Biology 12 Midterm 2
1/176
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
177 Terms
signal hypothesis
explains how proteins destined for secretion or membrane insertion are targeted to the ER
proof of signal hypothesis
swapping candidate sequences.
taking hypothesized ER signal sequence and removing it from an ER protein caused it to no longer be in the ER.
taking hypothesized ER signal sequence and adding it to a cytosolic protein caused it to be in the ER
necessary and sufficiency
necessary = without x, y won’t occur
sufficient = with only x, y would occur
an ER signal sequence is considered necessary and sufficient if only having that sequence causes the protein to localize to the ER and not having it causes the protein not to localize in the ER.
where is the ER signal sequence located
N-terminus/beginning of a newly synthesized polypeptide
nucleus
where DNA is stored in a cell, protects DNA
nuclear envelope
inner and outer parts of the nuclear membrane
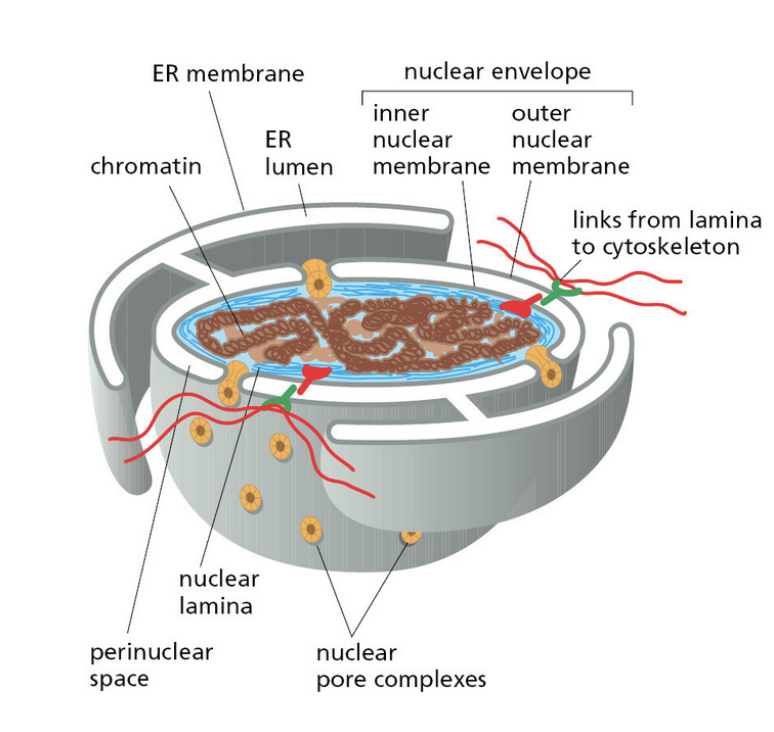
nuclear pores
the direct entry and exit out of the nucleus
cage like structure
what can freely enter nuclear pores
ions, small metabolites, and proteins up to 40kDa can diffuse freely through the nclear pore
nucleoporins
proteins that interact with one another through FG repeats, creating a gel-like mesh that acts like a filter/sieve inside the nucleopore.
what do nucleoporins prevent
nucleoporins prevent large molecules from freely diffusing (>40kDa)
Nuclear localization signal (NLS)
NLS is an amino acid sequence (either linear or 3D) that is necessary and sufficient for bringing a protein into the nucleus
removing NLS prevents protein from localizing to the nucleus (necessary)
adding NLS to a cytosolic protein causes it to localize to the nucleus (sufficient)
cargo
any molecule or particle that is actively transported from one location to another
importin
import receptor protein that binds to cargo with NLS and move it through the nuclear pore
contains low affinity binding sites for FG repeats in the nucleoporin
loosely associate with gel like mesh/sieve, aided by diffusion to get itself and cargo through the nuclear pore
G-protein
act like molecular switches
enzymes that can cleave/hydrolyze GTP (GTP → GDP)
G-proteins have GTPase activity that hydrolyzes itself by removing/cutting its phosphae group, leading to a conformation change in the G-protein (different conformation depending on if it is GTP or GDP bound)
can function on its own but usually get help by interacting with GEFs and GAPs
GEFs and GAPs
proteins that accelerate the conformation change of G-proteins
GTPase Activating Protein (GAP)
activates GTPase activity of G-protein which promotes GTP to hydrolysis to GDP
speeds up/facilitates the transition of a G-protein from GTP bound on state to GDP bound off state
Guanine exchange factor (GEP)
exchanges GDP for GTP
promotes onset of G-protein’s reactivity. G-protein has an intrinsic rate at which it replaces GDP with GTP but GEP accelerates this intrinsic rate
Ran (G-protein)
has weak GTPase activity (does not hydrolyze GTP to GDP quickly)
aided by GEFs and GAPs depending on where it is
Where is Ran GAP
Ran GAP is higher in concentration in the cytosol
Where is Ran GEF
Ran GEF is higher in concentration in the nucleus
Where does Ran encounter Ran GAP? What happens?
