Molecular Biophysics
1/152
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
153 Terms
What will happen to the incident and transmitted ray if there are a lot of cells?
The incident beam will scatter, and the transmitted beam is less intense.
Can use this to measure cell density
What is diffraction?
When the incident ray is scattered at an angle
How does fluorescence occur?
The incident ray promotes an electron in the sample from a ground state to an excited state
When the electron falls back down to its ground state fluorescence occurs
This is an absorbance method
What is the Boltzmann distribution equation
Useful for NMR
u = upper energy state
l = lower energy state
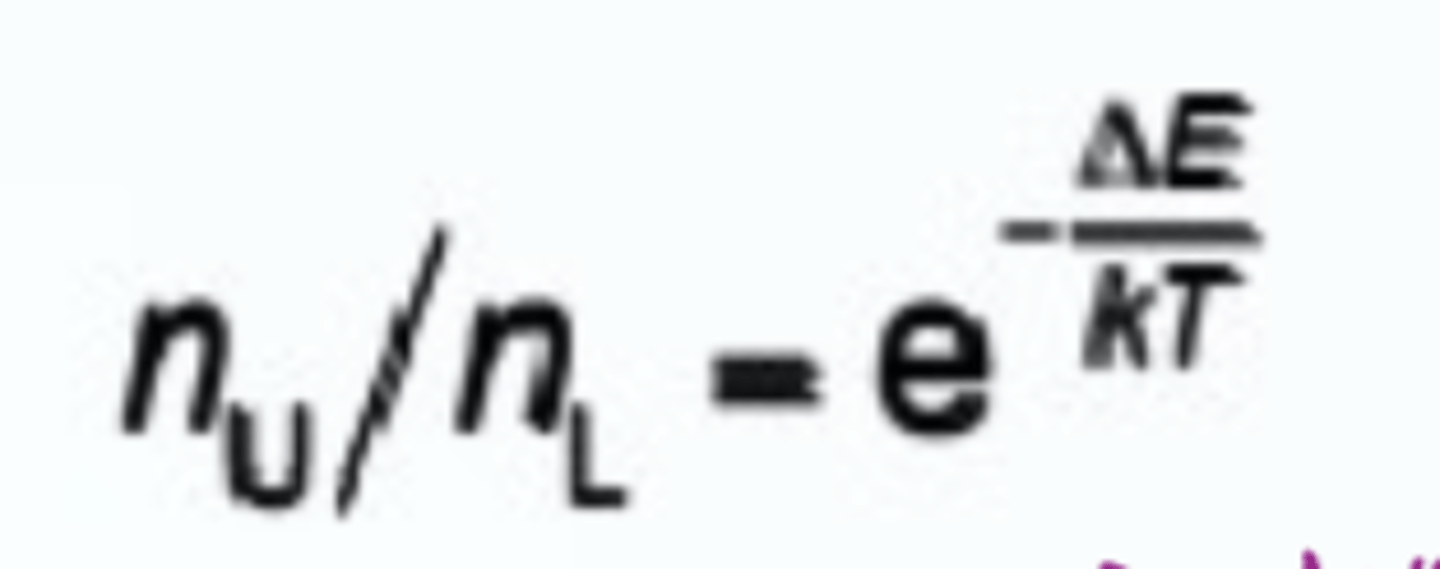
What is the nuclear overhauser effect equation
Useful for NMR
measures distances which can determine structures
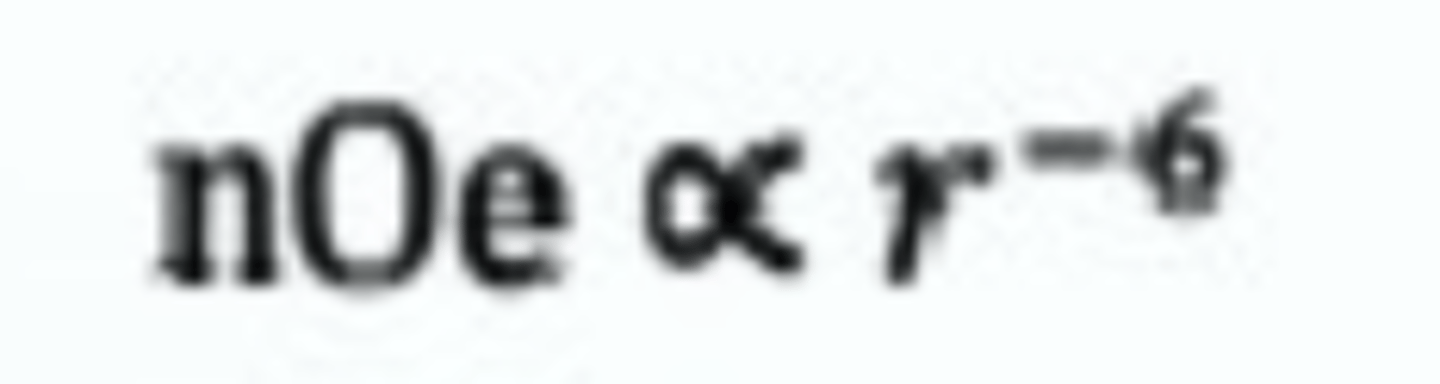
What is the Beer-Lambert law
A = absorbance
c = concentration
l = path length
epsilon = extinction coefficient

What is the Bragg equation

What is the equation for the association constant (Ka) involving concentrations
Where H is the host
G is the guest
and binding occurs with a 1:1 stoichometry

What is the equation for the association constant (Ka) involving rates
Where k on is the rate of binding
k off is the rate of dissociation

How does Ka relate to Kd
Kd is the inverse of Ka
If Ka = 1000
Kd = 1/1000
What is the equation for Gibbs free energy relating to Ka

What is the equation for Gibbs free energy from the 2nd law of thermodynamics

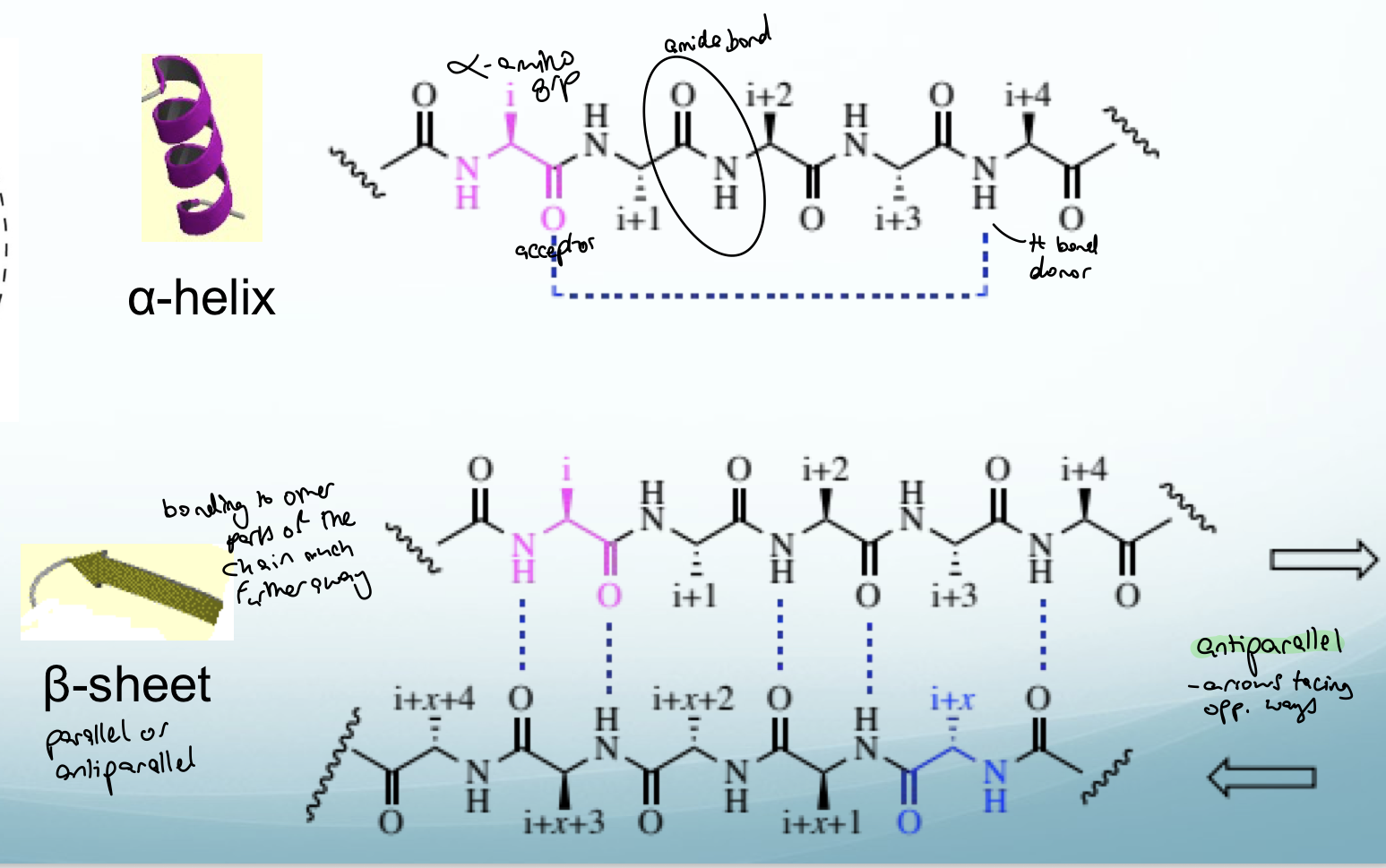
Describe the structure of proteins
1º structure: AA sequence - AAs connected by amide bonds (HN-C=O). C=O can form hydrogen bonds with NH on a different part of a chain. If the different parts of the same chain are close together then a condensed helix occurs (alpha helix). If the different parts of the same chain forming H bonds are far apart then an antiparallel structure forms (beta sheet)
2º structure: alpha helix or beta-sheet
3º structure: secondary structure undergoes quick folding to form the tertiary structure. This folding is important for the protein's function
4º structure: multiple tertiary structures fits together
What is the native protein state
Form where all secondary and tertiary structures are in place
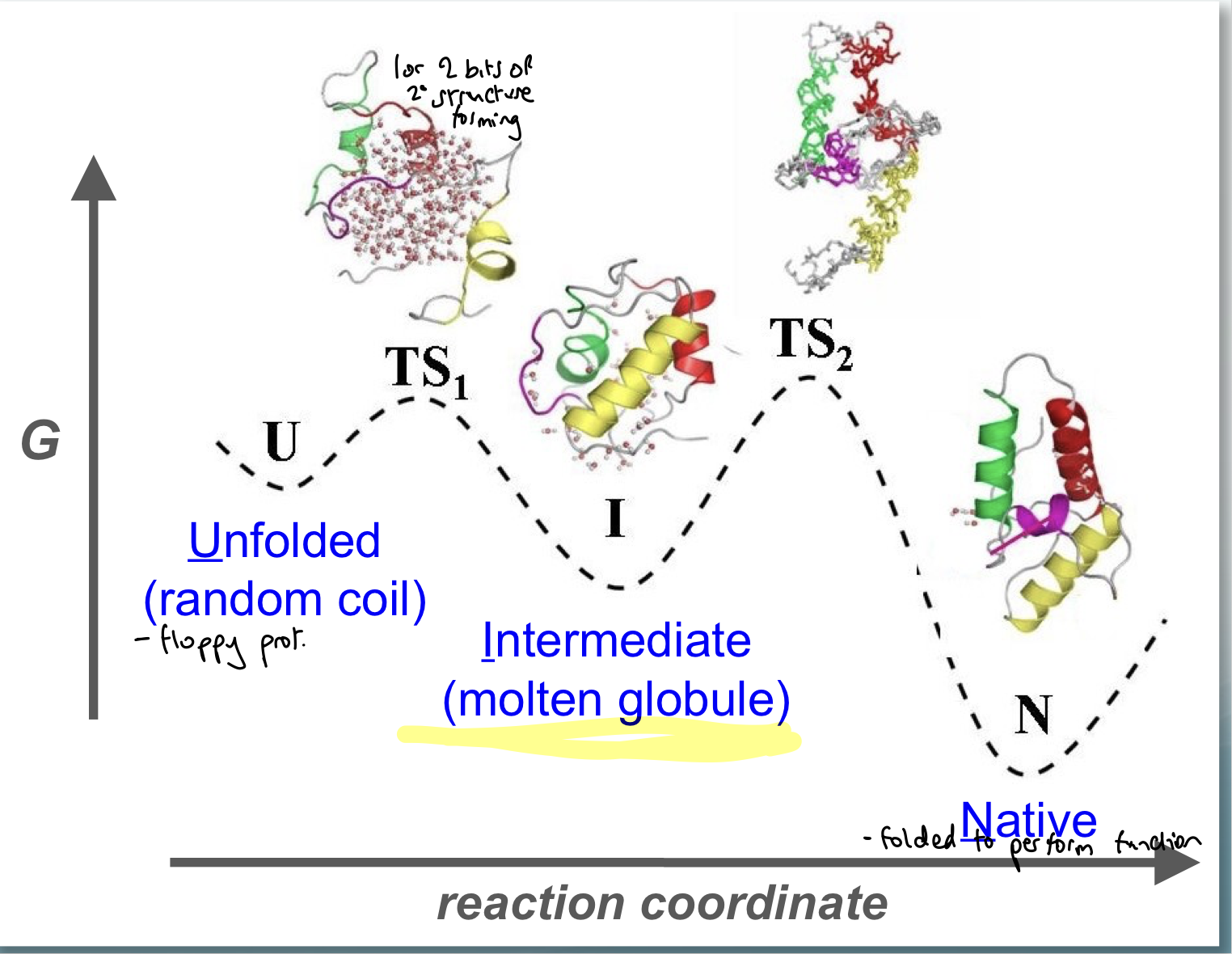
What underpins protein folding? Show E level diagram
Thermodynamics controls the conformation and stability of the native state
Kinetics determines how fast do proteins fold and unfold
Directed process:
unfolded is a random coil, which is floppy and has little secondary structure
intermediate (molten globule): can measure the rate of formation of this and work out the height of the energy barrier
native state
via 2 transition states
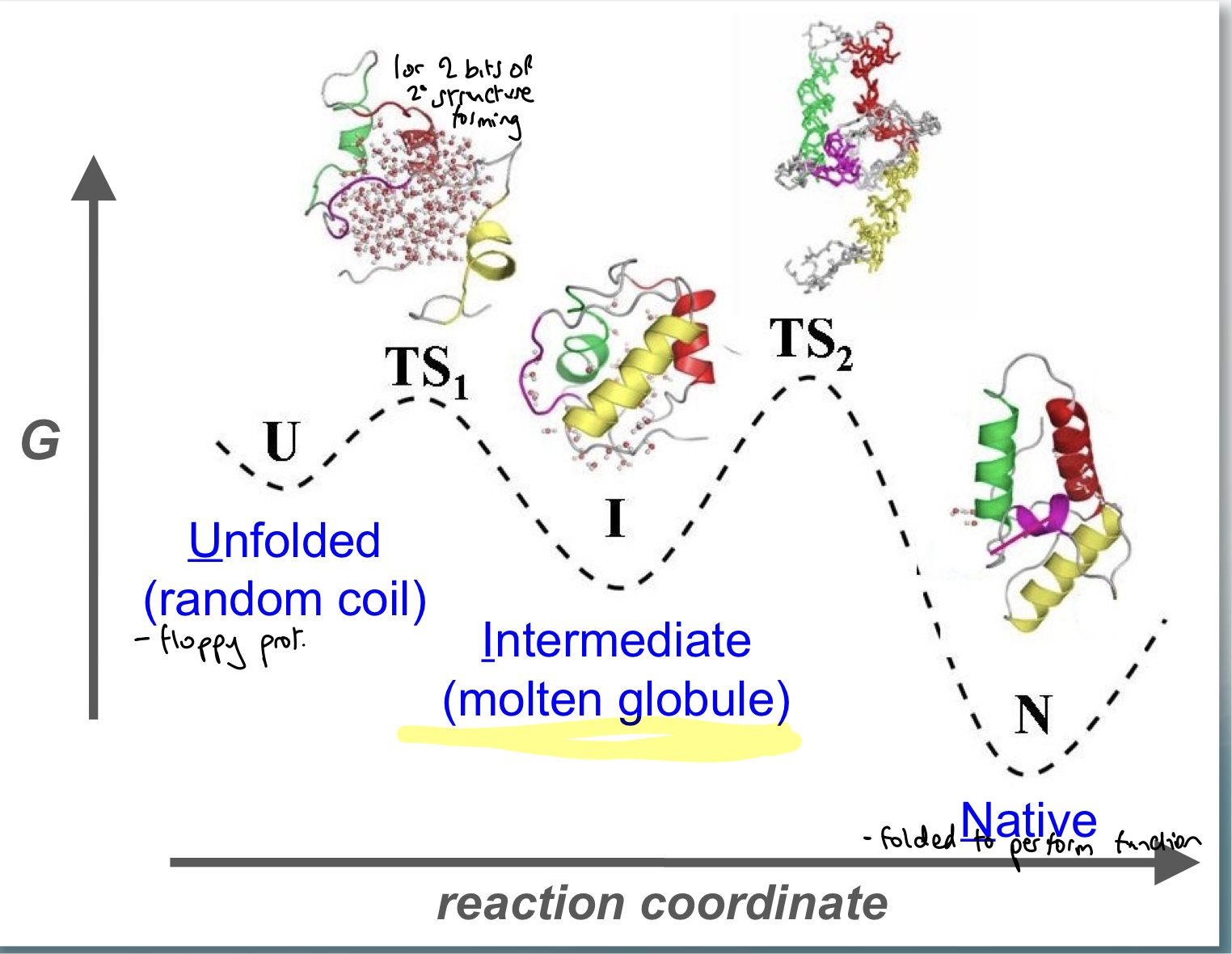
What can we use the molten globule form?
