Cell Structure and Function
1/104
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
105 Terms
all living organisms are composed of one or more cells
the cell is the basic unit of structure and organisation
all cells arise only from pre-existing cells
universal similarities bewteen cells:
DNA as the heritable material, RNA as a messenger and proteins as the workers
major cellular organelles - functions and arrangements within the cell
ATP as an energy source
the central dogma:
DNA → RNA → PROTEIN
what is cell theory
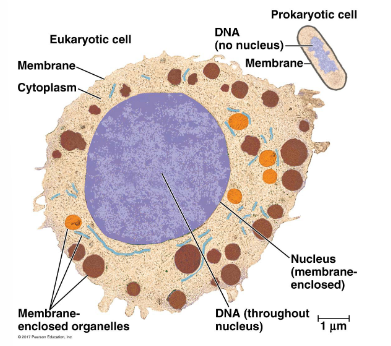
both have:
plasma membrane
cytosol
DNA
RNA
protein and ribosomes
eukaryotic cells have membrane-bound organelles and are much larger
prokaryote cells lack a membrane-bound nucleus
describe the similarities and differences between prokaryotes and eukaryotes
the cytoplasm is everything inside the plasma membrane except the nucleus
the fluid portion of the cytoplasm is the cytosol
water plus dissolved and suspended substances (e.g. ions, ATP, proteins, lipids)
major organelles include:
nucleus
endoplasmic reticulum (smooth and rough)
golgi apparatus
vesicles
these four make up the endomembrane system (along with plasma membrane, they work together to package, label and ship molecules)
mitochondria
ribosomes
what is the cytoplasm
the plasma membrane is a selectively permeable barrier controlling the passage of substances in and out of the cell
made up of a double layer of phospholipids with embedded proteins:
hydrophilic polar heads (phosphate)
hydrophobic lipid tails (fatty acids)
arranged as a double layer, tail to tail
much of our body is hydrophobic or ‘water loving’
fats are hydrophobic (‘water hating’)
fats in cell membrane provide a barrier to water
describe the plasma membrane
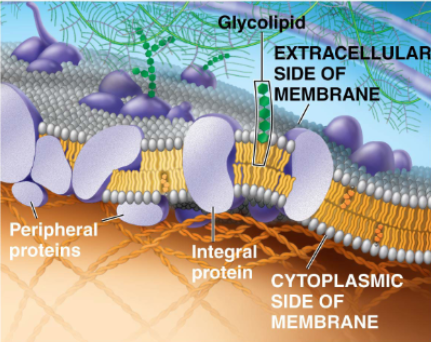
membrane proteins mediate movement of hydrophilic substances
are often amphipathic, meaning they have both hydrophilic and hydrophobic regions
integral proteins:
embedded (partially or fully) into the membrane
e.g. transmembrane proteins are integral membrane proteins that fully span the entire membrane, contracting both extracellular and cytoplasmic areas
peripheral membrane proteins:
are associated with the membrane, but not actually embedded within it
describe the plasma membrane proteins
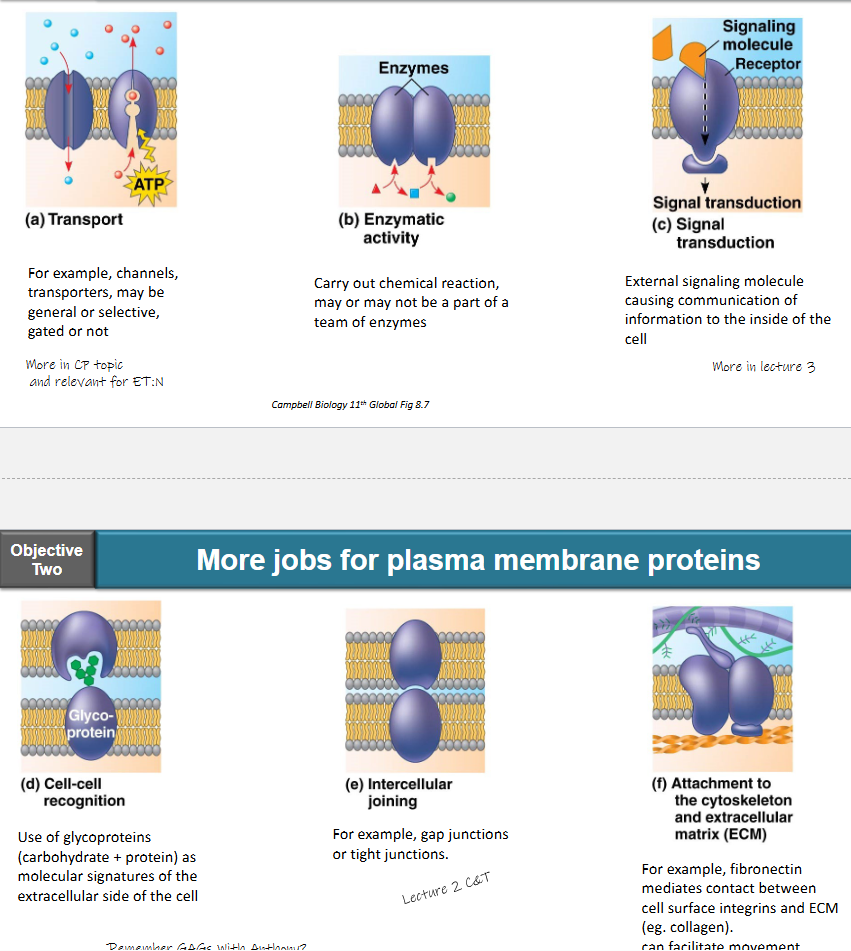
transport
e.g. channels, transporters
may be general or selective, gated or not
enzymatic activity
carry out chemical reaction, may or may not be a part of a team of enzymes
signal transduction
external signaling molecule causing communication of information to the inside of the cell
cell-cell recognition
use of glycoproteins (carbohydrate + protein) as molecular signature of the extracellular side of the cell
intercellular joining
e.g. gap junctions or tight junctions
attachment to the cytoskeleton and extracellular matrix (ECM)
e.g. fibronectin mediates contact between cell surface integrins and ECM (e.g. collagen)
can facilitate movement
what do the plasma membrane proteins do
membranes are not static
the membrane is a mosaic of molecules bobbing in a fluid bilayer of phospholipids
cell specific and dynamic repertoire of membrane-bound proteins present as required
describe the movement of membranes
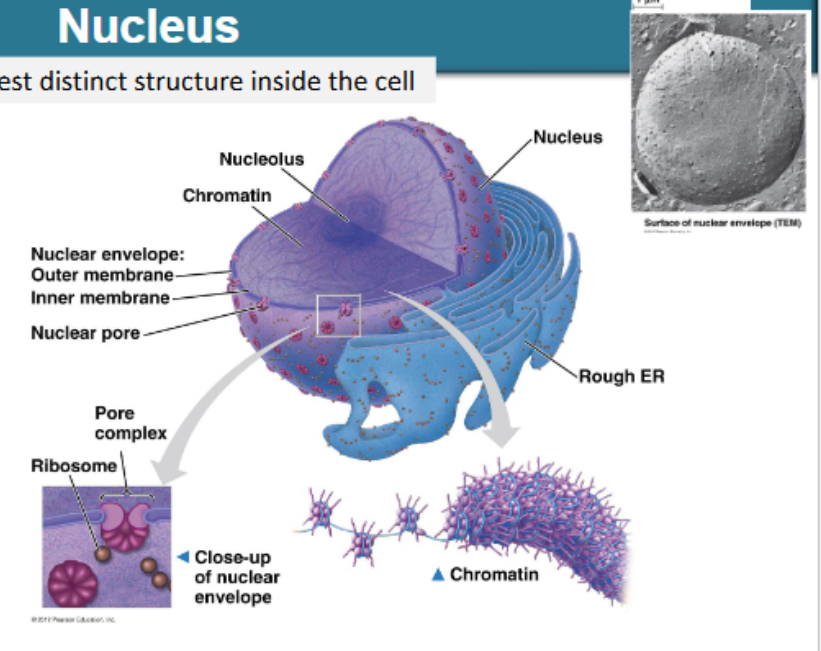
largest distinct structure inside the cell
enclosed by double lipid bilayer called nuclear envelope, continuous with rough ER
entry and exit through nuclear pores
nucleolus: rRNA production, assembly of small and large subunits of ribosomes
functions:
to house/protect DNA
make RNA
pores regulate movement of substances (e.g. protein and mRNA) in and out
molecule segregation to allow temporal and spatial control of cell function
describe the nucleus
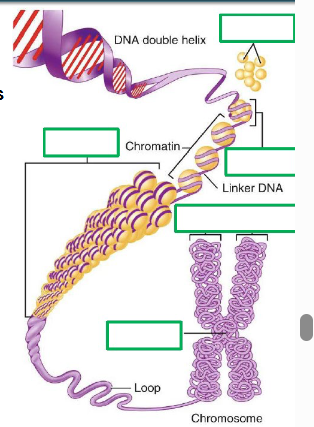
DNA wrapped 2x around group of 8 histones, to form nucleosomes - collectively known as chromatin
as the cell prepares for cell division, chromatin condenses to form chromatin fibres then condenses further into loops and then stacks as fully condensed chromosomes
most of the time, our DNA is present in our cells as chromatin and chromatin fibres
chromosome — comprises many genes, usually >1000
gene — a DNA segment that contributes to a phenotype/function
describe deoxyribonucleic acid (DNA) in the nucleus
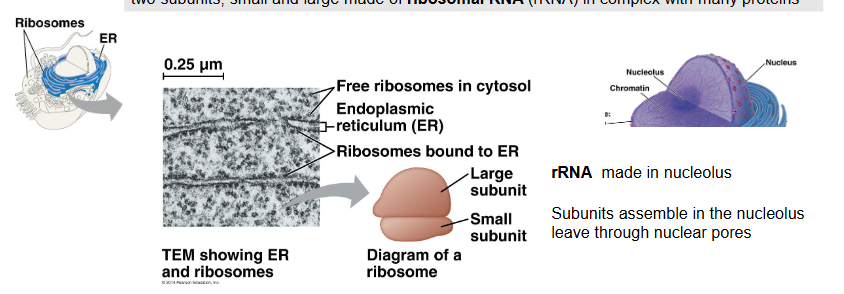
two subunits, small and large made of ribosomal RNA (rRNA) in complex with many proteins
rRNA is made in the nucleolus
subunits assemble in the nucleolus and leave through nuclear pores
function: protein production (translation), found in two places within the cell:
free in the cytoplasm — making proteins to be used in cytosol (non-endomembrane destinations)
attached to the RER — making non-cytosolic proteins/endomembrane
describe ribosomes
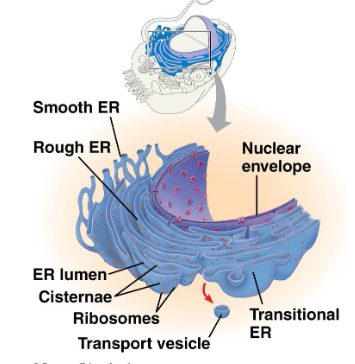
the ER is an extensive network of tubes and tubules, stretching out from the nuclear membrane
two types: rough ER and smooth ER
outline the endoplasmic reticulum
continuous with nuclear envelope dotted with attached ribosomes
proteins enter lumen within the rough ER for folding
rough ER membrane surrounds the protein to form transport vesicles destined for the Golgi