Ran encounters Ran GAP in the cytosol → Ran GDP (off)
Where does Ran encounter Ran GEF? What happens?
Ran encounters Ran GEP in the nucleus → Ran GTP (on)
What arrises from the fixed localization of Ran GAP and GEF?
concentration gradient of Ran GDP and GTP arise from the fixed localization of Ran GAP and GEF
properties of importin that allows nuclear import to happen
importin has high affinity for Ran GTP and low affinity for Ran GDP
importin releases its cargo when bound to Ran GTP
Process of Ran/importin for nuclear import
in the cytosol, importin binds to cargo with NLS
importin moves the cargo into the nucleus due to its low affinity binding sites which allow it to interact with the FG repeats in the nucleoporin
in the nucleus, importin binds to Ran GTP (nucleus has more Ran GEF)
importin binding to Ran GTP causes the release of cargo
importin Ran GTP exits the nucleus due to the conc gradient (less importin Ran GTP in the cytosol)
importin Ran GTP encounters Ran GAP in the cytosol (cytosol has more Ran GAP)
importin Ran GTP becomes importan Ran GAP
importin has low affinity to Ran GAP, importin dissociates from Ran, ready to repeat the cycle with another cargo + NLS
Process of Ran/exportin for nuclear export
in the nucleus, exportin binds to Ran GTP (nucleus has more Ran GEF)
Ran GTP promotes exportin to bind to cargo
exportin moves the cargo into the cytosol due to its low affinity binding sites which allow it to interact with the FG repeats in the nucleoporin and due to the conc gradient of Ran GTP
in the cytosol, exportin Ran GTP encounters Ran GAP in the cytosol (cytosol has more Ran GAP)
exportin Ran GTP becomes exportin Ran GDP
exportin has low affinity to Ran GDP, exportin dissociates from Ran and cargo, cargo released in the cytoplasm
Free exportin has NLS-like or import signals that allow it to interact with the FG repeats of the nucleoporin to get back into the nucleus
how do proteins reach the membrane?
rough ER → golgi complex → transgolgi network (directs protein to where they need to go)
pulse chase experiment
pulse: incubate cells with radioactive amino acids for short period (~3 minutes)
newly synthesized AAs will be radioactive
radioactive AAs were observed in the RER
chase: follow up with incubation with non-radioactive AA. “Chase”/locate radioactive AA from before
after 20 min, radioactive AA is in the golgi complex and secretory granules
after 120 minutes, radioactive AA is in secretory granules, some being secreted
suggests that secreted proteins follow trajectory of RER → golgi complex → secretion at plasma
places that the transgolgi network sorts proteins (4)
Plasma membrane - pathway for membrane proteins and constitutive secretion
Lysosomes - Mannose-6-phosphate tags on hydrolases recognized by M6P receptor
Secretory vesicles - Regulated exocytosis, only released upon a signal (e.x., neurotransmitters, hormones)
early/late endosomes - some cargo is redirected here
types of ribosomes
free ribosomes: floating in the cytosol, not attached to the rough ER (cytosol is the destination of produced protein)
ribosomes located in the RER (usually enters the ER lumen for secretion or embedded in the ER membrane)
cell free systems
enable studies of protein trafficking. Use differential centrifugation to separate microsomes and then use equilibrium density gradient centrifugation to separate rough and smooth ER as they have different densities
harness the fact that proteins can still be translated on the rough ER microsomes on a test tube
microsomes
small vesicles assembled from ERs
