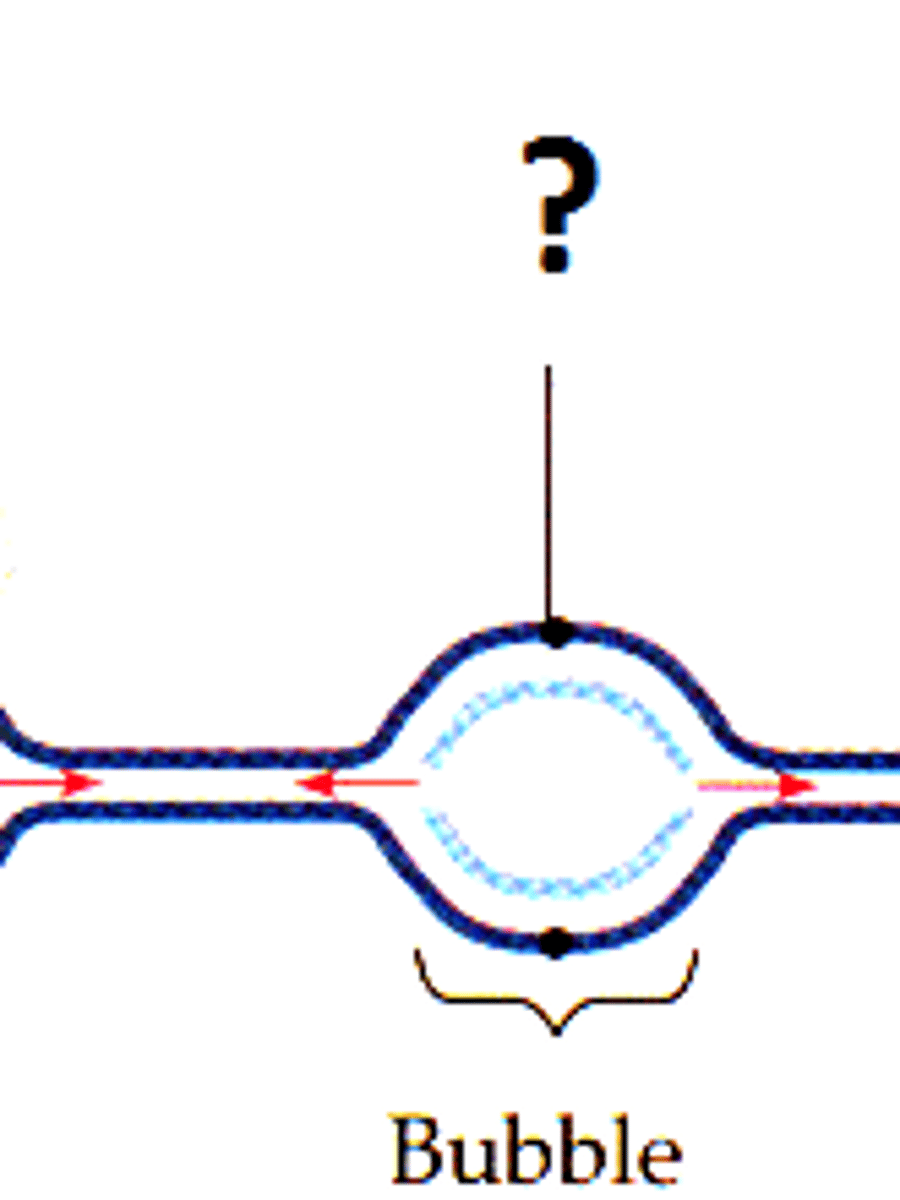
replication bubble
pre-replication protein complex opens the two strands of DNA at the origin of replication
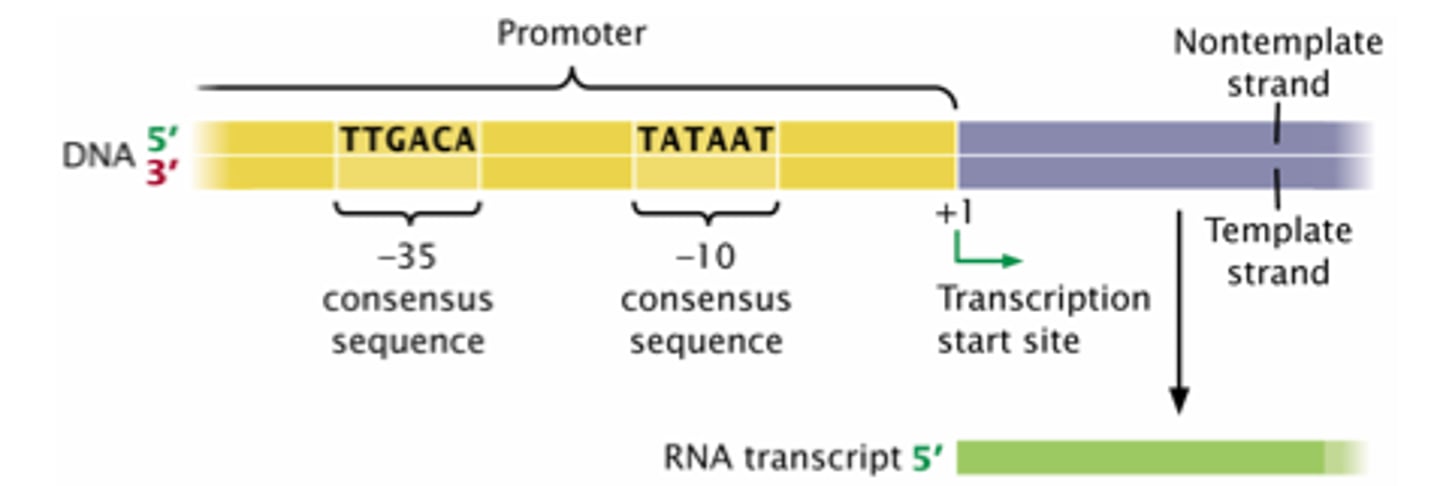
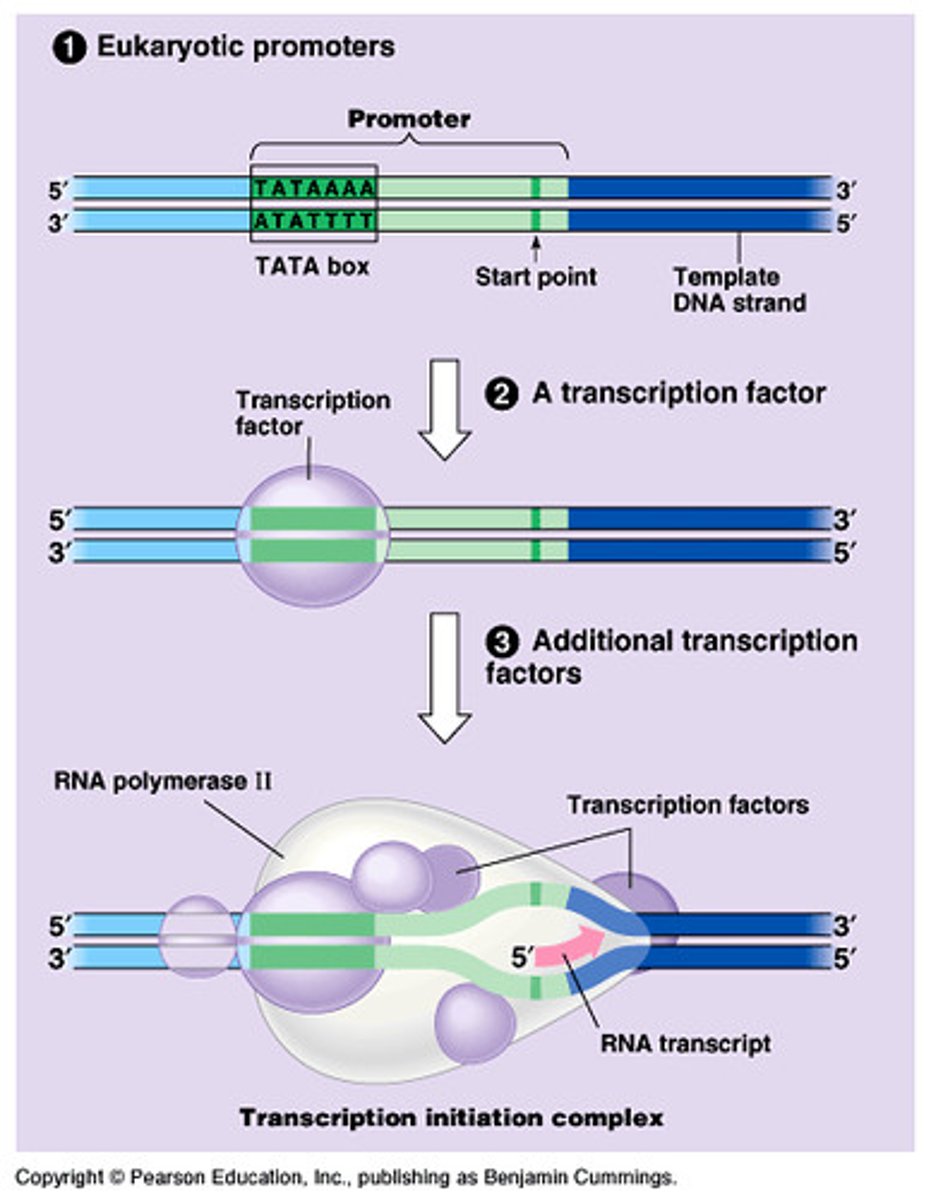
TATA box
crucial promoter DNA sequence; A-T bonds with only two hydrogen bonds are weaker, and easier to separate
bidirectional replication forks
When the replication bubble forms, replication proceeds as the replication forks move, unwinding double stranded DNA
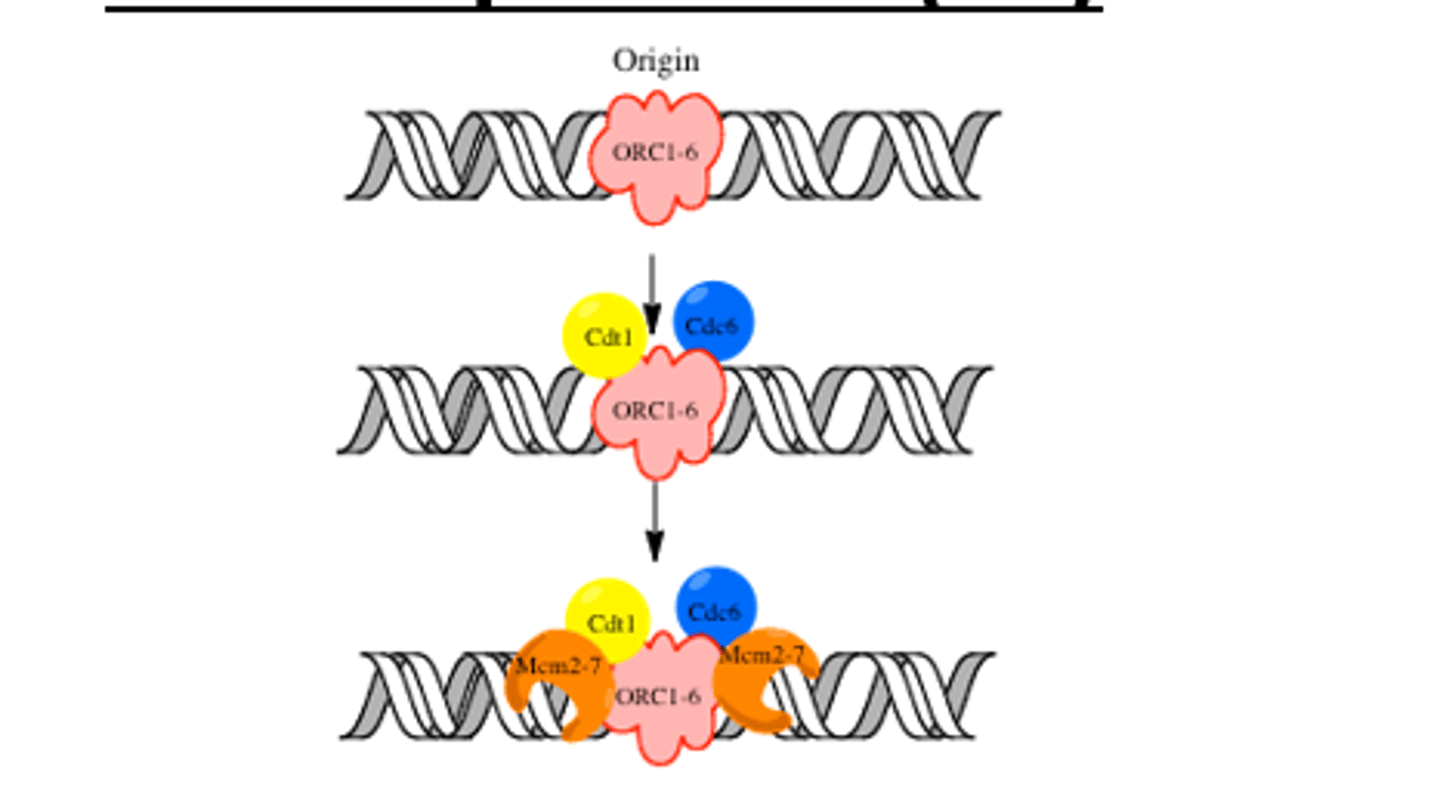
Pre-replication complex
a protein complex that forms at the origin of replication during the initiation step of DNA replication