can measure the rate of formation of this and work out the height of the energy barrier
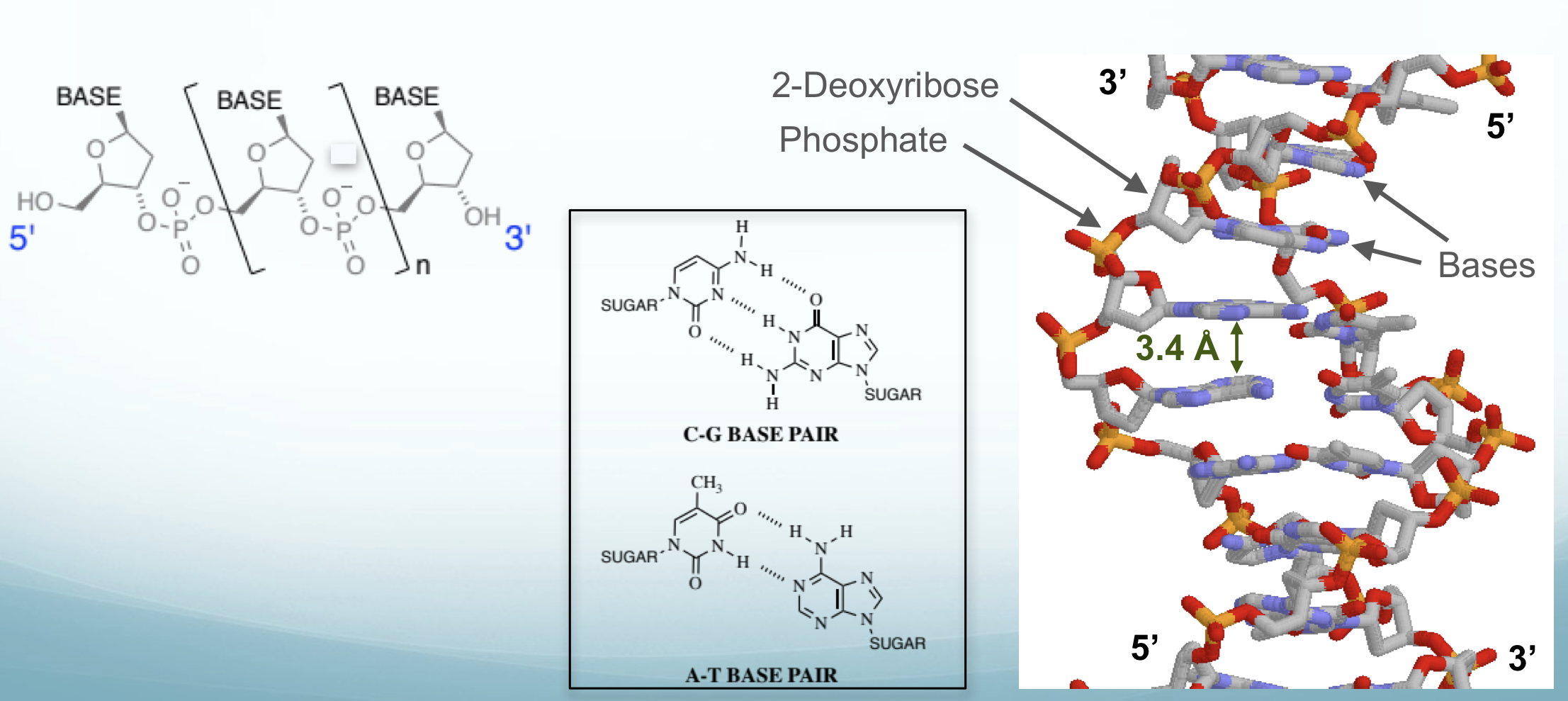
Describe the structure of DNA
Phosphate - Ribose Sugar - Base
Base could be A-T (2 H bonds), C-G (3 H bonds)
Helical polymer which forms a column of bases stacked upon each other (3.4 Angstroms apart)
different structures of DNA exist: left handed vs right handed (be aware)
Describe the structure of lipids
Glycerol molecule + a fatty acid tail
glycerol is the head and hydrophilic (polar group)
Fatty acid tails are long hydrocarbon chains and are hydrophobic
Self assemble into bilayers due to these properties (cell membrane)
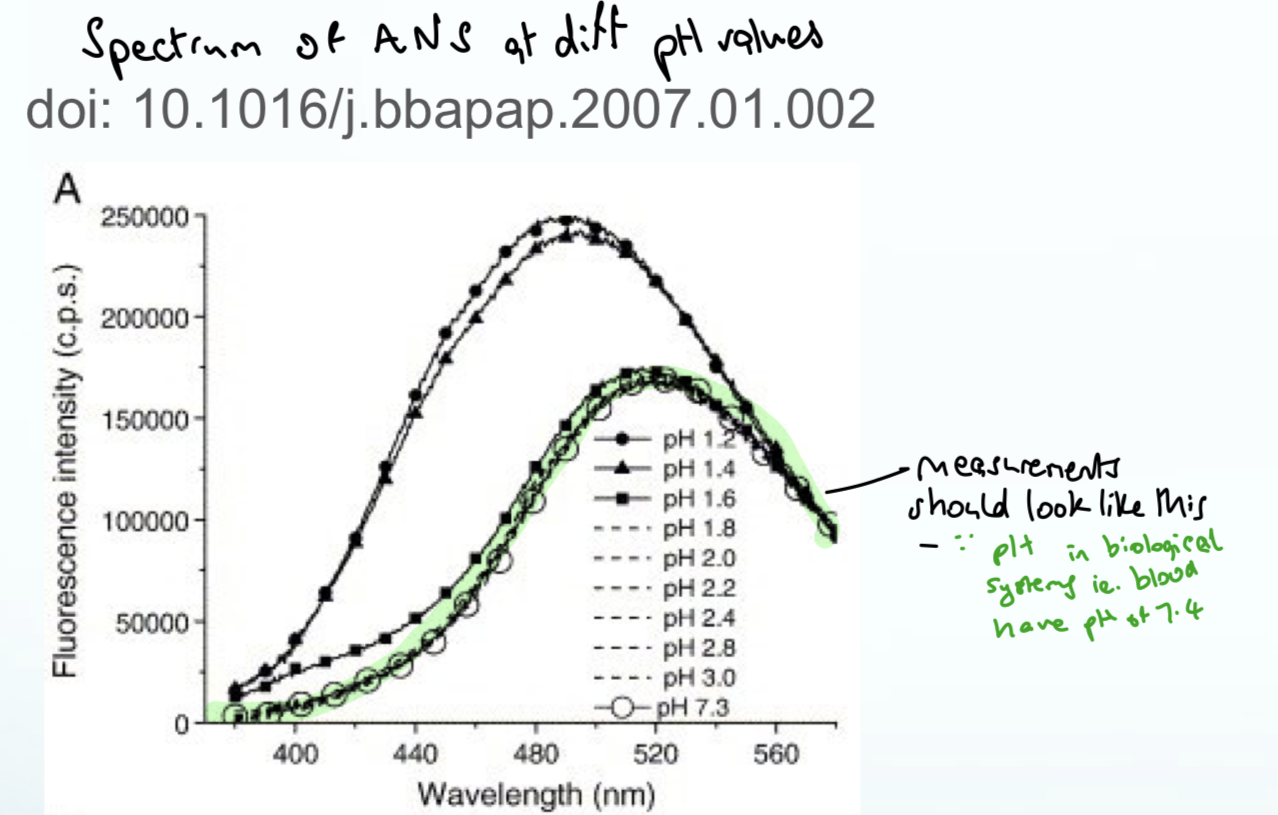
What is common between Trp, Tyr, Phe, His and describe what affects their absorbance
All are aromatics and have absorbances below 300
Aromatics absorb very well and peptide bonds can also absorb
Peptide bonds have low absorbances but because there are so many in a protein molecule, they dominate
With a protein or DNA it is common to see absorbances below 300
Abs (λmax)/nm and ɛ/M-1 cm-1 for Trp
Tryptophan
280 nm
5500 cm⁻¹
Abs (λmax)/nm and ɛ/M-1 cm-1 for Tyr
Tyrosine
274 nm
1400 cm⁻¹
Abs (λmax)/nm and ɛ/M-1 cm-1 for Phe
Phenylalanine
257 nm
200 cm⁻¹
Abs (λmax)/nm and ɛ/M-1 cm-1 for His
Histidine
211 nm
5700 cm-1
Abs (λmax)/nm and ɛ/M-1 cm-1 for peptide bond
170 - 220 nm
~1000 cm-1
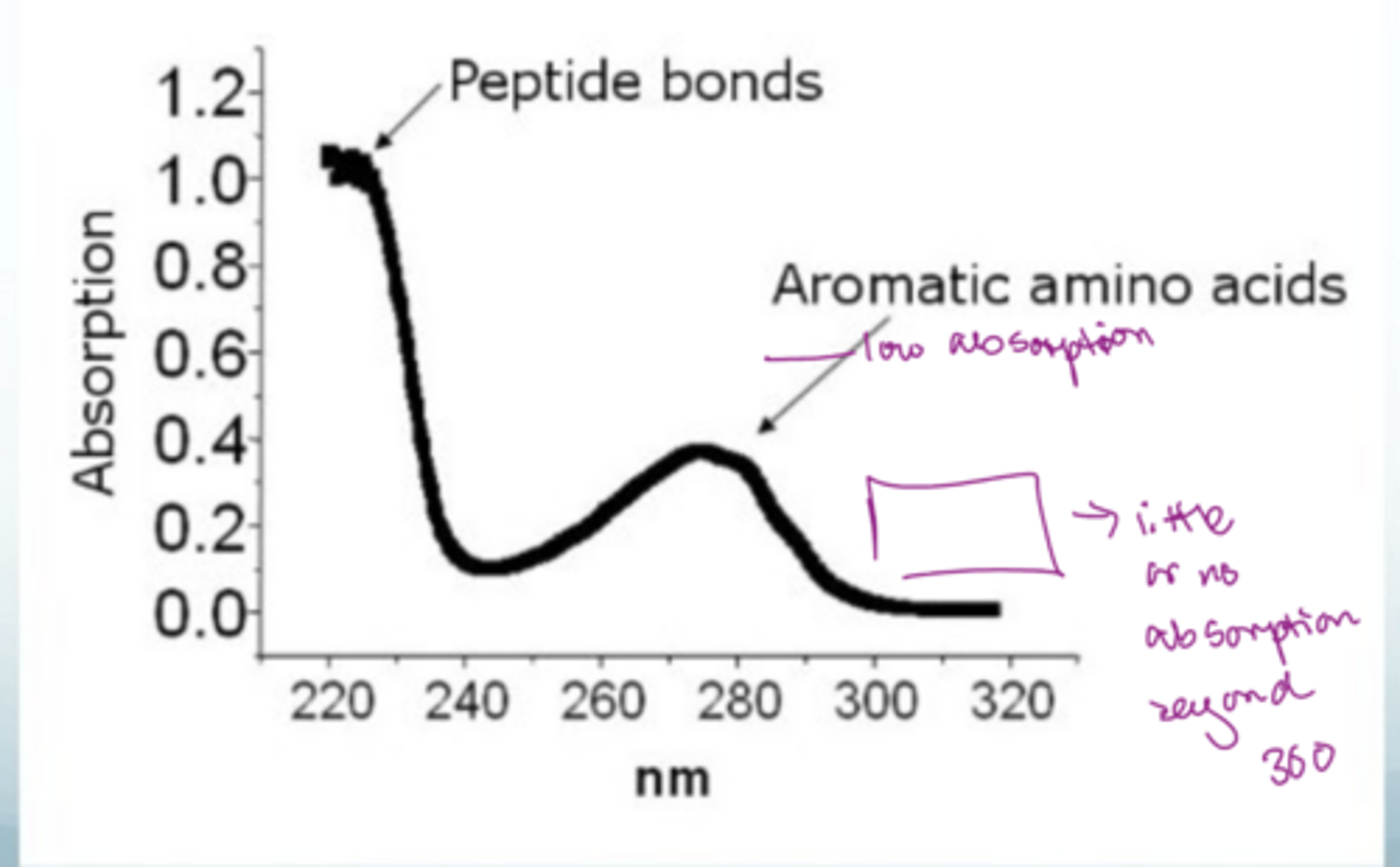
What does the absorption profile of proteins look like?
Peptide bonds dominate because there are so many but at a low nm
Aromatic amino acid have a second peak at a low absorption but a high nm
Little or no absorption beyond 300 nm
What is CD
Circular Dichroism
A form of spectroscopy that measures the difference in absorption when a sample absorbs left handed vs right handed circularly polarised light
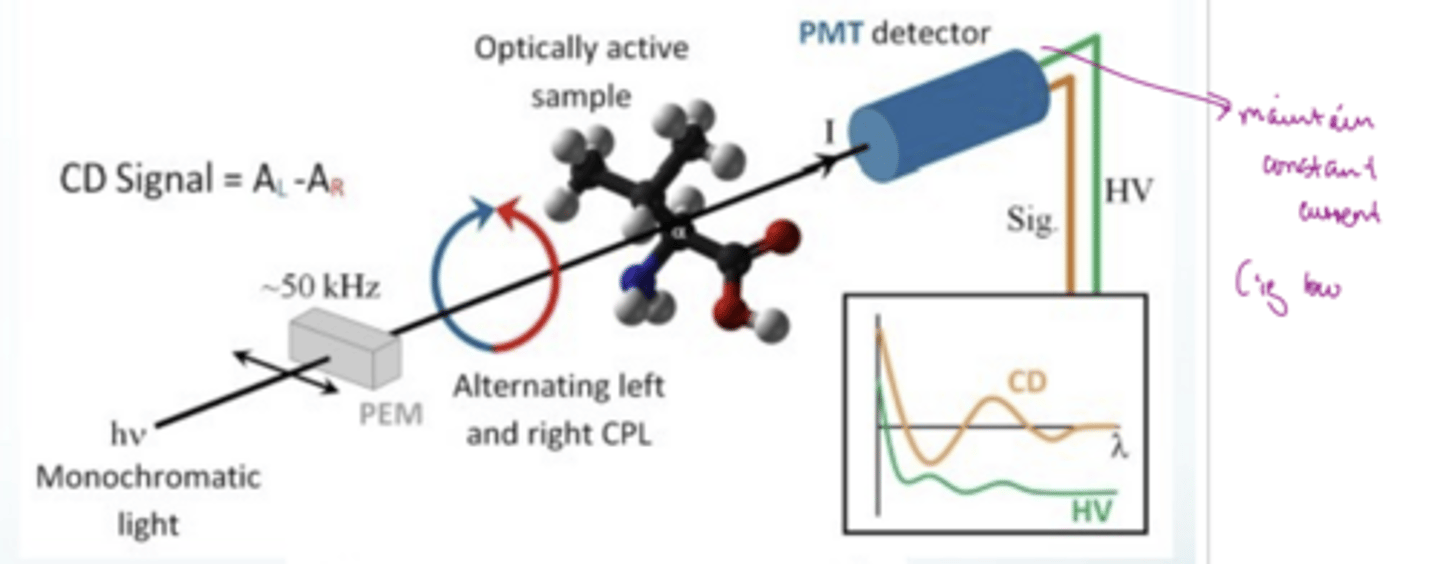
How does CD work?
CD measures the difference in absorption of left- and right-circularly polarised light by chiral molecules
1) produce light that can rapidly alternate between LCP and RCP
2) hit optically active sample (chiral (proteins) or achiral but bound in a chiral environment e.g. achiral ligand bound to a chiral protein)
3) hit detector which measure how much of each light is absorbed
The CD signal is the difference between the absorbance of LCP and RCP
can convert to change in epsillon if path length and concentration is known using the Beer-Lambert law
change in epsillon is much smaller than epsillon
pheta is molar ellipticity
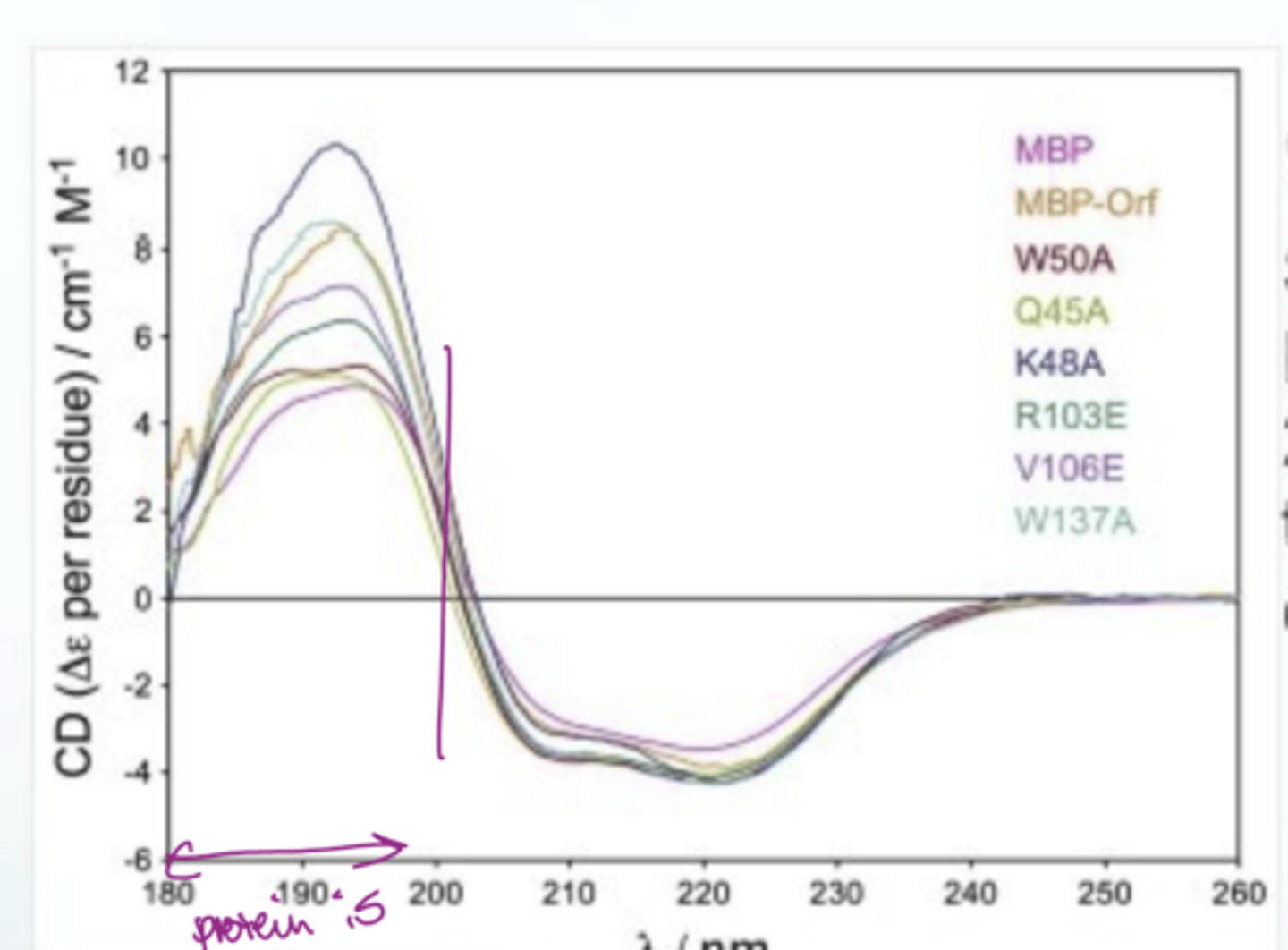
How do you interpret a protein CD spectrum to find out about protein folding?
• Key rule: [θ] is positive at 200 nm for a folded protein; negative if unfolded. This is the quickest folding check.
• α-helix: characteristic double minima at ~208 nm and ~222 nm; positive signal at ~193 nm
• β-sheet: minimum at ~218 nm; less negative than α-helix
• Random coil (unfolded): minimum at ~200 nm, generally featureless in the 208–222 region
![<p><span>•</span><span style="font-family: "Times New Roman"; line-height: normal; font-size: 7pt;"> <strong> </strong></span>Key rule: <strong>[θ] is positive at 200 nm for a folded protein</strong>; negative if unfolded. This is the quickest folding check.</p><p>•<span> <strong>α-helix:</strong> characteristic double minima at ~208 nm and ~222 nm; positive signal at ~193 nm</span></p><p class="MsoListParagraph"><span><strong>•</strong></span><span style="font-family: "Times New Roman"; line-height: normal; font-size: 7pt;"><strong> </strong></span><span><strong>β-sheet</strong>: minimum at ~218 nm; less negative than α-helix</span></p><p class="MsoListParagraph"><span>•</span><span style="font-family: "Times New Roman"; line-height: normal; font-size: 7pt;"> </span><span><strong>Random coil (unfolded)</strong>: minimum at ~200 nm, generally featureless in the 208–222 region</span></p><p class="MsoListParagraph"></p>](https://assets.knowt.com/user-attachments/9a4254b3-c063-4ec8-bb11-d06de53ea991.png)
What does a protein CD spectrum look like and how to interpret
Go to a wavelength of 190 and if pheta is above 0, the protein is folded
Tends to be noisy at very small wavelengths
region between 210-290 nm deviates since most transitions in this region are due to aromatic pi-pi*
What is quantum yield? (background info)
Fluoresence quantum yield (Φ): The ratio of photons absorbed to photons emitted through fluorescence
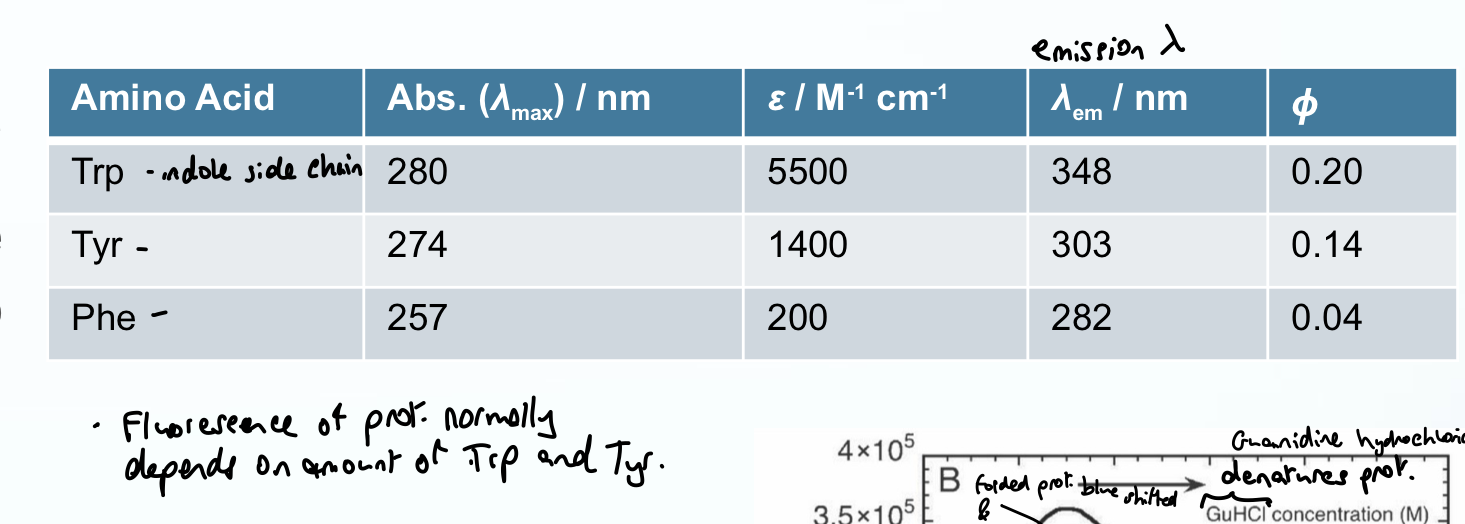
What values do AAs have for absorbance, emission wavelength, and quantum yield? (intrinsic fluoresence)
Abs < 300 nm
Emission wavelength > 300 nm (fluorescence)
Quantum yield: dependent but is ideal for studying fluorescence e.g. Quantum yield for Trp is 0.2, so for every 100 photons absorbed, 20 of them will fluoresce (0.04 is very poor so Phe is rarely studied)
What is the quantum yield of histidine?
