PSU BMB 211 EXAM 2
1/83
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
84 Terms
An electrostatic interaction might occur within a protein between what types of amino acid pairs at typical physiological pH?
+ and - charged
A hydrophobic interaction might occur within a protein between what types of amino acid pairs?
non polar and non polar
Secondary and higher orders of structure are not determined by
peptide bonds
If an aspartic acid residue were present in the interior of a globular protein, it would most likely be _________.
tightly associated with the R-group of a lysine residue
Alpha helices are stabilized primarily by what bonds between what
hydrogen bonds between main chain peptide bond component atoms
Tertiary structure is
the folding into a 3d piece
Fibrous proteins contain polypeptide chains ____ producing long fibers or large sheets
organized approximately parallel along a single axis
does a-keratin have polar amino acids?
no
In hemoglobin, a ____ protein, the space between the helices is filled efficiently and tightly with mostly amino acids with __side chains
globular, hydrophobic, polar
functional advantages of quaternary structures include
cooperativity, stability, bringing catalytic sites together, genetic economy and efficiency.
. In the tertiary structure of a protein, a hydrophobic interaction could form between the R-groups of which two amino acids? this question is essentialy asking for what
two non polar amino acids
The information needed for the structure of a protein is contained in
primary structure
. Hemoglobin differs from myoglobin because
hemoglobin is a tetramer, whereas myoglobin is a single polypeptide chain.
in allosteric interactions
changes that take place in one site of a protein cause changes at a distant site
what can denature proteins
heat, pH, detergents
. As catalysts, enzymes are
more effective than nonenzymatic cats
The substrate will only bind to the enzyme when the shapes fit together rigidly
false
In the induced-fit model of substrate binding to enzymes what happens
there is a conformational change in the enzyme when the substrate binds
The active site of an enzyme is the place where the following happen
The enzyme-substrate complex forms and the reaction occurs at the active site.
enzymes are flexible and can work well with different substrates?
Yes
The substrate-enzyme (E-S) complex
may break down to form free enzyme and substrate, or free enzyme and product (essentially may or may not work)
The rate of reaction depends on what
activation energy
catalizing a reaction does what
lowers the activation energy
does the catylist have an effect on delta g?
no
cofactors are what
a non-protein in chemical nature
Nicotinamide adenine dinucleotide is
. a coenzyme in oxidation-reduction reactions
. Which of the following statements about coenzymes is true?
a. They are commonly derived from vitamins.
b. They bind to the active site region on specific types of enzymes.
c. They can be metal ions, such as Zn(II).
d. NAD+, FAD and biotin are all examples of coenzymes.
e. All of these statements are true.
all of the above
which of the following is a mechanism of regulating enzyme activity?
a. Feedback inhibition by product.
b. Addition or removal of phosphate groups from of the enzyme.
c. Presence of activators.
d. Activation of zymogens.
e. All of these regulate enzyme activity
all of the above
Where do allosteric inhibitors bind on an enzyme?
they always bind at a site different from the active site
what type of inhibitor is most likely to inhibit regulatory subunits of an allosteric enzyme?
a competitive inhibitor
. For competitive inhibition is possible to overcome the effect of the inhibitor by increasing the concentration of substrate?
yes
A noncompetitive inhibitor binds where?
to the enzyme at a site other than the active site
does the Michaelis constant determine vmax?
no
What effect is seen on a Lineweaver-Burk graph when a competitive inhibitor is added?
the y intercept remains the same but the slope is changed
What effect is seen on a Lineweaver-Burk graph when a non-competitive inhibitor is added?
slope and y int are changes
Non-competitive inhibitors have what effect?
Changine the vmax
If an inhibitor changes the slope of the Lineweaver-Burk graph, but not the y-intercept, it is this type of inhibition:
competative
The rate of a zero-order reaction depends on the _____.
presence of a catalyst
Competitive Inhibition (what is it, vmax and km)
mimics the substrate and competes for the enzymes activation site, v max remains unchanged, km is higher
Non-Competitive Inhibition (what is it, vmax and km)
binds to the allosteric site (an additional site) on both the free enzyme and the e-s complex, decreases vmax, km unchanged
Un-Competitive Inhibition (what is it, vmax and km)
binds to the e-s complex, reduces vmax and kmax
competative graph
x
uncompetative graph
parallel
non- comp
touch at x axis and continue on
v=
vmax [s]/km+[s]
lineweaver burke the y intercept is what and the slope is what
y intercept is the km, slope is km/vmax
Name the positively charged amino acids
Lysine, Arginine, and Histidine
Name the negatively charged amino acids
Aspartatic acid and Glutamic acid
A polypeptide has a high pI value. What type of amino acids are likely to be present?
basic/negatively charged
Given that pK1 = 2.35 and pK2 = 9.69 for alanine. What is its isoelectric point?
6.02
Secondary and higher orders of structure are determined by all EXCEPT
hydrophobic interactions
ionic bonds
van der Waals forces
hydrogen bonds
peptide bonds
peptide bonds
An electrostatic interaction might occur within a protein between what type of amino acid pairs at typical physiological pH?
±
What remains intact when a protein is denatured
primary structure
The expression of the Michaelis constant is equal to
Where
K1 = the rate constant for ES formation
K2 = the rate constant for ES dissociation
K3 = the rate constant for product formation
(K2 + K3)/K1
In the Lineweaver-Burk double reciprocal plot the slope is equal to __________.
Km/Vmax
In the Lineweaver-Burk double reciprocal plot the vertical intercept is equal to __________.