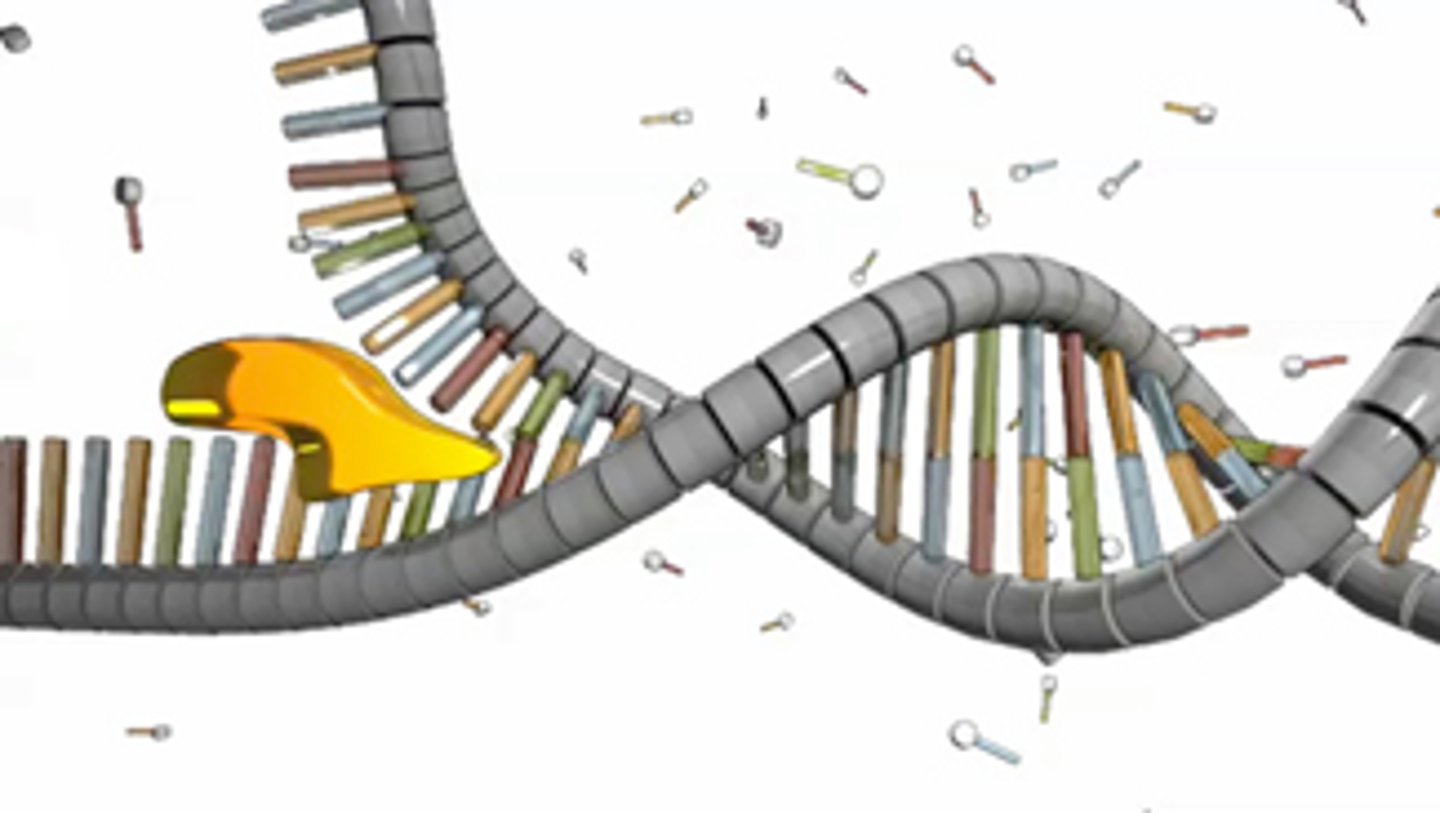
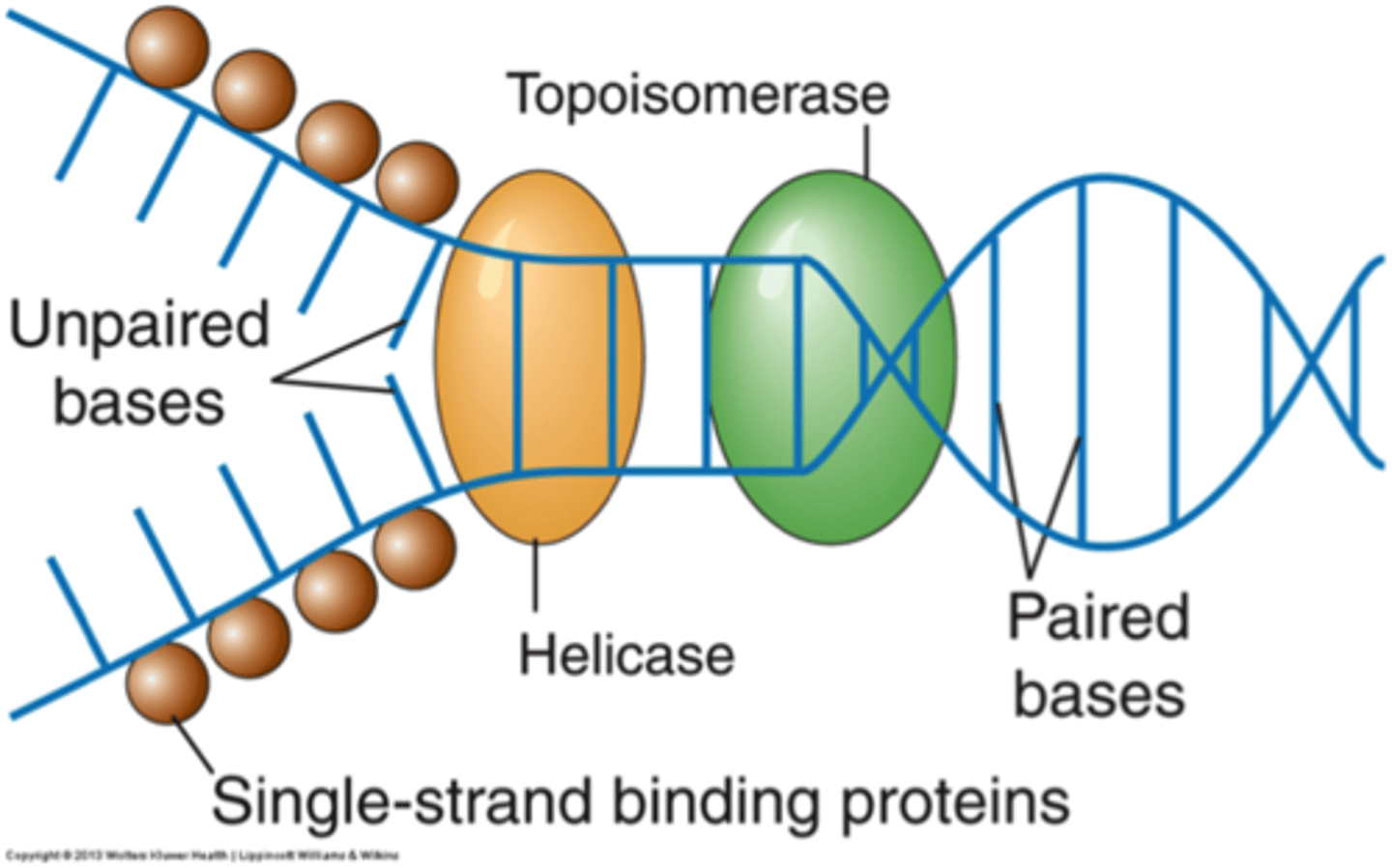
Helicase
Uses ATP as energy to break the H-bonds; After the replication bubble is created, this enzyme unzips the double-stranded DNA helix by breaking the hydrogen bonds between the bases
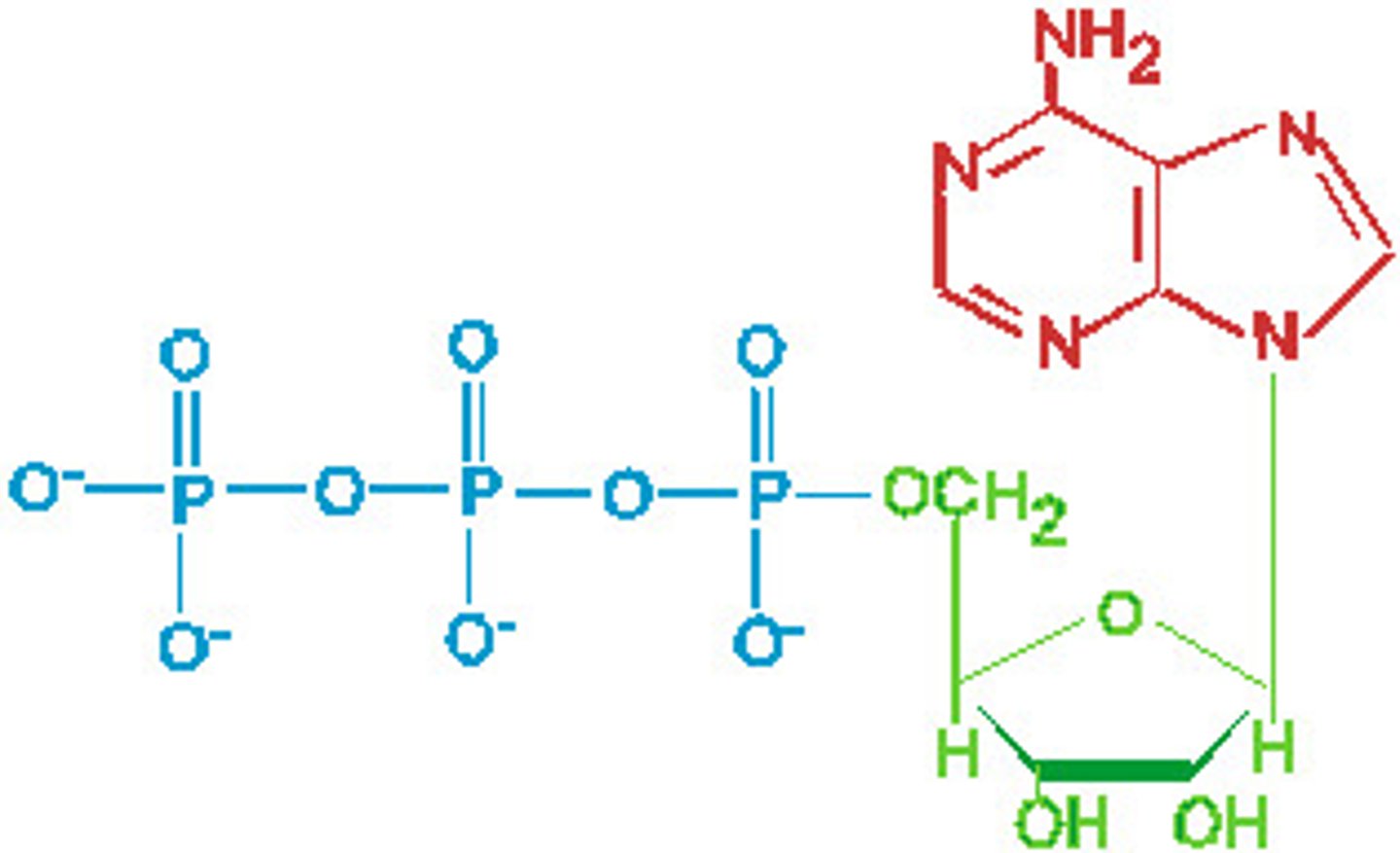
Nucleoside triphosphate pool
essential, tightly regulated, and dynamic building blocks for RNA/DNA synthesis
Single stranded binding protein (SSBP)
Bind the DNA strands and keep them from reannealing (prevent hydrogen bonds to form between the bases)
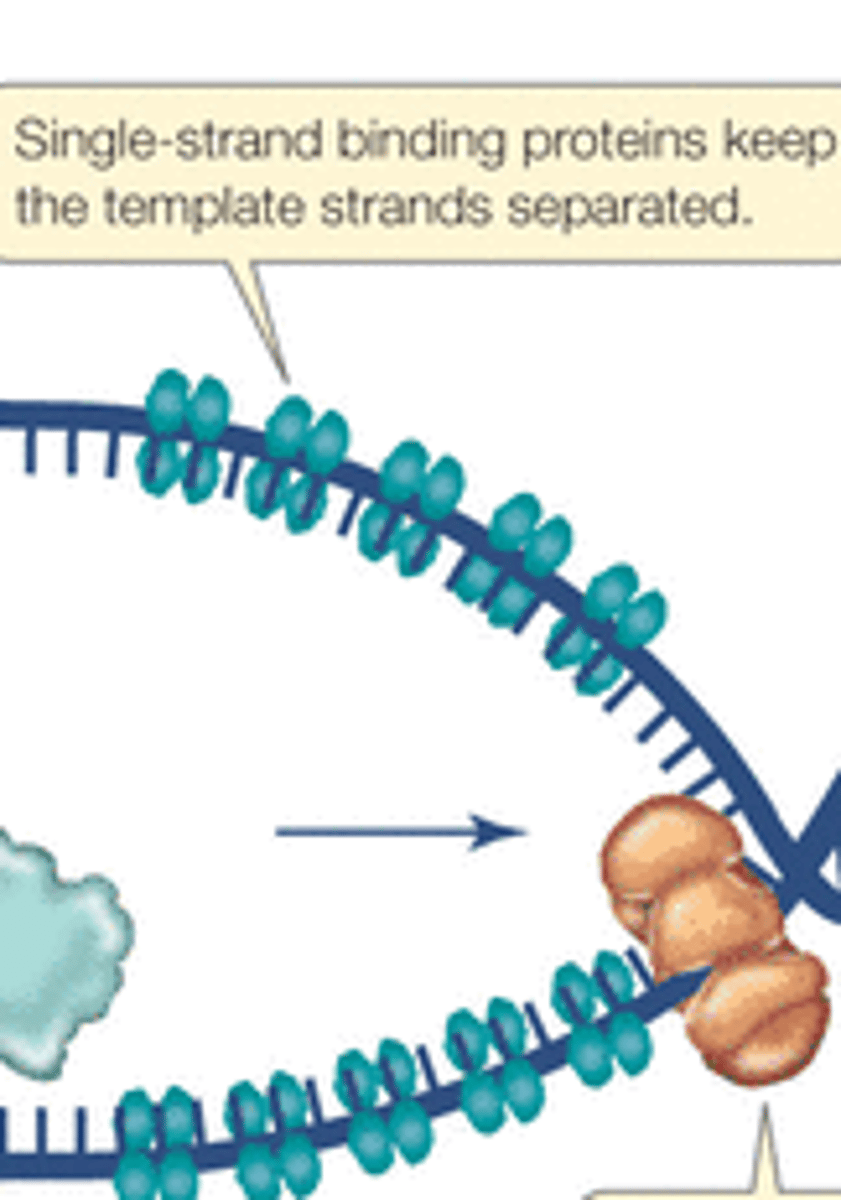
Why need SSBPs?