ER signal sequence of emerging polypeptides experiment
if proteins with ER signal sequence is synthesized without microsomes, proteins are not incorporated into microsomes if microsomes are added after synthesis (adding protease still destroys the proteins)
if proteins with ER signal sequence is synthesized with microsomes, they are incorporated during synthesis and are protected from protease
thus, timing of microsome presence matters in order to confer some kind of protection from the protease
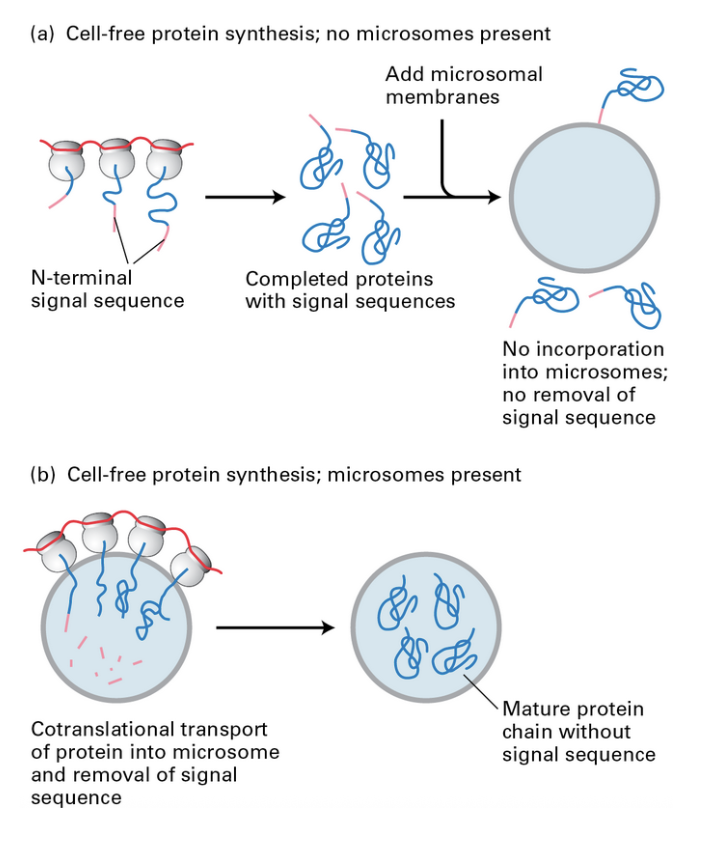
experiment with or without ER signal sequence (+ protease)
in the presence of microsomes during synthesis
proteins synthesized without ER signal sequence were degraded when protease was added (suggests they were not incorporated into microsome due to lack of ER signal sequence)
were then degraded when detergent was added, proving that it was the microsomes initially preventing the degradation
proteins synthesized with ER signal sequence were NOT degraded when protease was added (suggests they were incorporated into microsome due to presence of ER signal sequence)
proves necessary and sufficiency again
what does co-translational translocation do
directs proteins (such as the integral membrane proteins) into the RER
key players of co-translational translocation
ER signal sequence
G-proteins SRP and SRP receptor
SRP: signal recognition particle, recognizes the signal sequence
SRP receptor: binds with SRP to receive its signal
process of co-translational translocation
Ribosome translates the ER signal sequence
SRP recognizes and binds to the ER signal sequence along with the ribosome it is emerging from
this binding causes a conformational change in the ribosome which pauses the translation
this pausing of translation ensures that the protein is translated into the ER lumen and not the cytosol
the whole SRP ribosome complex binds to the SRP receptor on the ER membrane
translocon (similar to a protein channel) is usually closed but its association with SRP, SRP receptor, and ribosome contributes to its opening
SRP and SRP receptors are found in its GTP bound state
SRP binding to SRP receptor stimulates the GTPase of the SRP and SRP receptor G-proteins which hydrolyzes GTP to GDP
hydrolysis of SRP and SRP receptor causes a conformational change that causes SRP to dissociate from the ribosome and SRP receptor, allowing translation to continue
also causes ribosome to lock into place on the open translocon
unpaused translation now translates the polypeptide chain into the ER since the translocon is now open
signal peptidase snips the signal sequence as the sequence has already served its purpose helping with translocation
disassociated complex allows SRP to keep doing this with other ribosomes as they pickup a GTP
two ways integral membrane proteins go to the ER membrane
N-terminal ER signal sequence
Signal anchor sequences (SA) with positively charged amino acids on either side to help orient
Stop-transfer anchor (STA) sequence
brings polypeptide from ER lumen to cytosol
first transmembrane domain of a protein with a N-terminal ER signal sequence
causes the opening of a lateral gate in the translocon and then makes its way to the side
Signal anchor (SA) sequence
acts as an N-terminal sequence, brings polypeptide from cytosol to ER lumen
similar to the STA, SA transfers itself laterally into the membrane through the lateral gate of the translocon
orientation/topology of the polypeptide is driven by chain of positively charged amino acids
due to leaflet asymmetry, which side of the membrane is more positive?