1/Vmax
In the Lineweaver-Burk double reciprocal plot the horizontal intercept is equal to __________.
-1/km
In the michaelis menten plot the x axis is equal to __________.
S
In the michaelis menten plot the y axis is equal to __________.
V0
In the lineweaver burk plot the x axis equal to __________.
1/s
In the lineweaver burk plot the y axis equal to __________.
1/v
In competitive inhibition, increasing the concentration of substrate __________
increases the overall rate of the reaction
The pKa values for tyrosine are as follows:
pK1 2.20
pK2 9.11
pKR 10.07
What is the pI of tyrosine?
(choose the closest answer to the one you calculated)
9.59
5.65
6.14
9.11
5.65
A hydrophobic interaction might occur within a protein between what type of amino acid pairs?
polar/nonpolar
Alpha helices are stabilized primarily by:
electrostatic interactions between R-groups.
hydrogen bonds between the main chain peptide bond component atoms.
hydrophobic interactions between the α-carbons of the main chain.
hydrogen bonding between the R-groups.
hydrogen bonds between the main chain peptide bond component atoms.
In allosteric interactions
proteins that consist of a single polypeptide chain form aggregates.
disulfide bonds are broken.
changes that take place in one site of a protein cause changes at a distant site.
metal ions always bind to the protein.
changes that take place in one site of a protein cause changes at a distant site.
In competitive inhibition, Vmax remains unchanged but KM increases.
True
False
true
Increasing the substrate concentration can overcome non-competitive inhibition.
True
False
false
In a Michaelis Menten Plot, decreasing [S] has what effect on Vo ?
Has no effect
Decreases Vo
Increases Vo
Cannot be determined
Decreases Vo
In a Michaelis Menten plot ______ is represented on X-axis and ______ is represented on Y-axis.
[S], Vo
For an enzyme kinetics experiment, the Lineweaver-Burk plot of the data shows Y-intercept is 0.25. What is the Vmax of the enzyme under investigation.
4
In Michaelis–Menten kinetics, what does a low Km value indicate? Check all that apply
Low product formation
High reaction rate
High substrate affinity
Low enzyme concentration
Low substrate affinity
substrate concentration at Vmax
substrate concentration at Vmax/2
High reaction rate, High substrate affinity, substrate concentration at Vmax/2
In Michaelis Menten graph, the which of following hold true once the reaction reaches Vmax? Select all the answers that are correct. (select all that apply)
substrate concentration increases
Enzyme active sites are all saturated with substrate
No more substrate is available to bind to enzyme
Reaction follows first-order kinetics
reaction follows zero-order kinetics
The substrate concentration equals KM
Enzyme active sites are all saturated with substrate, reaction follows zero-order kinetics
after pka1 what is deprotonated, what isn’t
COOH is deprotonated, NH3+remains protonated
Why should the core of most globular and membrane proteins consist almost entirely of α-helix and β-sheets?
Hydrogen bonded structures must be kept away from water solvent.
Highly polar N−H and C=O moieties of the peptide backbone must be neutralized in the hydrophobic core of the protein.
Hydrogen bonding only occurs in the core of proteins.
Trapped water stabilizes the helix and sheet structures.
None are true.
Highly polar N−H and C=O moieties of the peptide backbone must be neutralized in the hydrophobic core of the protein.
Vmax of a reaction is is 100 umol/min and Km is 5mM. What is the velocity of this reaction when [S] is 20 mM?
a) cannot be determined
b) 40
c) 80
d) both b and c are right
e) 20
80
Which of the following is a coenzyme?
NADP+
Zn++
Cu++
Mg 2+
Oxytocin
NADP+
Which of the following is not a type of oxidoreductase?
Peroxidase
Hydroxylase
Reductase
Dehydrogenase
Peptidase
Peptidase
what is competitive inhibition
an inhibitor molecule binds directly to the enzyme’s active site
what is noncompetitive inhibition
an inhibitor binds to an allosteric site (a location other than the active site), decreasing enzyme efficacy
what is uncompetitive inhibition
Binds exclusively to the enzyme-substrate (ES) complex, preventing product formation.
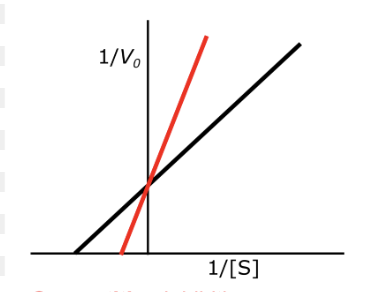
draw non competitive inhibition
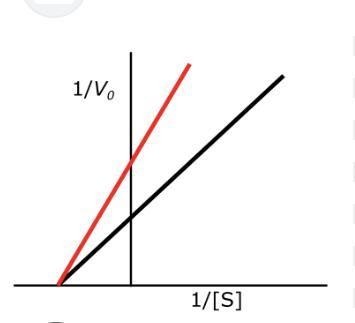
draw un competitive inhibition
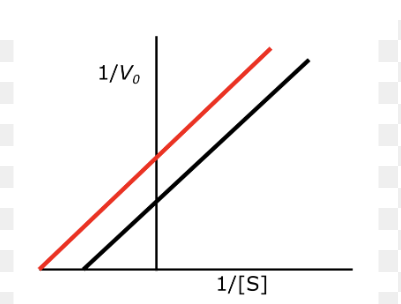
draw competitive inhibition
x shape (y intercept is same, x is changed)