major function is production of:
secreted proteins
membrane proteins
organelle proteins
describe the rough endoplasmic reticulum
extends from the rough ER
lacks ribosomes: doesn’t make proteins
synthesises lipids, including steroids and phospholipids
stores cell-specific molecules
functions of smooth ER vary greatly from cell to cell
very cell/tissue-type specific
examples:
liver: houses enzymes for detoxification and for glucose release
muscle: calcium ions
describe the smooth endoplasmic reticulum
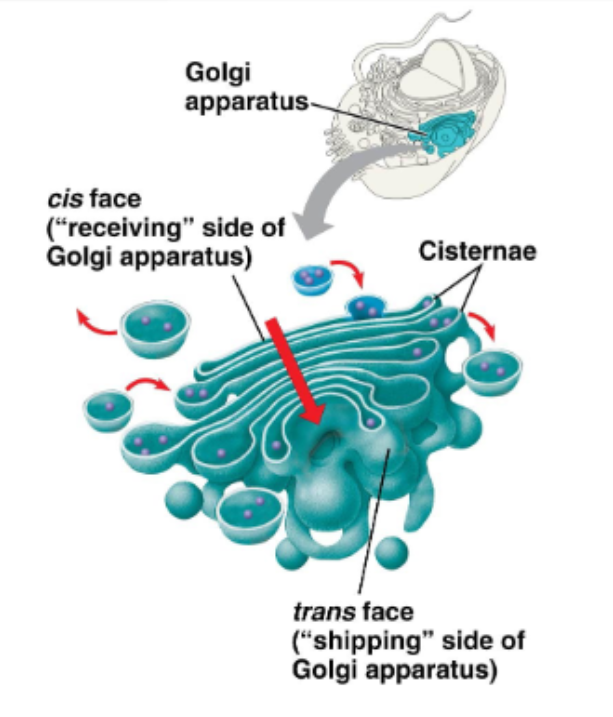
the ‘warehouse’ of the cell
this complex is made up of 3 to 20 flattened membranous sacs called cisternae, stacked on top of one another (like ‘pita bread’)
functions:
modify, sort, package, and transport proteins received from the rough ER using enzymes in each cisternae
formation of:
secretory vesicles (proteins for exocytosis)
membrane vesicles (PM molecules)
transport vesicles (molecules to lysosome)
describe the Golgi apparatus - receiving and modifying
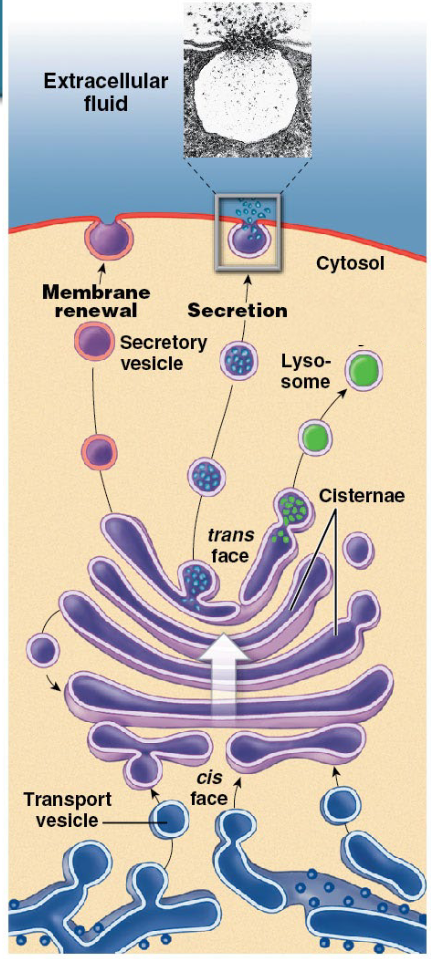
each sac or cisternae contains enzymes of different functions
proteins move cis to trans from sac to sac
mature at the exit cisternae
travel to destination within vesicles
modifications occur within each sac (formation of glycoproteins, glycolipids, and lipoproteins)
describe the Golgi apparatus: to destination
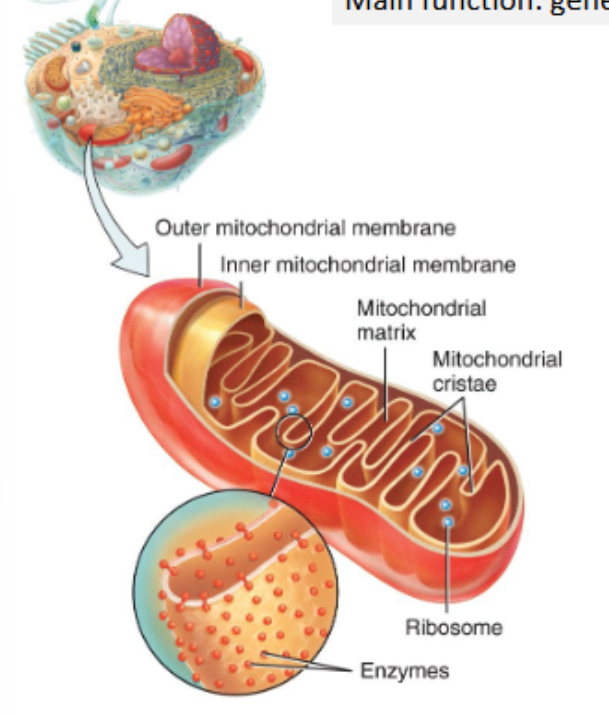
main function: generation of ATP through cellular respiration
mitochondria are made up of:
outer mitochondrial membrane
inner mitochondrial membrane, with folds called cristae f
fluid filled interior cavity, called the mitochondrial matrix
despite all of these membranes, mitochondria are not part of the endomembrane system
the more energy a cell requires, the more ATP it must take, and the greater the number of mitochondria present
mitochondria carry a separate small (37 genes) genome encoding mitochondrial-specific products
describe the mitochondria
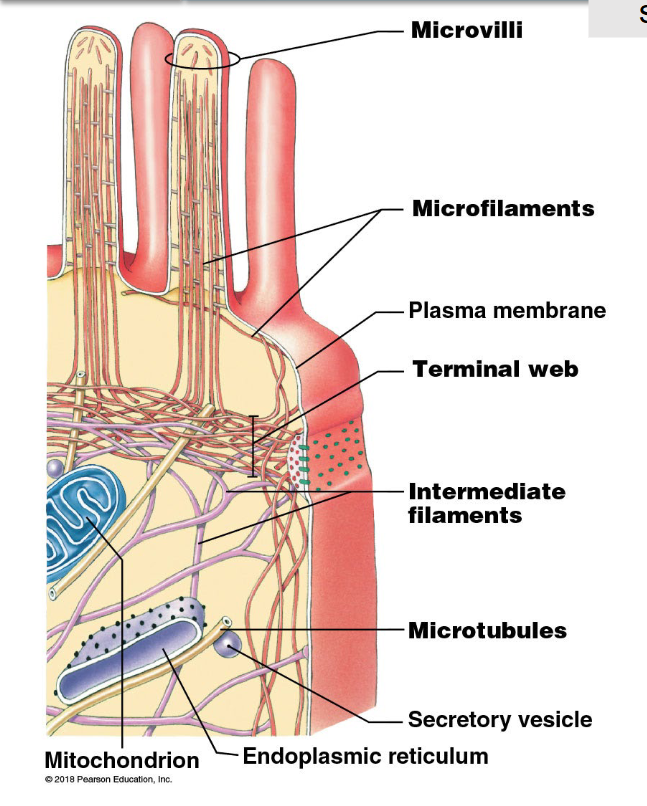
structural support system of the cell
fibres or filaments that help to maintain the size, shape, and integrity of the cell:
act as scaffolding across the cell
involving in intracellular transportation and cell movement
three types of fibres (from smallest to largest):
microfilaments
intermediate filaments
microtubules
describe the cytoskeleton
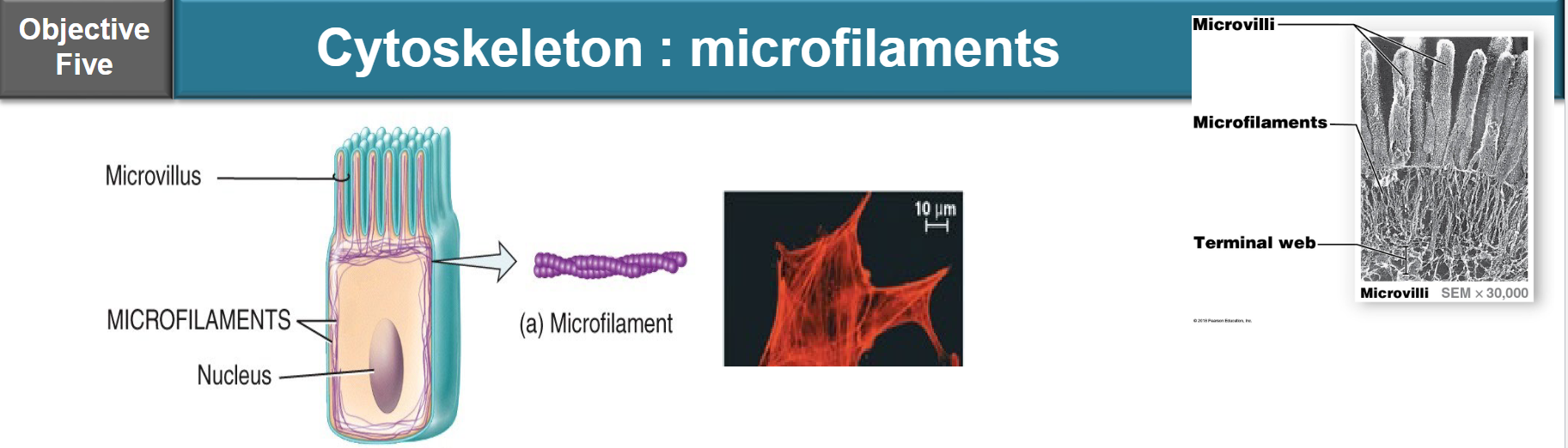
diameter: ~7nm
comprised of actin molecules assembled in two long chains, twisted around each other
found around the periphery and lining the interior of cell
function:
bear tension and weight by anchoring cytoskeleton to plasma membrane proteins, and promote amoeboid mobility if required (e.g. macrophage)
assembled and disassembled as required — they are dynamic
describe the microfilaments in the cytoskeleton
diameter: 8-12nm
comprised of diverse range of different materials; one example: keratin
found in the cytoplasm of the cell
function:
bear tension and weight throughout cell, e.g. during cell anchoring
acts as a scaffold for cellular organelles, e.g. the nucleus
usually the most permanent of cytoskeletal structures — they are less dynamic
describe the intermediate filaments of the cytoskeleton
diameter: tubular structure, 25nm with central lumen of 15nm diameter
comprised of tubulin dimers (alpha and beta), coiled, to form a tube
extends from centriole into cytoplasm/nucleus
functions:
support cell shape and size
guide for movement of organelles
e.g. vesicles from Golgi to membrane
chromosome organisation — cell division
support and movement of cilia/flagella
assembled and disassembled as required — are dynamic
describe the microtubules in the cytoskeleton
ATP powers cellular work - it is our energy currency
the hydrolysis of ATP to ADP and inorganic phosphate releases energy
outline the mitochondria as the ATP factory
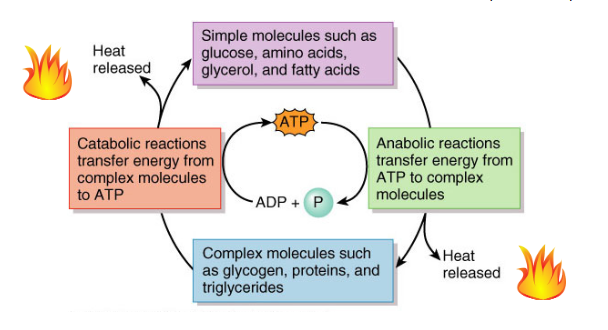
ATP cycle: the transfer of energy between complex and simple molecules in the body, with ATP as the mediator
many cellular processes require energy in the form of ATP — they are not spontaneous
simple molecules such as glucose, amino acids, glycerol, and fatty acids → anabolic reactions transfer energy from ATP to complex molecules → complex molecules such as glycogen, proteins, and triglycerides → catabolic reactions transfer energy from complex molecules to ATP
describe the ATP Cycle
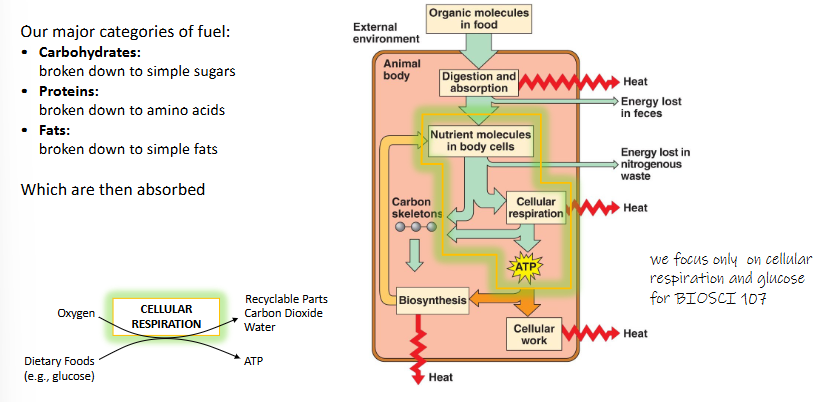
our major categories of fuel:
carbohydrates: broken down to simple sugars
proteins: broken down to amino acids
fats: broken down to simple fats
which are then absorbed
how is fuel needed to generate ATP
glucose in food/intestines → glucose in bloodstream ← storage for harder times (facilitated by glucagon)
glucose in bloodstream → into a cell (faciliated by insulin)
cellular respiration ← cell → storage for harder times (glucose cross-linked together, called glycogen, in liver and skeletal muscle)
cellular respiration → cellular work
describe the use of glucose in different parts of the body as it moves around
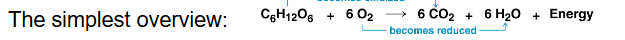
the controlled release of energy from organic compounds to produce ATP
conversion of glucose to ATP is due to 4 main steps:
glycolysis
pyruvate oxidation
citric acid cycle (or Krebs cycle)
oxidative phosphorylation
the simplest overview: C6H12O6 + 6O2 → 6CO2 + 6 H2O + Energy
outline the process of cellular respiration
glycolysis (glucose → pyruvate) → cytosol
pyruvate oxidation and Kreb’s cycle (acetyl CoA)→ mitochondrial matrix
oxidative phsophorylation (electron transport and chemiosmosis) → across inner membrane
where does cellular respiration occur
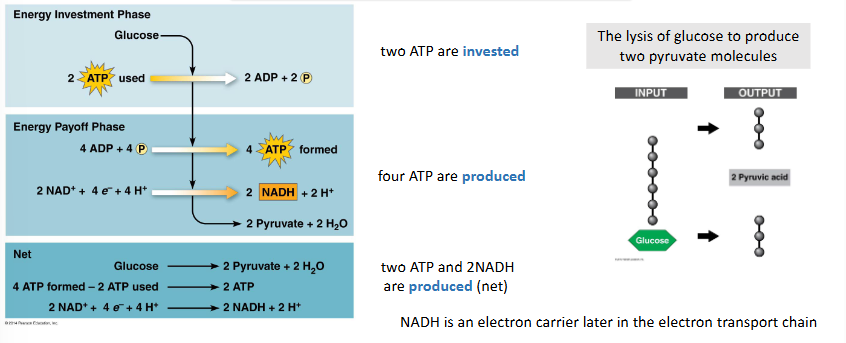
glycolysis invests and produces ATP
occurs in the cytosol and oxygen is not required
two ATP are invested
the lysis of glucose to produces two pyruvate molecules
four ATP are produced
two ATP and 2NADH are produced (net)
NADH is an electron carrier later in the electron transport chain
describe step 1 of cellular respiration: glycolysis
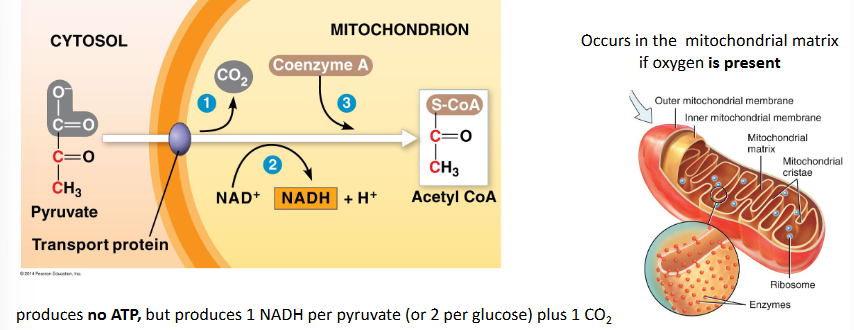
pyruvate oxidation to form acetyl CoA
this step links glycolysis to the citric acid cycle
occurs in the mitochondrial matrix if oxygen is present
produces no ATP, but produces 1NADH per pyruvate (or 2 per glucose) plus 1 CO2
the 2 carbon acetyl CoA molecule is able to enter the nitric acid cycle
describe step 2 of cellular respiration: pyruvate oxidation
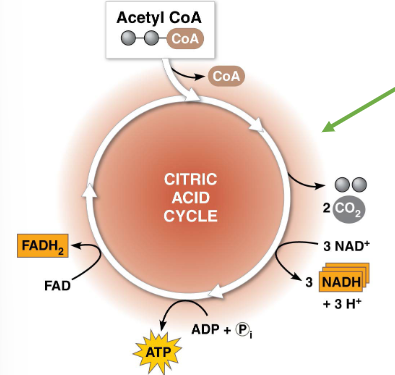
occurs in the mitochondrial matrix if oxygen is present
results in (per glucose molecule):
2 ATP
6 NADH
2 FADH2
4 CO2
requires oxygen — it is an aerobic process
FADH2 and NADH are electron donors in the electron transport chain
describe step 3 of cellular respiration: citric acid/Kreb’s cycle
citrate → fatty acid synthesis
α-Keto-glutarate → amino acid synthesis and neurotransmitter
oxaloacetate → amino acid synthesis
malate → gluconeogenesis
a series of reactions: product of one reaction is the substrate for the next
the citric acid cycle completes the extraction of energy from glucose
outline the citric acid cycle intermediates are used in other metabolic pathways
ATP genereated by direct transfer (from a substrate) of a phosphate group to ADP via substrate phosphorylation
what is substrate phosphorylation
ATP is generated from the oxidation of NADH and FADH2 and the subsequent transfer of electrons and pumping of proteins
what is oxidative phosphorylation
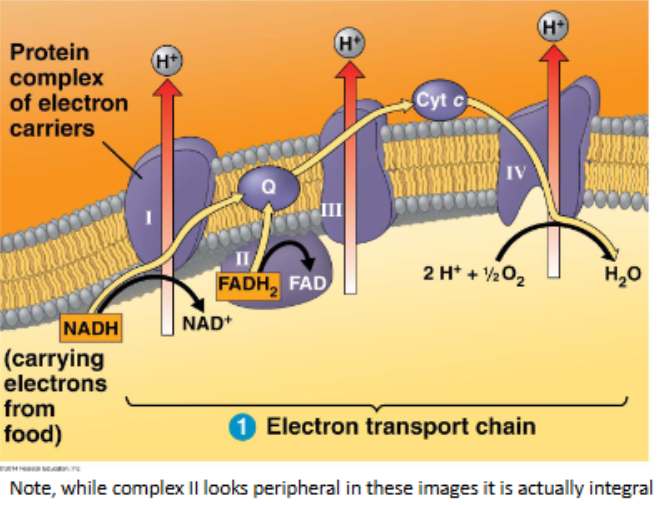
the electron transport chain
occurs at proteins within the inner membrane
requires oxygen — it is an aerobic process
NADH and FADH2 are oxidised to donate electrons
electrons transfer from protein-to-protein along the chain in a series of redox reactions
at each transfer, each electron gives up a small amount of energy which enables H+ ions to be pumped into the intermembrane space
oxygen ‘pulls’ the electrons down the chain, and is then the final electron acceptor where it is reduced to water
NADH and FADH2 from earlier steps are used here
chemiosmosis
the hydrogen ions in the intermembrane space rush down their concentration gradient (chemiosmosis) through ATP synthase
this causes the ‘turbine’ within ATP synthase to turn
the rotation of the ATP synthase turbine enables the phosphorylation of ADP to generate ATP
this results in the production of 26 or 28 ATP (per glucose)
ETC and chemiosmosis = oxidative phosphorylation
this is much more efficient than substrate phosphorylation
the bulk of ATP production occurs here
‘fall’ of electrons down the chain enables movement of H+ ions into intermembrane spcae and generates a proton gradient which ‘drives’ the ATP synthase turbine
step 4: oxidative phosphorylation: the electron transport chain and chemiosmosis
we can derive energy from more than just glucose
fats, proteins, and more complex carbohydrates generate ATP also
monomers enter glycolysis and the citric acid cycle at different points
outline how cellular respiration is versatile
phosphofructokinase is the ‘gate-keeper’ for glycolysis; it catalyses step 3 — where glycolysis becomes irreversible
inhibited by citrate and ATP
i.e. products of cellular respiration
stimulated by AMP
AMP accumulates when ATP is being used rapidly
how is cellular respiration controlled
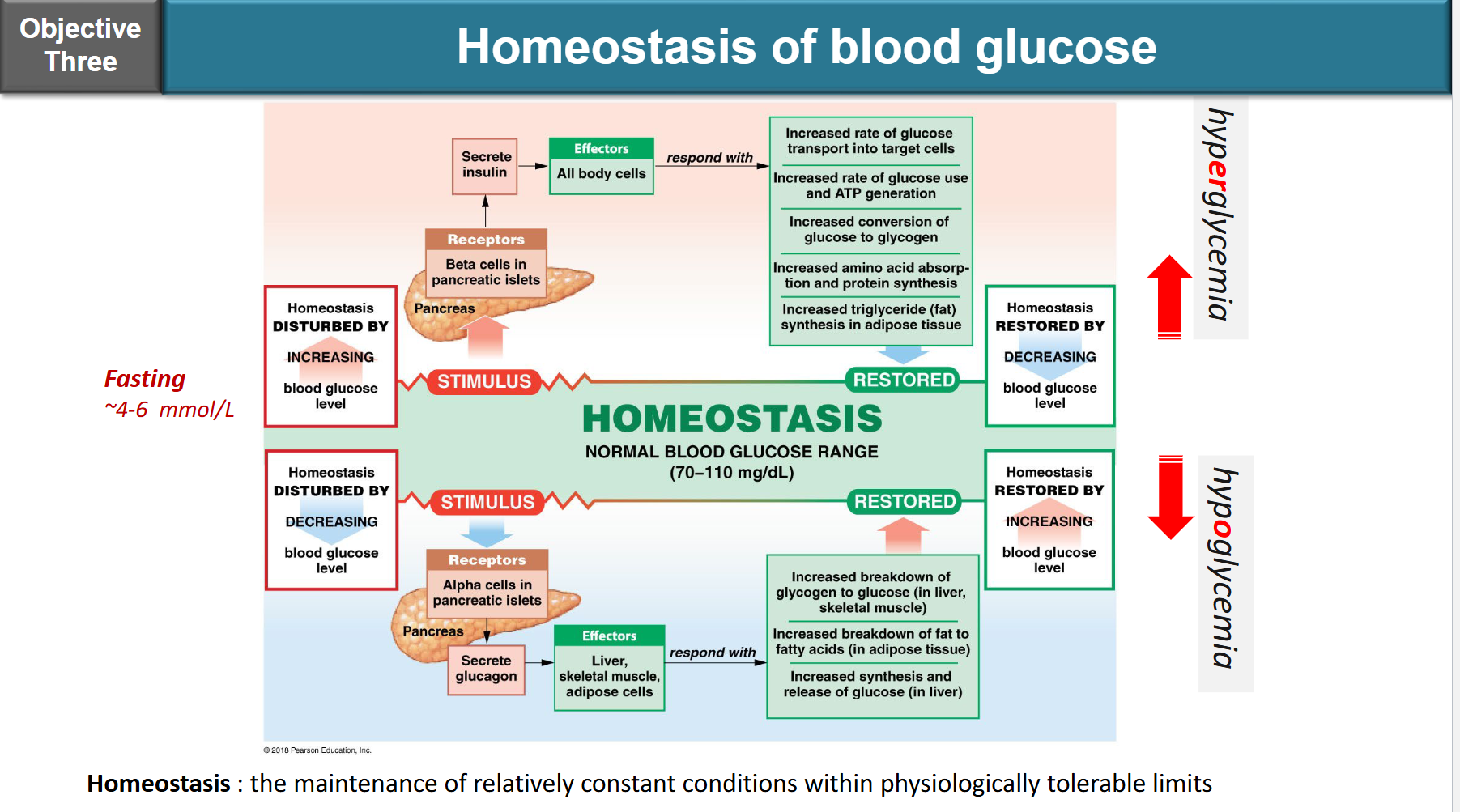
insulin:
produced by beta cells of islets and langerhans in pancreas
function: promote glucose uptake into cells (for ATP production or storage in liver)
glucagon:
prodcued by alpha cells of Islets of Langerhans in pancreas
function: stimulates the breakdown of glycogen to increase blood sugar levels
outline insulin and glucagon
no glucose in cells
no ATP from glucose
no glycogen stored for harder times
diabetes mellitus:
the ability to produce or respond to hormone insulin is impaired
results in abnormal metabolism of carhydrates and elevated levels of glucose in the blood
what happens if you lose the function of insulin
type 1 or insulin-dependent diabetes:
body does not produce insulin, as beta cells of pancreas are destroyed, often this is autoimmune, or genetic or through environmental factors
affects 5-10% of diabetics, and onset usuaully occurs in children or adolescents
requires insulin replacement
type 2 or non-insulin-dependent diabetes:
body produces insulin, but receptors are non function (insulin resistance)
most (>90%) diabetics are type II, usually adults over the age of 40