Histidine has no measurable fluorescence
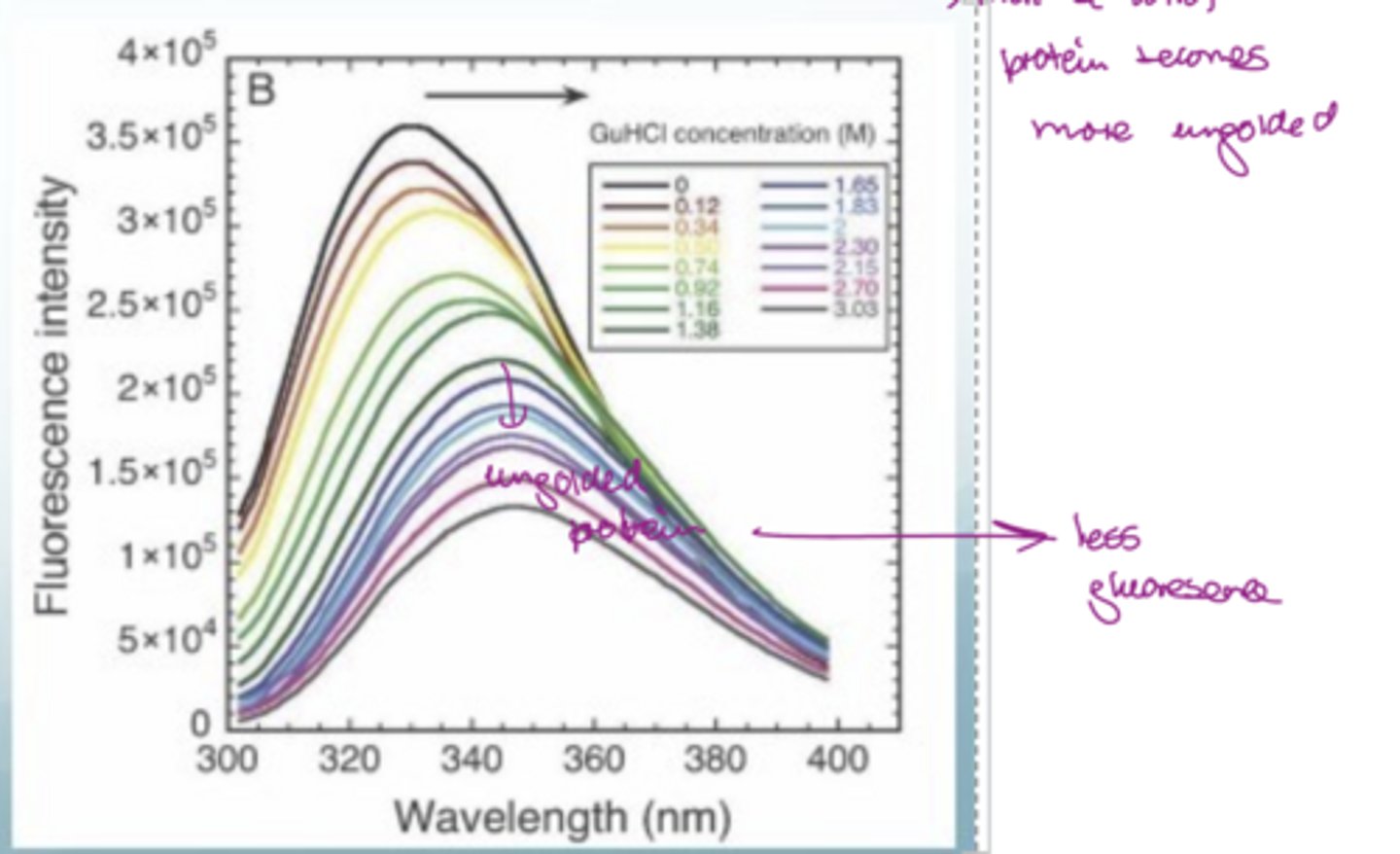
What does GuHCl do to proteins?
Denatures them
What happens to Trp fluorescence in low polarity environments (relative to water)?
Undergoes a hypsochromic (i.e. blue) shift
has a higher quantum yield
so lower emission maximum
The lowest curve has less fluorescence so this is an unfolded protein
What is ANS and how does it bind to proteins? Why does the binding curve plateau?
1-anilino-naphthalene-8-sulfonic acid: an extrinsic reagent
excellent for comparing structural states and mutants
ANS binding is driven by electrostatic (sulfonate to Lys/Arg - AA side chains) and hydrophobic interactions
ANS has a sulfonate group (always deprotonated at pH 7.4, negatively charged) and an aromatic ring (hydrophobic)
Binds to partially exposed hydrophobic regions on proteins (especially molten globule states) due to phenyl ring
↑ fluorescence intensity and causes blue shift
Fluorescence signal and ANS conc plateaus since the ANS binding sites are saturated (lots of AMS bound to prot)
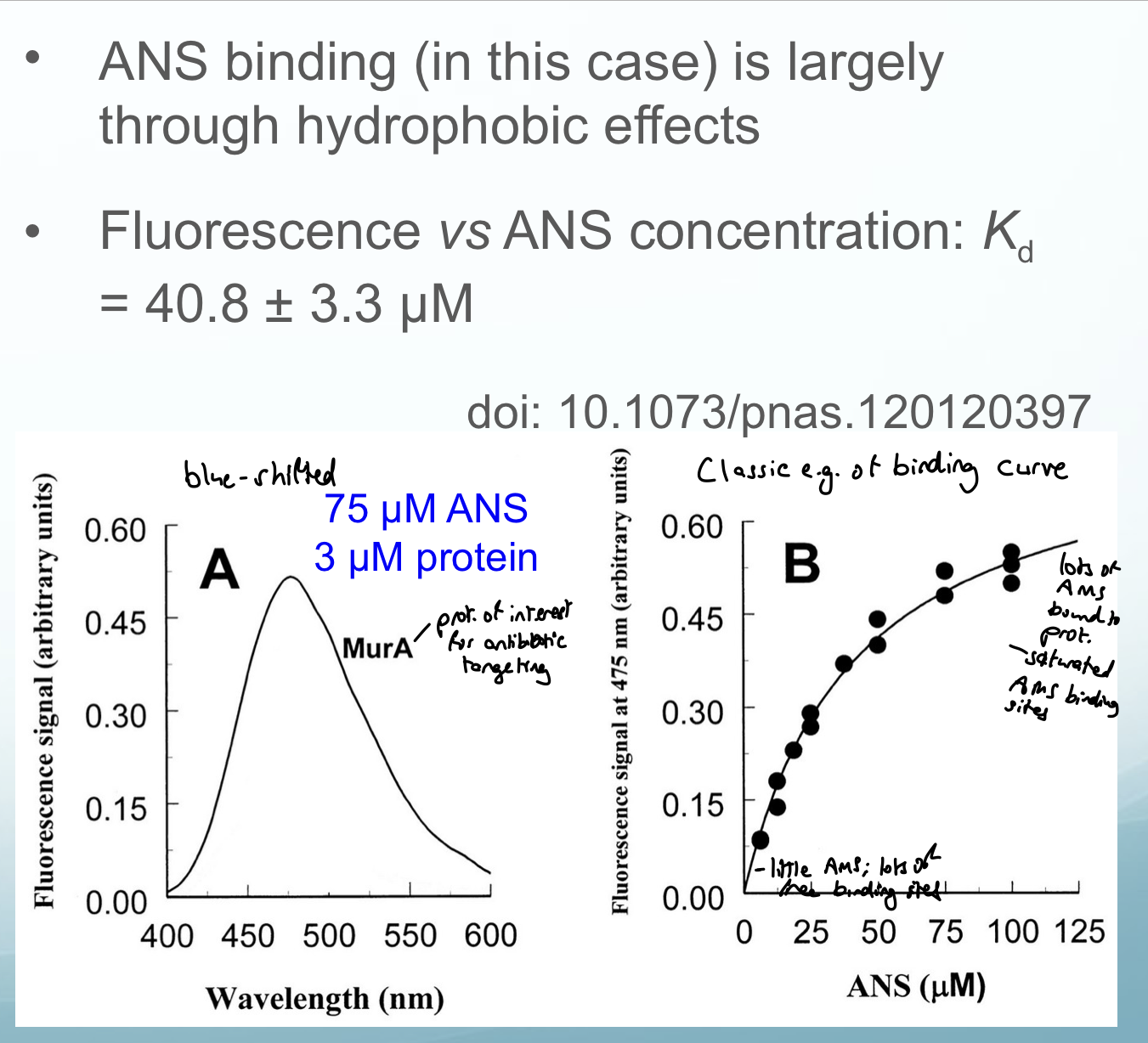
Explain interactions of ANS
ANS binding is driven by electrostatic (sulfonate to Lys/Arg) and hydrophobic interactions (aromatic ring)
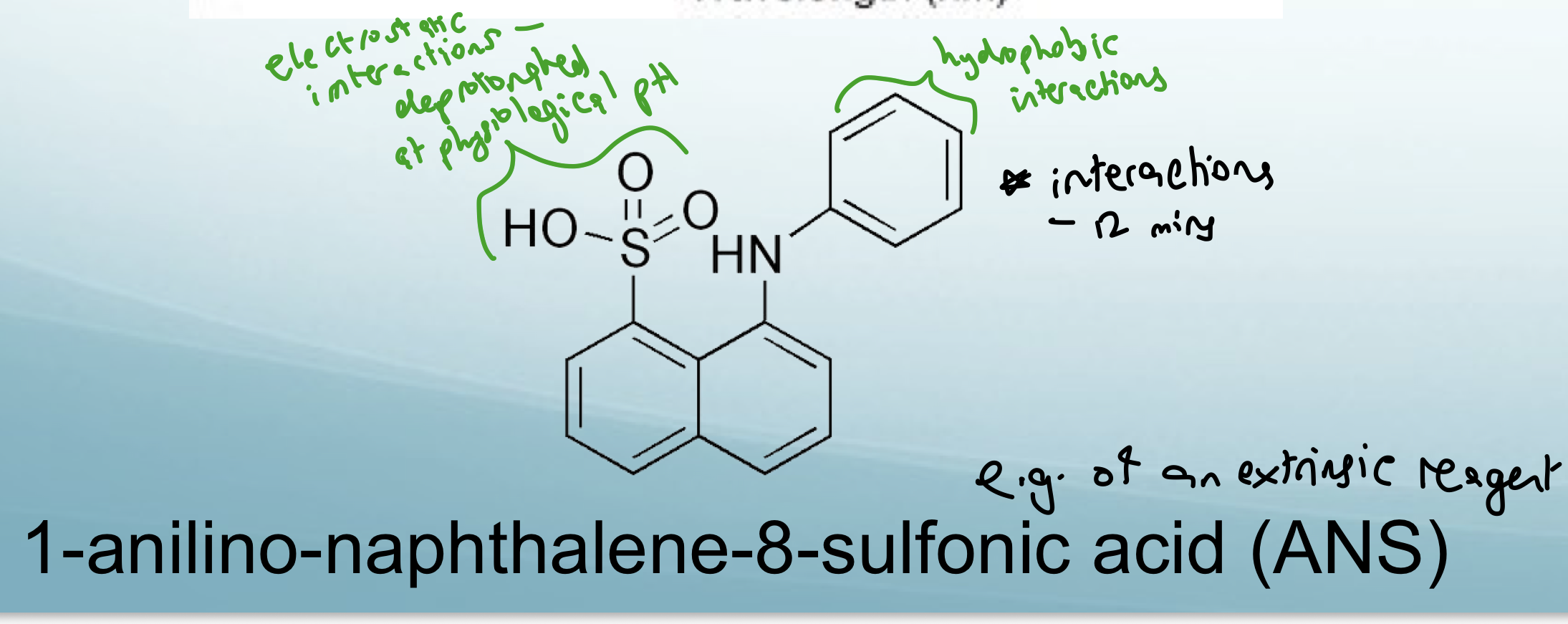
What should ANS spectrum look like in physiological systems?
Curve should peak at around 500 nm because pH is 7.4 in physiological systems
How can ANS binding tell you how a protein is folded?
fully folded protein hides hydrophobic patches → low ANS fluorescence
fully unfolded protein exposes them all → maximum ANS fluorescence
molten globule (intermediate) → high ANS signal (partially exposed)
What is GFP?
green fluorescent protein
can express GFP in the genome or tag it onto a sequence that encodes the protein of interest
Expressed using natural mechanisms
useful for tagging and monitoring
How do nucleic acids compare to aromatic AAs in terms of absorption?
Much better at absorption
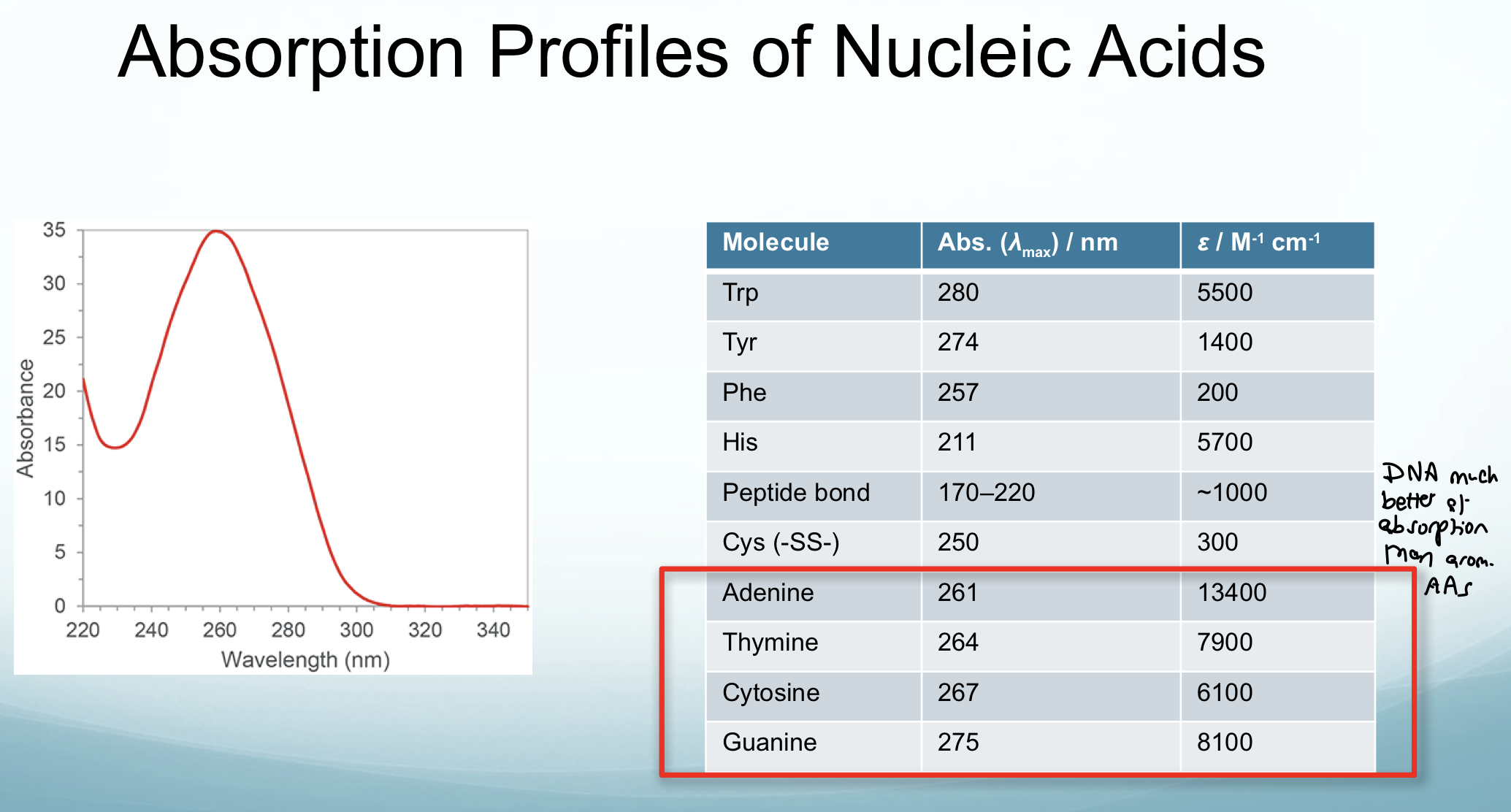
What is DNA melting?
Often a thermal process which causes the DNA double helix (native conformation) to dissociate into its single strands (random coil conformation)
Why does DNA melting occur?
Cells need to access single stranded DNA for reading in transcription
has to be able to unwind in DNA melting
Is also used in PCR to make templates (thermal process)
What is the Tm?
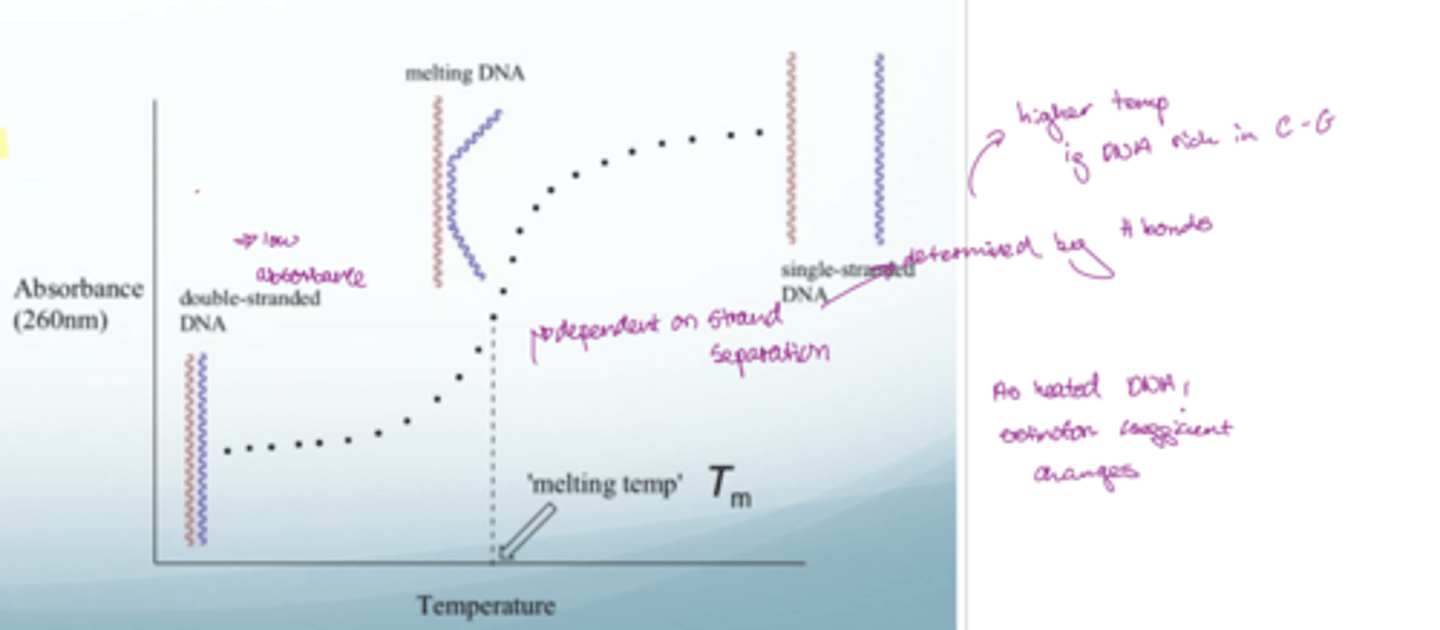
Melting temp of DNA: the tempeature at which half the DNA is single stranded (where unfolding is happening most rapidly e.g. if were to differentiate and then observe at what temperature the peaks occur)
Dependent on H bonds e.g. Tm will be higher if DNA is rich in C-G
If one person has a complementary sequence and the other person is a slight variant, the person with the complementary sequence has a higher Tm
How can Tm be measured using absorbance?
UV-Vis
dsDNA has a low absorbance
ssDNA has high absorbance
dsDNA is hypochromic (stacked bases reduce absorption).
On melting: hyperchromicity — A260 increases as bases unstacked.
Sigmoidal curve vs T; midpoint = Tm.
How can Tm be measured using the extinction coefficient?