The complementary strands want to reanneal after the hydrogen bonds between them are broken
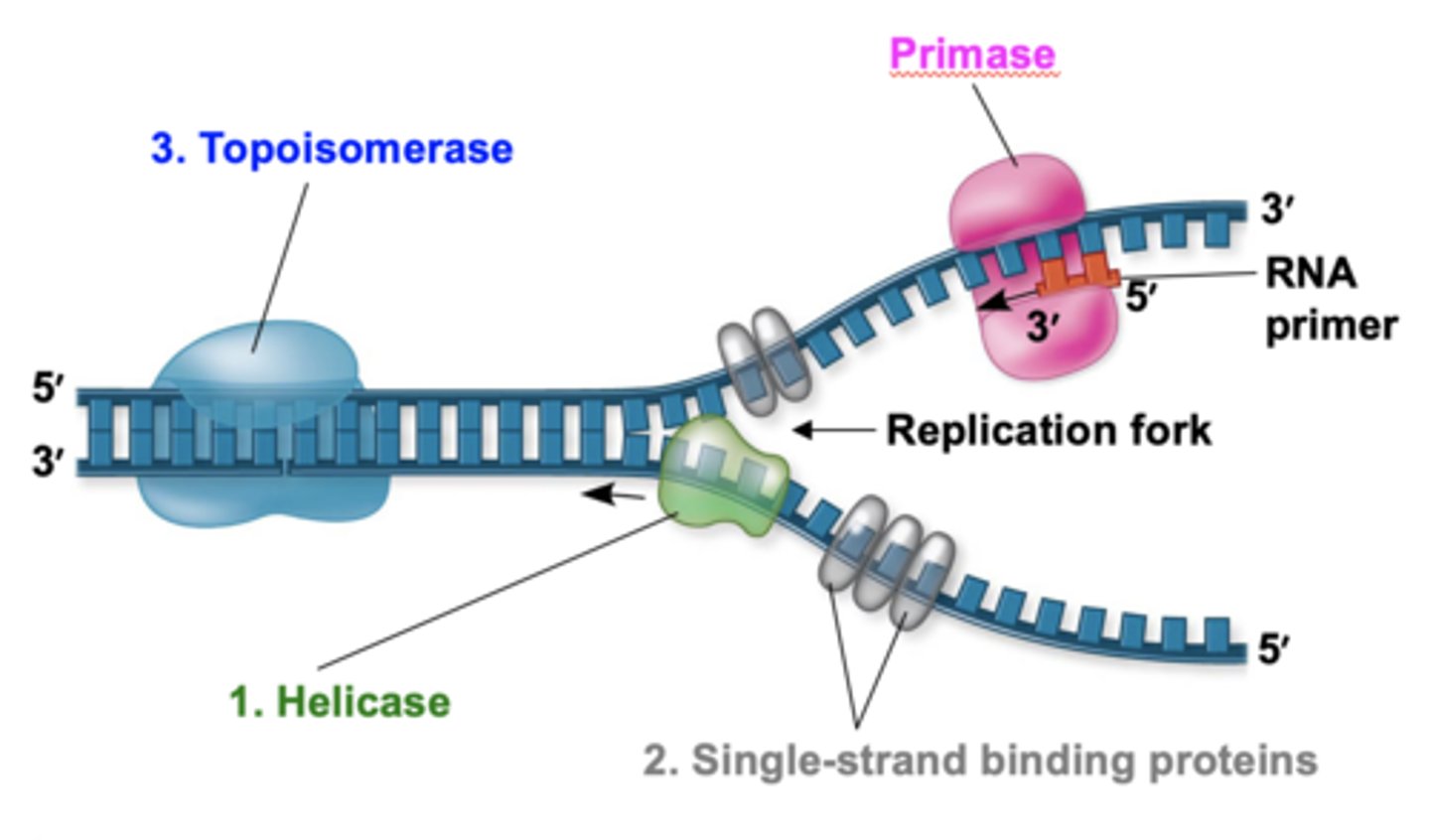
template/parental strand
a strand of DNA that is used to synthesize a complementary strand of DNA or RNA
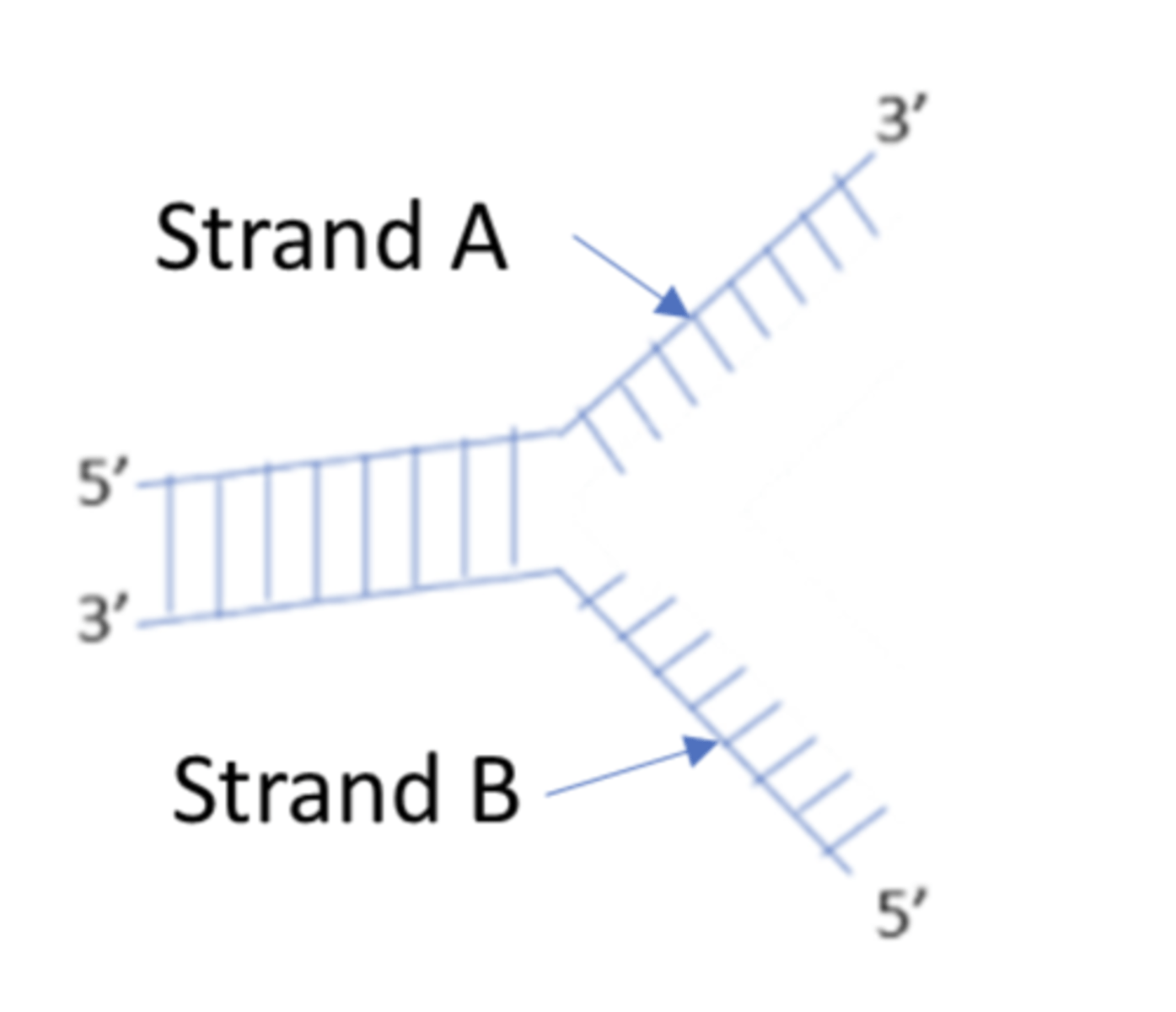
primer
A short segment of DNA that acts as the starting point for a new strand:
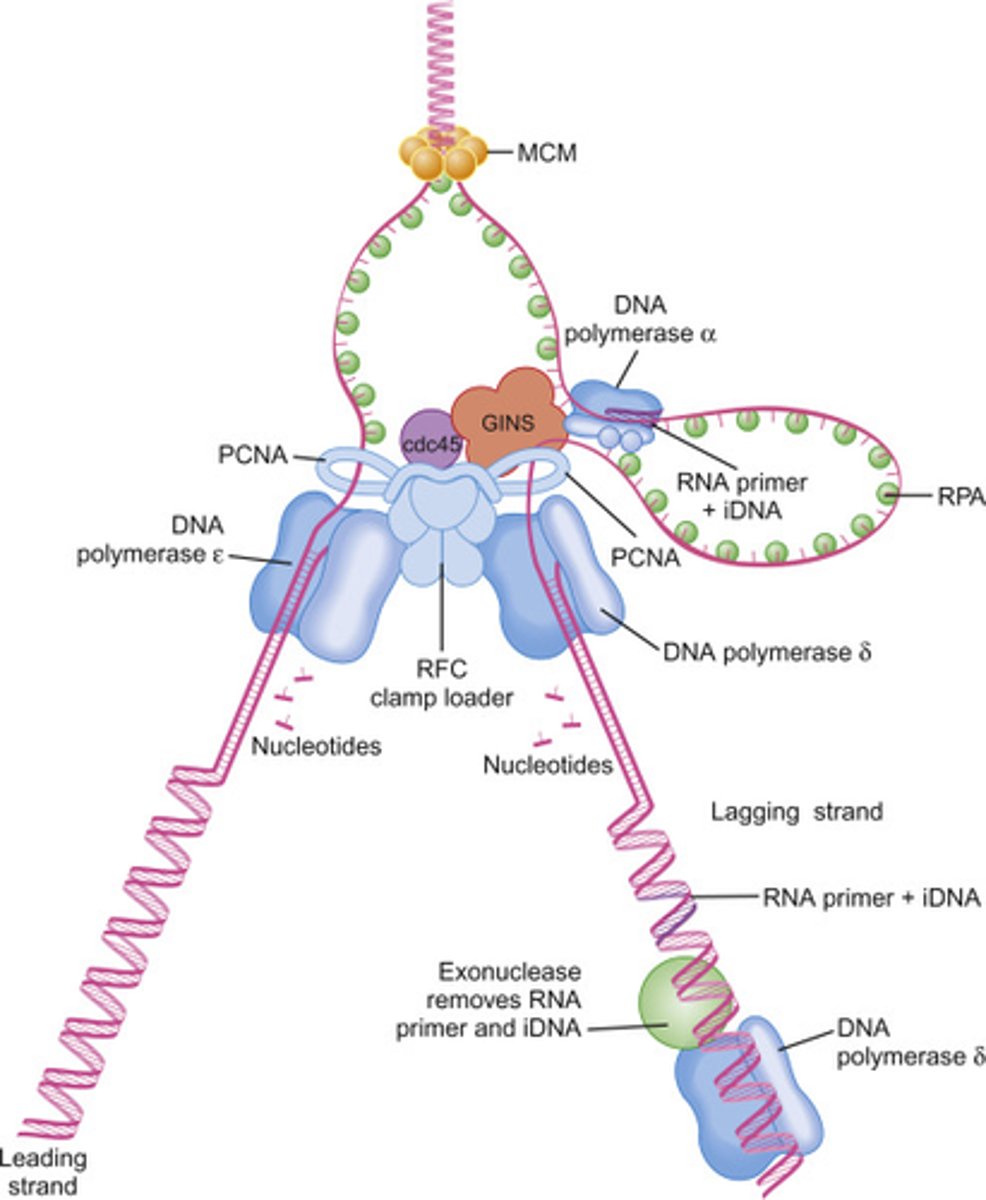
eukaryotic DNA replication is a RNA-DNA hybrid
Short RNA- DNA stretch made by DNA Pol (Alpha)/primase; can start DNA synthesis from scratch
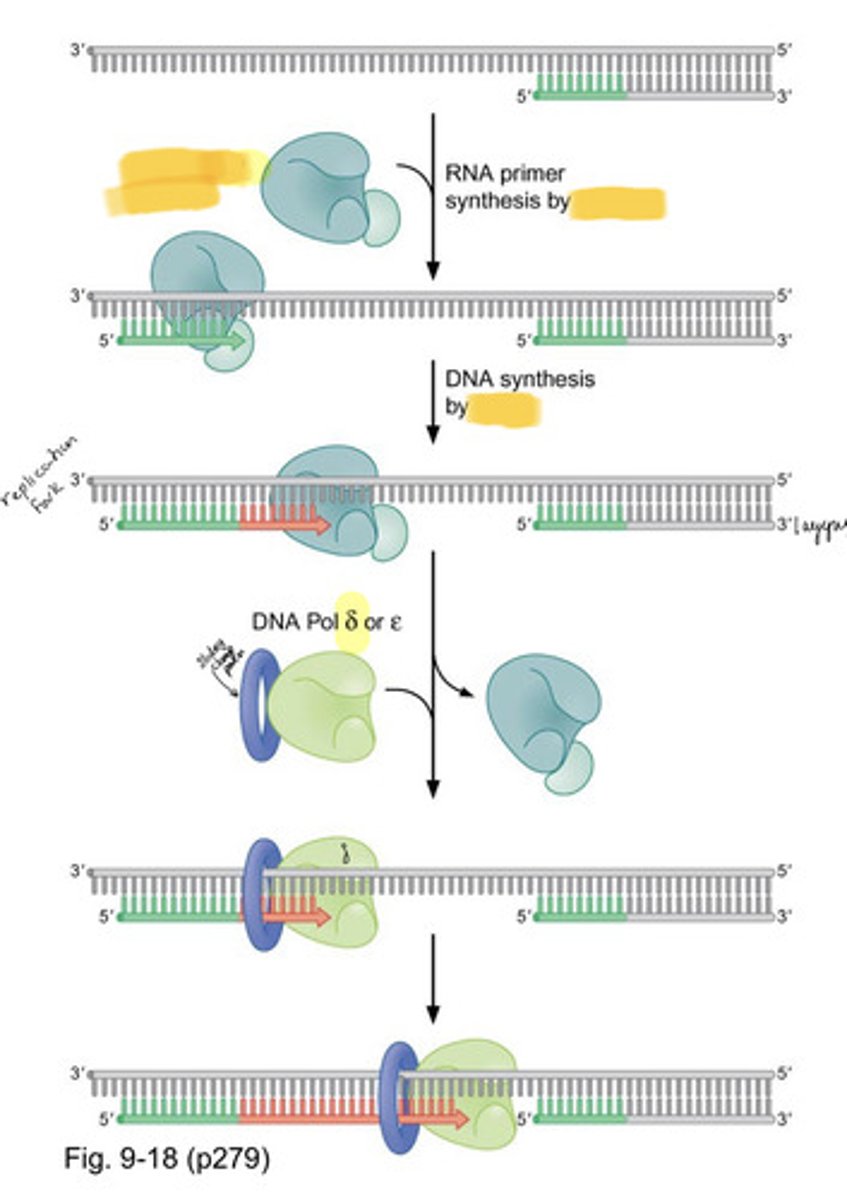
DNA Pol (Alpha)/primase
the only DNA Pol which also has a subunit with primase activity
can only polymerize in the 5' to 3' direction
- synthesizes a short RNA-DNA hybrid (10 RNA nucleotides followed by20 - 30 DNA nucleotides)
-Details poorly understood
In bacteria, the enzyme "Primase"
is a type of RNA polymerase
uses nucleoside triphosphates to make a short RNA stretch complementary to the 3' end of the template
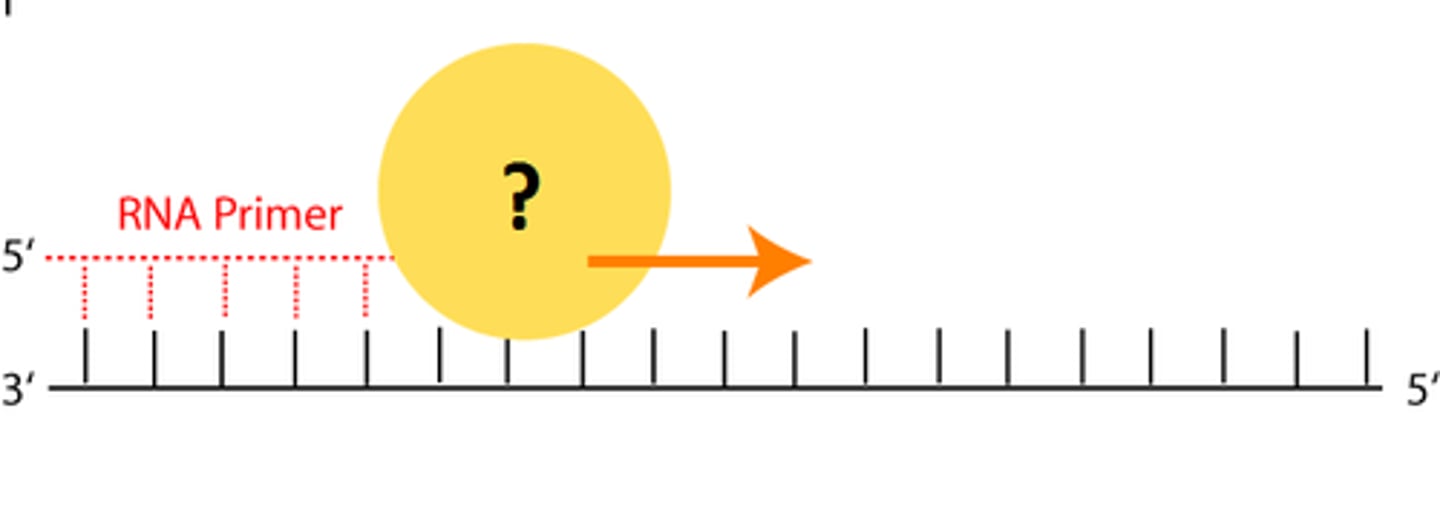
RNA polymerases
can start new chains from scratch
RNA chains are made 5' to 3'
delta and epsilon DNA polymerase
- cannot start DNA replication from scratch, need alpha DNA polymerase to set a primer with a free hydroxyl
- use it 3'-5' exonuclease activity to proofread any mistake it makes during replication
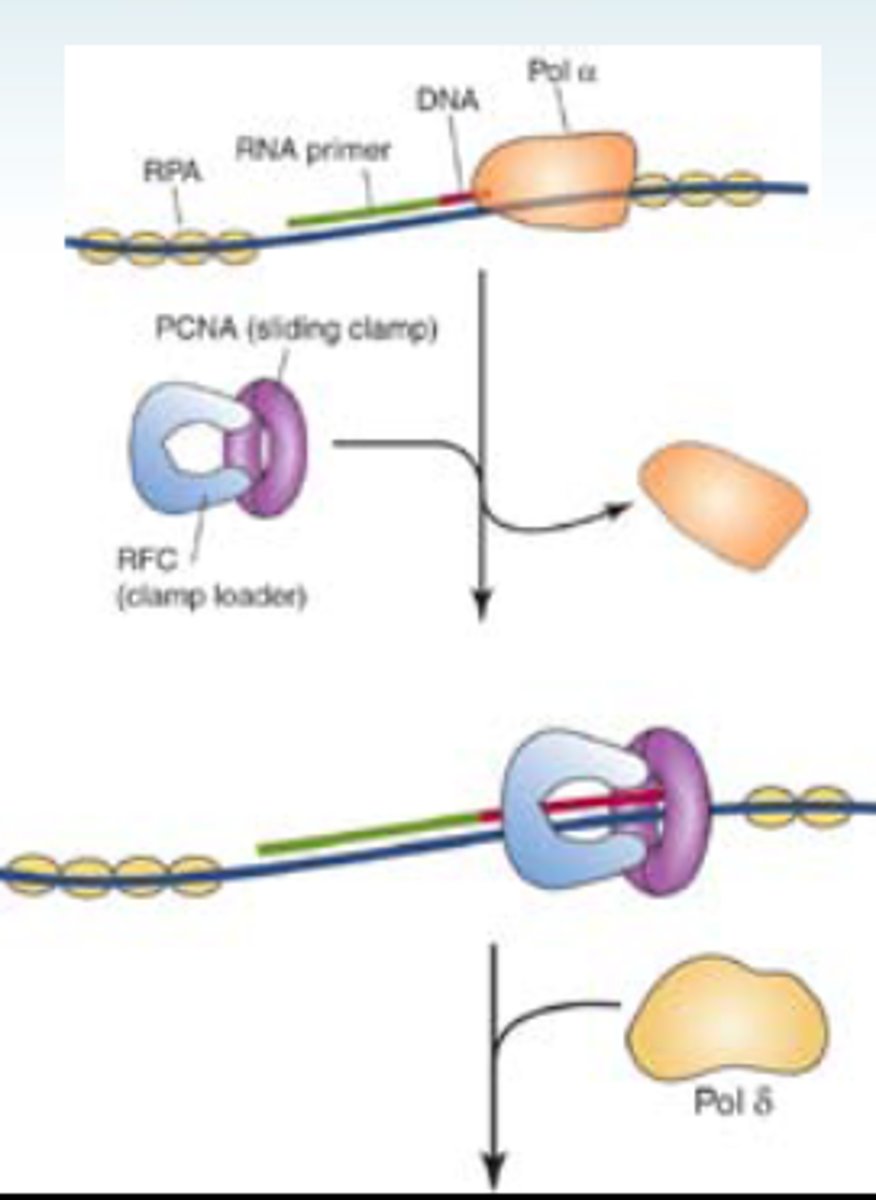
RNA/DNA primer
primase end of alpha DNA polymerase adds 10 RNA bases in primase pocket, then the DNA polymerase end builds off that and adds 20-30 DNA bases
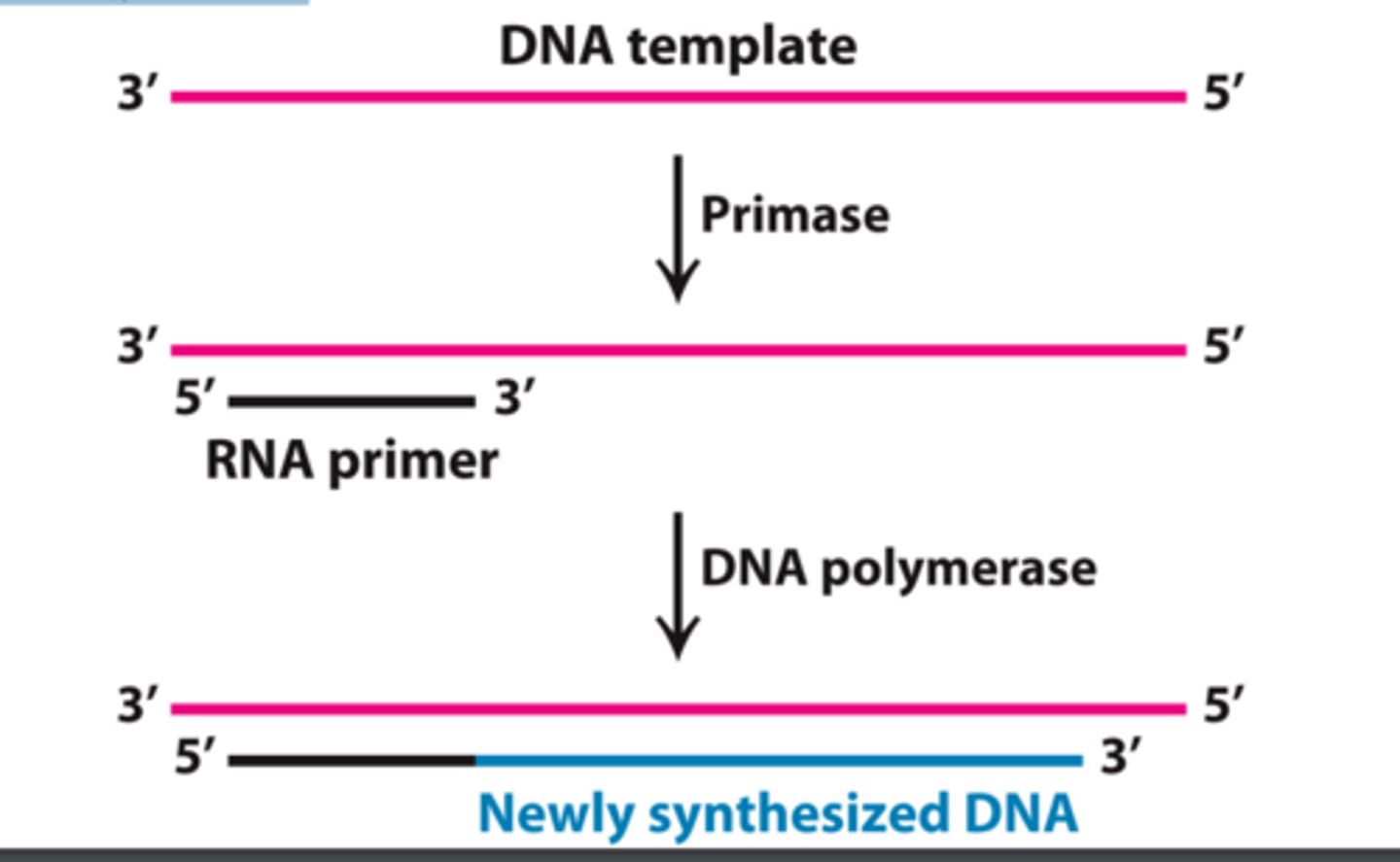
primase
An enzyme that joins RNA nucleotides to make the primer using the parental DNA strand as a template.