can be linked to other pathologies and obesity
outline diabetes mellitus
diabetes mellitus is caused by a lack of functional insulin
as a result, levels of glucose in the blood build up, well beyond normal homeostatic limits
increased blood glucose alters the volume and osmolarity of blood, with subsequent pathological consequences
two of the symptoms of this diseases are:
significantly increased hunger
significant weight loss
these two symptoms seem to be in opposition to each other: if the patient is constantly hungry and eating, why would they then lose weight?
what are contradictory symptoms of diabetes mellitus
cells need to be able to respond as a cell, and as part of a whole tissue
they respond to signals from other cells and from the environment
these signals are often chemical
why do cells communicate
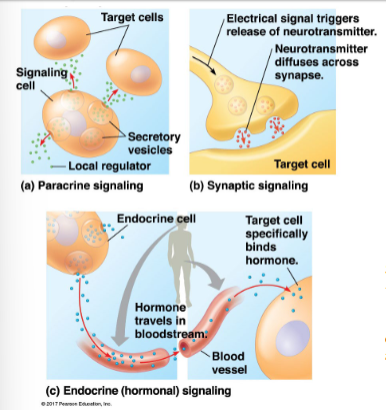
secreted signals can be local or long distance
local signaling:
signals act on nearby target cells
growth factors such as fibroblast growth factor — FGF1 (paracrine)
neurotransmitters such as acetylcholine - ACh (synaptic)
can act on the signalling cell (autocrine)
long distance signaling:
signals act from a distance
hormones secreted from endocrine cells travel via circulatory system to act on target cells
e.g. insulin secreted from pancreatic beta cells enter bloodstream and travels and is detected by various body cells
outline the differences between local signaling and long distance signaling
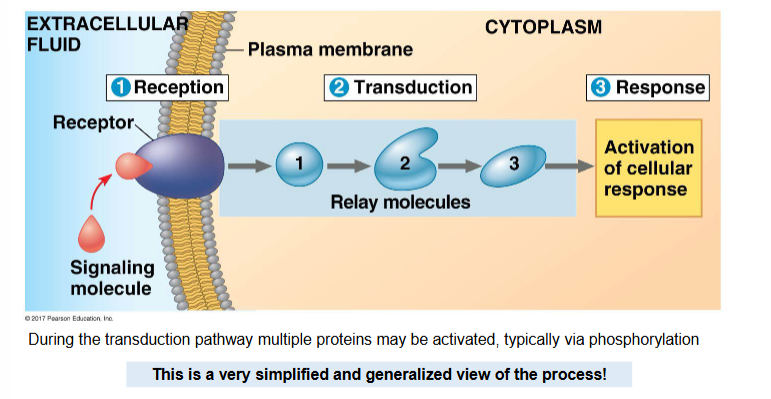
reception
signalling protein (primary messenger) binds to a receptor protein
results in shape and/or chemical state change in the receptor protein
transduction
altered receptor activates a another protein, e.g. G-protein/adenylyl cyclase
the activated protein (often an enzyme) may cause a relay of changes
relay molecules known as ‘secondary messengers’, e.g. cAMP, IP3
multiple other proteins may be activated
each activated protein causes a series of changes, this is often via phosphorylation — known as a phosphorylation cascade
response
all of the activated protens cause one or more functions to occur in the cell
this is where the cell actually does something
outline the three main steps of cell signalling
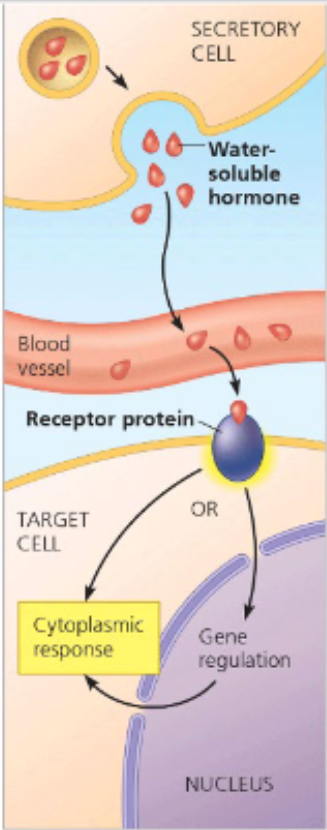
the human body will simulataneously send out many different chemicals and molecules, all aimed at eliciting specific responses BUT only the target receptors will interact with that signal (ligand) and use it to activate signal transduction pathways
specifity comes from the 3D molecular shape of the proteins involved — structure determines function
exquisite control is possible: only certain cells at certain times will have particular receptors (i.e. dynamic), meaning that while the signal might be widespread the transmission of the signal occurs only where it is needed
describe the specifity of receptors
receptors for water soluble molecules are membrane bound
e.g. G protein couped receptor, receptor tyrosine kinase, ligand-gated ion channel
receptors for lipid soluble molecules are not membrane bound
can be located in the cytoplasm or inside the nucleus
e.g. lipid soluble hormones such as testerones, estrogen, progesterone, thyroid hormones bind to receptors within the cytoplasm and move to nucleus as a complex
outline where receptors are located
transmembrane proteins — pass PM 7 times
hundreds of different GPCRs exist
many different ligands
diverse functions:
e.g. development, sensory reception (vision, taste, smell)
GPCRs couple with G proteins
G proteins are molecule switches which are either on or off depending on whether GDP or GTP is bound
(GTP: guanosine triphosphate, similar to ATP)
describe G-protein coupled receptors (GPCRs)
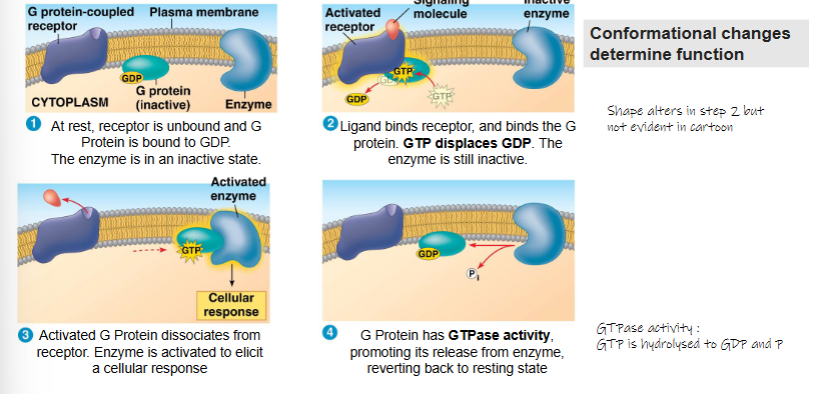
At rest, reeptor is unbound and G Protein is bound to GDP. The enzyme is in an inactive state
Ligand binds receptor, and binds the G protein. GTP displaces GDP. The enzyme is still inactive
Activated G Protein dissociates from receptor. Enzyme is activated to elicit a cellular response
G Protein has GTPase activity, promoting its release from enzyme, reverting back to resting state
describe the process of G-protein coupled receptors (GPCRs) being activated
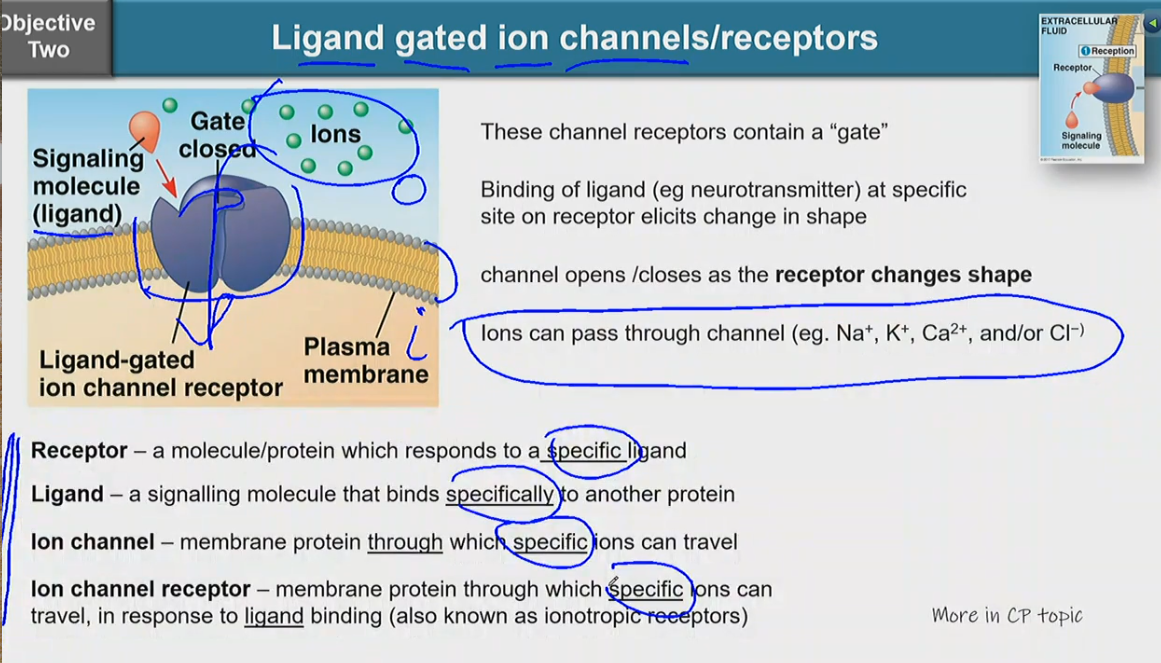

these channel receptors contain a ‘gate’
binding of ligand (e.g. neurotransmitter) at specifc site on receptor elicits change in shape
channel opens/closes as the receptor changes shape
ions can pass through channel (e.g. Na+, K+, Ca2+, and/or Cl-)
receptor — a molecule/protein which responds to a specific ligand
ligand — a signalling molecule that binds specifically to another protein
ion channel — memmbrane protein through which specific ions can travel
ion channel receptor — membrane protein through which specific ions can travel, in response to ligand binding (also known as ionotropic receptors)
describe ligand gated ion channels/receptors
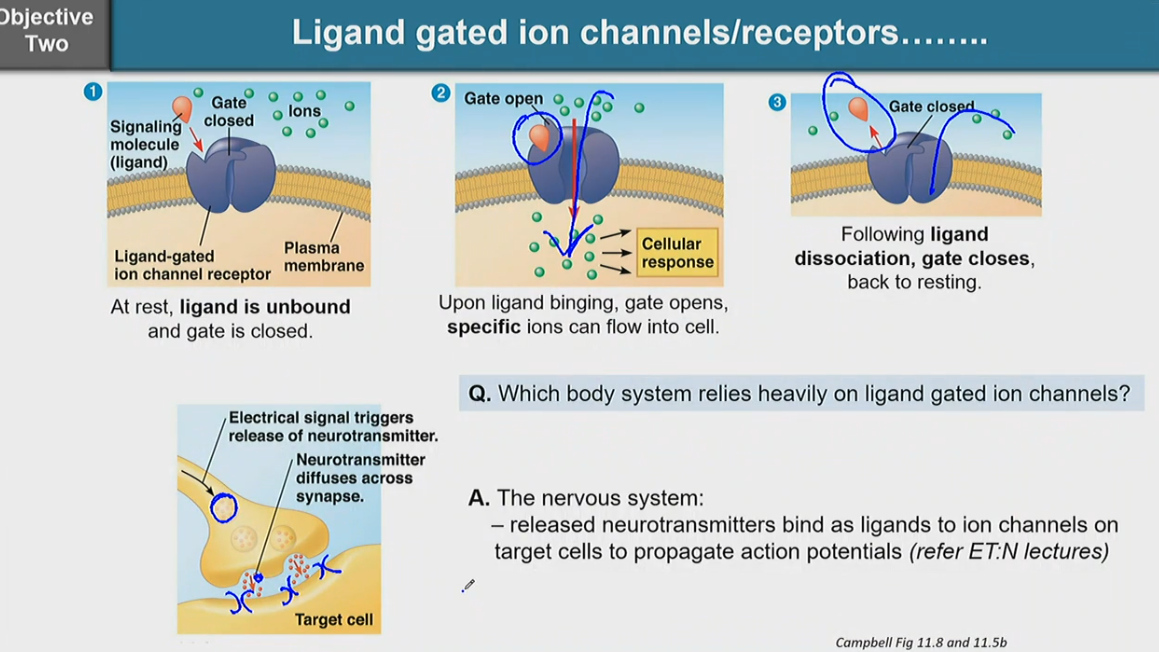
at rest, ligand is unbound and gate is closed
upon ligand binding, gate opens, specific ions can flow into cell
following ligand dissociation, gate closes, back to resting