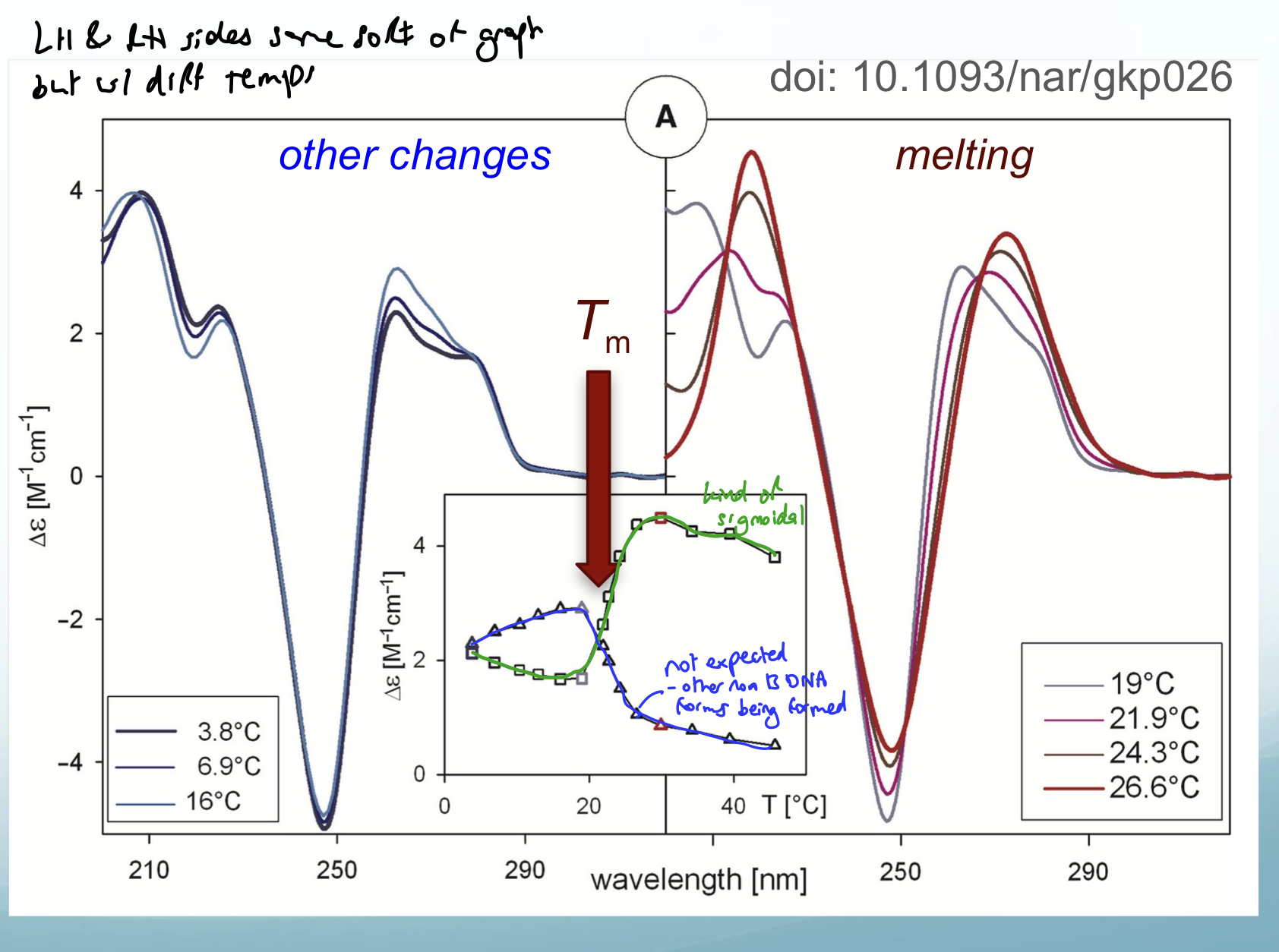
CD
Useful for telling you if other structures of DNA helix is present e.g. A form or Z form
CD signal of DNA changes as helix unwinds with ↑ temp.
Also monitors A→B→Z-form transitions.
Need molecule that absorbs and is chiral (nucleic acids tick both).
More sensitive than UV-vis
Why is dsDNA hypochromic (decrease in intensity of absorption (εmax)) compared to ssDNA?
Because of proximity of transition dipoles in pi-stacked base pairs - affects E
What is FRET, and how does it occur?
Förster Resonance Energy Transfer
Involves E transfer from donor → acceptor ( through-space dipole–dipole coupling (NOT emission/re-absorption)_
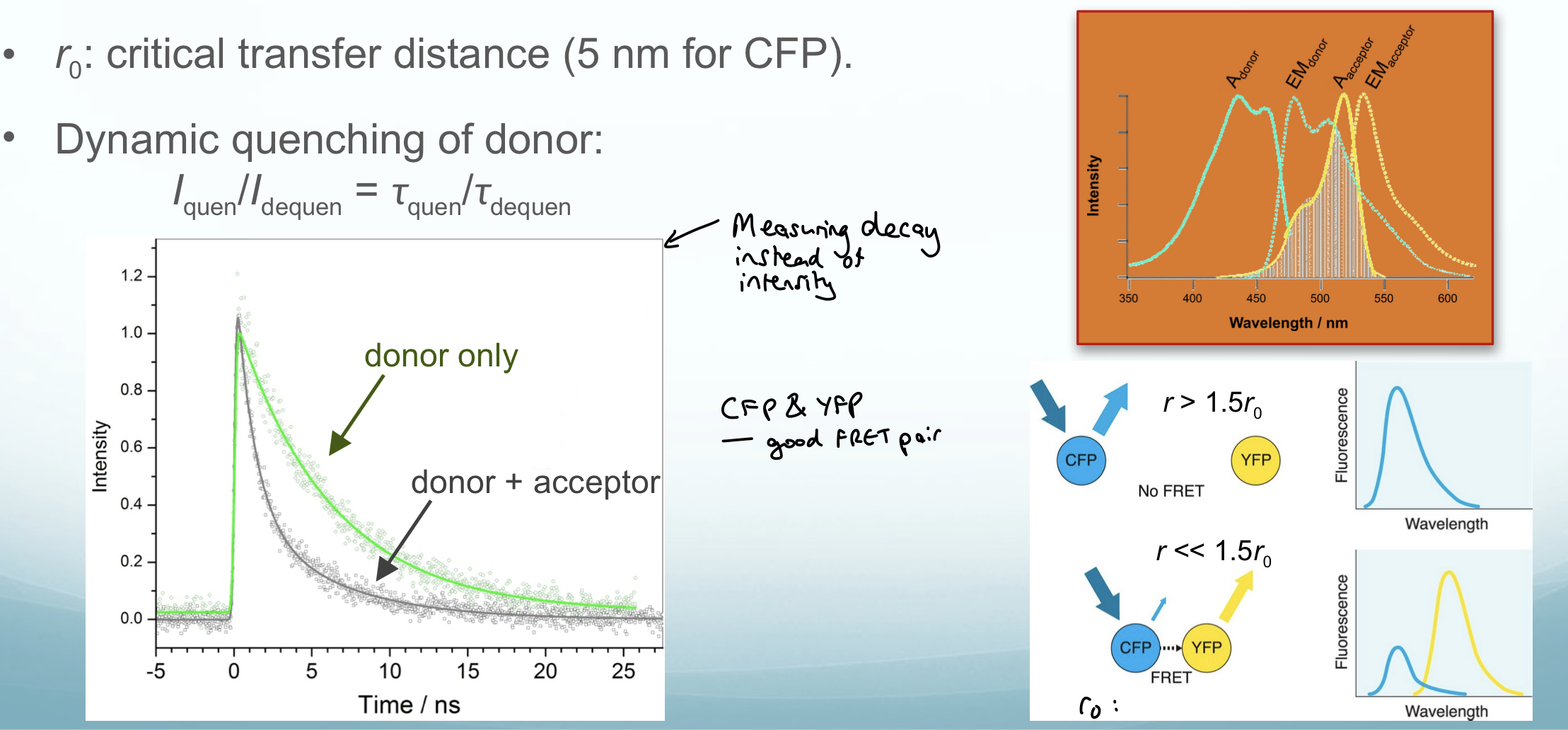
Detected by: quenching of donor fluorescence (decrease in intensity/lifetime) + appearance of acceptor emission
Occurs if the donor emission overlaps with the acceptor excitation
e.g. if close enough in the spectrum, the emission of GFP will excite RFP
only effective at short distances - overlap is essential
• Rate of energy transfer kET ∝ r⁻⁶ (very steep distance dependence: r ≤ 5 nm; r₀ ≈ 5 nm for CFP)
How can Tm be measured using Fluorescence
FRET
Even more sensitive than CD
One strand labelled with donor, complementary strand with acceptor.
In dsDNA: strands close → FRET seen.
On heating above Tm: strands separate → loss of FRET.
Sigmoidal curve midpoint = Tm.
When the distance between fluorophores is less than the critical distance, FRET occurs
Produces a spectrum where the donor peak is a lot smaller than the acceptor peak and there is overlap in wavelength
Measuring fluorescence decay gives a better signal: noise ratio than if fluorescence intensity was measured since samples scatter light
What property of DNA enables things to slide between the bases
Bases are stacked in the middle of the double helix structure so things can slide between them
absorbance is affected by the space between them
What FRET do you get if single stranded DNA is tagged with a fluorophore (one strand per fluorophore)
No FRET
only donor emission
What FRET do you get if double stranded DNA is tagged with a fluorophore (one strand per fluorophore)
FRET
donor and acceptor emission so 2 peaks
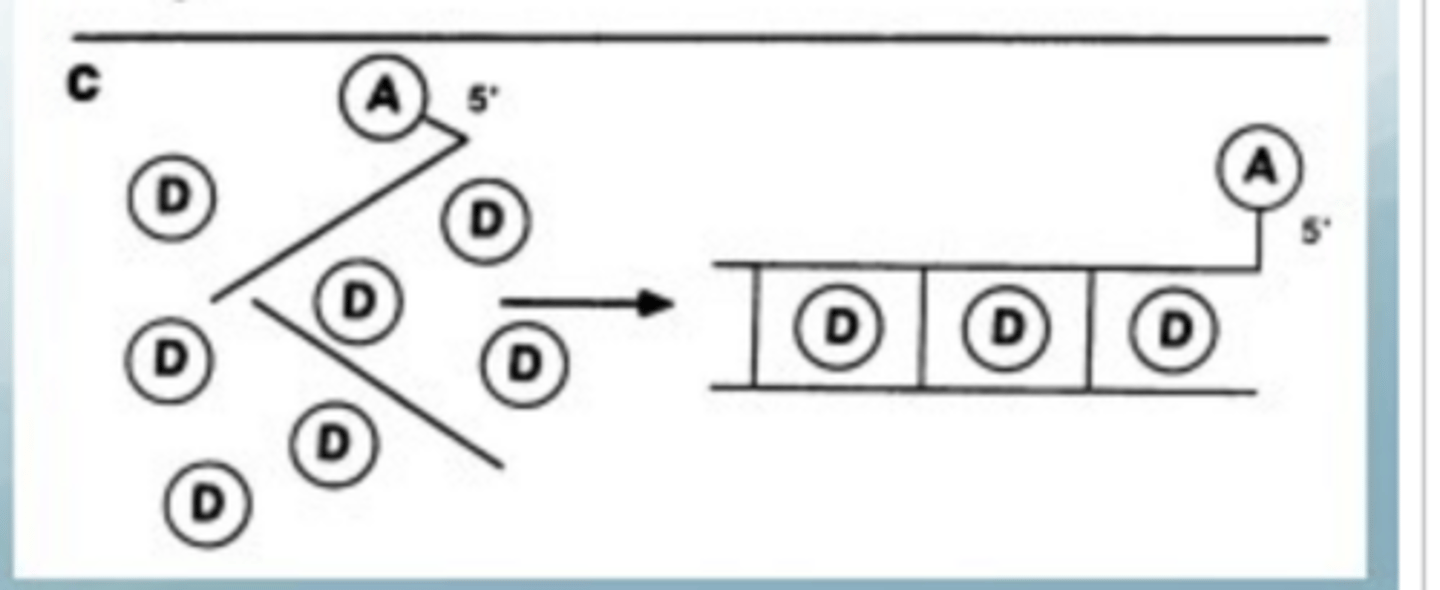
What is intercalating FRET (iFRET)? State one application
Excess of donor so can slide in between DNA bases when the DNA strand is tagged with the acceptor
used to look at mutations
only 1 strand needs to be tagged
Donor = intercalating dye (flat aromatic, inserts between base pairs of dsDNA; cationic → interacts with phosphates)
Acceptor = fluorophore on 3' end of probe strand
• Donor can only intercalate when dsDNA is present → FRET only occurs when probe has hybridised to its target