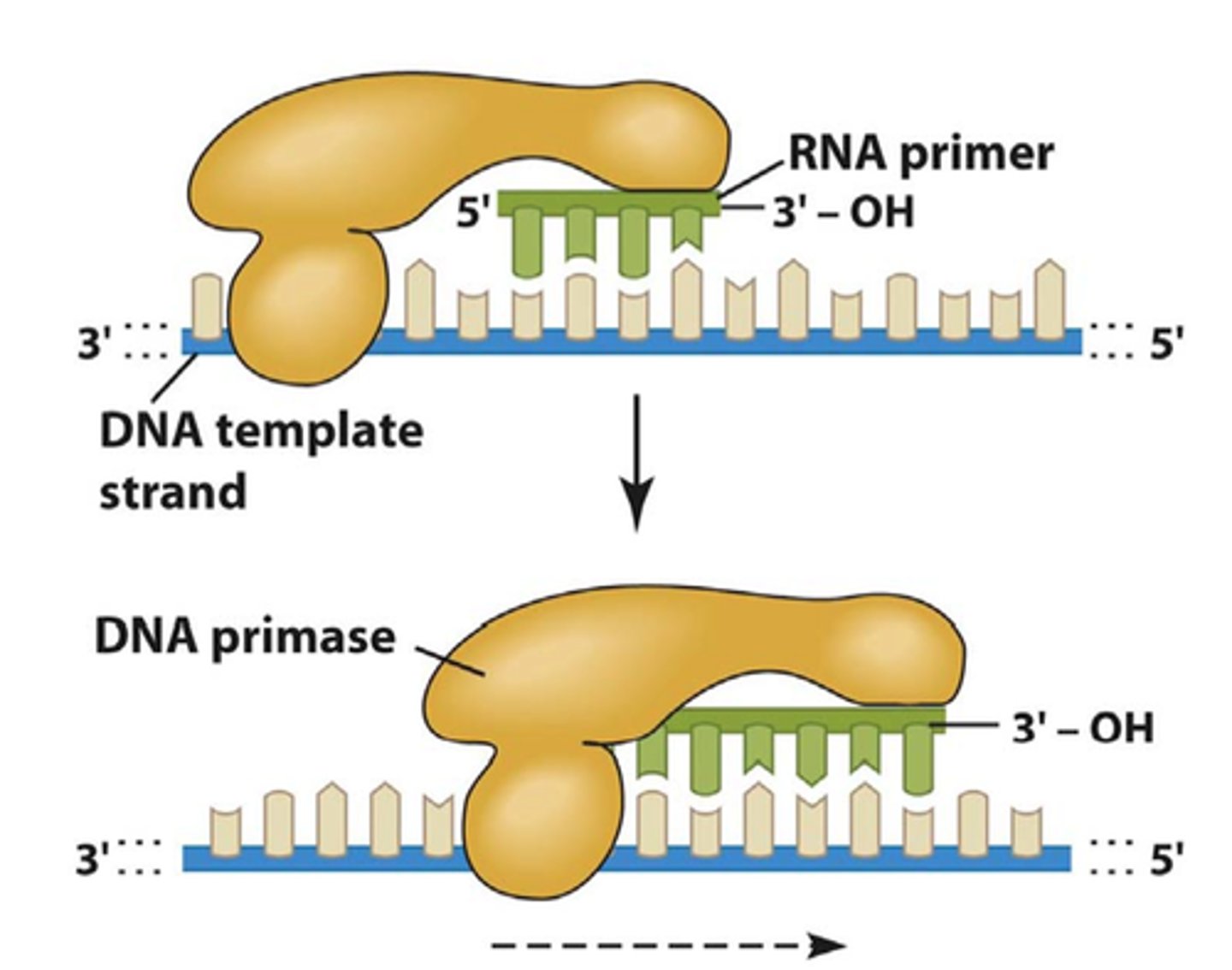
primase pocket
specialized, conserved binding sites within primase-polymerase enzymes that bind DNA templates and nucleotides, with distinct pockets used for initiation (I-site) and elongation (E-site)
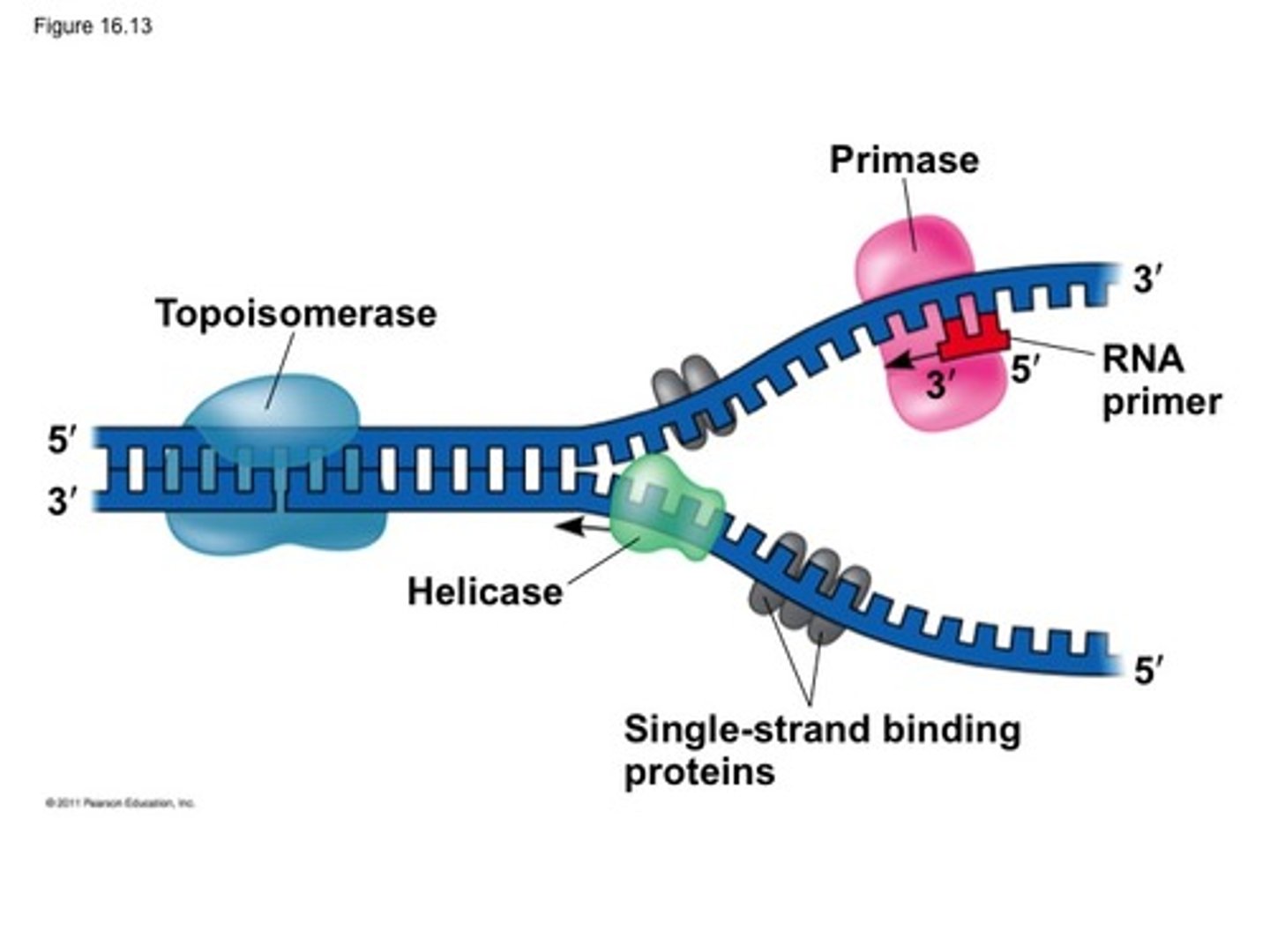
How many nucleotides are added in DNA replication
α DNA pol adds 50-100 pairs total, and δ/ε extend it further by 100-10000 nucleotides

Nucleotides added to DNA chain
must be in the form of nucleoside triphosphates (dNTP)
- deoxyAdenosinetriphosphate (dATP)✓
- deoxyGuanosinetriphosphate (dGTP)✓
- deoxyCytidinetriphosphate (dCTP)✓
- deoxyThymidinetriphosphate (dTTP)✓
- Uridine triphosphate (dUTP)
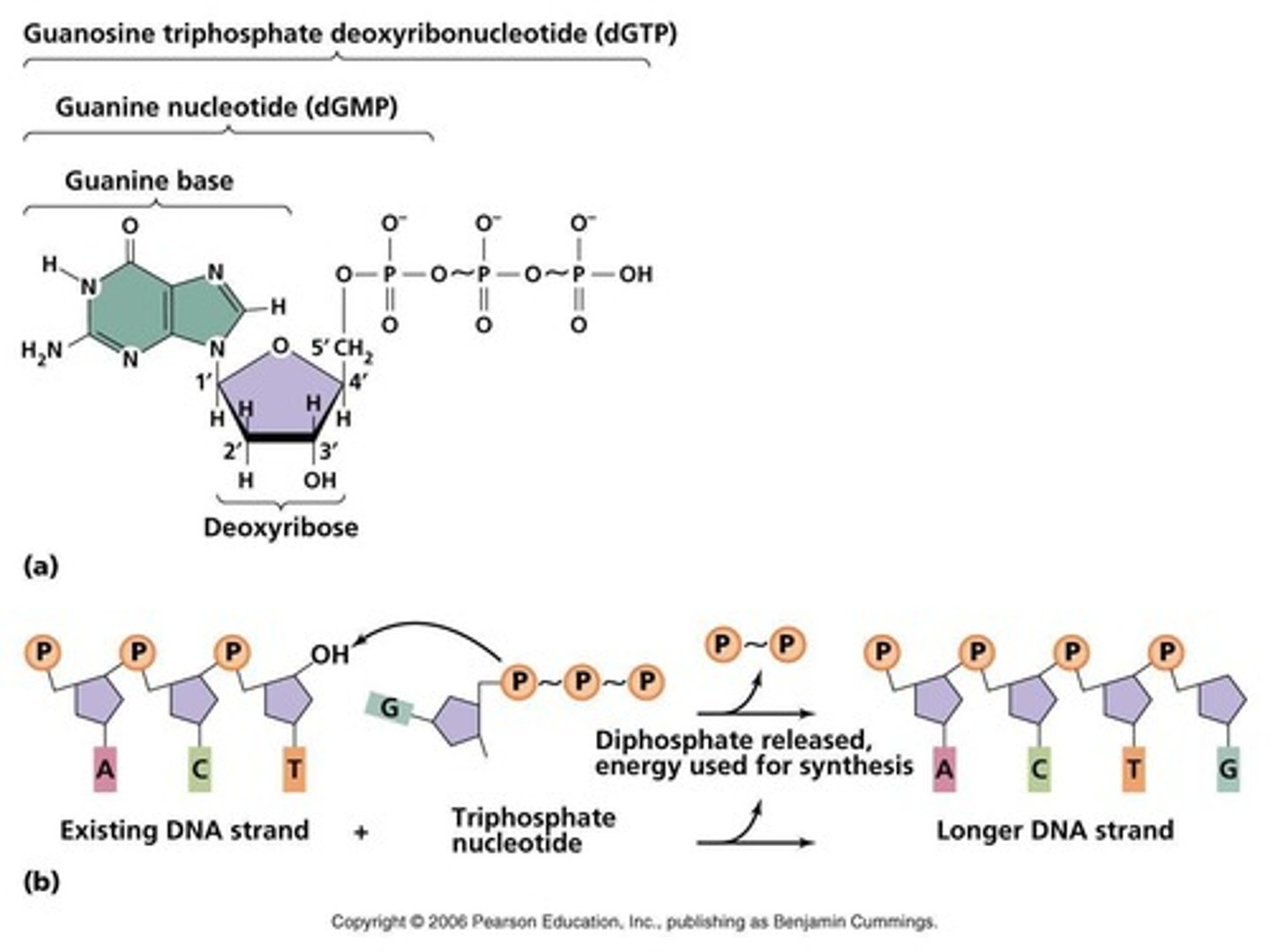
energy to create the phosphodiester bonds
Hydrolysis of the nucleoside triphosphate bonds to release two phosphate groups + Energy