the nervous system heavily relies on ligand gated ion channels
the nervous system releases neurotransmitters and bind as ligands to ion channels on target cells to propagate action potentials
describe the process in which ligand gated ion channels/receptors work
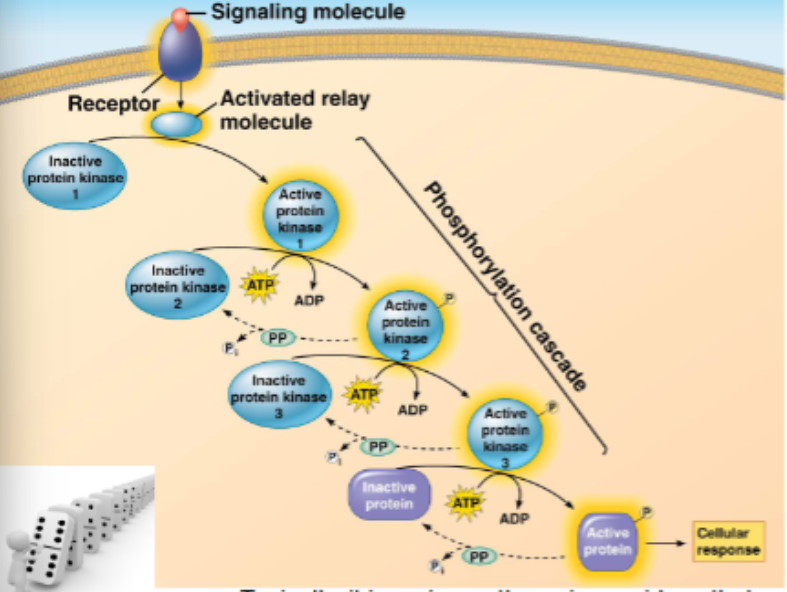
signals relayed from receptors to target molecules via a ‘cascade’ of molecular interactions
protein kinases are enzymes that transfer a phosphate group from ATP to another (specific) protein (kinases phosphorylate), typically, this activates the protein
series of protein kinases each adding a phosphate to the next kinase
phosphates are enzymes that dephosphorylate (remove the phosphate) rendering the protein inactive, but recyclable
typically, it is serine or threonine residues that are phosphorylated
this means that mutations affecting these residues could be detrimental
describe transduction pathways
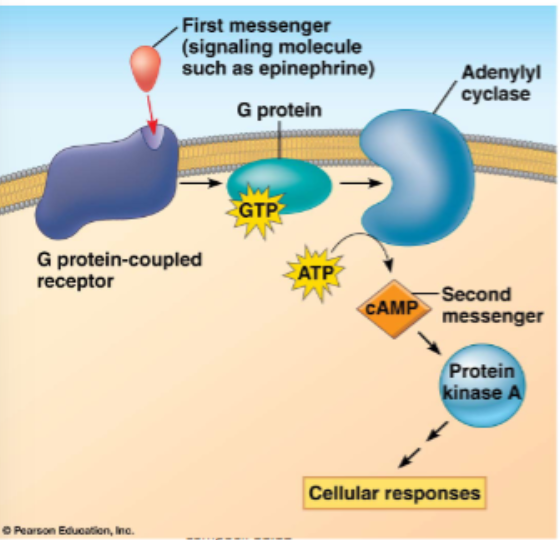
sometimes another small molecule is included in the cascade, these are second messengers
e.g. cAMP and calcium ions
recall earlier GPCR slide, plus:
the activated enzyme is adenylyl cyclase converts ATP to cAMP
cAMP acts as a secondary messenger and activates downstream proteins, for example, PKA which phosphorylates other proteins
outline cAMP as a second messenger
![<ul><li><p>low [Ca2+] inside cell (typically ~100nm)</p></li><li><p>very high [Ca2+] outside the cell (more than 1000-fold higher)</p></li><li><p>maintenance of concentration via calcium pumps is important </p><ul><li><p>out of cell</p></li><li><p>into ER</p></li><li><p>into mitochondria</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/6de4e3b2-f59a-4713-ae4d-1b8badd41d05.png)
low [Ca2+] inside cell (typically ~100nm)
very high [Ca2+] outside the cell (more than 1000-fold higher)
maintenance of concentration via calcium pumps is important
out of cell
into ER
into mitochondria
outline calcium as a secondary messenger
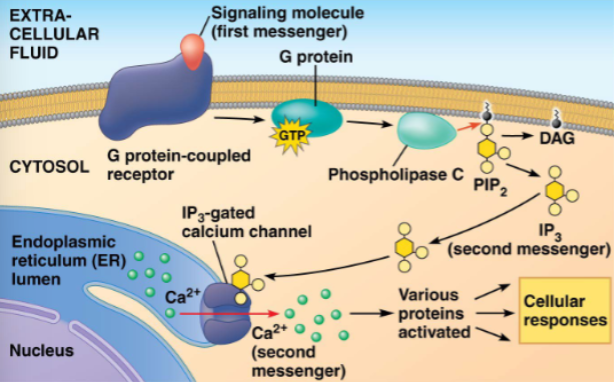
here, the activated proteinis phospholipase C which then cleaves PIP2 (a phospholipid) into DAG and IP3
IP3 diffuses through cytosol and binds to a gated channel in the ER
calcium ions flow out of the ER down a concentration gradient and activate other proteins toward a cellular response
describe the role of Ca2+ and IP3 in GPCR signalling
amplifies the response
provides multiple control points
allows for specificity of response
temporal
spatial
despite molecules in common
allows for coordination with other signalling pathways
why are there so many steps to transduction
examples of a cellular response include activation or regulation of:
gene expression
alteration of protein function to gain or lose an activity
opening or closing of an ion channel
alteration of cellular metabolism
regulation of cellular organelles or organisation
rearrangement/movement of cytoskeleton
a combination of any of these
the transduction of a signal leads to the regulation of one or more cellular activities
what are examples of cellular responses
all of the signals are for a limited time: activation usually promotes the start of deactivation, so that signalling is of short period of time, ensuring homeostatic equilibrium
it means the cell is ready to respond again if required
cAMP is broken down by phosphodiesterase (PDE)
caffeine blocks the action of pDE
inhibition of specific PDE’s can also be a therapeutic approach
e.g. viagra — inhibits a specific cGMP — degrading PDE
outline the importance of a response being turned off
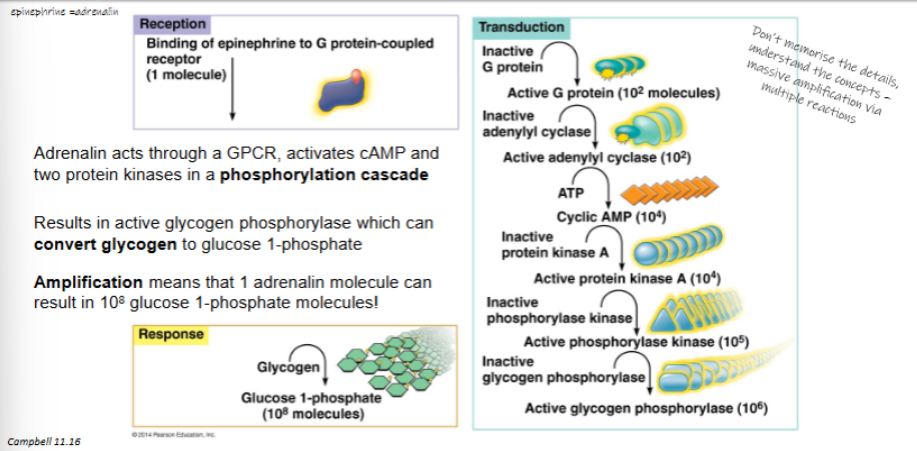
adrenalin acts through GPCR, activates cAMP and two protein kinases in a phosphorylation cascade
results in active glycogen phosphorylase which can convert glycogen to glucose 1-phosphate
amplification means that 1 adrenalin molecule can result in 108 glucose 1-phosphate molecules
outline how adrenalin stimulates glycogen breakdown
glycogen is a long term energy store in liver and skeletal muscle
glycogen breakdown results in glucose 1-phosphate
glucose 1-phosphate is then converted to glucose 6-phosphate which can then be used in glycolysis to generate ATP
outline how a large amount of ATP is generated quickly
angiotensin-converting enzyme 2 (ACE2) is the cellular receptor for the coronavirus (SARS-CoV-2)
surface spike glycoprotein (S protein)
here, ACE2 in our respiratory tract is the lock, and the S-protein on the virus is the key
outline how receptors can be deceived
the process of going from DNA to a functional product (typically a protein)
what is gene expression
an organisms’s hereditary information
what is a genotype
actual observable or physiological traits
what is a phenotype
our genotype and its interaction with the environment
what determines our phenotype
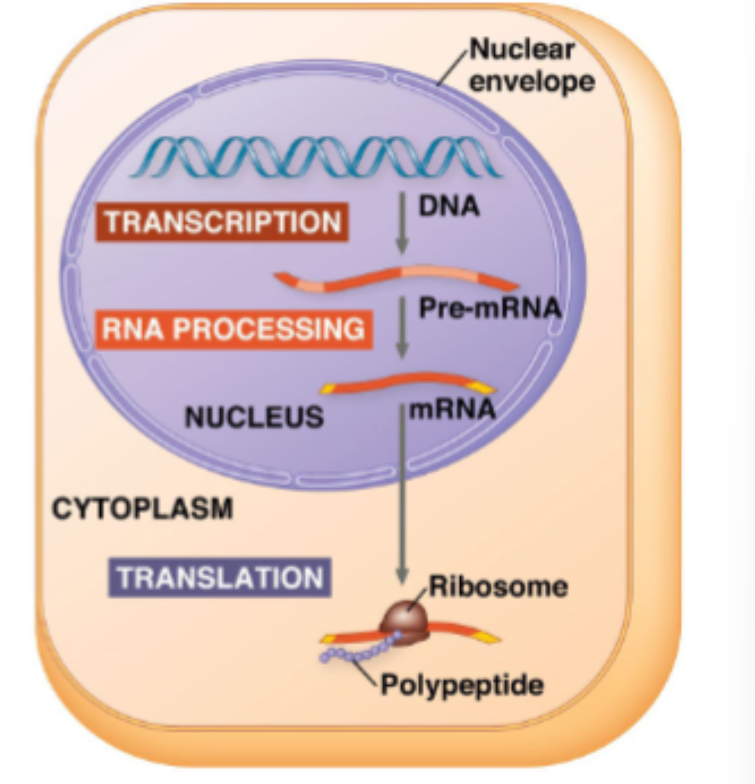
DNA (deoxyribonucleic acid) is the heritable material that is used to store and transmit information from generation to generation
RNA (ribonucleic acid) acts as a messenger to allow the information stored in the DNA to be used to make proteins
proteins carry out cellular functions
three main steps:
transcription of RNA from DNA
processing of the pre-mRNA transcript
translation of the mRNA transcript to a protein