The cytosolic leaflet is more negative
The lumenal leaflet is more positive
if sequence of positive AA is on the N-terminus of the polypeptide, N terminus will be in the cytosol sine the cytosol is more negative
is the integral membrane domain orientation maintained throughout the secretory pathway?
yes, the topology of the integral membrane domain remains consistent throughout the entire secretory pathway
exoplasmic
outside of the cell or luminal (interior of a membrane bound organelle)
NEVER cytosolic
cytosolic
always facing the cytosol, never the outside of the cell or the lumen/interior of a membrane bound organelle
what does the rough ER do for protein folding?
serves as a site of quality control of protein folding and also adds post-translational modifications
what are the 5 principle modifications that newly synthesized polypeptides in the membrane and lumen of the ER undergo?
disulfide bonds
proper folding
addition and processing of carbohydrates
specific proteolytic cleavages
assembly into multimeric proteins
how strict is the quality control process of the ER
incredibly strict, ensuring that only correctly folded modified proteins move to the next step in the secretory pathway.
failure of quality control leads to proteins that cause neurodegenerative diseases
cells with an accumulation of misfolded proteins attempt to fold itself through an unfolding protein response or will literally trigger apoptosis if unfixable
disulfide isomerases (PDI)
PDIs are like a disulfide bond partner. It forms and rearranges disulfide bonds within a protein when its folding in the ER lumen
through redox reactions, PDIs exchange disulfide bonds to ensure protein is connecting with itself in the right way and at the right time
PDIs can also correct the incorrect disulfide bonds
sugar modification/glycosylation
contribute to protein folding, stability, trafficking, and interactions
two types of sugar modifications/glycosylation
N-linked (ER)
O-linked (golgi)
N-linked glycosolation
happens in the ER and helps with protein folding
more common
sugar covalently links to the N of asparagine of R-group
O-linked glycosolation
occurs as more refined processing in the golgi
sugar covalently linked to O of serine/threonine of R-group
oligosaccharyl transferase
catalyzes the transfer of sugar to protein from a large precursor (dolichol) onto the asparagine of polypeptide emerging from translocon (for N-linked glycosylation)
how does glucose subunits of N-linked sugars allow for quality control in the ER?
unfolded polypeptide chain enters the ER
oligosaccharyl transferase performs N-linked glycosolation to the polypeptide
glucose chain has 3 glucoses, 2 are trimmed so 1 remains
polypeptide with 1 glucose associates with calnexin (chaperone protein, helps polypeptide achieve their 3D structure) which helps the protein fold
glucosidase cleaves the final glucose (prevents reattaching to calnexin)
no glucose left causes the protein to exit the ER as it is done folding
what happens if a protein is not done folding after entering the ER?