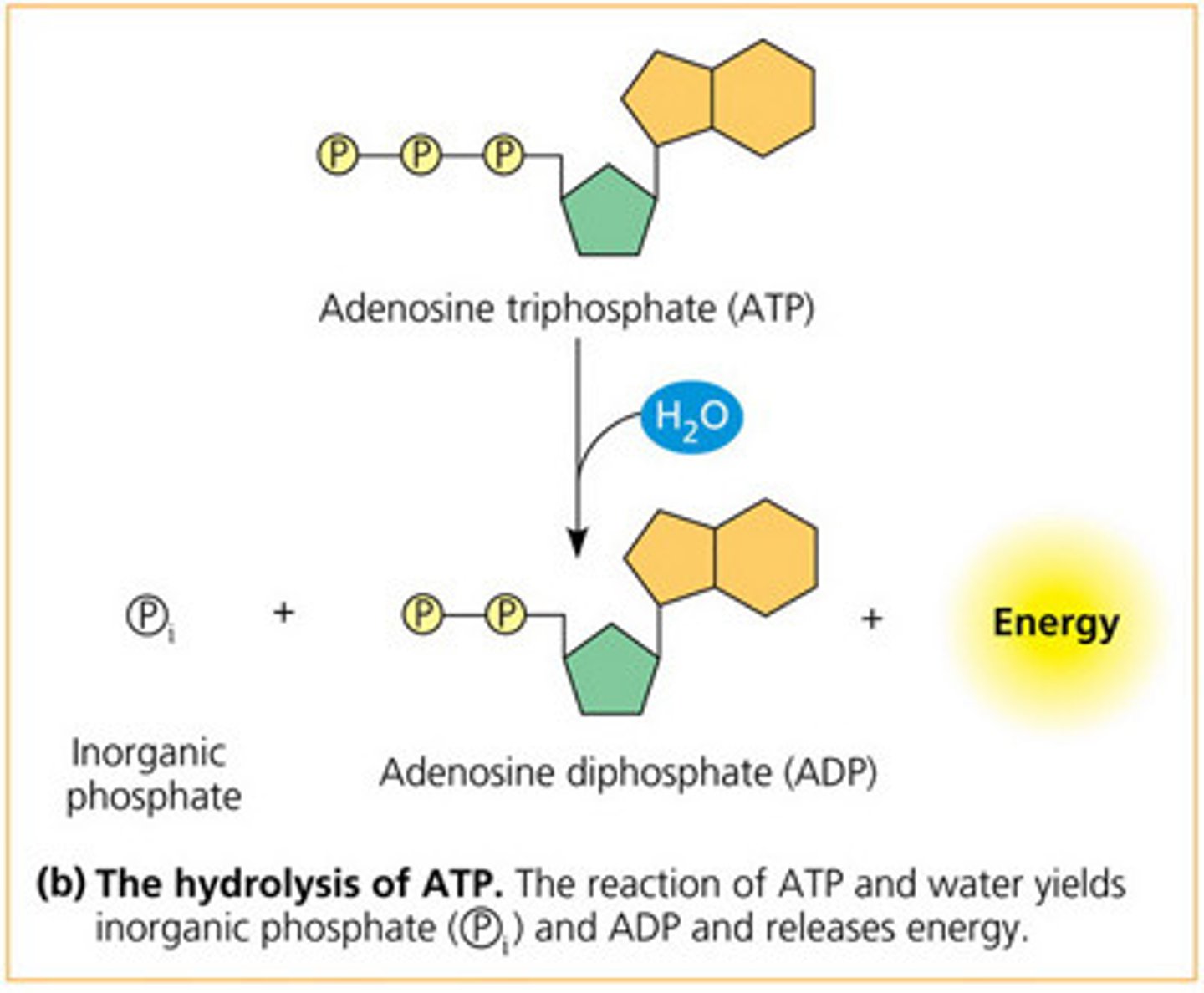
leading/continuous strand
Can only add nucleotides in the 5' to 3' direction
The bases must be complementary
Polymerization towards the replication fork
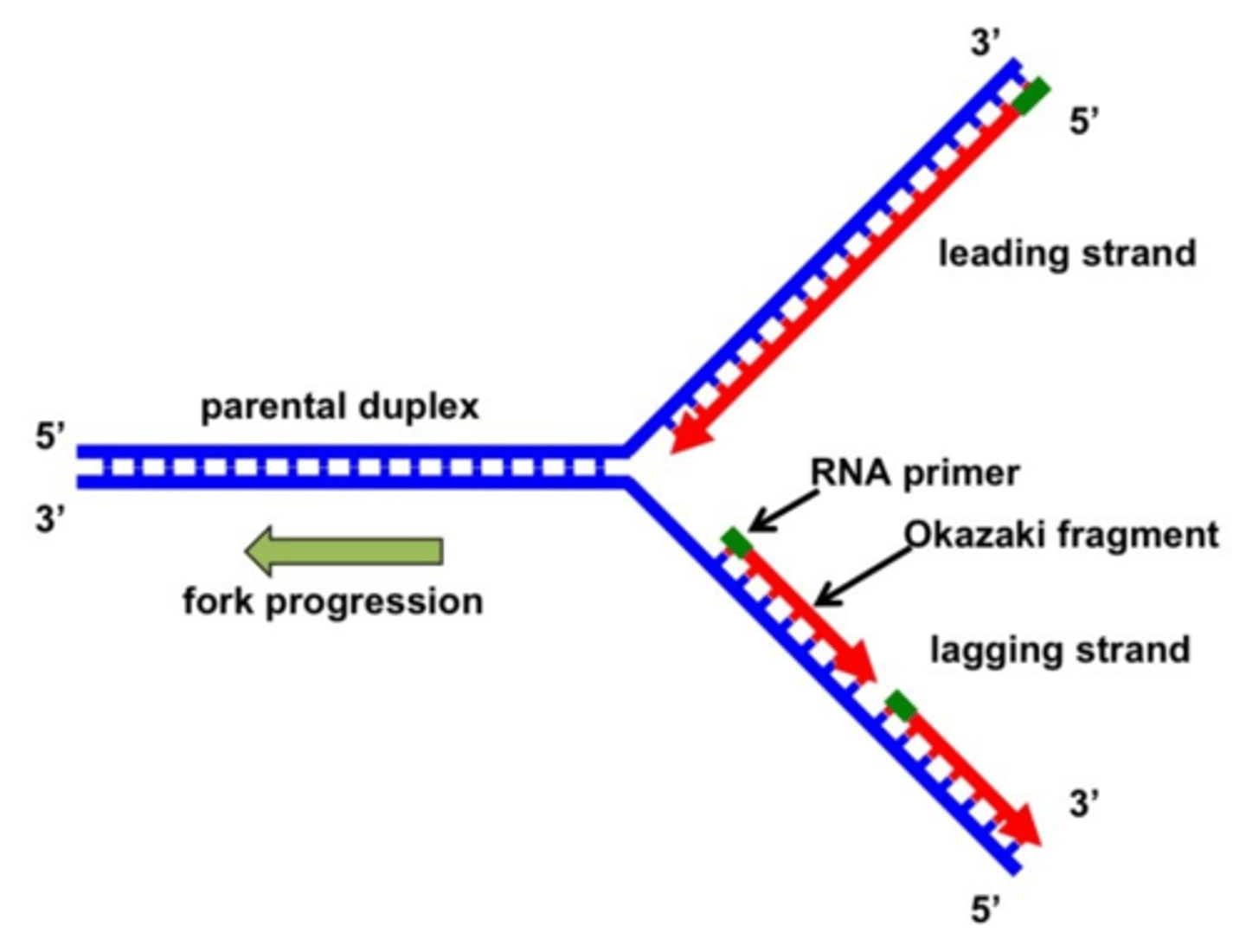
lagging/discontinuous strand
Can only add nucleotides in the 5' to 3' direction
The bases must be complementary
Polymerization AWAY from the replication fork
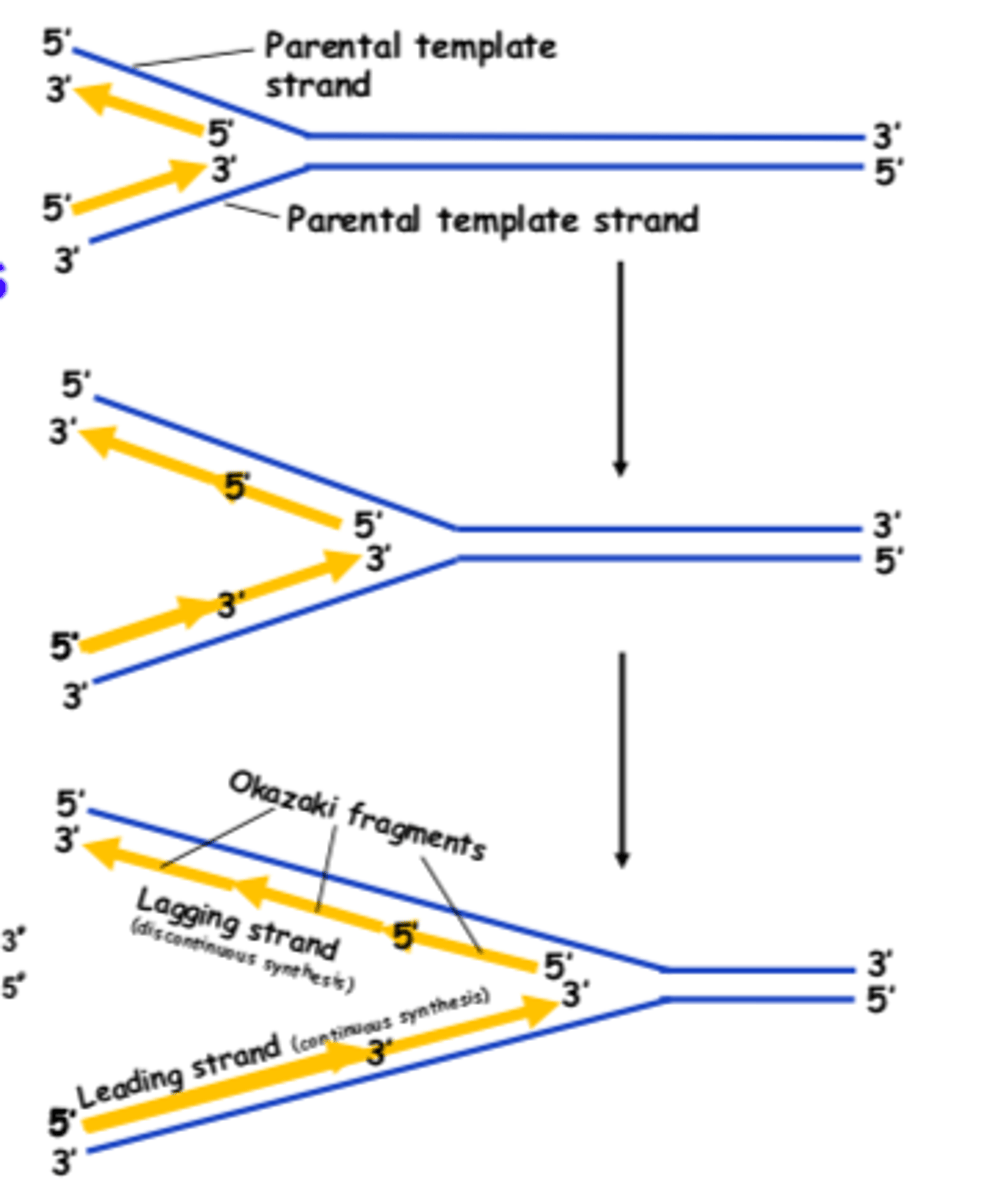
How many primers are needed for the leading strand
one, primer is placed and then DNA replication continues as DNA strands unravel
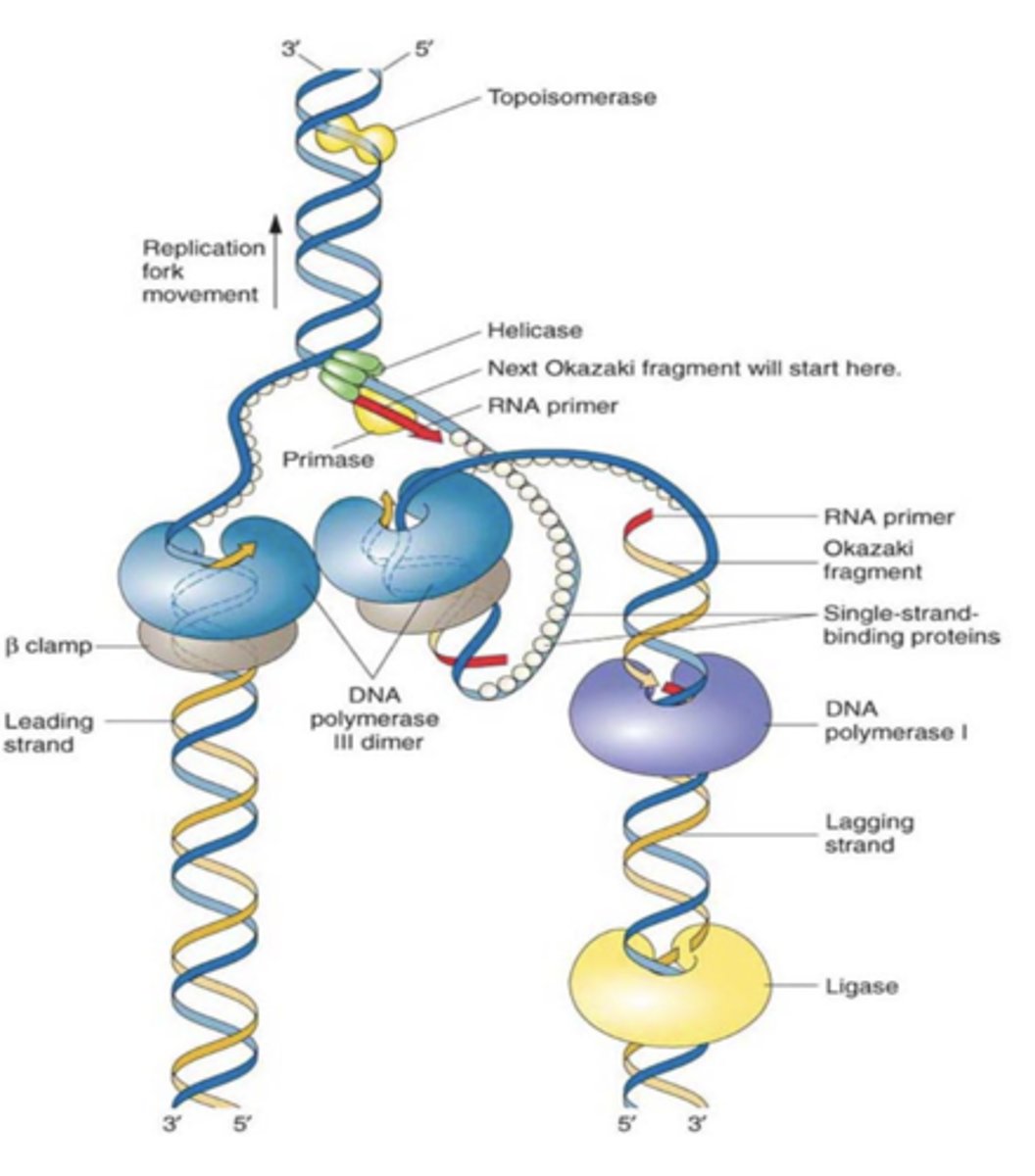
Topoisomerase
Enzyme that functions in DNA replication, helping to relieve strain in the double helix ahead of the replication fork; relieve DNA supercoiling
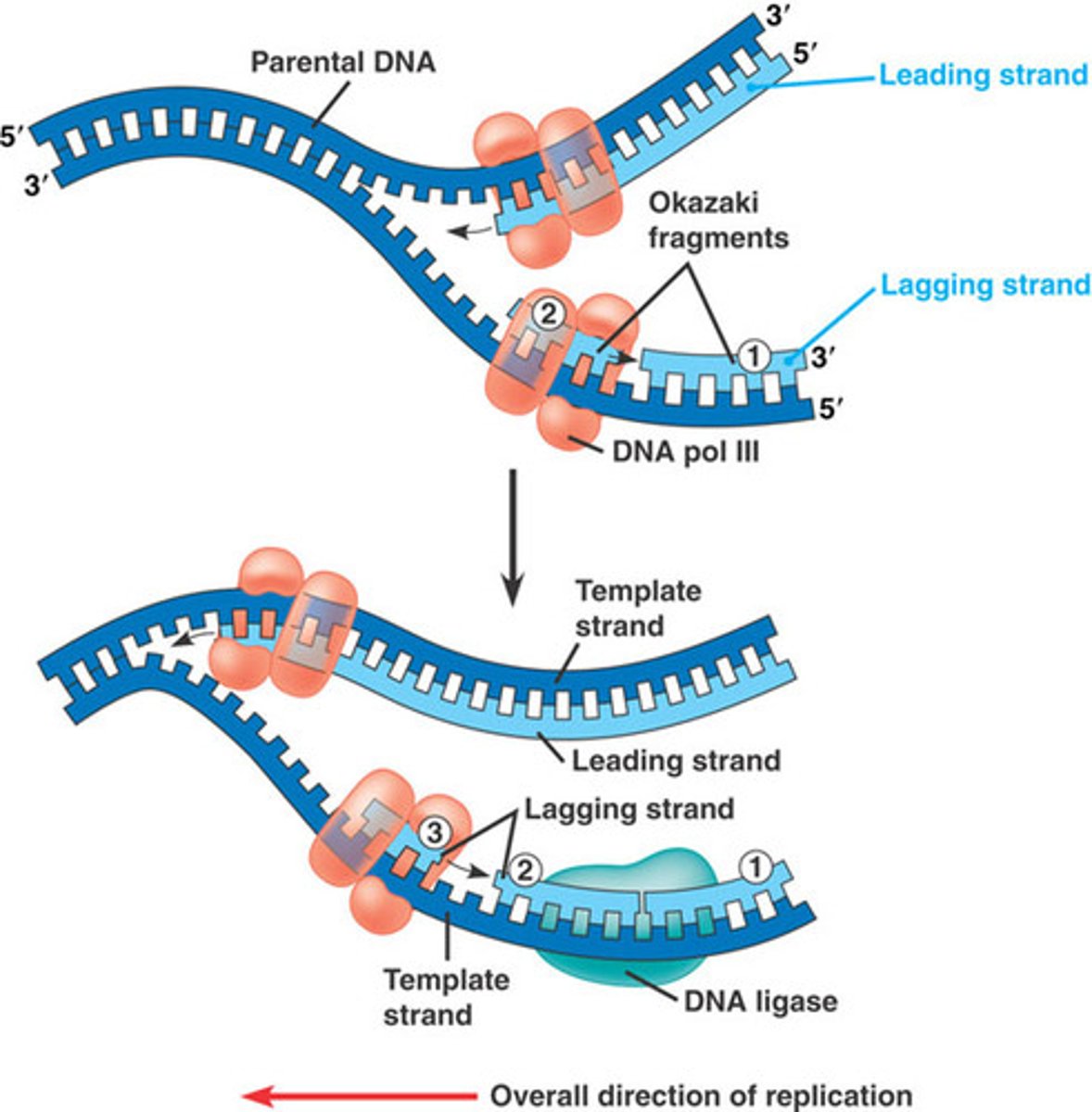
okazaki fragments
how lagging strand is synthesized