where does gene expression happen and what happens during the process of gene expression
three steps:
initiation: polymerase binds to promoter
elongation: moves downstream through the gene, transcribing RNA
termination:detaches after terminator reached
RNA uses the nitrogenous base Uracil, in place of Thymine and it is single stranded, while DNA is double stranded
outline transcription
assembly of multiple proteins required before transcription can commence
TATA box typically ~25nt upstream found in the promoter region
assembly of several transcription factors including the TATA box binding protein (TBP) bind to DNA
RNA Pol II can now bind along with more transcription factors to form the transcription initiation complex, and so transcription begins
initiator tRNA = tRNA carrying methionine (Met)
small ribosomal subunit with initiator tRNA already bound binds 5’ cap of mRNA
small ribosome subunit scans downstream to find translation start site (AUG)
hydrogen bonds form between initiator anticodon and mRNA
large ribosomal subunit then binds — completing the initiation complex
energy (GTP — guanosine triphosphate) is required for assembly
outline the initiation process of transcription
10-20 nucleotides exposed at a time when DNA unwound
elongation: complementary RNA nucleotides added to 3’ end of growing transcript (3’OH of transcript binds with 5’ phosphate of incoming nucleotide) — It forms a phosphodiester bond
codon recognition:
base pairs with complementary anticodon GTP invested to increase accuracy/efficiency
peptide bond formation: '
a large subunit rRNA catalyses peptide bond formation
removes it from tRNA in P site
translocation:
moves tRNA from A to P site
tRNA in P site moves to E and is released
energy is required
empty tRNAs are ‘reloaded’ in the cytoplasm using aminoacyl-tRNA synthetases
double helix reforms as transcript leaves the template strand
termination: after transcription of the polyadenylation signal (AAUAAA) nuclear enzymes release the pre-mRNA and RNA polymerase then dissociates from the DNA
ribosome reaches a stop codon on mRNA
mRNA stop codon in the A site is bound by a release factor
release factor promotes hydrolysis
bond between p-site tRNA and last amino acid is hydrolysed, releasing polypeptide
ribosomal subunits and other components dissociate
hydrolysis of two GTP molecules required
ribosome components can be recycled
fidelity (proofreading) is less than for DNA replication
the pre-mRNA transcript is now ready for further processing
outline the elongation and termination process of transcription
capping: a modified guanosine nucleotide is added to the 5’ end
tailing: 50-250 adenosine nucleotides (polyA) are added to the 3’ end
capping and tailing are thought to facilitate export, confer stability and facilitate ribosome binding in cytoplasm
splicing: introns are removed from the transcript, typically making mRNA much smaller than Pre-mRNA
definitions to know:
exons: regions that remain in mature RNA (includes UTR)
UTR: untranslated regions of 5’ and 3’ ends of mRNA
introns: intervening regions that do not remain in mature RNA
outline the second step of mRNA processing — capping, tailing, and splicing
splicing occurs at the spliceosome, within the nucleus
spliceosome: a large complex of proteins and small RNAs
introns are removed from the transcript and exons are rejoined to form mature mRNA
alternative splicing is a process by which different combinations of exons are joined together, this results in the production of multiple forms of mRNA from the same pre-mRNA population
alternative splicing allows for multiple gene products from the same gene
~20,000 genes, there could be many times that number of proteins
where does splicing occur
protein sequence determine its final structure
structure determines function
DNA mutations can affect ability of the protein to function
outline how protein sequence determines the function
mature mRNA transcript exits nucleus and is bound by the ribosome
codons are translated into amino acids
tRNA molecules within the cytosol with specific anticodons carry corresponding amino acids
hydrogen bonds form between mRNA and antidcodon of the appropriate tRNA
the amino acid is added via peptide bonds to the growing polypeptide chain
outline translation
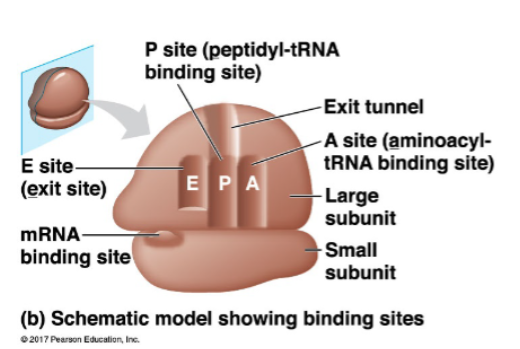
tRNA and mRNA are held within a ribosome to enable the formation of the polypeptide
mRNA binding site on small subunit
A site: holds ‘next in line’ tRNA
P site: holds tRNA carrying the growing polypeptide
E site: tRNAs exit from here
outline the ribosome binding sites for mRNA and tRNA
tRNA is the physical link between the mRNA and the amino acid sequence of proteins
what is the role of tRNA
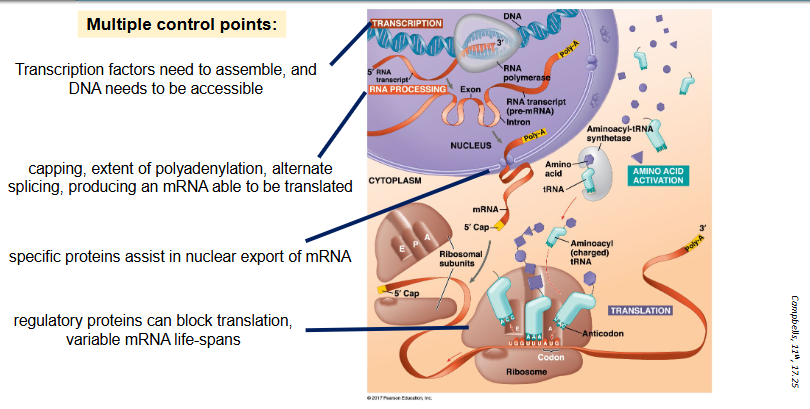
multiple control points:
transcription factors need to assemble, and DNA needs to be accessible
capping, extent of polyadenylation, alternate splicing, producing an mRNA able to be translated
specific proteins assist in nuclear export of mRNA
regulatory proteins can block translation, variable mRNA life-spans
outline why gene expression is tightly regulated
to achieve the right thing at the right time in the right place (this is temporal and spatial control)