if a protein is not done with folding/if anything looks off, glucosyl transferase detects the misfolds and reattaches a signal glucose
this allows the polypeptide to associate with calnexin again to repeat the folding process
there is an extremely high concentration of glucosyl transferase present in the ER to ensure that misfolded proteins do not evade the cycle and leave
what does glucosyl transferases detect
they detect misfolded proteins by detecting exposed hydrophobic patches that are supposed to be folded inside (since they are hydrophobic)
manose molecules of N-linked sugars
serves as timers for protein folding
mannose subunit encounters mannosidase which trims off the manose subunits
once chewed up, it serves as a signal that the protein is not properly folded and should be exported for degredation
the longer the protein is in the ER lumen, the more likely it is to encounter a mannosidase (proteins that have been around for a long time but still not properly folded/still has a glucose attached are more likely to encounter a mannosidase just because they’ve been around for longer)
unfolded protein response (UPR) — 4 ways
triggered by an accumulation of misfolded proteins
increase protein folding capacity through translational upregulation of ER chaperones (ex. produce more calnexins so there are more subunits facilitating protein folding)
reducing translation
degradation of misfolded proteins
programmed cell death as a last resort
accumulation of misfolded proteins contribute to various diseases, including neurodegeneration
golgi complex
a membrane-bound eukaryotic organelle that acts as the cell's post office, responsible for modifying, sorting, and packaging proteins and lipids for secretion or delivery to other organelles
golgi complex structure
stacked membrane structure consisting of cisternaes
cisternae
each individual stack of te golgi complex
cis cisternae
cisterna in the golgi nearest to the ER
closest to the nuclear envelope which is contiguous with the RER
trans cisternae
cisternae furthest from the ER
medial cisternae
cisterna between the cis and trans cisternae
order of the cisternae as you progress from ER to plasma membrane
cis → medial → trans cisternae
golgi compartment protein composition
different cisternae have their own protein composition/identities which causes it to do different things
this is due to specific enzymes that localize to different regions of the golgi complex
ex. cis and medial cisternae have different sets of enzymes which help process the protein in different ways
modification to n-linked oligosaccharide in each golgi compartment
as the oligosaccharide sugar tree makes its way from the cis to trans golgi, it encounters different enzymes. The sugar tree changes as it goes from the cis to trans golgi as it encounters different enzymes
two models of how cargo proteins move through the golgi
vesicular transport model
cisternal maturation model
vesticular transport model
cargo gets shuttled from the CGN to the TGN in vesicles. Cargo is what is present in the vesicles
cisternal maturation model
cisternae matures from cis to trans. Vesicles from trans cisternae carrying golgi resident proteins move retrograde and fuse with cis cisternaes. Cis cisternae becomes medial cisternae as the enzymes in the medial cisternae transport in vesicles from the medial to the cis cisternae
which model of cargo moving through the golgi is most supported
the field largely believes in the cisternal maturation model although cells technically performs both.
evidence for the cisternal maturation model
Staining secreted cargo proteins show that they are localized in the cisternae stacks, not in the vesicles
however, staining golgi processing enzymes show that they can sometimes be found in a vesicle, suggesting that it is the localized enzymes themselves that move between the cisternaes
how do proteins move from the ER to the golgi?
make vesicles
sort proteins
put correct proteins in the vesicle
ensure vesicles target and fuse with the right target membrane
vesicles from ER must fuse to the golgi in the right place in the right way
yeast analysis revealed major steps in the secretory pathway
yeast allowed for a genetic approach to studying secretory pathways. Results of mutations allowed scientists to deduce what each gene was in charge of in the secretory pathway
why did scientists choose to work with yeast?
easy to perform large scale genetic screens to isolate temperature sensitive mutations
proteins functional at permissible temps
proteins non-functional at restrictive temps
yeast has essential cellular processes shared across the tree of life
yeast experiment to understand secretory pathways
mutated a lot of yeast randomly
identified mutants where secretory pathways were damaged
cataloged set of mutants that affected secretory pathways
all the mutants they identified in yeast were nearly identical to human mutants
Coat proteins (COPs)
specialized proteins that assemble on membrane surfaces
COPs bend the membrane to form vesicle spheres
COPs simultaneously have mechanisms to sort content within vesicles
directionality of vesicular traffic
anterograde: ER to golgi (nucleus to plasma membrane to extracellular)
retrograde: golgi to ER (plasma membrane to nucleus)
what is transported anterograde
secreted protein, membrane protein, golgi proteins
what is transported retrograde
enzymes for cisternal maturation that
proteins that contribute to the maintenance of compartment identity
ex. ER resident protein could leak out and make its way to the golgi. Retrograde transport moves these leaked proteins back
KDEL sequence
sequence located on the c-terminus of ER resident proteins that gets recognized by the KDEL receptor of the COPI assembly
allows COPI assembly to bring ER resident proteins back to the ER
difference in pH between the lumen of the golgi and ER causes KDEL receptor to associate/dissociate, dropping off the proteins
cargo receptor
a type of protein that has some sort of recognition site for the appropriate cargo
ex. cargo/protein meant to leave the ER will have an ER exit signal that enables it to associate/interact with a specific cargo receptor
this is how sorting of the right cargo for the vesicle to contain happens
COPII coated vesicles
moves vesicles anterograde (ER to golgi)
COPI coated vesicles
moves vesicles retrograde (golgi to ER)
important for cisternal maturation
COPII coat protein
drives the assembly of various proteins, including cargo receptor
how do COPII coated vesicles move things from ER to golgi?