• Only the probe needs to be labelled — target genomic DNA can be unmodified
Application: screening genomic DNA populations for specific sequences or SNPs
If target contains mismatch: lower Tm (weaker hybridisation) → separate sigmoidal curve at lower temperature in first-derivative plot
How can iFRET be used to m
Intercalator acts as donor; loss of intercalation at Tm removes FRET. Only probe needs labelling.
What happens to nucleic acids below the Tm?
double stranded
helical (normal DNA structure)
What happens to nucleic acids above the Tm?
DNA becomes single stranded
H bonds break
base stacking disrupted
strands become random coils
What is nuclear spin characterised by?
spin quantum number I
What does nuclear spin give the nucleus?
a magnetic moment (μ)
this magnetic moment can align with or against an external magnetic field (B₀) with a magnetic quantum number between ml (-l and +l)
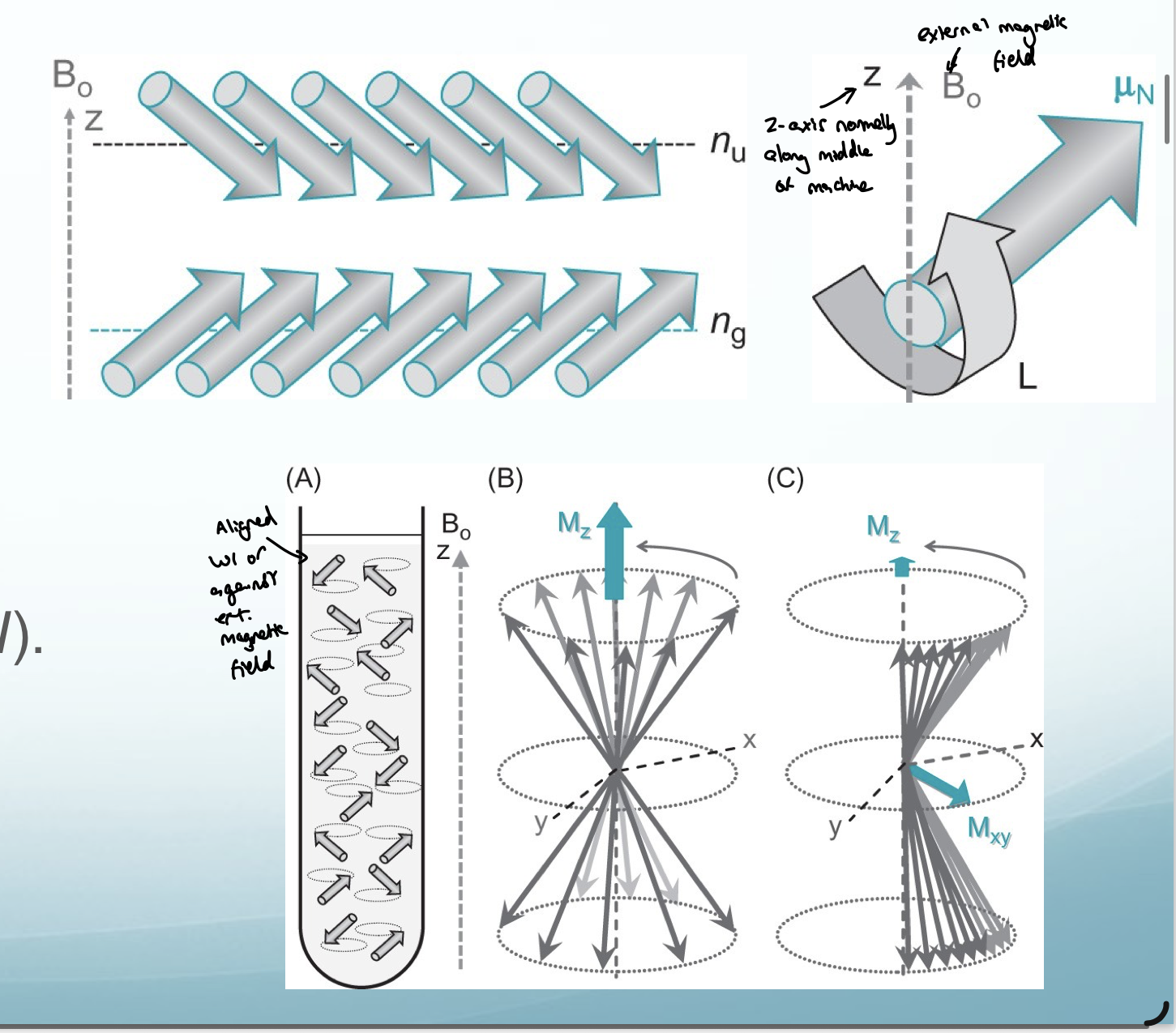
What is the spin lattice relaxation time
T1 - z direction
What is the spin-spin relaxation time
T2 - x,y direction
what is the relationship between T1 and T2
T2 is less than T1 for biomacromolecules
T1 and T2 are similar for small organic molecules
What is tau(c)
the correlation time for random motions in solution
What are some practical decisions for biomolecular NMR
small proteins need 2D
large proteins need 2D/3D NMR but the maximum size limit isn't actually very much
need a certain amount of H1
some atoms can be isotopically enriched
need to assign chemical shifts to structural components
partial assignments may be sufficient for dynamics or ligand binding
What does 2D NMR give info about
connectivity or proximity
what does the 2D NMR spectrum of a folded protein look like
a broad spread across the whole spectrum with similar intensities is indicative of a folded protein
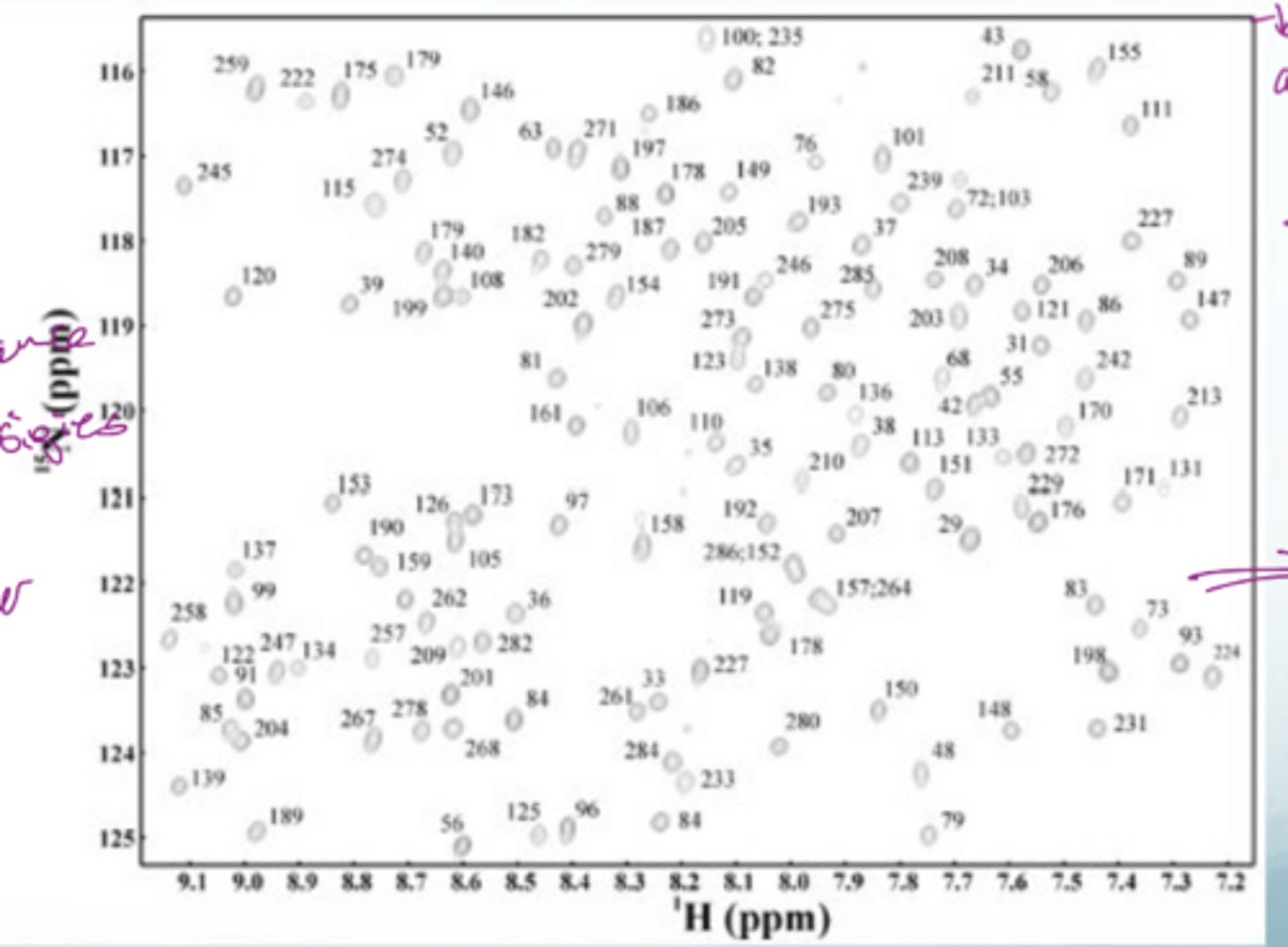
What are the 2 dimensions in 2D NMR
t1 and t2
What can the data be thought as in 2D NMR
a stack of 1D files, each differing by a change in t1
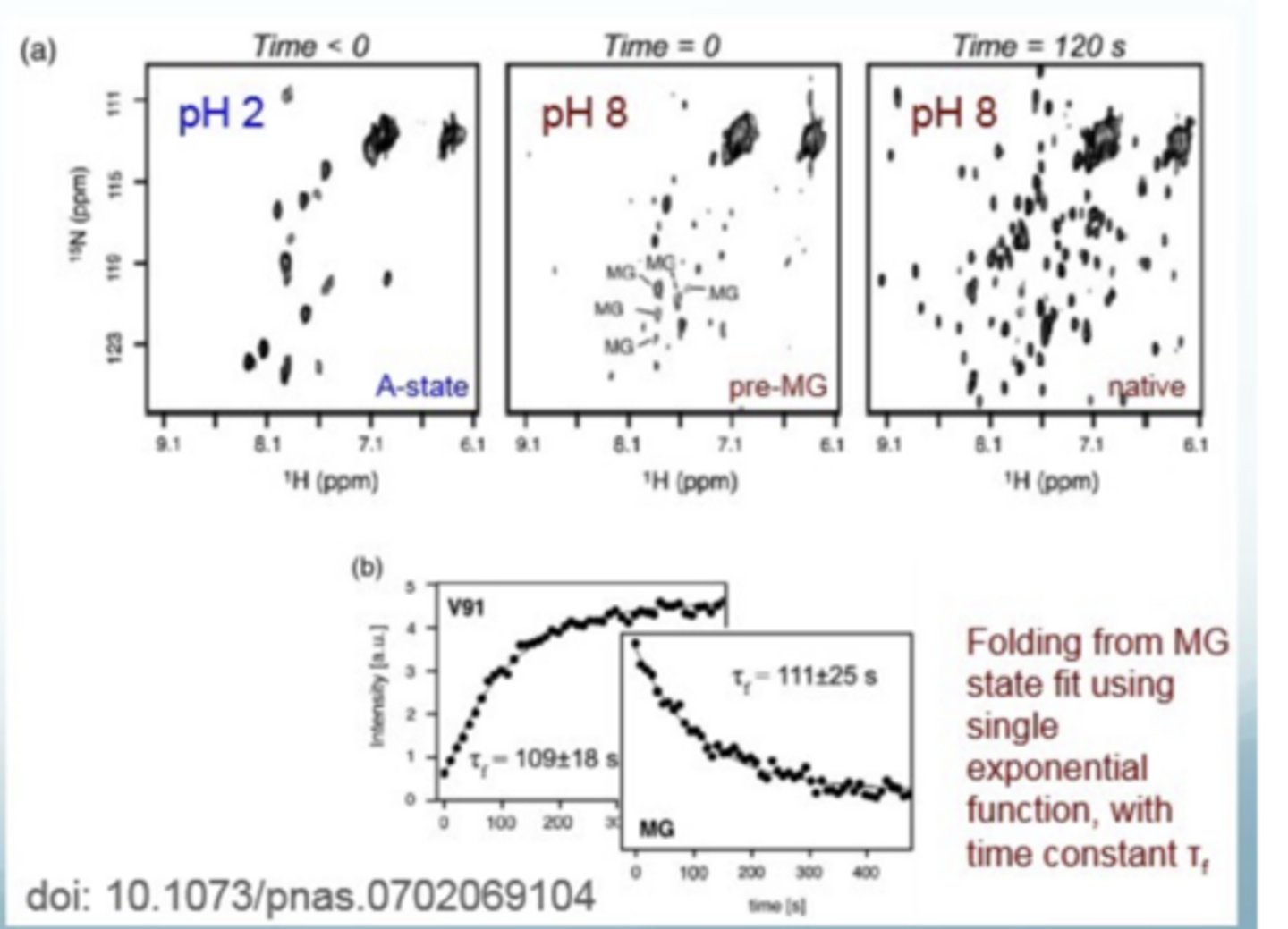
What does a HMQC spectrum look like
at pH 2, proteins are in molten globular state
at pH 8, states are beginning to fold into native states
at pH 8 after some time, the proteins are fully folded and have reached the native state
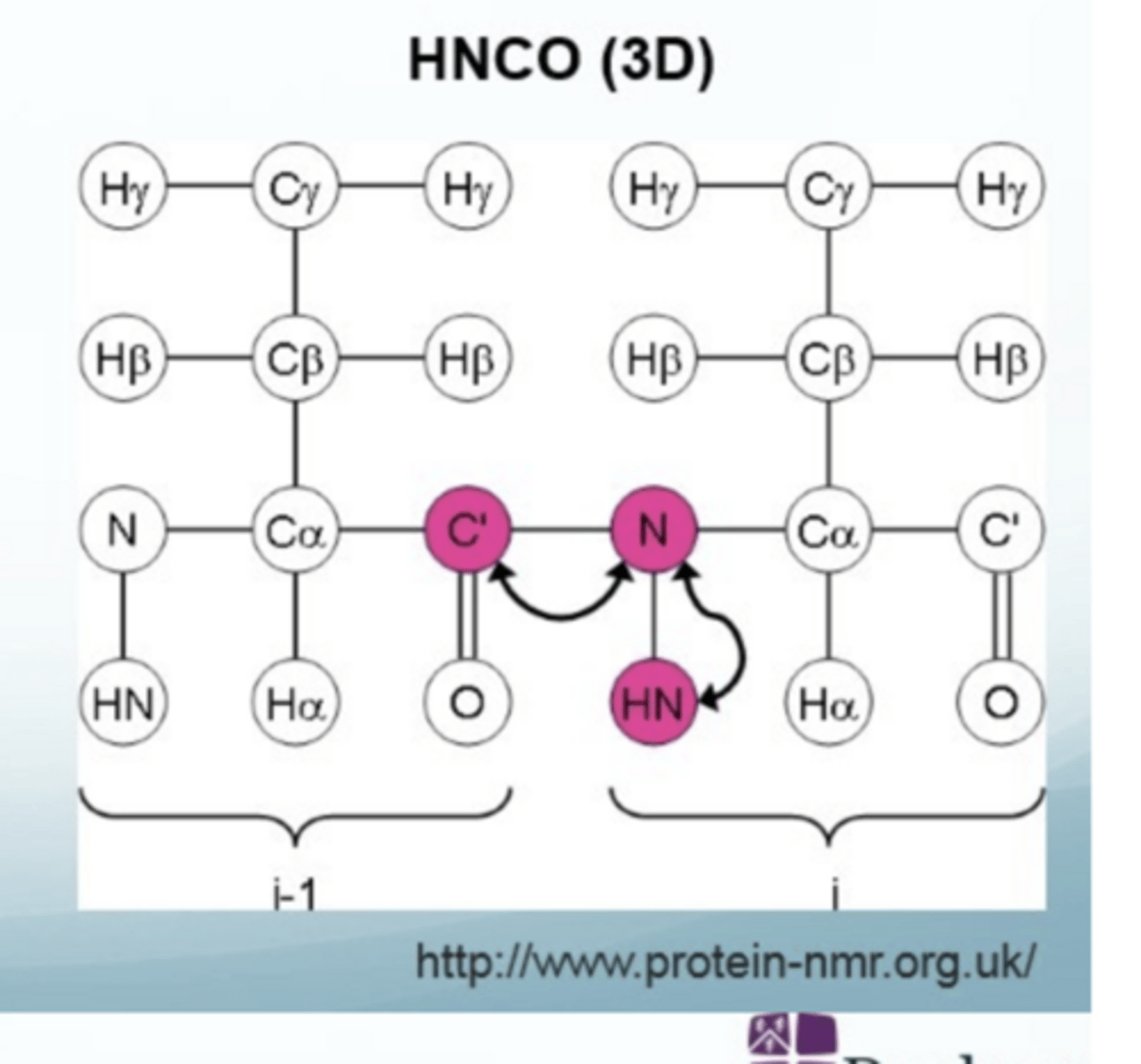
What is HNCO
3D method of biomolecular NMR
magnetisation transferred from the H amide to the N and then to the C=O and back again
confirms secondary structure (from C=O shifts) and aids backbone assignment
What is HNCA
3D method of biomolecular NMR
magnetisation transferred from the H amide to the N and then to the alpha C and back again
useful for backbone assignment
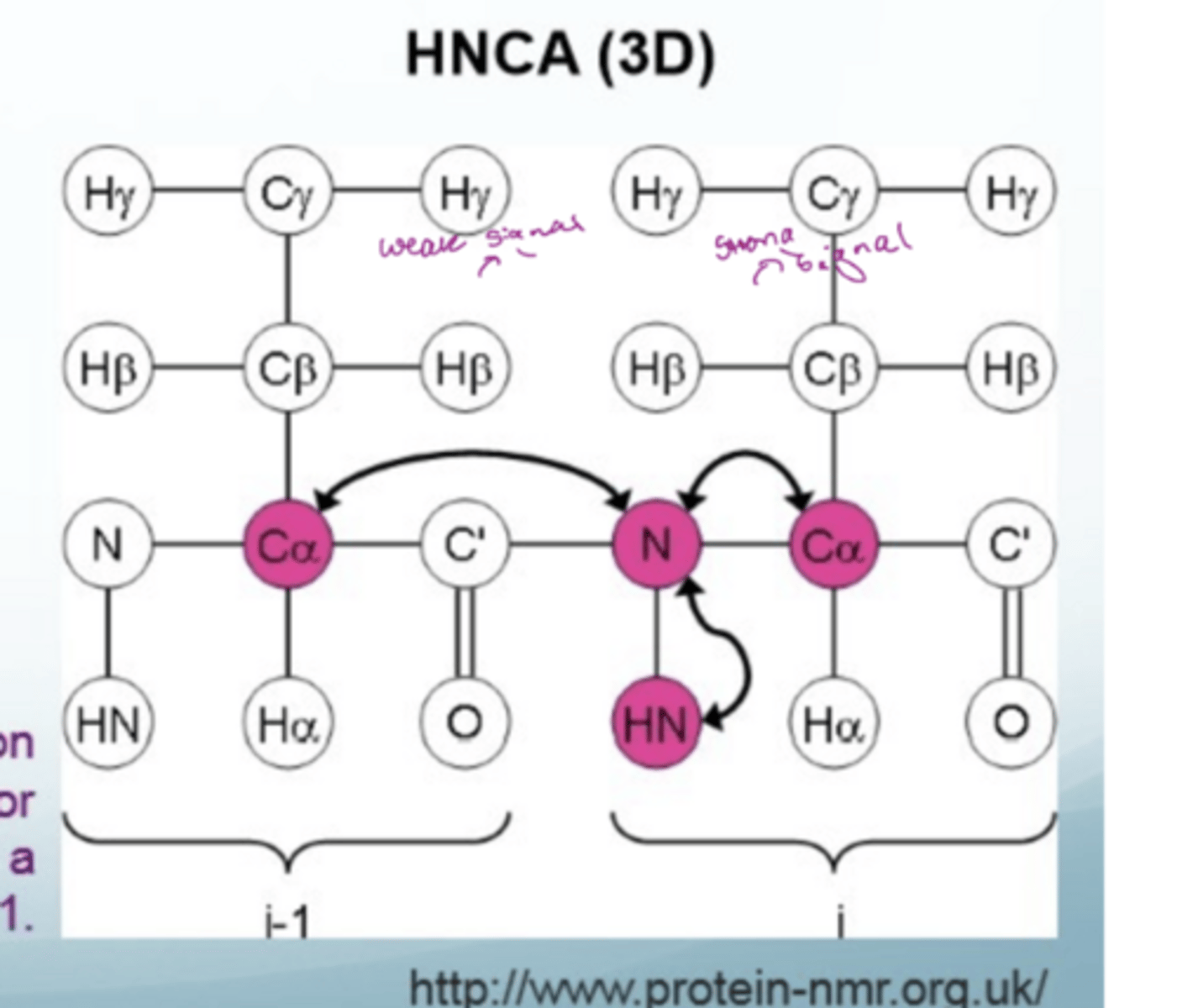
What is an example of a 3D NMR spectrum
each slip correlates to each other