Small fragments of DNA produced on the lagging strand during DNA replication; primer + 100-200 DNA nucleotides
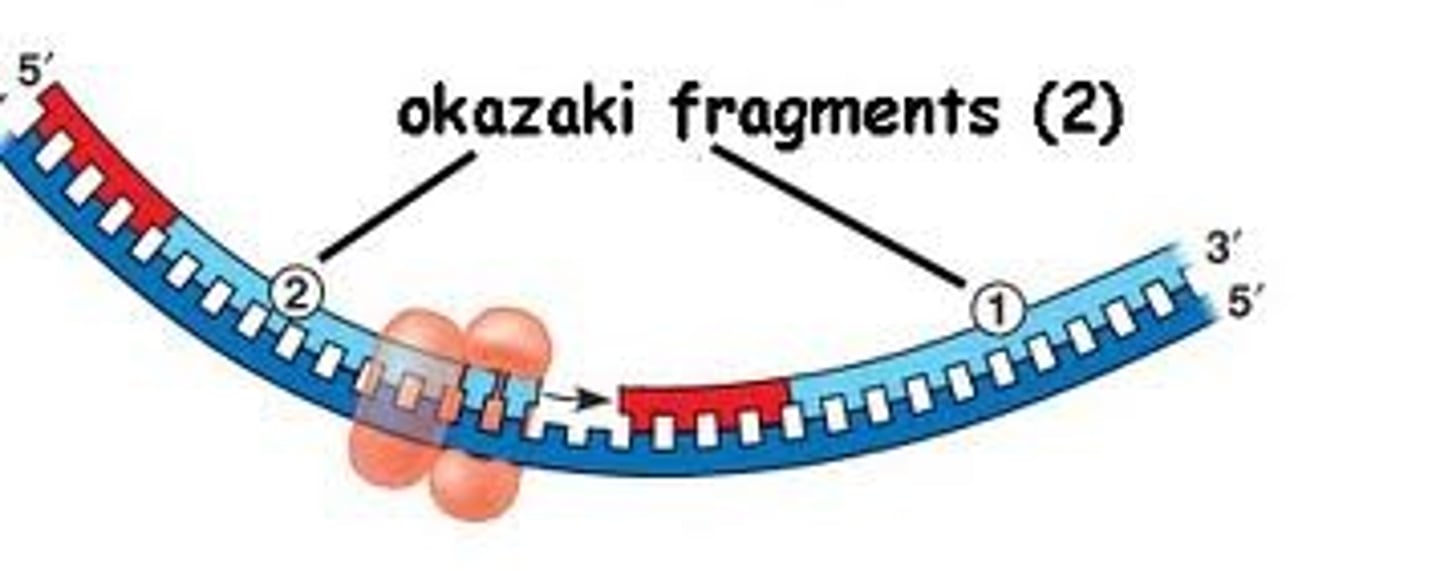
Why does the lagging strand synthesize incrementally
for efficiency; because the lagging strand is built away from the replication fork, the primer is not able to lay where the DNA is still wound
must work in fragments as DNA unwinds
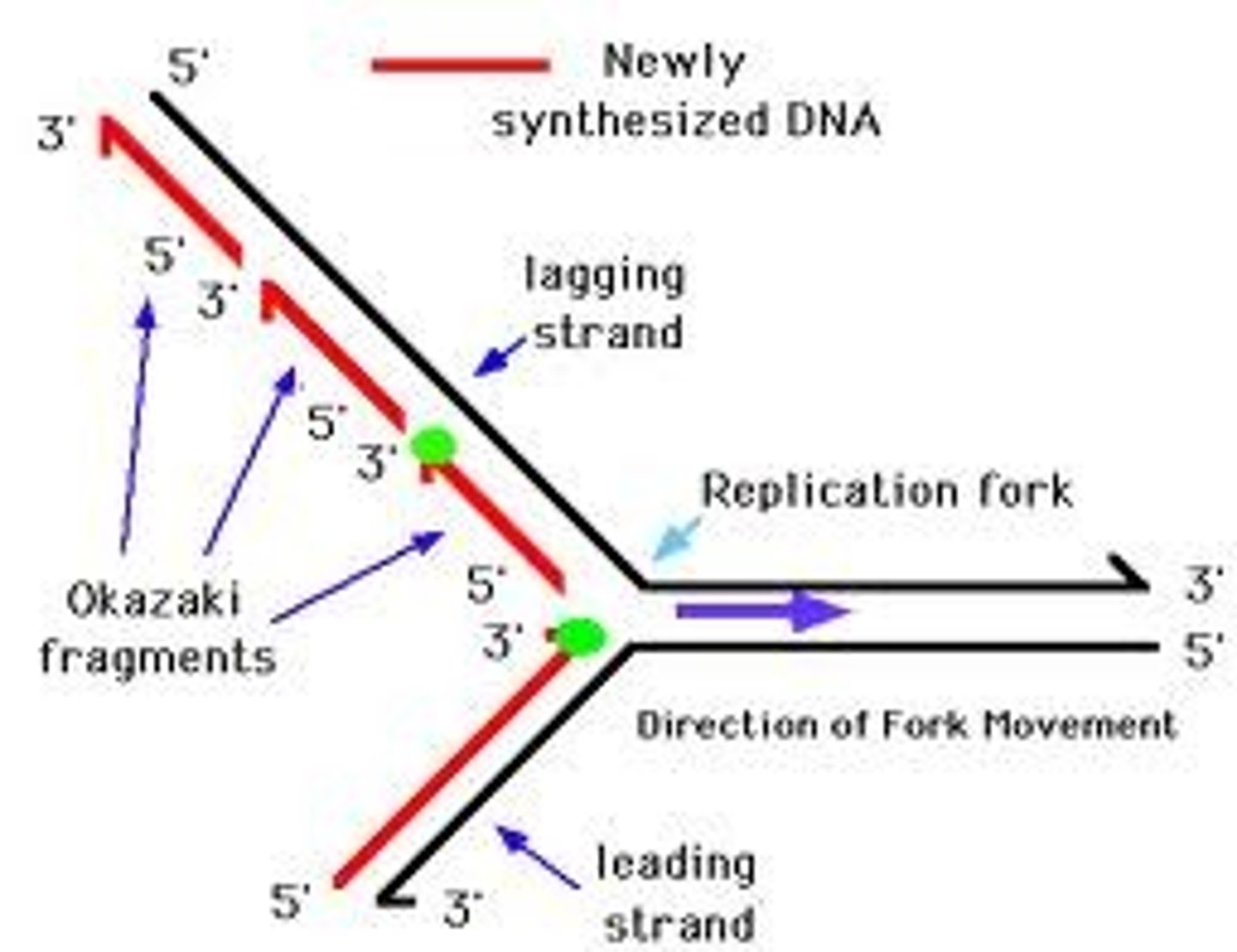
proteins/enzymes used in DNA replication
work until they cant anymore/are old, from where they are sent to the degradation pathway and harvested for amino acids
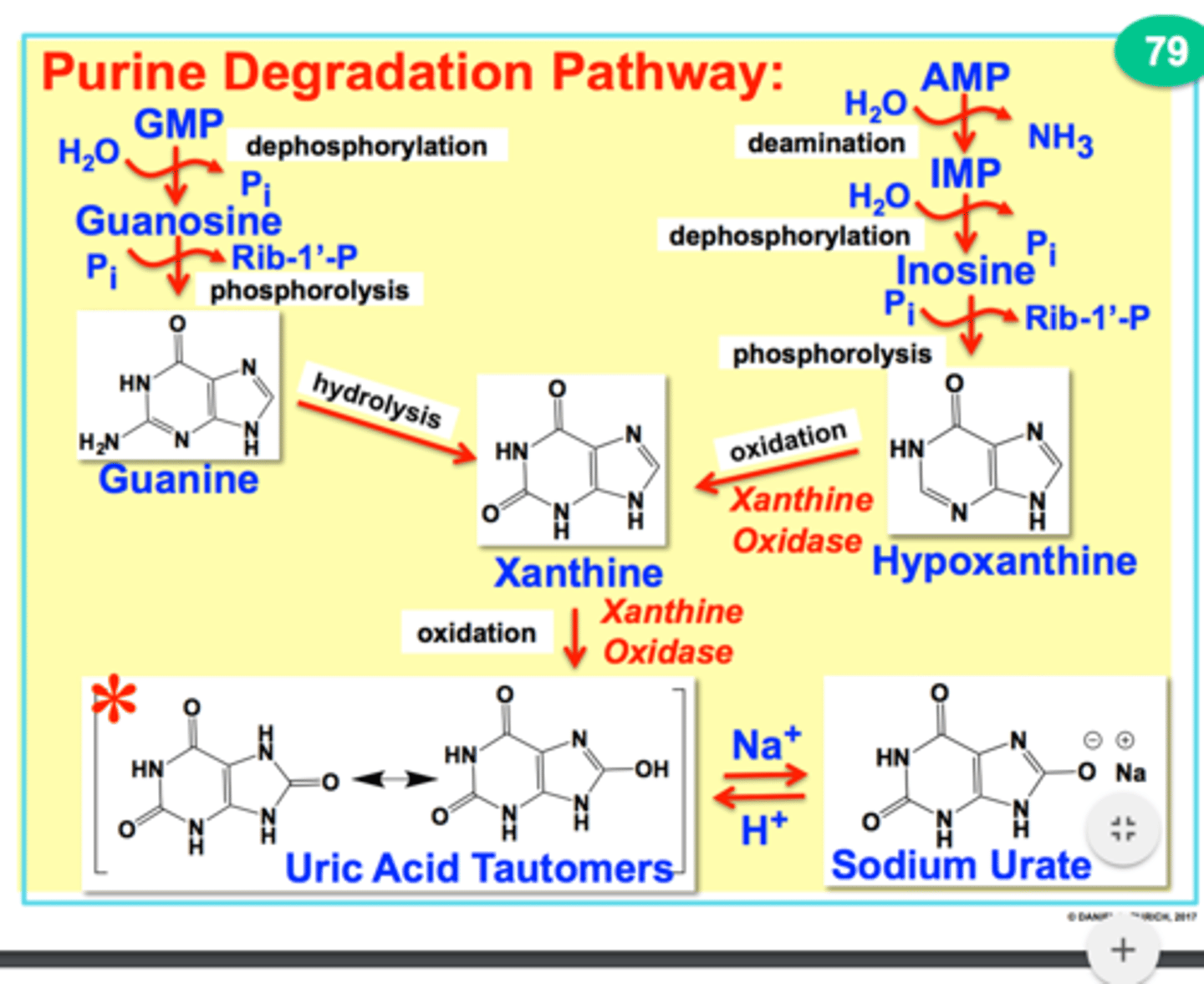
ribonuclease enzyme
RNase H removes the RNA part of the primers
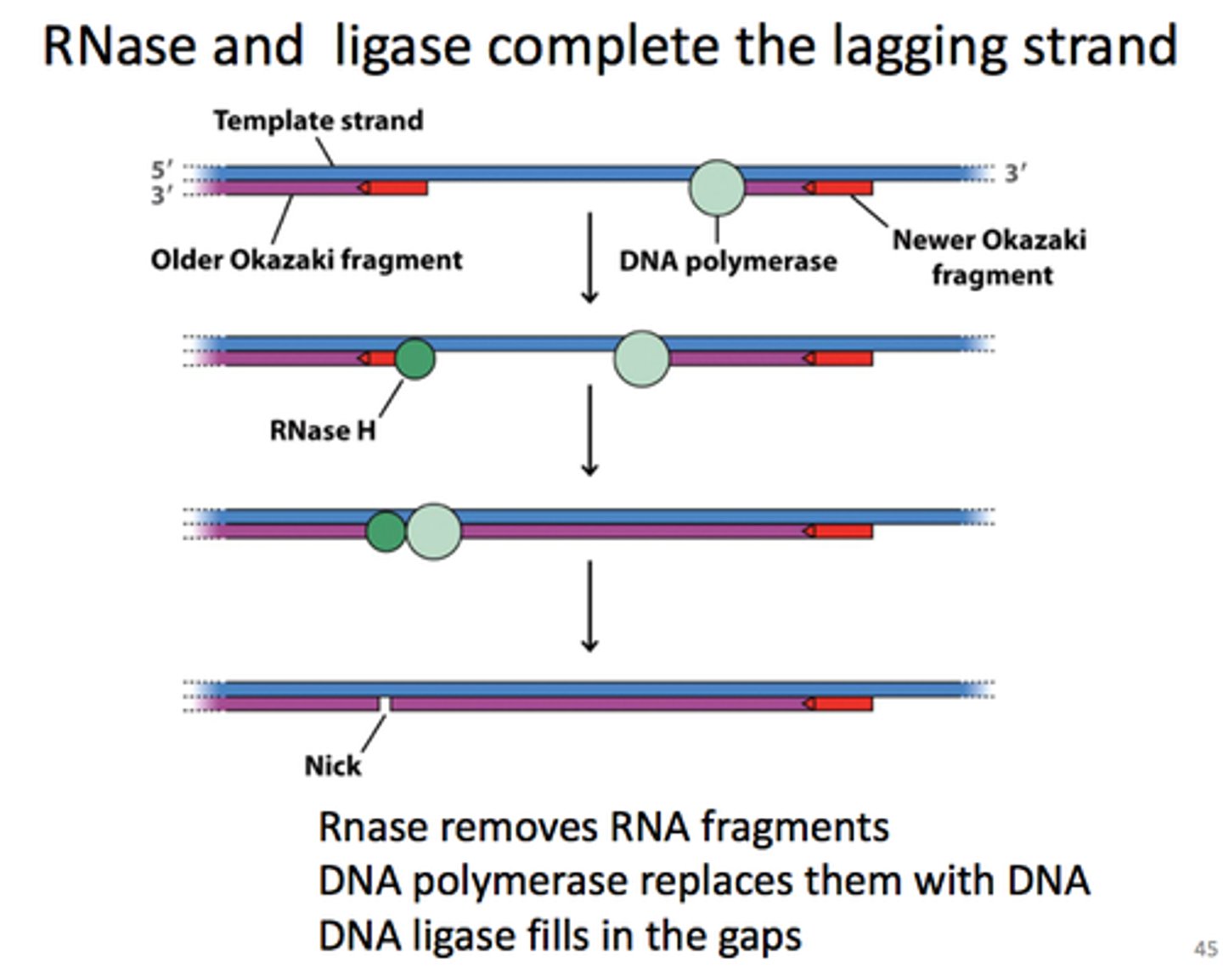
DNA polymerase delta's special job
adds DNA nucleotides to replace the RNA nucleotides that was removed (for both leading and lagging strand)