G-protein Sar1 hangs out in the cytoplasm in a GDP bound state
Sar1 GDP encounters GEF in the ER membrane which turns it into Sar1GTP
Sar1 GTP kicks out an amphipathic helix which looks like a transmembrane domain
this amphipathic helix allows Sar1 GTP to insert itself into the ER membrane
insertion of Sar1 GTP causes membrane curvature, enhanced by association with other coat associated proteins
Sar1 GTP in the membrane kickstarts a multimeric assembly
Sar1 GTP recruits proteins for COPII coat
this includes recruiting cargo receptor which sorts cargo proteins
after GTP hydrolysis, Sar1 GTP becomes Sar1 GDP which dissociates from the vesciel and causes the loss of the COPII coat → allows vesicle to fuse to the golgi as vesicle doesn’t stay fixed as a vesicle
how do COPI coated vesicles move things from golgi to ER?
similar to the COPII coating process, except uses Arf1 instead of Sar1
G-protein Arf1 hangs out in the cytoplasm in a GDP bound state
Arf1 GDP encounters GEF in the golgi membrane which turns it into Arf1 GTP
Arf1 GTP kicks out an amphipathic helix which looks like a transmembrane domain
this amphipathic helix allows Arf1 GTP to insert itself into the golgi membrane
insertion of Arf1 GTP causes membrane curvature, enhanced by association with other coat associated proteins
Arf1 GTP in the membrane kickstarts a multimeric assembly
Arf1 GTP recruits proteins for COPI coat
this includes recruiting KDEL receptor which binds to proteins with KDEL sequences
after GTP hydrolysis, Arf1 GTP becomes Arf1 GDP which dissociates from the vesicle and causes the loss of the COPI coat → allows vesicle to fuse to the ER as vesicle doesn’t stay fixed as a vesicle
trans-golgi network
acts as a sorting station that determines where proteins go
places that trans-golgi network sorts proteins
secretion
target to other sites in the cell that’s not the plasma membrane (ex. endosome/lysosome)
clathrin
a coat complex
clathrin-coated vesicles help mediate transport from TGN to endosome/lysosome
also helps move things from plasma membrane into endosome via endocytosis
clathrin assembly
clathrin proteins self-assemble into a triskelion shape (cage/basket like structure which creates the vesicle shape)
adaptor proteins
connects triskelions with each other and with the right membrane compartments + cargo receptors
Adaptor protein 2 (AP2) associates clathrin triskelions with appropriate coat assembly/cargo sorting factors.
dynamin
a g-protein involved in endocytosis (bringing in something from the membrane back to the cell)
how does dynamin facilitate endocytosis?
when dynamin is in its GTP bound state, it assembles at the neck between the vesicle and plasma membrane
it uses GTP hydrolysis to mechanically constrict the neck, ultimately leading to pinching of the neck/excision of the vesicle
what happens if dynamin has a non-hydrolzyable GTP
if dynamin is bound to a non-hydrolyzable GTP, dynamin gets locked.
this forms an elongated neck as dynamin continues to assemble but cannot hydrolyze GTP for the excision
lysosomal proteins
proteins destined to head to the lysosome
how do lysosomal proteins end up in the lysosome?
the sugars of lysosomal proteins/proteins destined for the lysosome are modified to receive a mannose 6 phosphate (M6P) in the cis-golgi
special M6P receptor in the TGN recognizes and binds to the M6P modification
adaptor protein connects the M6P receptor with the Arf1 GTP complex
once associated, the adaptor helps to recruit the clathrin coat
difference in pH alters Arf1 GTP association with the M6P receptor, causing the release of protein
Rab
a g-protein that coordinates the fate and trajectory of proteins by identity marking the transport vesicles
how does the Rab g-protein work?
Rab in GTP bound state helps recruit a series of proteins that helps specify what is in the vesicles and where they need to go
ex. helps to organize specific tethering complexes/proteins that helps proteins make contact with the correct target membrane
v-SNARE
vesicle SNARE
t-SNARE
target membrane SNARE