if blob is strong (big and black), there is a strong shift for the alpha carbon
What is NOESY
useful for backbone assignment and structure determination
get proximity effect between neighbouring residues- determine secondary structure of proteins
measures changes in intensity depending on spatial arrangement
energy transfer is dipole-dipole bases relaxation through space
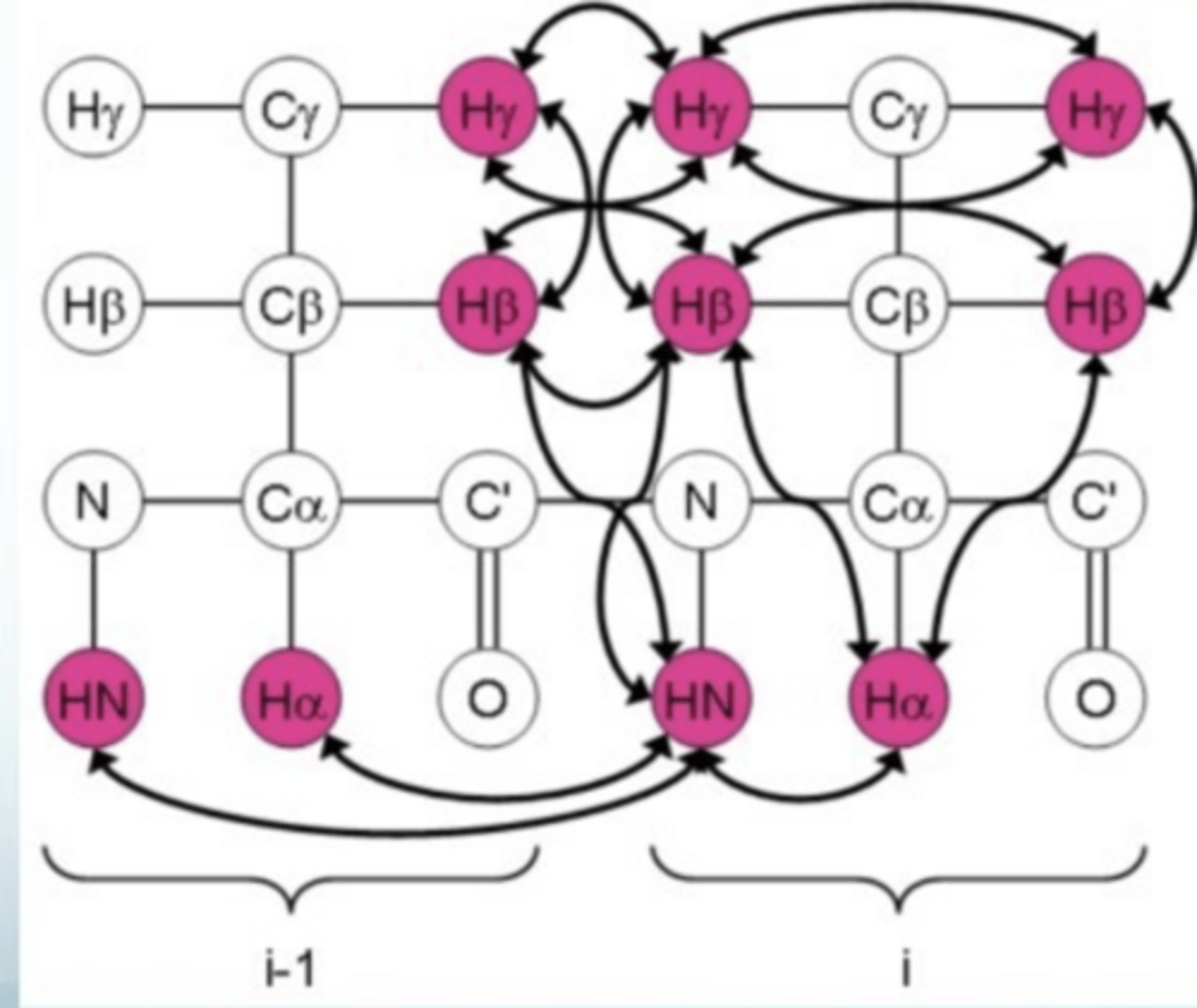
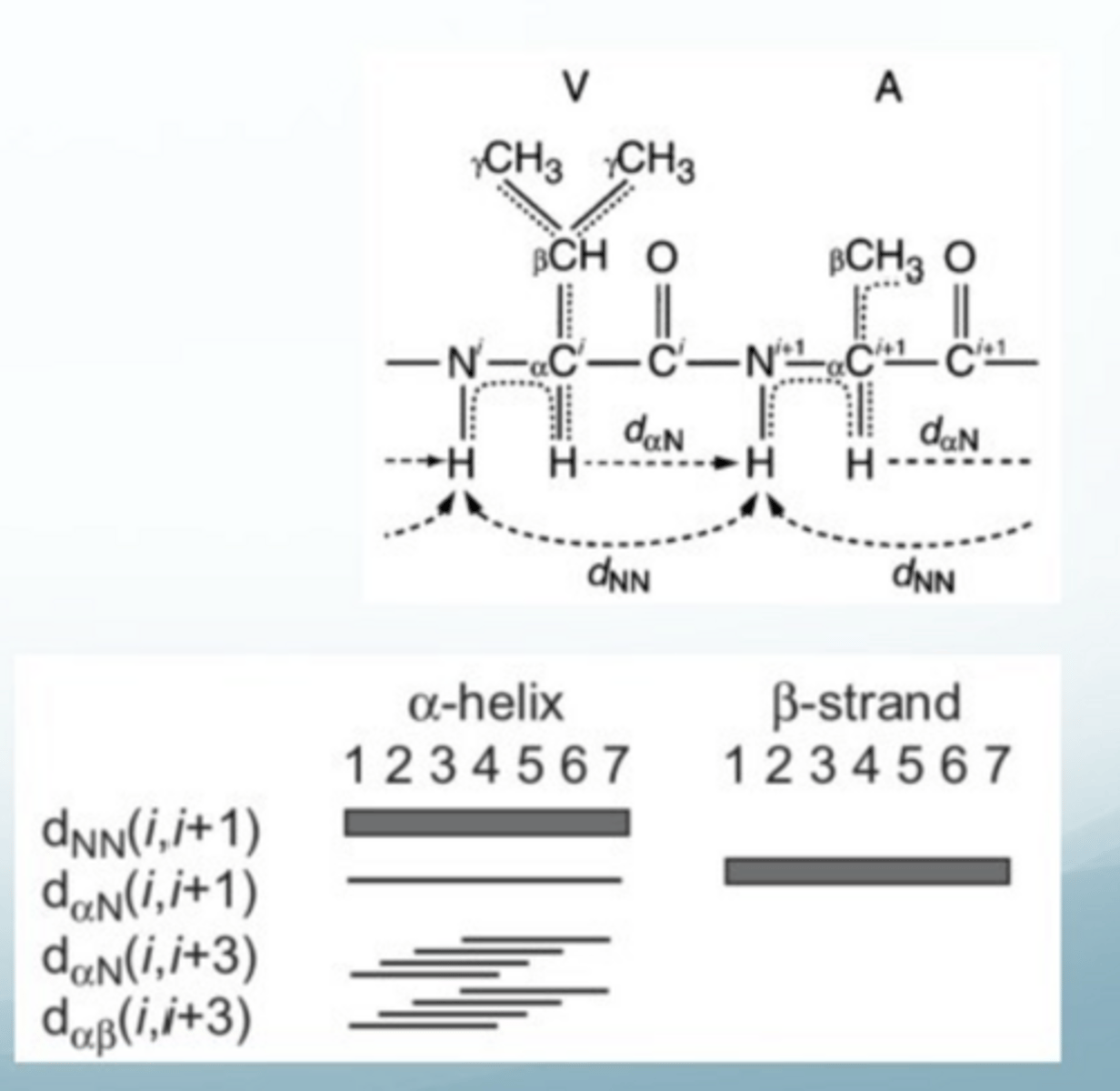
how can you determine the secondary structure of a protein through NOESY data
strong nOe between NH of residue i and NH of residue i + 1 for an alpha helix, but not in beta helix
strong nOe between proton on alpha C or residue i and the NH residue of i+1 in beta sheet
can be used to set distance restraints but need a reference distance for residues
What is H/D exchange
H atoms involved in H bonding exchange slowly particulary inside a folded protein
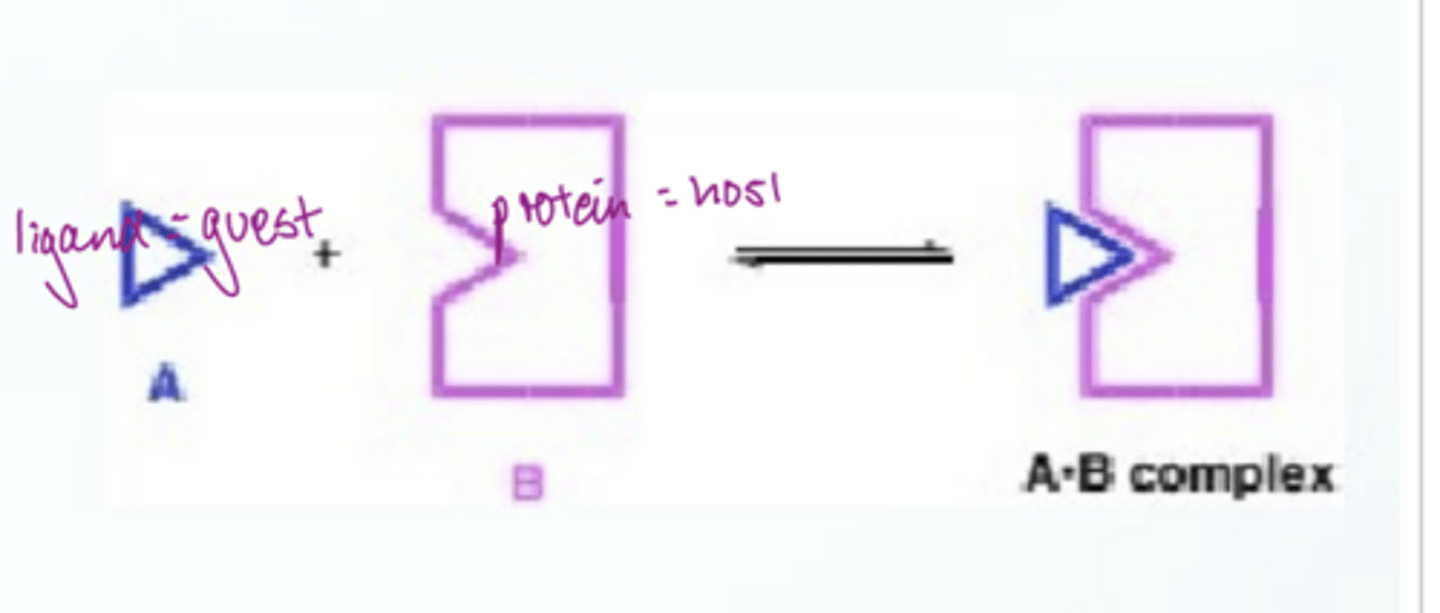
How is host-guest binding measured
using K: K can be measured by monitoring any observable property that changes as the host and guest concs are varied
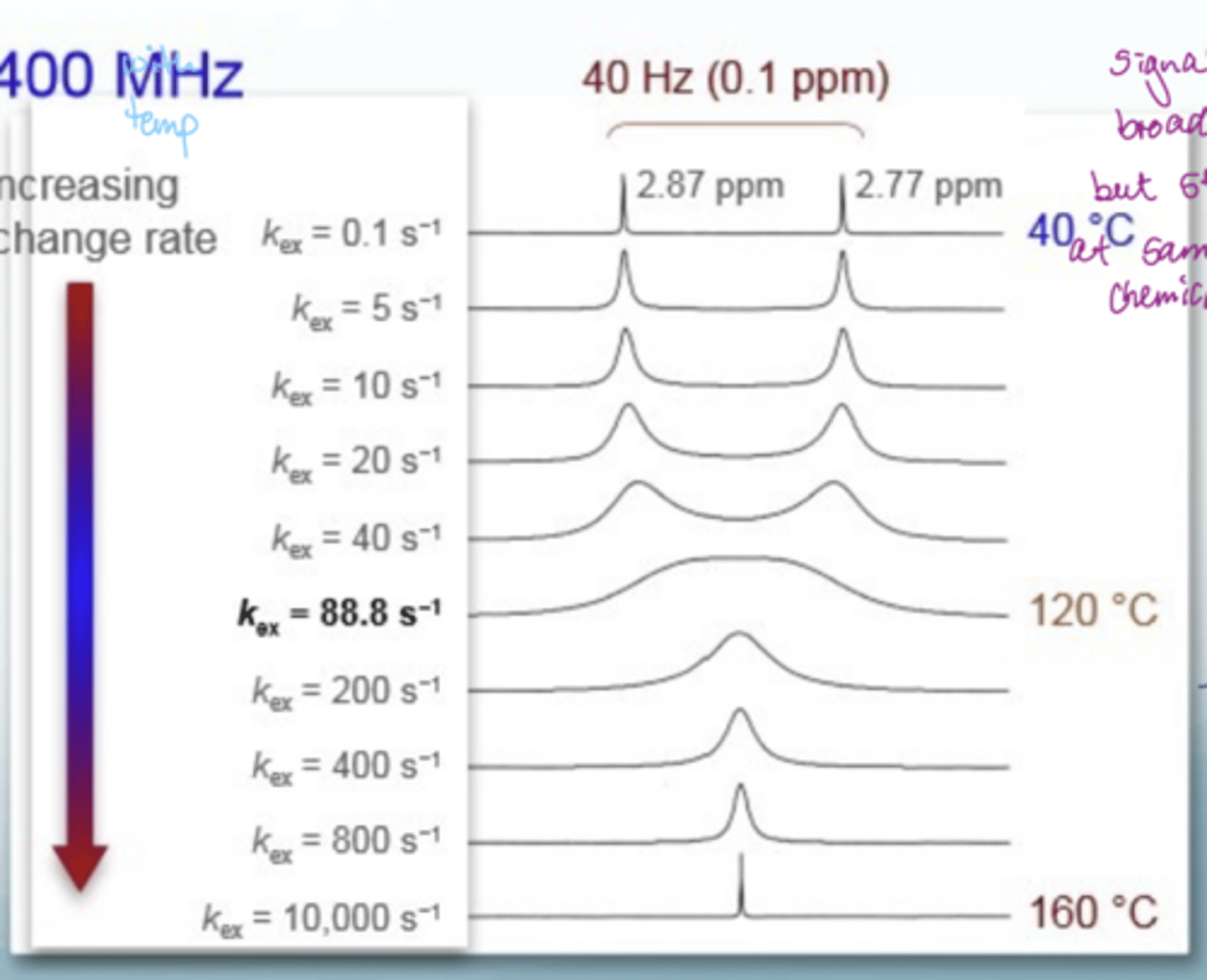
What does the chemical exchange profile of DMF look like
at low temps, there are 2 sharp peaks due to methyl groups (no significant rotation around CN so bonds are distinct)- slow exchange
as temp increases, signals broaden but stay at the same chemical shift - intermediate exchange
at high temps, there is rotation around the amide bond so get fast exchange and see 1 signal- fast exchange
a difference in peak intensity reflects equilibrium intensities
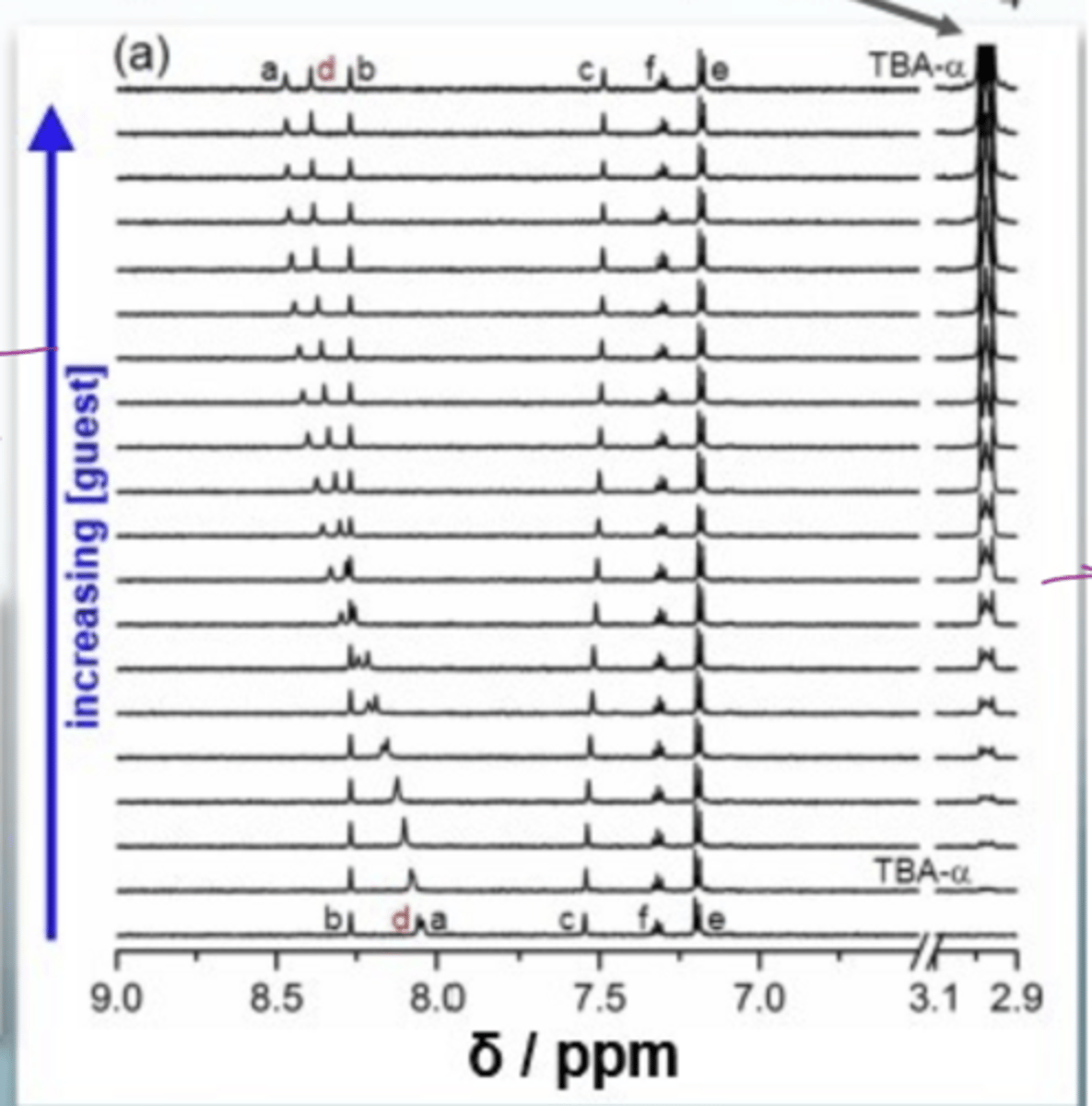
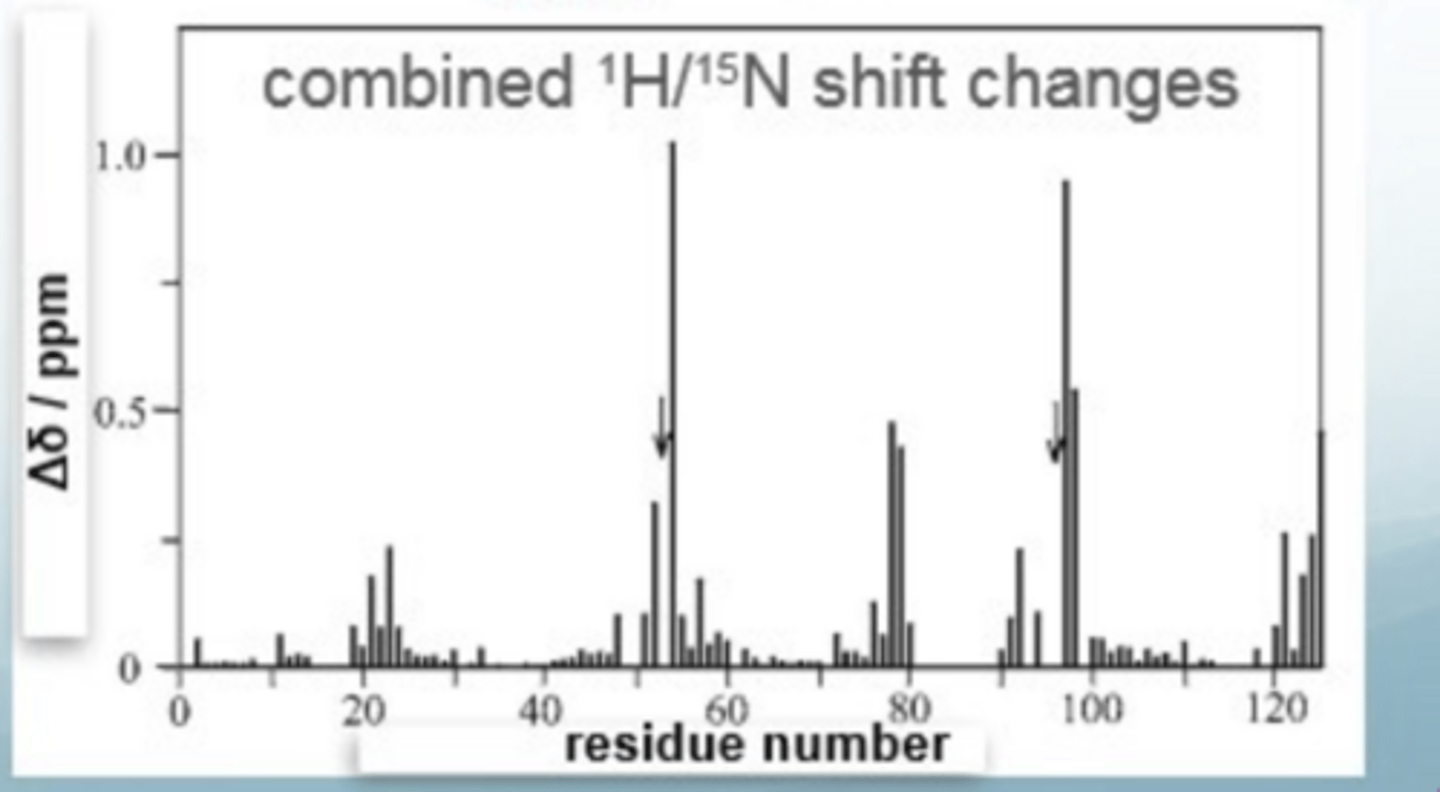
What does the NMR spectrum of protein- ligand binding looks like
at the bottom, there are weak signals since there is a small conc in guest
the magnitude of shift change tells you how close it is to the binding site, NOT how strongly it binds (K)
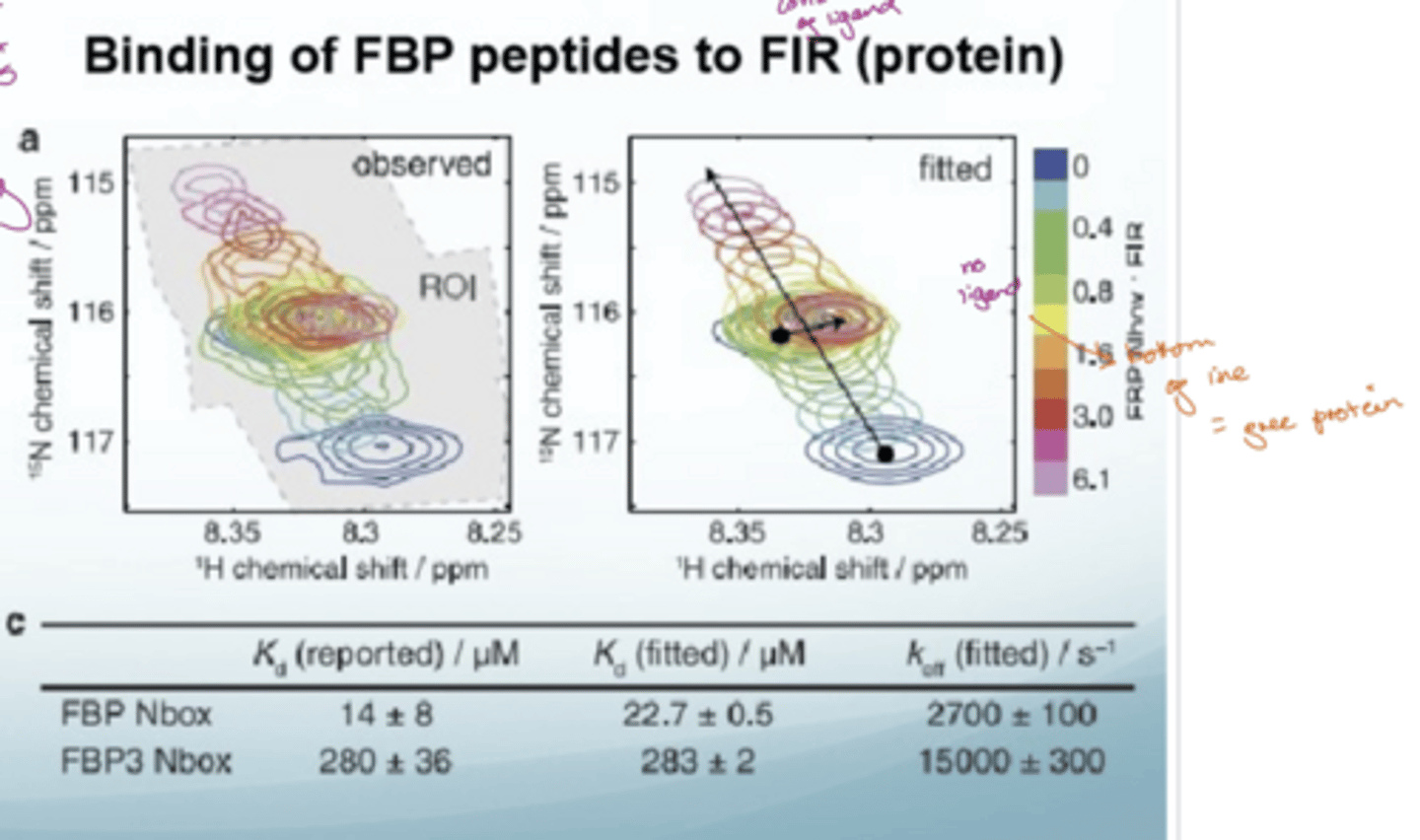
Where are resonances that move likely to be located
close to the binding site
a dot = free protein
top of arrow = high conc of ligand
LHS = noise
RHS = no noise depicted by smoother circles
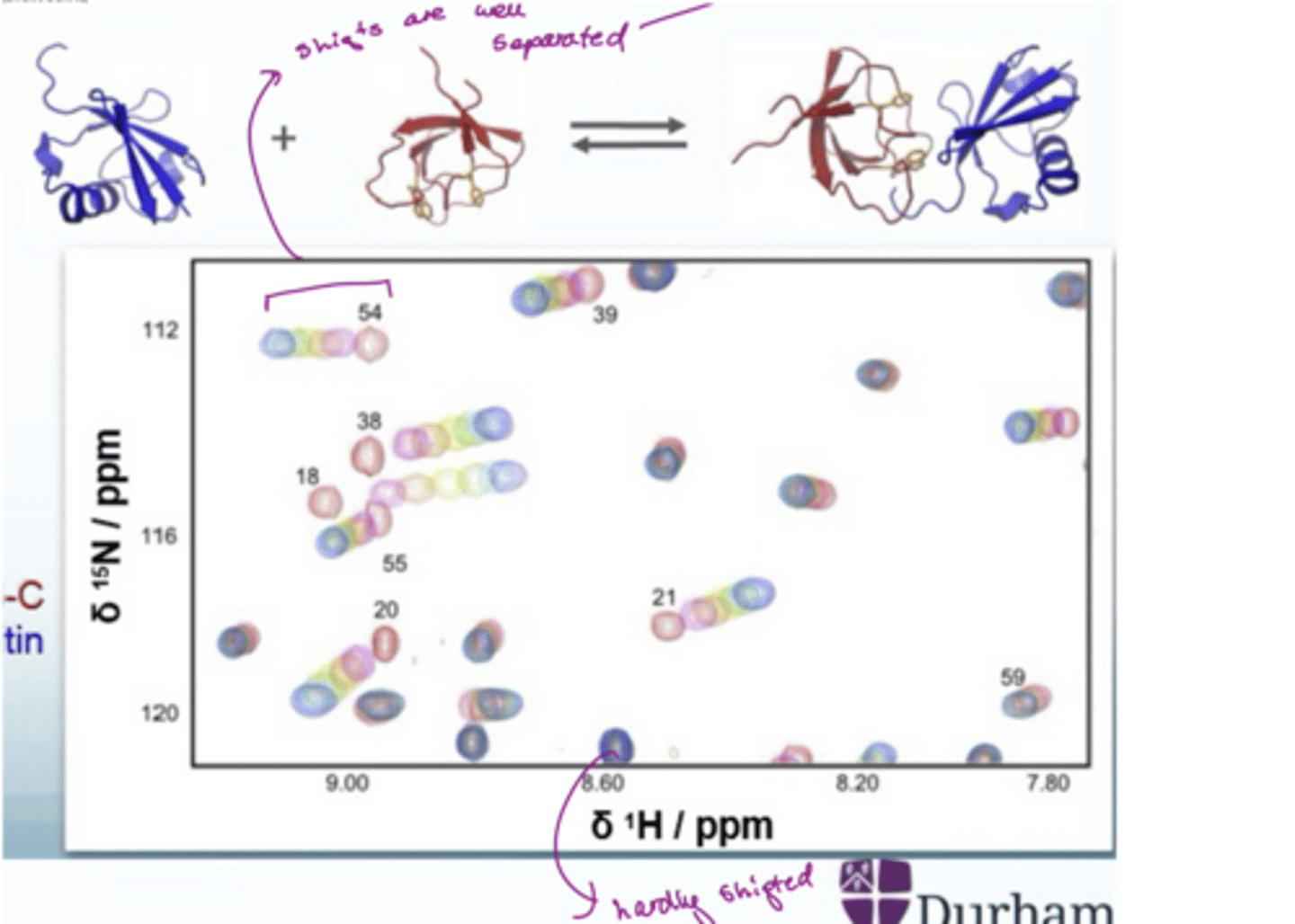
How to interpet a protein-ligand NMR
If there shifts are well separated e.g. 54, then there is fast exchange, indicating the residue is close to the binding site (there is a progressive change in chemical shift
If it seems like there is only one spot (the one at the bottom), the residue hardly shifted so far away from the binding site
Residues with big shift changes are closer to the binding site
Why is protein-ligand NMR useful
the interactions at the binding site of the protein e.g. which residues are involved in binding, are being revealed by chemical shift changes