- cannot bind these new nucleotides with Okazaki fragments because they have monophosphate ends
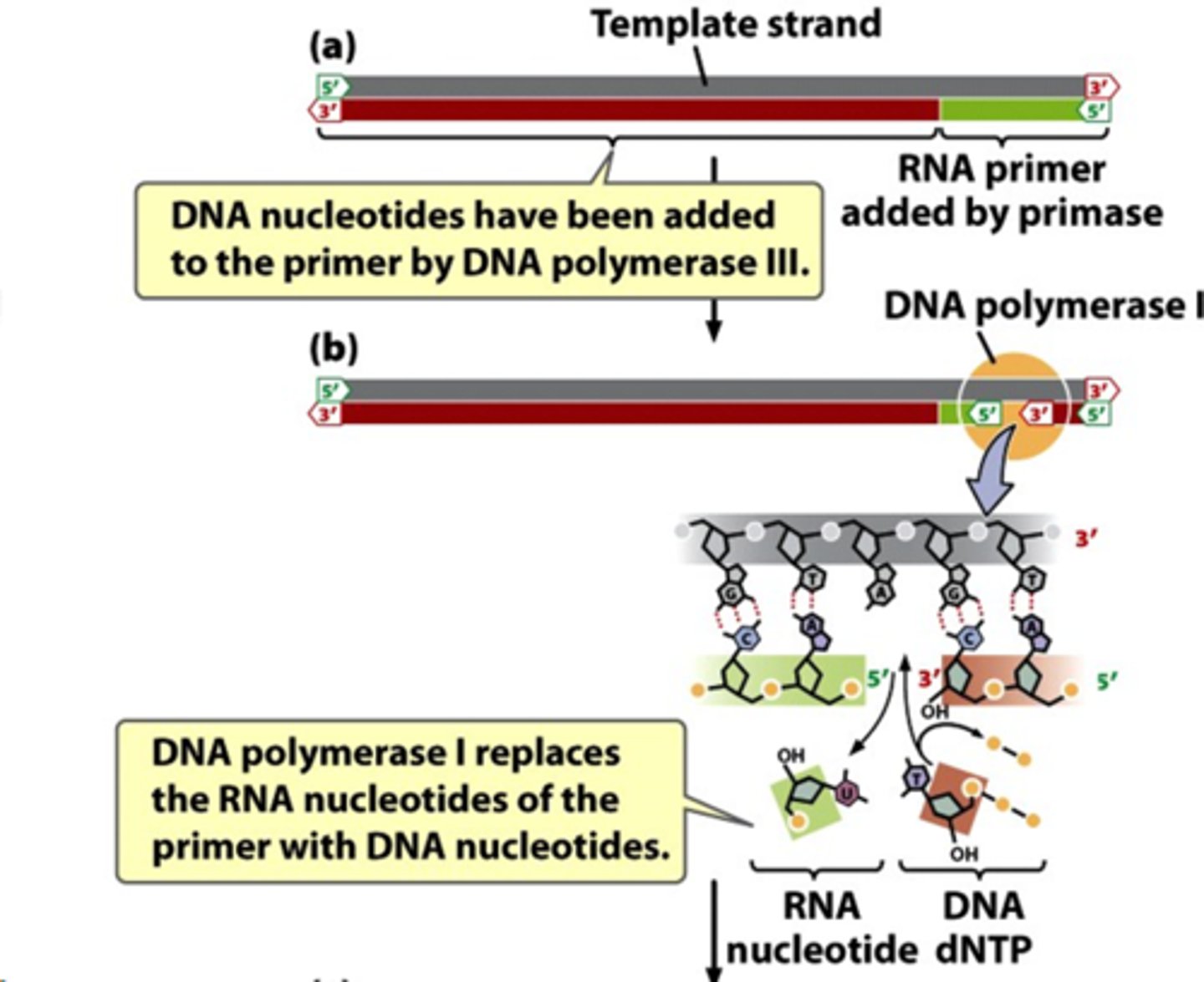
DNA ligase
Catalyzes the formation of phosphodiester bonds; joins okazaki fragments
makes diphosphate on okazaki fragment end (using ATP or NAD)
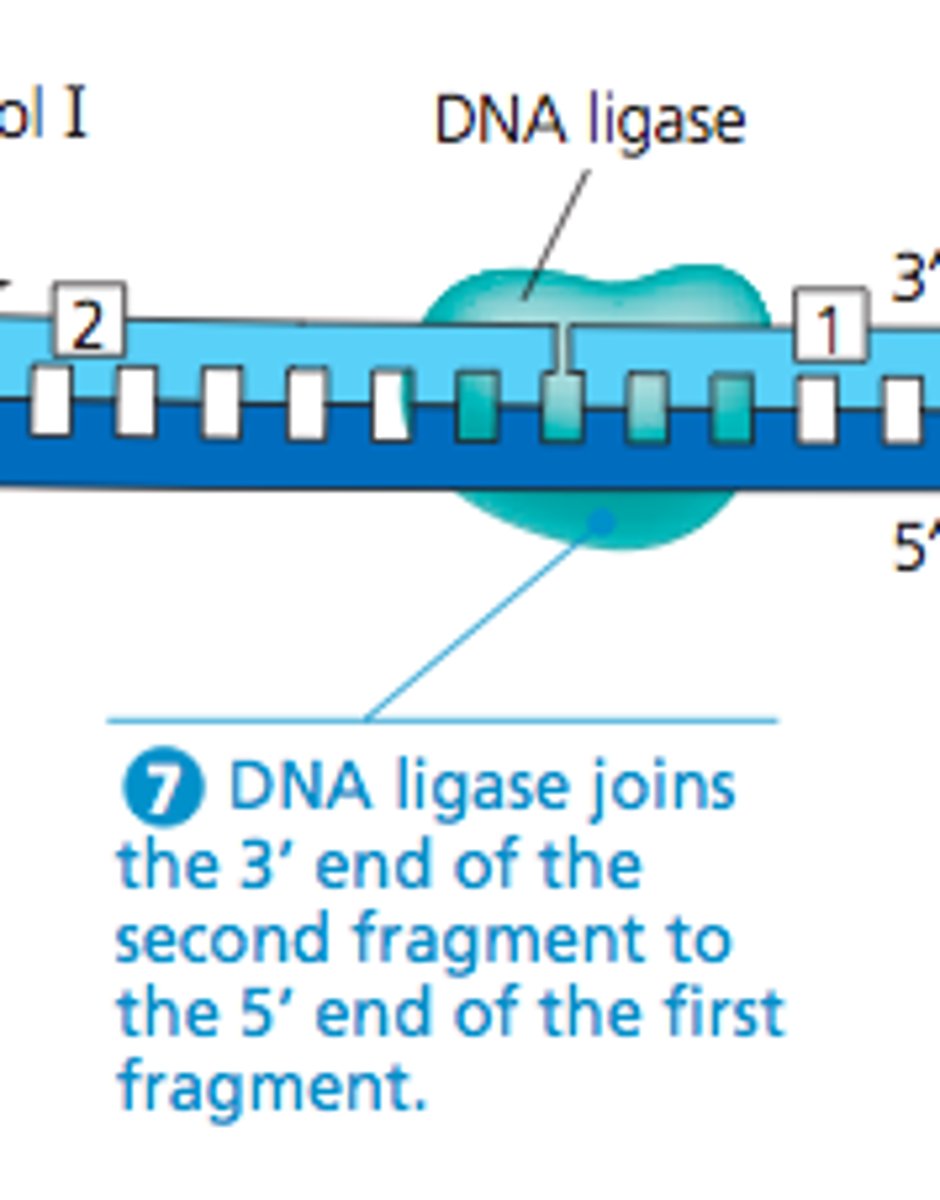
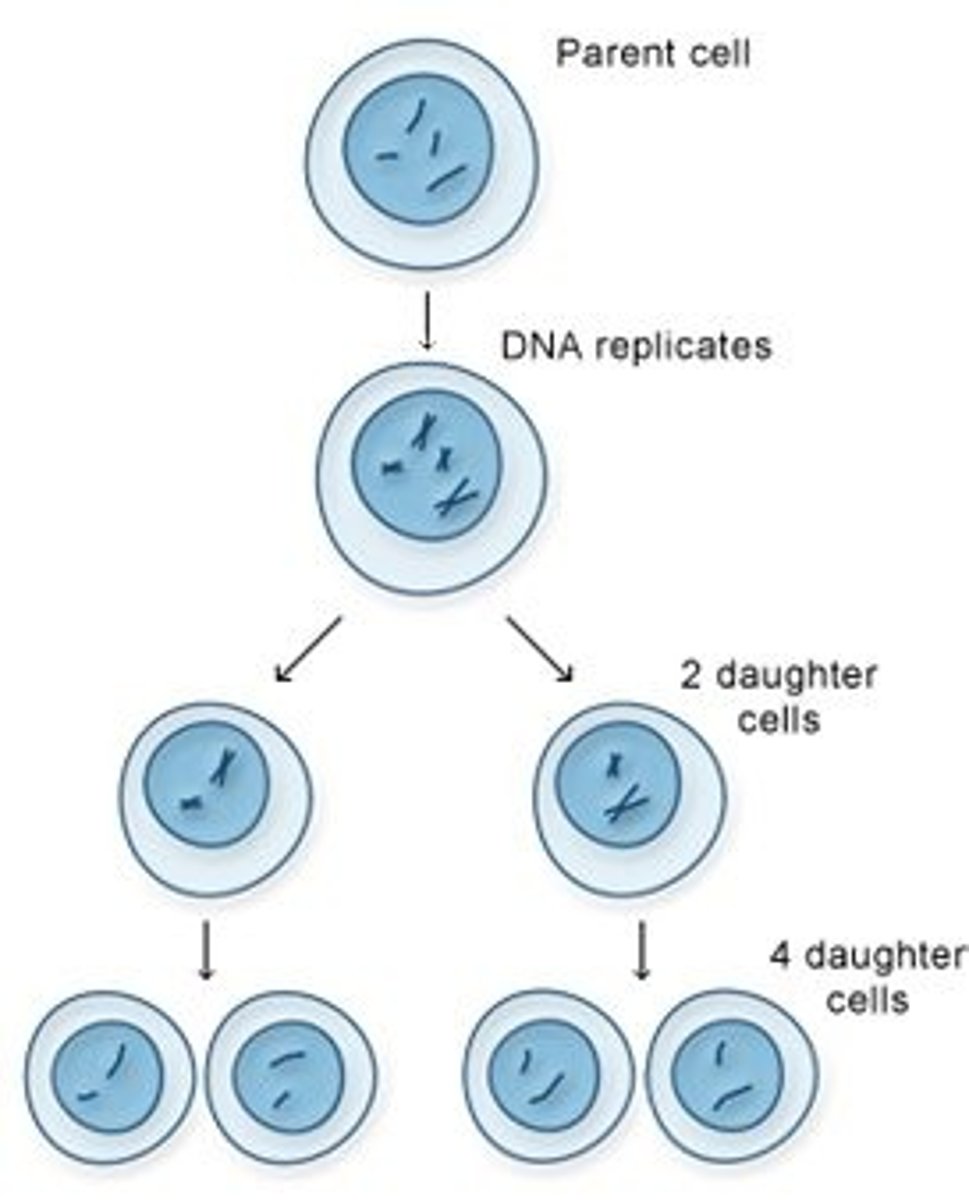
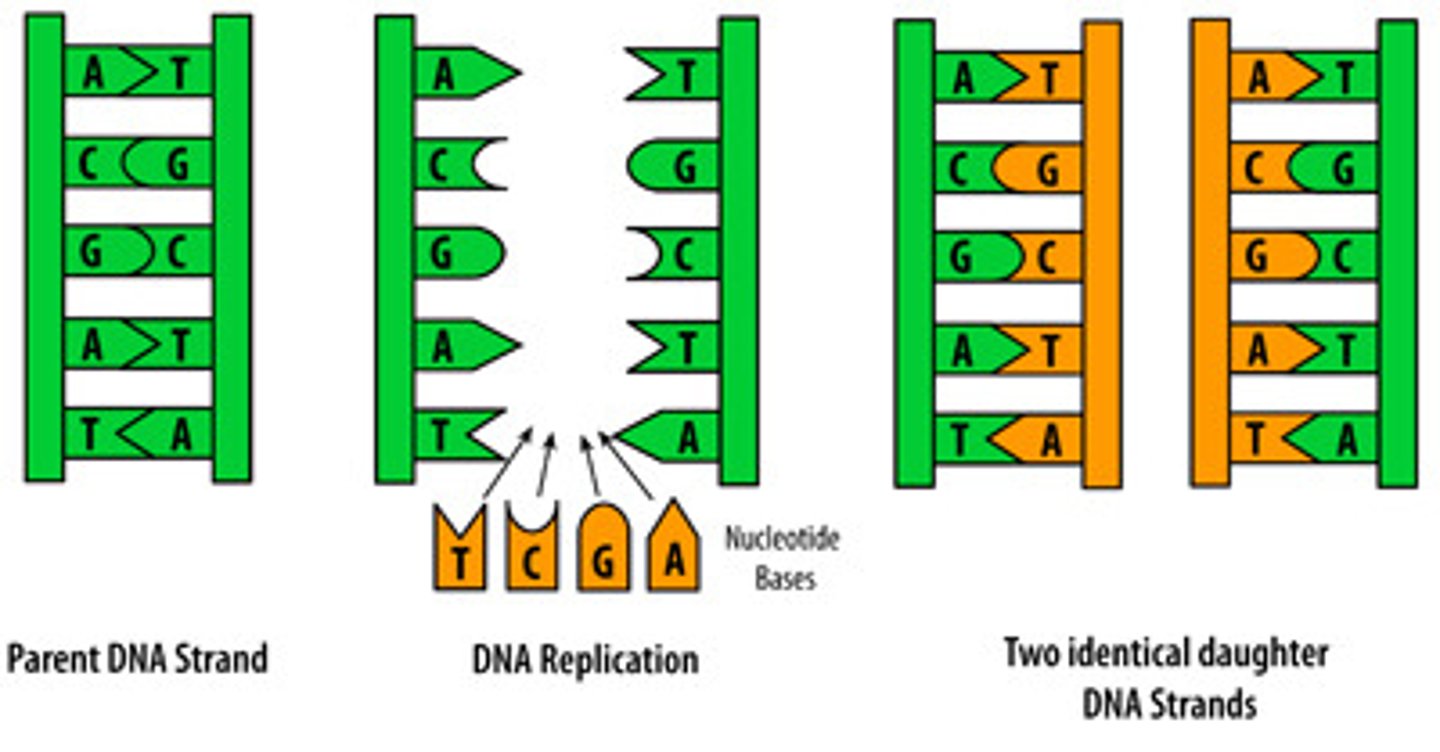
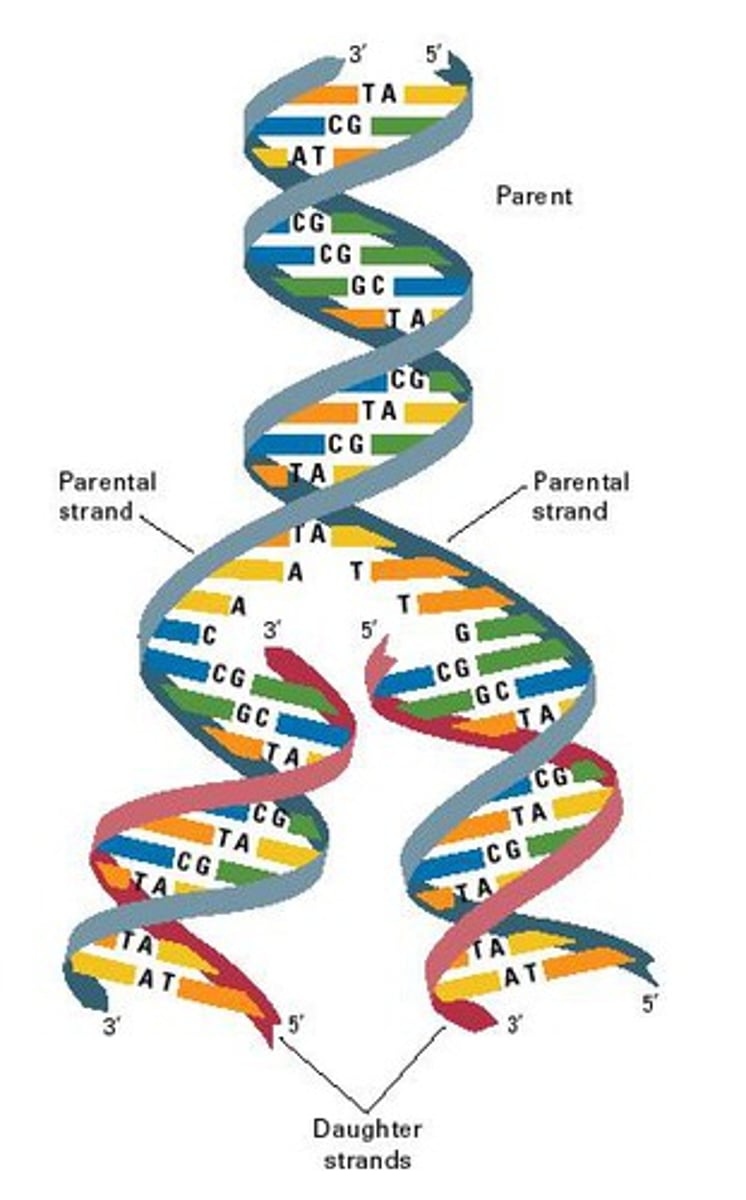
result of DNA replication
two identical strands that each contain one old strand and one new strand
Each of these strands will end up in two daughter cells by the end of mitosis
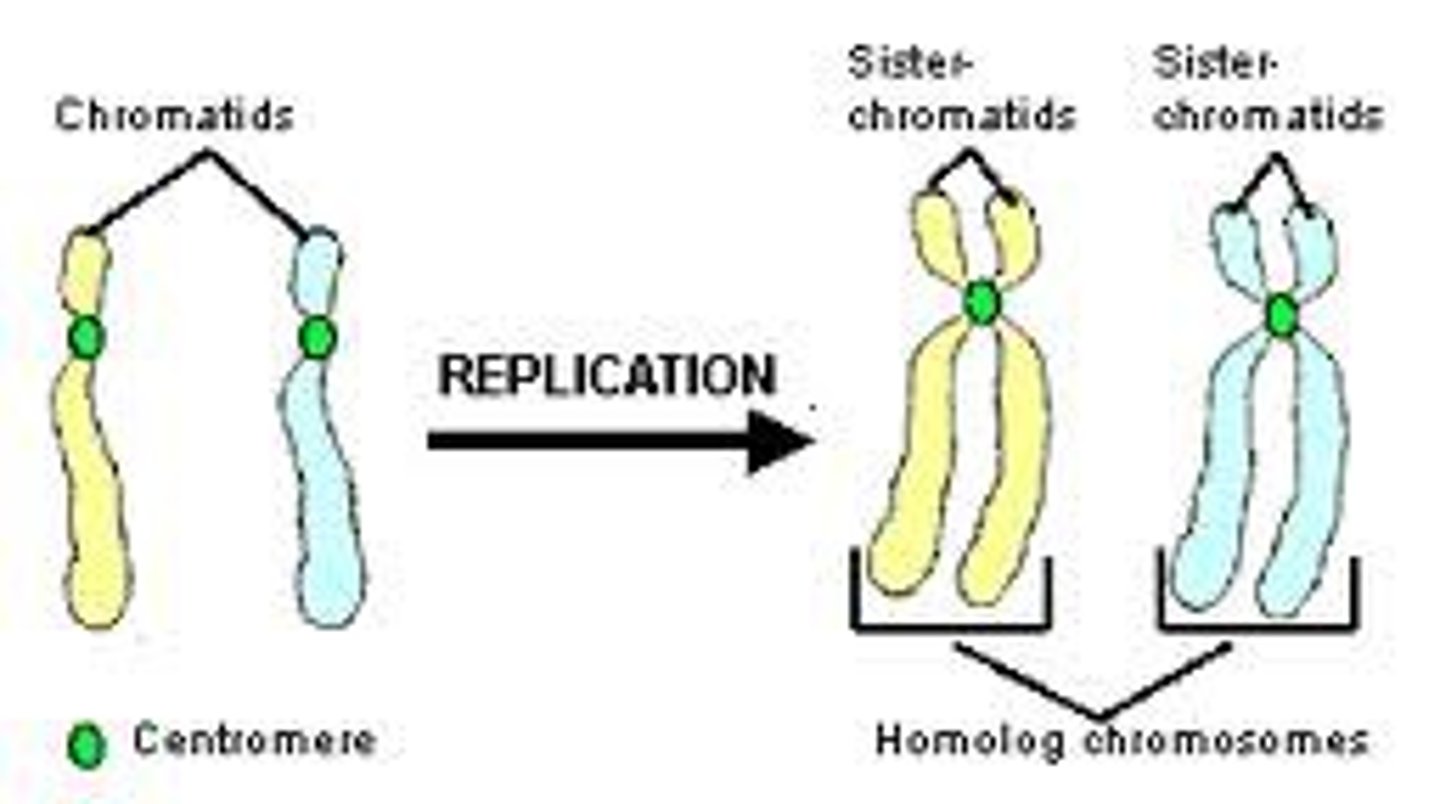
homologous chromosomes
Chromosomes that have the same sequence of genes and the same structure; slightly different DNA sequences
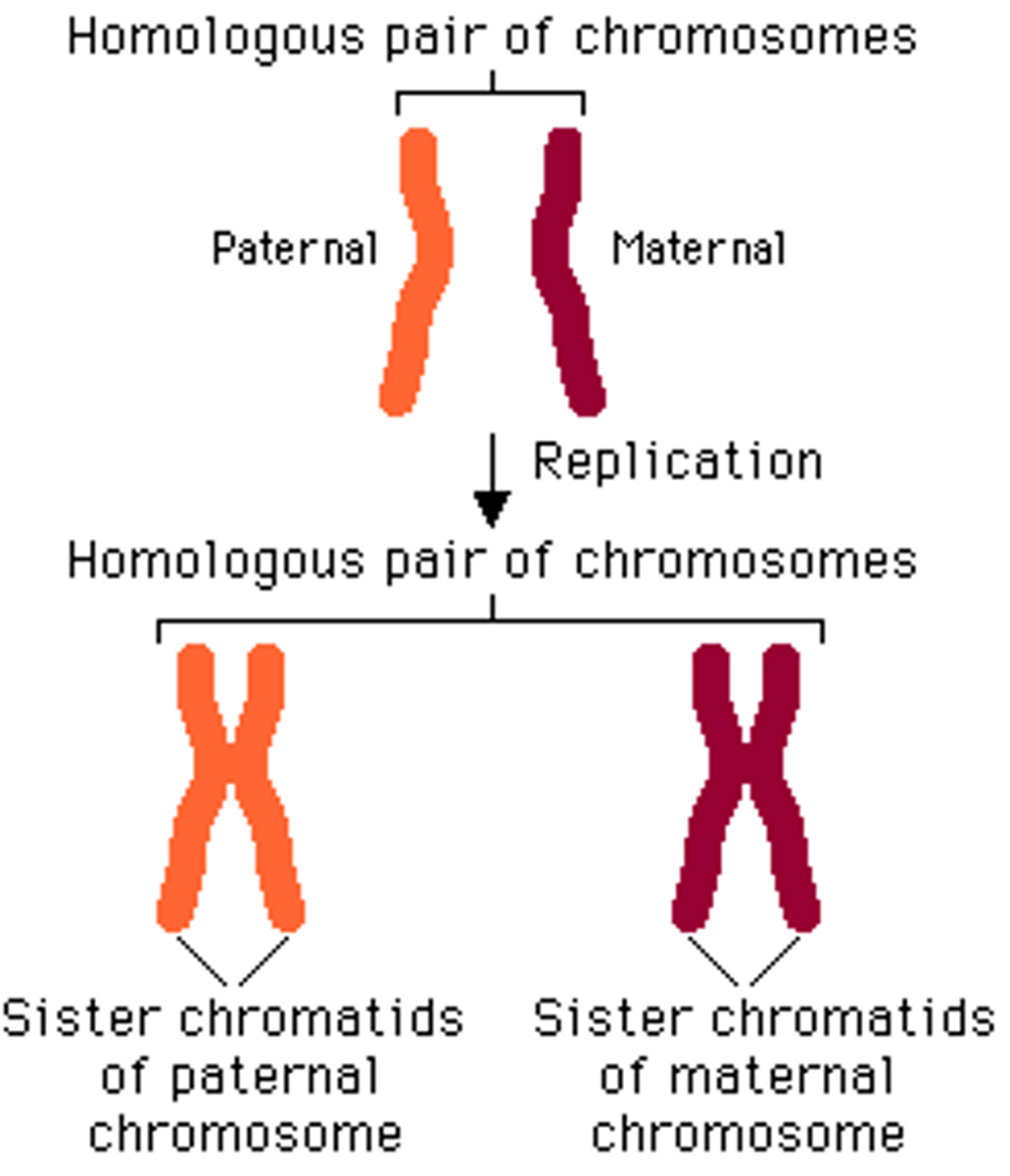
Sister chromatids