housekeeping (commonly used) proteins are continuously produced
protein and mRNA are present in large quantities (e.g. Tubulin)
typically, have longer ‘half-life’ in cells
other proteins are produced in response to stimuli as required
cell signaling (e.g. ligand binding to cell surface receptor, or activiating an intracellular receptor)
signal transduced and may enter nucleus to activate transcription
results in the production of a short-lived protein to carry out the required function
why is control of gene expression important
the side chains (R groups) determine the properties of each amino acid
they collectively determine the final structure and function of the protein
there are twenty standard (coded for) amino acids
amino acid properties
protein sequence (primary structure) is determined by DNA sequence
peptide bonds are covalent bonds between amino acids (relatively strong)
the polypeptide starts to form secondary structures as soon as it leaves the ribosome
describe the primary structure of amino acids
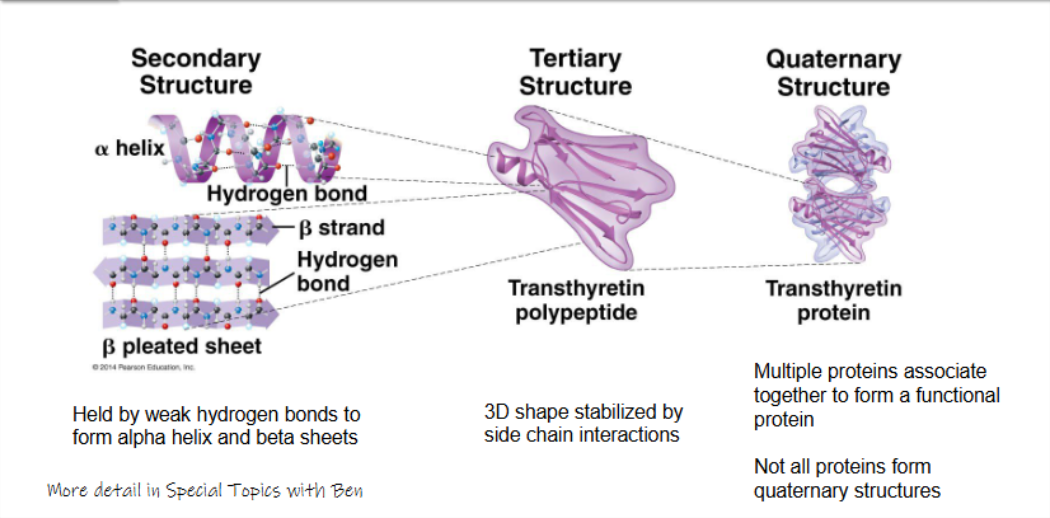
secondary structure:
held by weak hydrogen bonds to form alpha helix and beta sheets
tertiary structure:
3D shape stabilised by side chain interactoins
quaternary structure:
multiple proteins associate together to form a functional protein
not all proteins form quaternary structures
describe the secondary, tertiary, and quaternary structures of amino acids
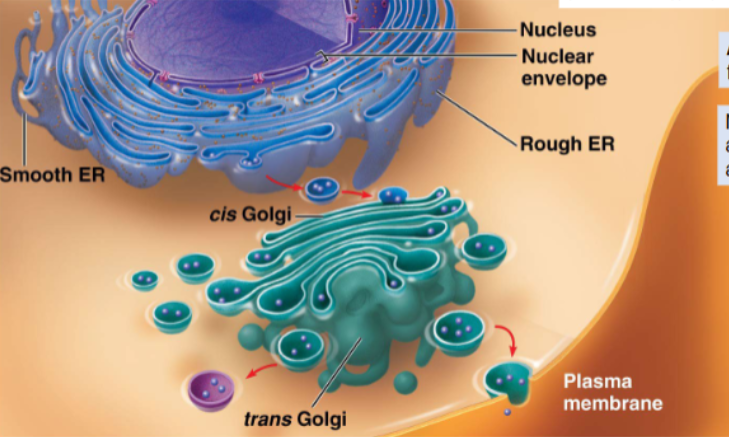
all translation commences on free ribosomes
many proteins are processed and sorted through the RER and Golgi — but not all
proteins destined to function in the cytosol — complete translation on free ribosomes
proteins that go through the endomembrane system — complete translation at fixed ribosomes on the RER
outline protein processing and sorting
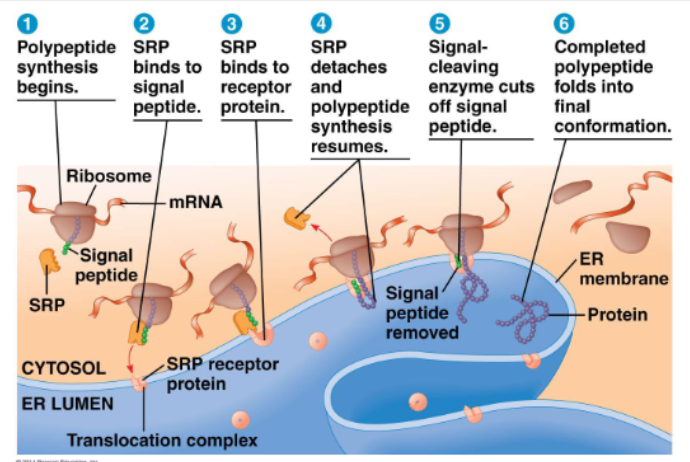
signal peptide:
at N terminus of the protein (~20aa)
SRP: signal recognition particle
polypeptide synthesis begins
SRP binds to signal peptide
SRP binds to receptor protein
SRP detaches and plypeptide synthesis resumes
signal-cleaving enzyme cuts off signal peptide
completed polypeptide folds into final conformation
at step 6:
a secretory protein such as insulin is solubilised in lumen, while a membrane protein remains anchored to the membrane
both then go to the Golgi via vesicles for further maturation
describe how signal peptides direct ribosomes to RER
translation is now complete, but the protein may not yet be functional
common (there are 100s) post translational modifications include:
phosphorylation (addition of a phosphate group)
methylation (addition of a methyl group)
acetylation (addition of an acetyl group)
biotinyation (addition of biotin)
carboxylation (addition of a carboxylic acid group)
carbohydrate addition (particulary for membrane bound proteins, e.g. glycoproteins)
cleavage
ubiquitination
some occur within the Golgi, others in the cytosol
can confer activity — e.g. via phosphorylation or enzyme cleavage
or ability to interact with other molecules — e.g. biotinylation, methylation of histones
or direct to particular locations — e.g. ubiquitination for proteasome degradation
outline post-translational modifications to proteins
human cells are diverse and have different destinies
a cell has three possible destinies:
live and function without dividing
grow and divide
die
various signals tell a cell which path to take
cell diversity and cell destiny
somatic cell division: mitosis — diploid (2n) to diploid (2n)
reproductive cell division: meiosis — diploid (2n) to haploid (1n)
what are the two different types of cell division
G1: growth or gap phsae 1
most cellular activities are occurring here
duration variable — cell type specific
S: synthesis of DNA
DNA replication occurs strands are separated at the hydrogen bonds holding the nucleotides together new strand of DNA is synthesised opposite each of the old strands
G2: growth or gap phase 2
checks for correct DNA synthesis prepares for the mitotic phase (synthesis of the proteins and enzymes required, gathering of reactants)
replication of centrosomes is completed
outline the interphase of the eukaryotic cell cycle
mitotic phase = mitosis plus cytokinesis
prophase:
mitotic spindle forms
two sister chromatids join together at the centromere to form the chromosome
fragments of nuclear envelope, condensed chromosome, and spindle tracks visible
metaphase:
condensed chromosomes aligned
anaphase:
separated chromosomes
telphase and cytokinesis:
nuclear envelope forming
cleavage furrow
describe the mitotic phase
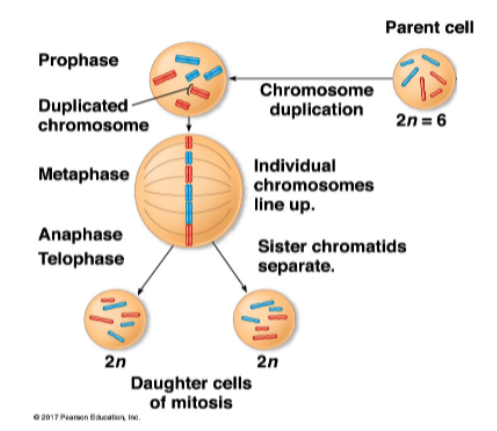
during interphase, DNA replicates
during prophase, DNA condenses
two identical chromatids per chromosome
these are called sister chromatids
during metaphase, chromosomes ‘line’ up
during anaphase, sister chromatids separate before the nuclear envelope refors in telphase
daughter cells are ‘identical’ to parent cell
human diploid cells have 46 chromosomes, 23 from each parent
what is a sister chromatid
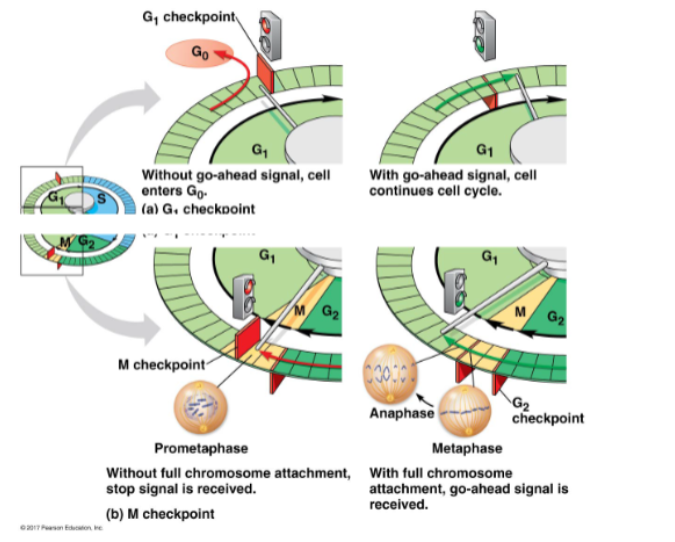
G1 checkpoints:
is the DNA undamaged?
is cell size and nutrition ok?
appropriate signals present?