What is another example of protein-ligand NMR
G98 is a free protein because there is a big shift change and progressive contours, which means there is fast exchange
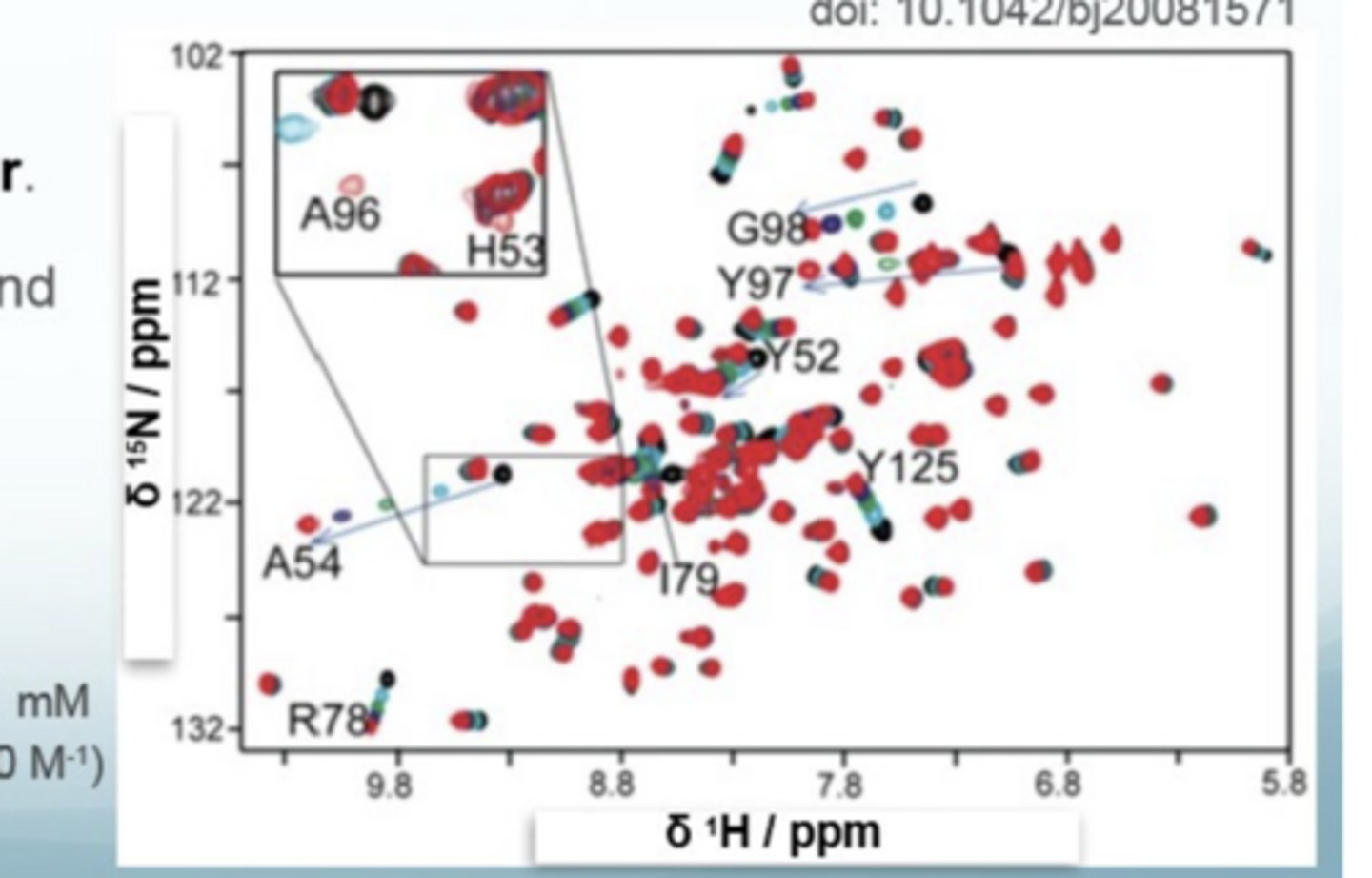
What do you always say after progressive change in shift in protein-ligand NMR
there is fast exchange
What do you say if the shift change has a large magnitude
large magnitudes = close to the binding site
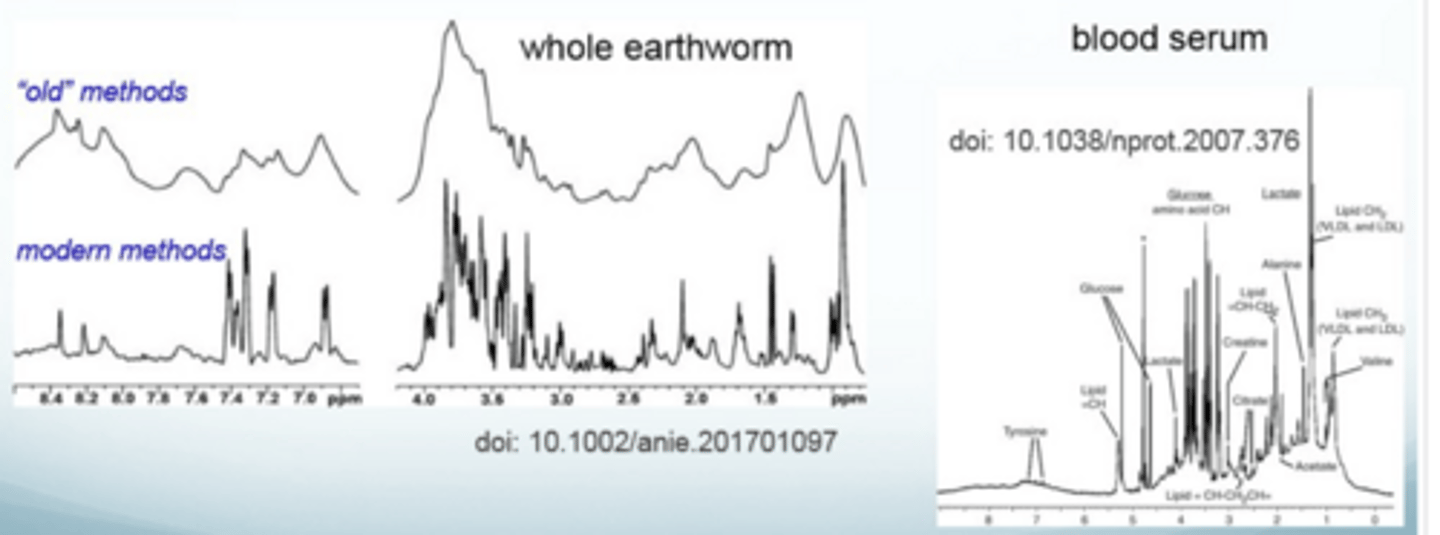
What does 1D NMR spectra look like
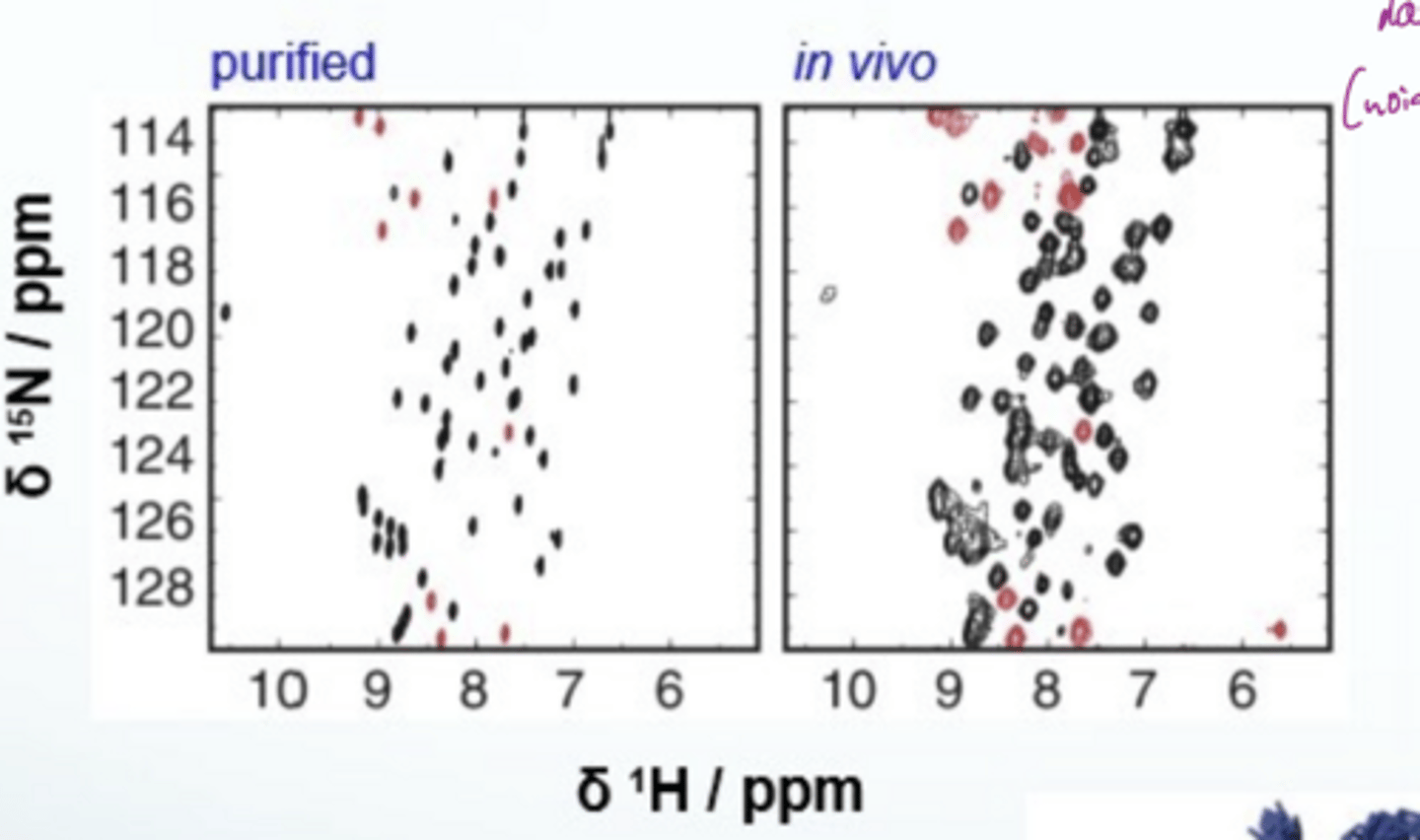
What does 2D NMR spectra look like
in vivo spectra are hard to interpret since there is lots of noise
What is isothermal titration calorimetry
measures heat exchange from induced ligand-protein binding
the guest (ligand) solution is titrated into a solution of the host (protein) in the sample cell
power is supplied to maintain the reference and sample cell temperature
gives info about delta H and Ka, and therefore delta S
can be used to determine the binding stoichiometyr
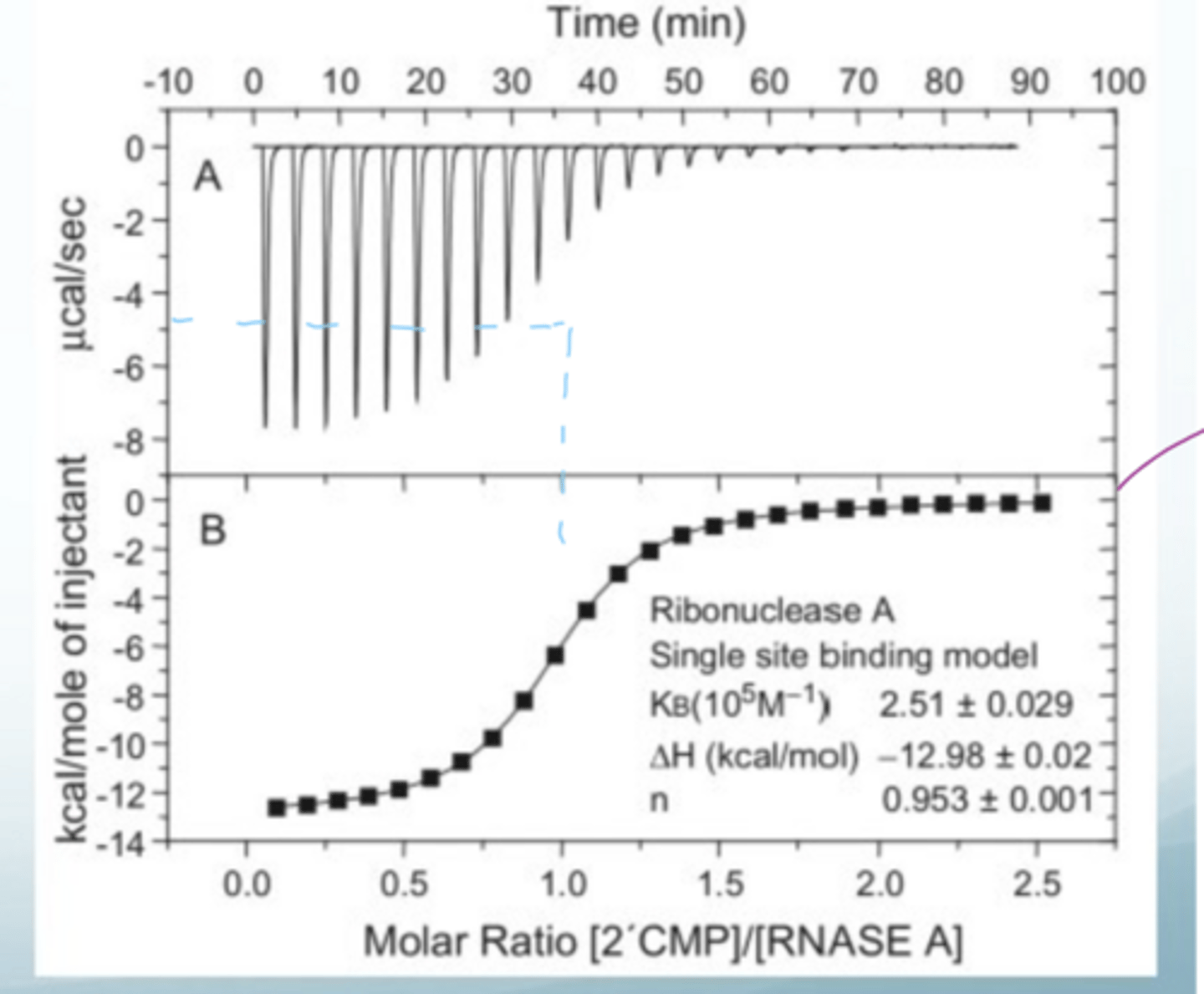
What does the data look like from isothermal titration calorimetry and how to interpret
the first peak is unusual since there are often bubbles in the syringe
before the first injection there is no ligand at all, so binding is small since [ligand] is small but [protein] is massive
if you integrate under the peaks, you get the amount of heat released per injection in kilo J (the bottom graph)
the bottom graph is in kJ and you look half way in the curve (-5) and go down to 1, which means there is 1:1 binding
the control would be to inject a buffer instead of a ligand
![<p>the first peak is unusual since there are often bubbles in the syringe</p><p>before the first injection there is no ligand at all, so binding is small since [ligand] is small but [protein] is massive</p><p>if you integrate under the peaks, you get the amount of heat released per injection in kilo J (the bottom graph)</p><p>the bottom graph is in kJ and you look half way in the curve (-5) and go down to 1, which means there is 1:1 binding</p><p>the control would be to inject a buffer instead of a ligand</p>](https://knowt-user-attachments.s3.amazonaws.com/7b40c9fd-4771-48d1-a946-bc25e09ef3e2.png)
What could they ask you in exam about isothermal titration calorimetry
BE AWARE: COULD BE IN KILO CALORIES RATHER THAN KILO JOULES
integrate data to get the sigmoidal curve
at low/ medium conce, the most heat is produced (maximal binding)
at high concs all binding sites are saturated
if know the Kd, know delta G and therefore delta H and delta S
Also know the stoichiometry
LIMITATION: binding must be strong enough to produce enough heat
Interpret this example for ITC
at -5 kcal, the molar ratio is 1, so the stoichiometry is 1:1 (bottom graph)