Identical copies of a chromosome; full sets of these are created during the S subphase of interphase.
DNA replication must be error free.
Errors can cause mutation in genes which can lead to diseases
must be quick and accurate
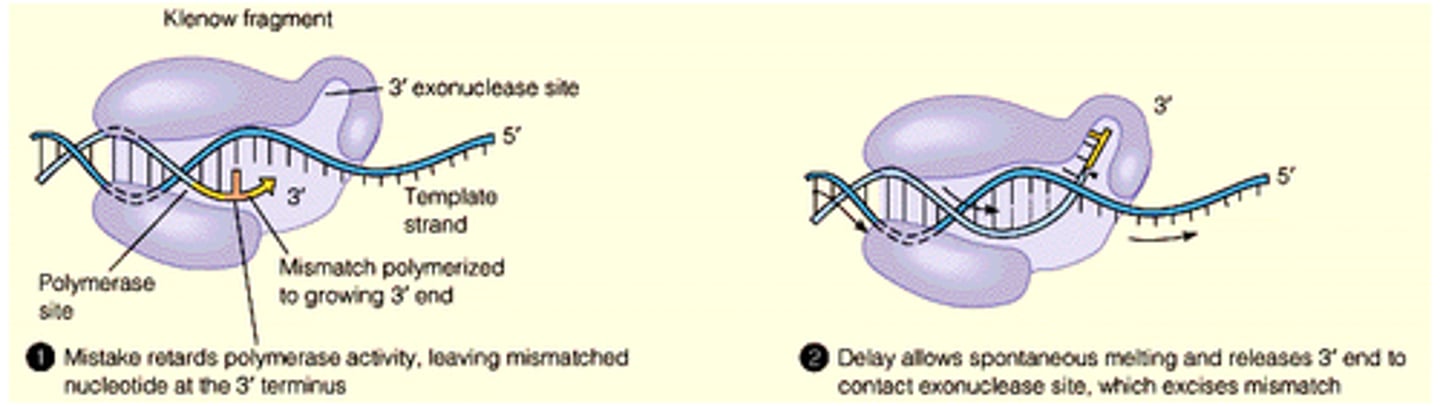
DNA polymerase has proofreading abilities
DNA polymerase has 3' to 5' exonuclease activity
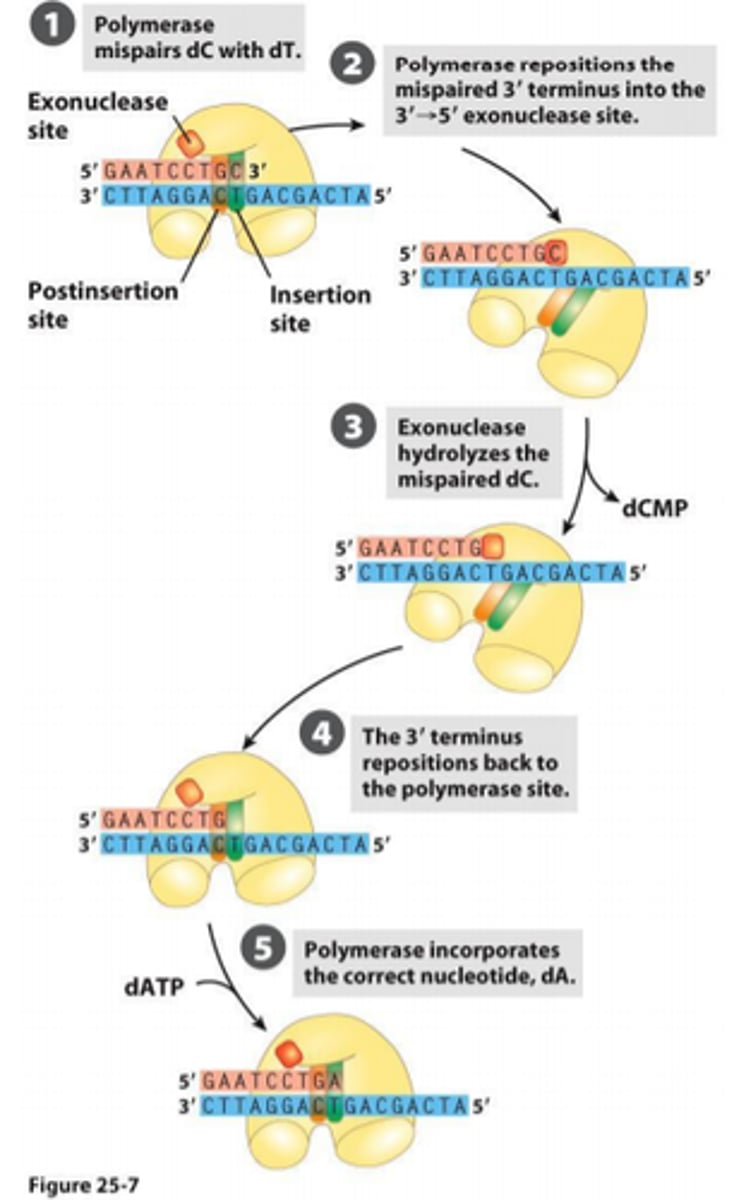
During DNA replication, is it impossible for the wrong base to be inserted?
Not impossible - biochemical process, disfavored Keq(energetically less good)
there is an equilibrium constant for inserting the wrong base
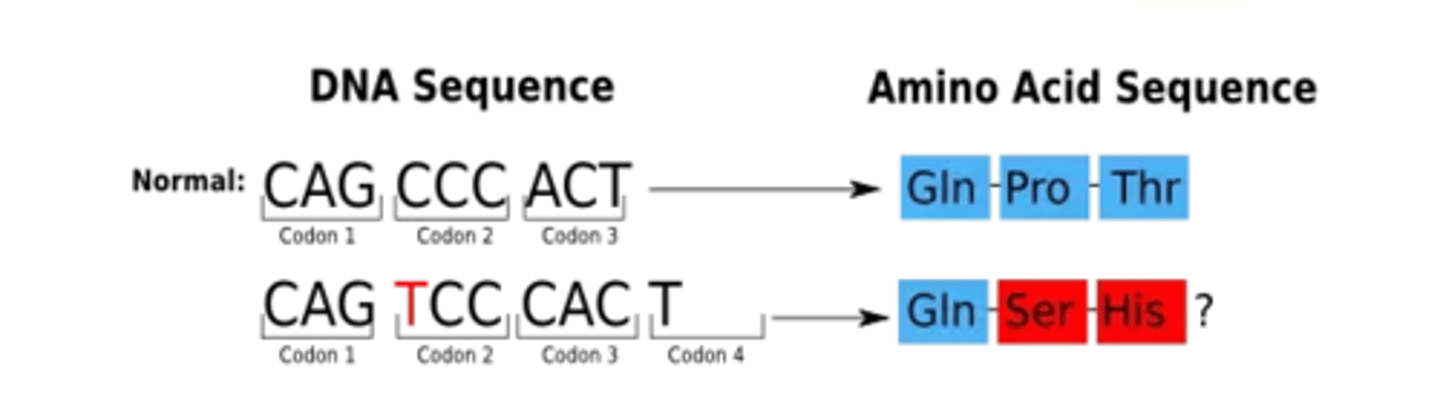
Keq
original accuracy + proofreading = 10^-6
mismatch repair
original accuracy + proofreading + mismatch = 10^-9
What happens if an error is detected?
bases around error are cut out and then beta DNA polymerase fills in correct sequence