if not — may exit to G0
M checkpoints:
are all chromosomes attached to spindles
multiple signals required to pass G1 and M checkpoints
occurs in the gonads (ovaries and testes)
produces gametes which are haploid (a single set of 23 chromosomes)
fertilisation then restores the diploid number of chromosomes (2n)
produces cells genetically different from the parent cell
there are two stages of meiosis:
meosis I:
prophase I (synapsis and crossing over, tetrads form)
metaphase I (pairs of homologous chromosomes)
anaphase I (sister chromatids remain attached)
telophase I
meiosis II:
prophase II
metaphase II
anaphase II
telophase II
outline meiosis
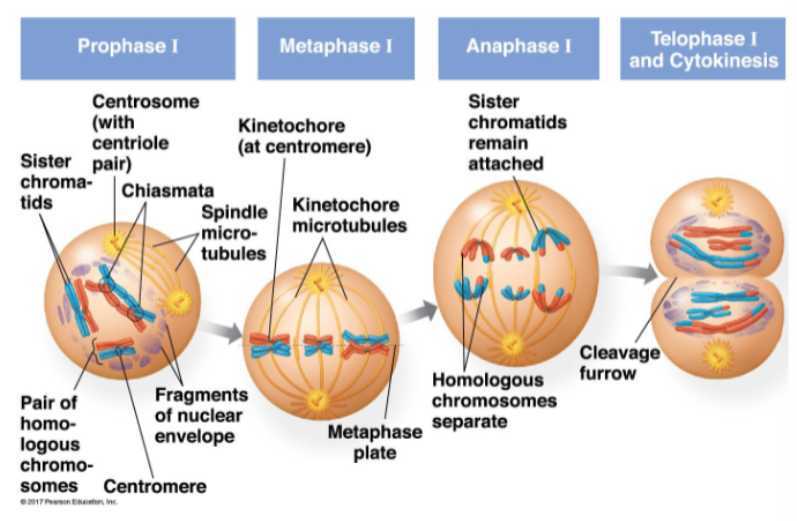
meiosis I separates homologous chromosomes
synapsis: two sister chromatids of each pair of homologous chromosomes pair up
the 4 chromatids are called a tetrad
non-sister chromatids within these tetrads may then cross over: causes recombination
prophase I:
sister chromatids present
centrosome (with centriole pair) present
crossing over occurs at the chiasmata
spindle micro-tubules present
fragments of nuclear envelope visible
pairs of homologous chromosomes present
metaphase I:
kinetochore (at centromere)
kinetochore microtubules
metaphase plate
anaphase I:
sister chromatids remain attached
homologous chromosomes separate
telophase I and cytokinesis:
cleavage furrow
meiosis I — separates homologous chromosomes
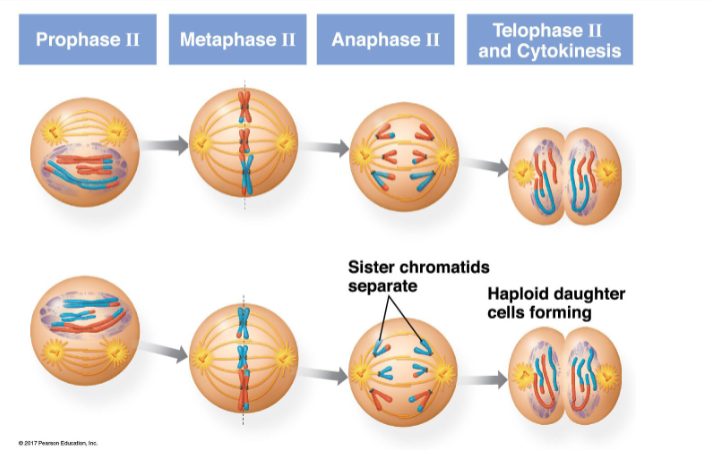
very similar to mitosis, except not preceeded by DNA replication
sister chromatids separate in anaphase II
haploid daughter cells form in telphase II and cytokinesis
meiosis II — separates sister chromatids
mitosis:
prophase:
chromosome duplicated
metaphase:
individual chromosomes line up
anaphase/telophase:
sister chromatids separate
meiosis
prophase I:
crossing over at chiasma
chromosome duplication results in pair of duplicated homologs
metaphase I:
pairs of homologous chromosomes line up
anaphase I/telophase I:
homologs separate
sister chromatids separate
outline the differences between mitosis and meiosis during each of their processes
mitosis
DNA replication: occurs during interphase before mitosis begins
number of divisions: one, including prophase, prometaphase, metaphase, anaphase, and telophase
synapsis of homologous chromosomes: does not occur
number of daughter cells and genetic composition: two, each genetically identical to the parent cell, with the same number of chromosomes
meiosis:
DNA replication: occurs during interphase before meiosis I but not meiosis II
number of divisions: two, each including prophase, metaphase, anaphase, and telophase
synapsis of homologous chromosomes: occurs during prophase I along with crossing over between nonsister chromatids; resulting chiasmata hold pairs together due to sister chromatid cohesion
number of daughter cells and genetic composition: four, each haploid (n); genetically different from the parent cell and from each other
compare the properties of mitosis to meiosis
sources of genetic variation:
independent assortment at metaphase I (2³³ > 8 million possible combinations)
crossing over at prophase I (~1-3 crossover events per pair)
fusion between two gametes (> 233 times 233 combinations)
where do sources of variation occur from
mutations can affect the structure and function of a protein
altered DNA sequence can have major effects on resulting protein function
germ line — passed on to future progeny
local/somatic — during cell division, not whole body — local effects
large scale alterations — chromosomal rearrangements
small scale alterations — one or a few nucleotides altered
small scale mutations can be:
substitutions — where one base is replaced by another - can have minimal or major effect
insertions/deletions — can have major effect if within coding sequence - can cause a frameshift
what is the effect of DNA sequence changes
substitutions can be:
silent
missense
nonsense
insertions or deletions (indels):
cause frameshift if 1 or 2 nt
can maintain frame if 3 nt
outline substitutions and indels in protein coding regions
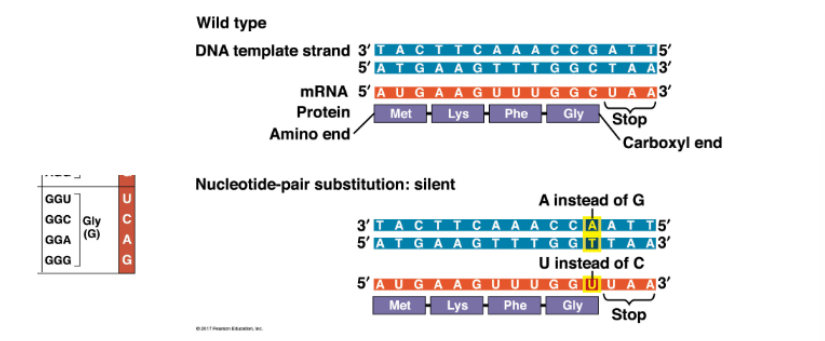
GGC codon becomes GGU codon, but still codes for Glycine — so no effect on protein
name an example of a silent mutation
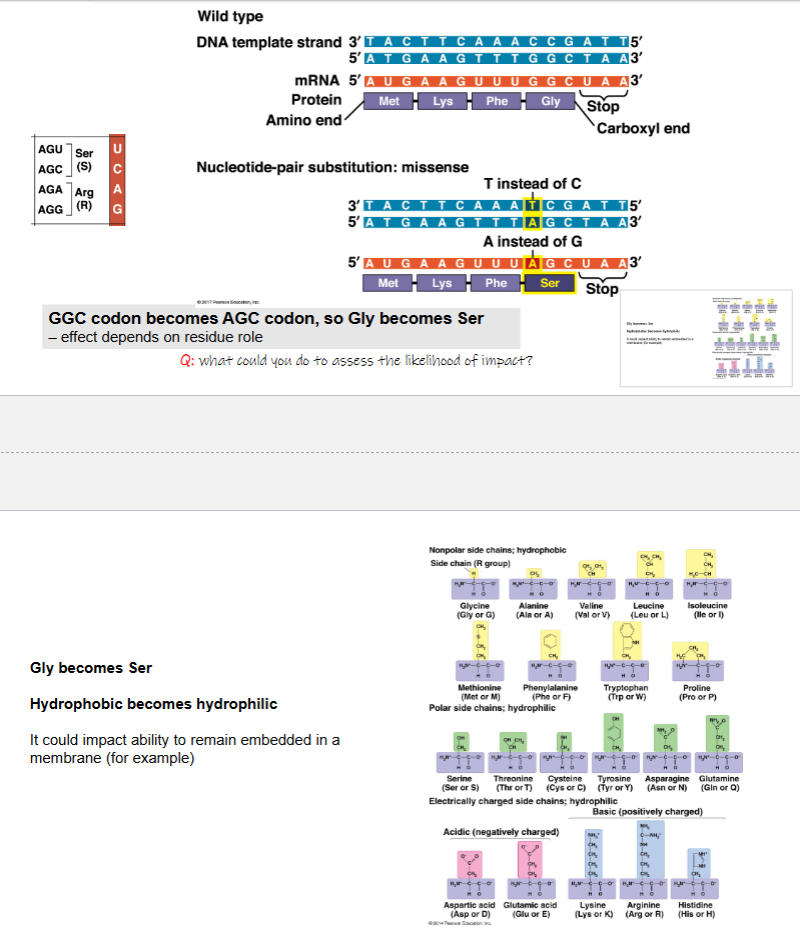
GGC codon becomes AGC codon, so Gly becomes Ser — effect depends on residue role
Gly becomes Ser
hydrophobic becomes hydrophilic
it could impact ability to remain embedded in a membrane
what is an example of a missense mutation
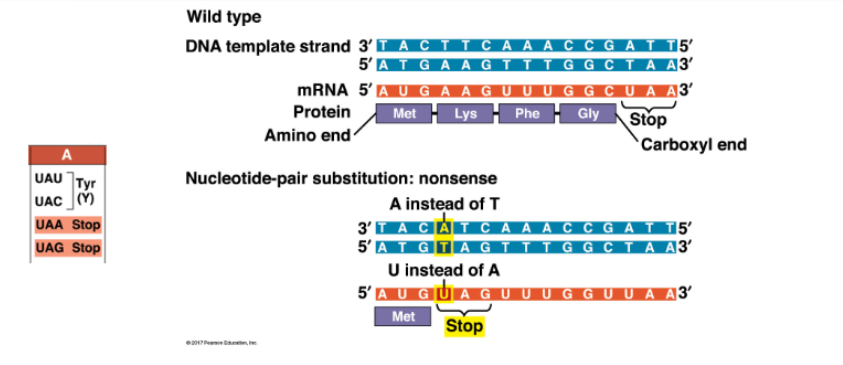
AAG codon becomes UAG codon, so Lys becomes a STOP — truncated protein
what is an example of a nonsense mutation
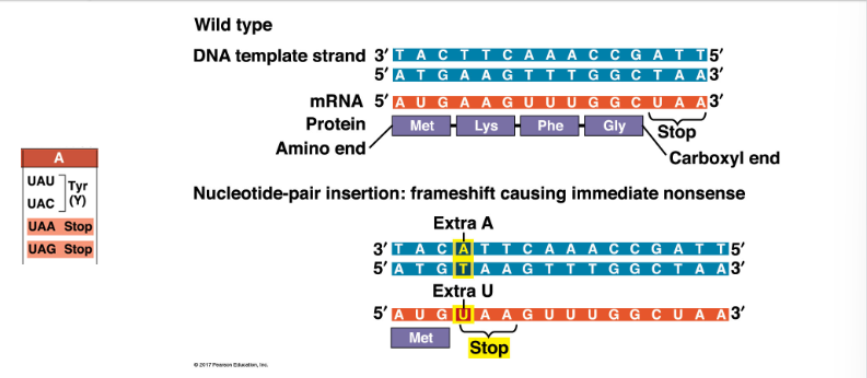
AAG codon becomes UAA codon, so Lys becomes STOP — truncated protein
what is an example of a frameshift mutation via insertion
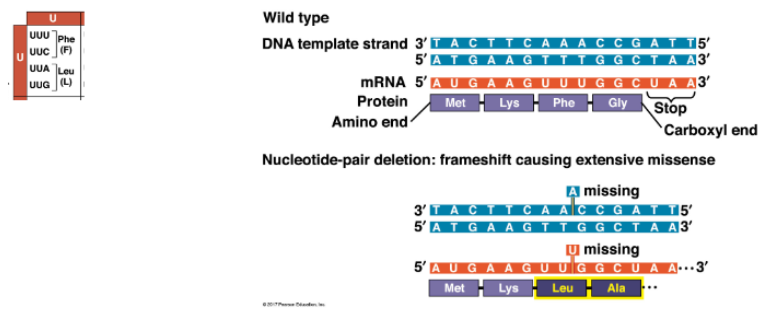
UUU codon becomes UUG, so Phe becomes Leu, plus downstream residues
protein is completely altered from point of frameshift, can have catastrophic effect
what is an example of a frameshift mutation via deletion
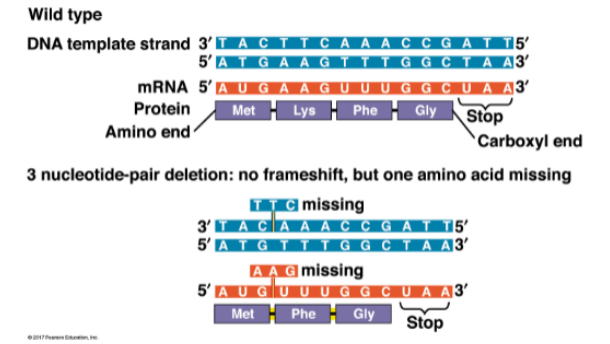
AAG codon is lost (Lys), but downstream residues are intact — frame is maintained
what is an example of a 3 nucleotide-pair mutation