quantitative data matches qualitative data well
What is induced circular dichroism
some samples e.g. berenil are achiral so no CD on its own
Bind to DNA (chiral) to get a CD signal as it has benefitted from being in a chiral environment
flat line in top graph= no binding
see signals as DNA conc increases
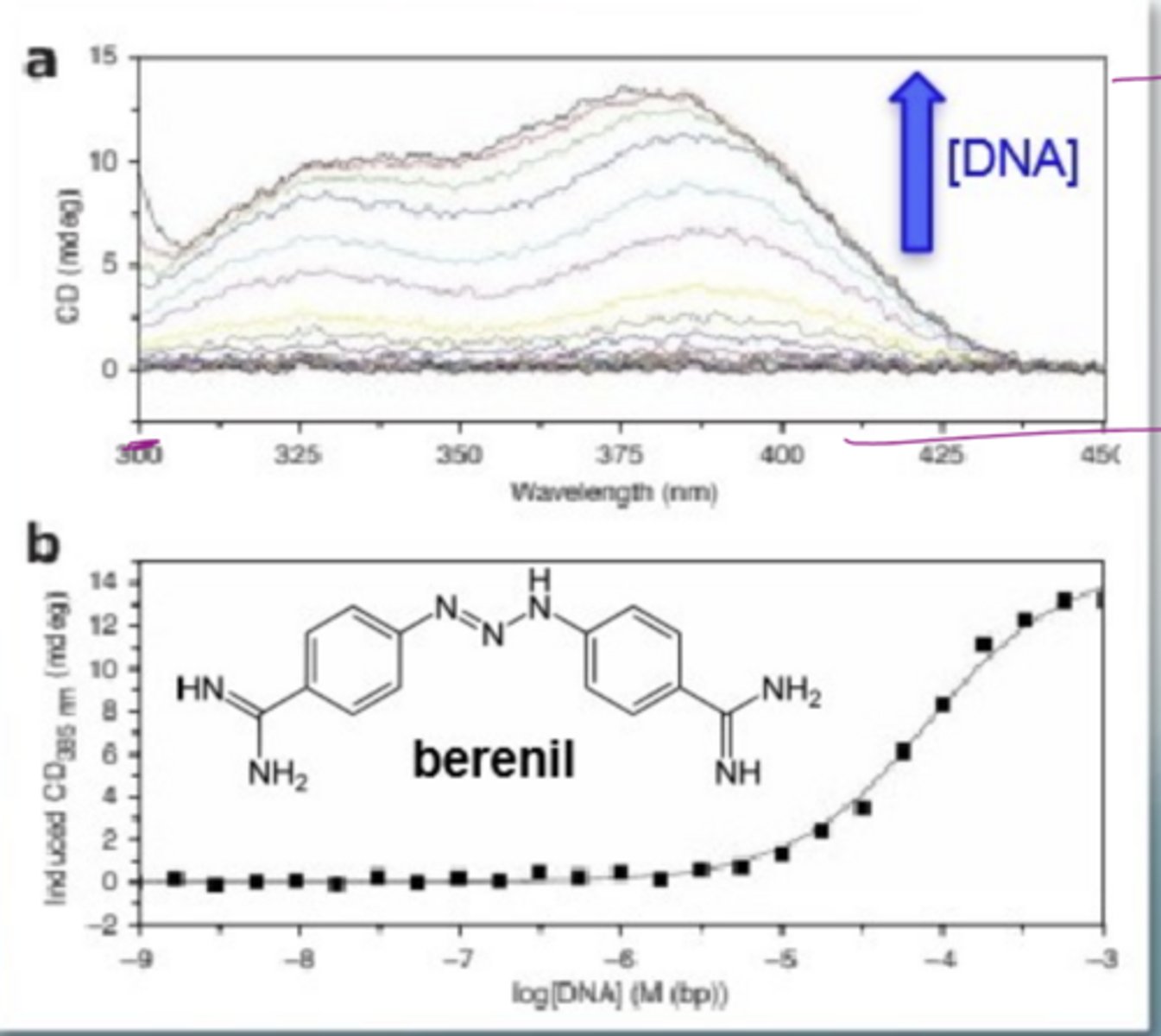
How does berenil bind to DNA
it is an intercalator
at pH 7, both ends of berenil are charged so can interact with the negative charge on the DNA phosphate backbone
it is planar so can slide inbetween the stack of bases
it can either strengthen the interactions (and increase Tm) or disrupt the structure and lower the Tm (like adding salt to water)
can use the changes in Tm to determine binding
What are sphingolipids
there is a fatty acyl group attached to the amine of the amide
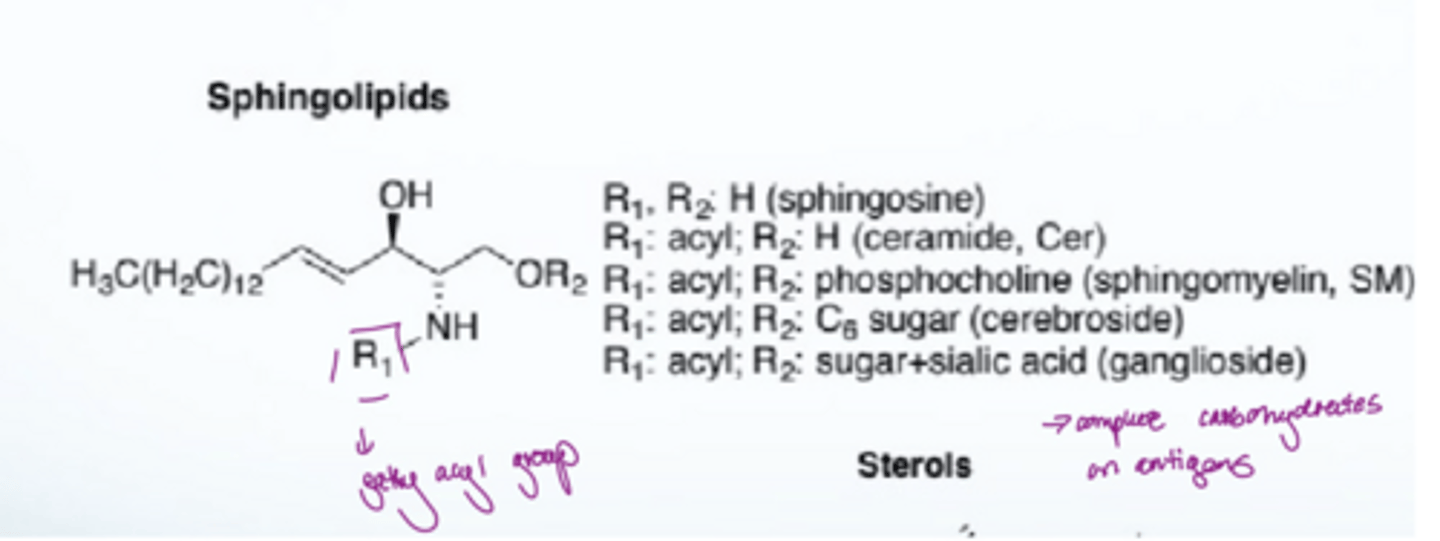
What are sterols
They are lipids that do not consist of glycerol or fatty acids. They are composed of four connecting rings of carbon and hydrogen, with a polar OH group
the polar OH group means sterols cannot dissolve in water
they are responsible for regulating the fluidity of the lipid menbrane
What are liposomes
closed-lipid bilayer spheres
they are used as models of the phospholipid bilayer but they are much simpler
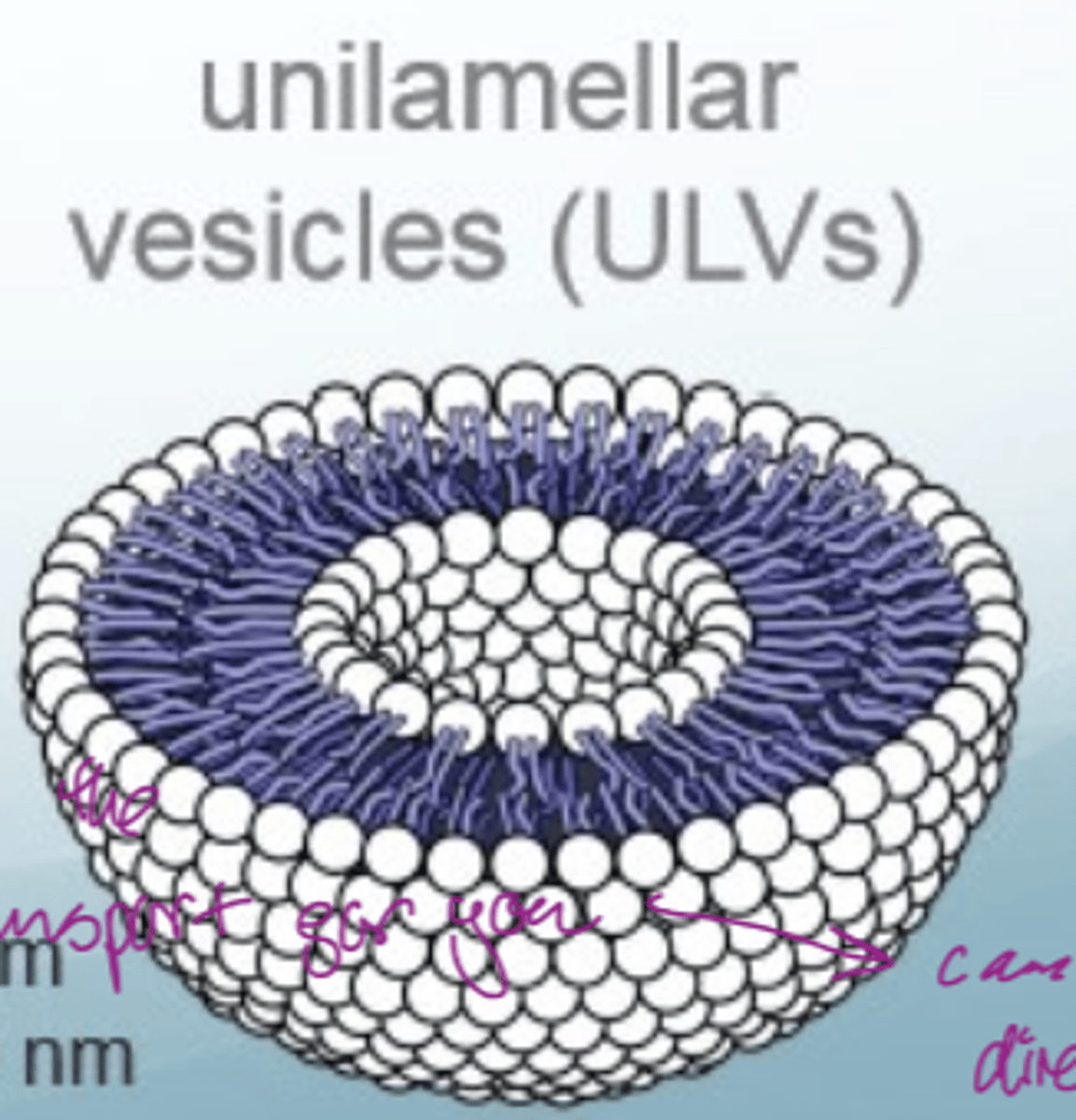
What are cell membranes made out of
a complex mixture of sterols, (glyco)lipids, (glyco)proteins
fluid mosaic model
have transport mechanisms since need to transport polar molecules into cells without touching the inside of the bilayer (cannot directly diffuse across)
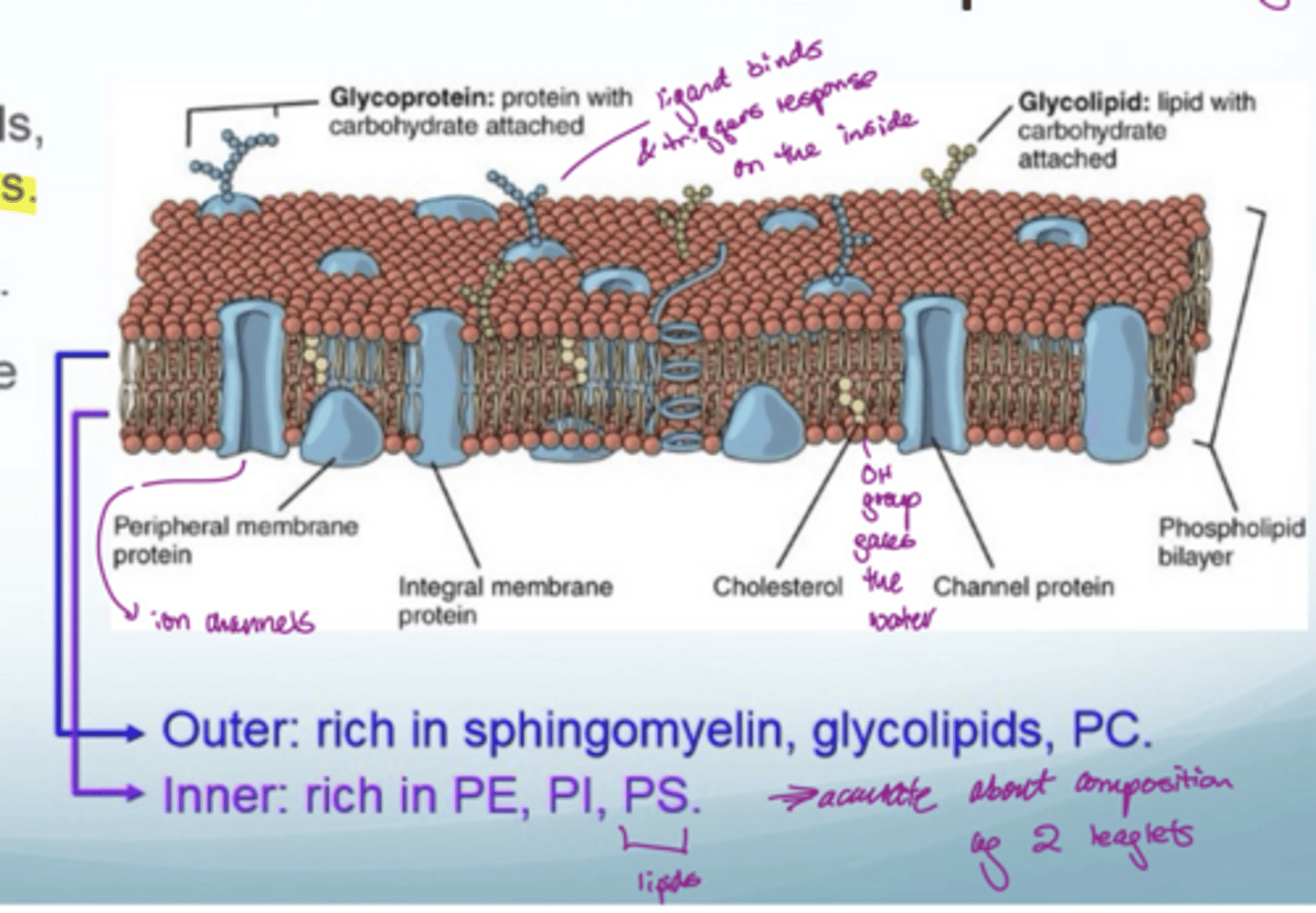
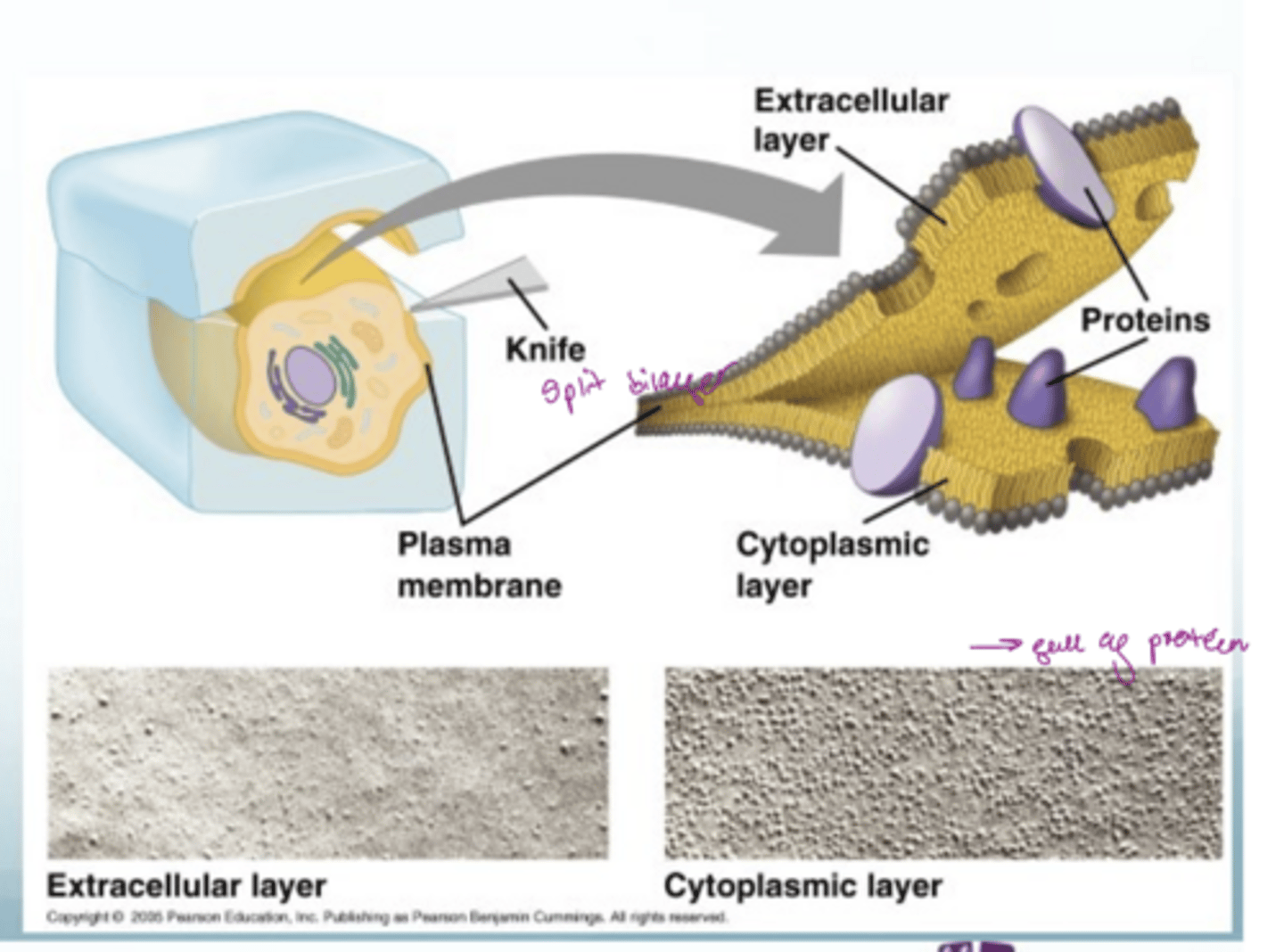
How can you separate the lipid bilayer for analysis
use freeze fracture
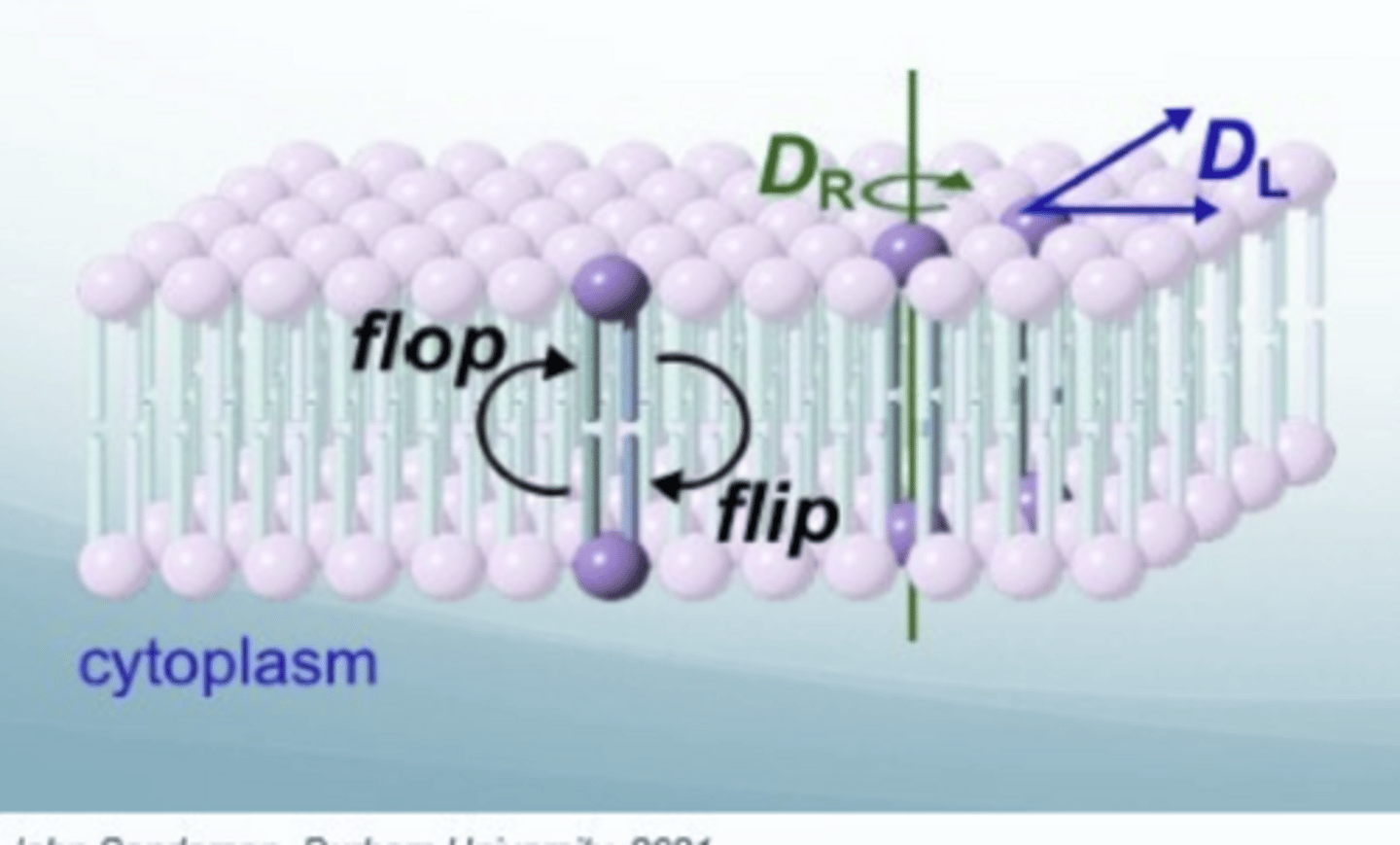
What are the three types of diffusion across the cell membrane
Rotational diffusion: D(R)
Lateral diffusion: D(L) - diffuse in the plane of the bilayer
Transbilayer diffusion: flip (move to the inside) /flop (move to the outiside) - phospholipid moves between leaflets
what are the typical rates of D(R) for proteins and lipids
proteins: 10^4 s^-1
lipids: 10^6-10^7 s^-1
what are the typical rates of D(L) for proteins and lipids
proteins: 1 - 5 micro m^2 s^-1
lipid: 20-30 micro m^2 s^-1
important for cell signalling (membrane remodels in response to cellular events)
what are the typical rates of flip flop for proteins and lipids
proteins do not flip flop
lipids flip flop slowly (every 200 - 1500 min)
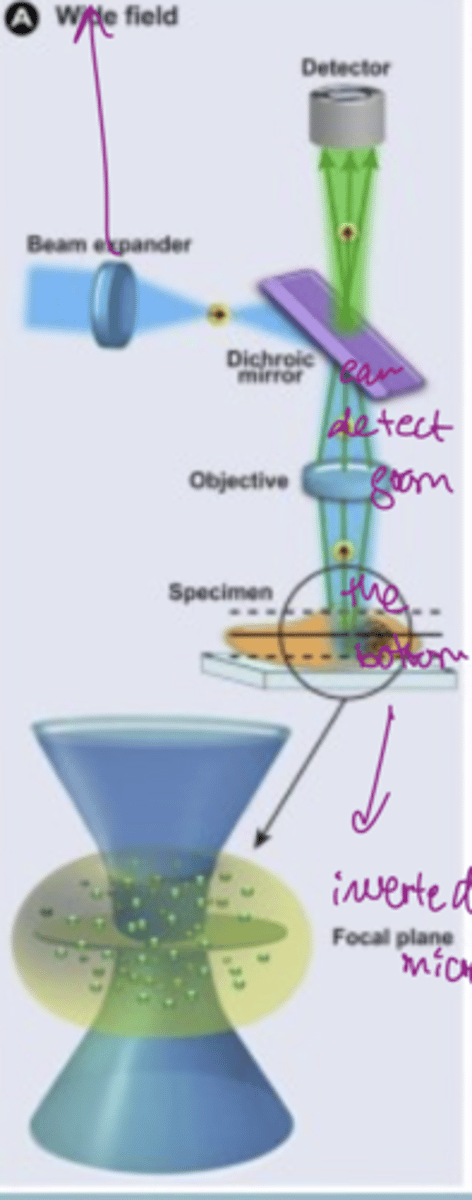
How does a dichroic mirror work
only transmits fluorescence (some wavelengths) and reflects others
the focal volume determines the resolution
can detect from the bottom of the slide (inverted microscopy)