BIS 2A Midterm 2
1/34
Earn XP
Description and Tags
yea idk
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
35 Terms
Glycolysis
Takes place in cytoplasm, is a coupled reaction that gives phosphate to glucose and turns 1 molecule of glucose into 2 molecules of pyruvate.
Fermentation (Circumstantial)
Occurs if no ETC or oxygen. Allows glycolysis to continue by regenerating NAD+.
Pyruvate Oxidation
Converts pyruvate into acetyl-CoA.
Citric Acid Cycle/Kreb’s Cycle
Finishes breaking down carbon and loads up electron carriers.
Electron Transport Chain (ETC)
Builds proton concentration gradient. Takes electrons from NADH and FADH2 and passes them along protein complexes.
ATP Synthase
Occurs in inner mitochondrial membrane. Uses H+ gradient like a turbine to make ATP from ADP + Pi. This is where most ATP is made.
Cellular Respiration Cycle
Glycolysis Splits Glucose > Pyruvate Oxidation makes Acetyl-CoA > Kreb’s Cycle loads NADH/FADH2 > ETC Builds H+ Gradient > ATP Synthase makes ATP
Non-Cyclic Photophosphorylation
Main oxygenic pathway in plants. Electrons start in water and end in NADPH, with the output of ATP, NADPH, and oxygen.
Cyclic Photophosphorylation
Electrons start and end in chlorophyll. Only makes ATP.
(Light-dependent reactions) 1. Light hits Photosystem II
Light excites electrons in chlorophyll, gives electrons more energy.
(Light-dependent reactions) 2. Water is Split
Water is broken apart to replace lost electrons, making O2, electrons/H+ in the process.
(Light-dependent reactions) 3. Electron Transport Chain
Excited electrons move through chain of proteins in membrane, building a proton gradient as energy is released and pumps H+ into membrane.
(Light-dependent reactions) 4. ATP Synthase makes ATP
Photophosphorylation occurs as ATP synthase uses H+ flowing back through it to make ATP.
(Light-dependent reactions) Step 5. Photosystem I makes NADPH
Electrons get re-energized in Photosystem I and are used to reduce NADP+ into NADPH. NADPH is used later to help make sugar.
Calvin Cycle
Occurs in chloroplast stroma and uses CO2, ATP, and NADPH to make sugar. It is light independent.
(Calvin Cycle) 1. Carbon Fixation
CO2 attached to RuBP by rubisco, forming 3-PGA. Carbon gets captured and put into organic molecules.
(Calvin Cycle) 2. Reduction
ATP and NADPH are used to turn 3-PGA into G3P. Cells spend energy to make more useful carbon molecule.
(Calvin Cycle) 3. Regeneration
Some G3P is used to rebuild RuBP so the cycle can continue.
Photosynthesis Basics
Light reactions make ATP and NADPH, then Calvin Cycle use ATP and NADPH to turn CO2 into sugar.
Nucleotide contains:
Pentose sugar, nitrogenous base, and phosphate group.
Nucleoside contains:
Pentose sugar and nitrogenous base.
DNA is a double helix with two strands that run in opposite direction, this is called:
Antiparallel.
DNA is read and built in the ____ direction:
5’ » 3’
The two strands of DNA are held together by:
Hydrogen bonds.
DNA strands are built by making:
phosphodiester bonds between nucleotides. This bond forms between 3’ OH on growing DNA strand and 5’ phosphate of incoming nucleotide.
DNA can only be built by adding to the:
3’ end.
DNA Replication:
Initiation, Elongation, and Termination.
Initiation of Replication is done by:
Helicase, breaks the bonds between two strands while Topoisomerase prevents supercoiling of the strands as it’s unzipped.
Elongation of Replication
Primase lays down the starter piece “primer” and DNA polymerase III adds new nucleotides to the primer in 5’ » 3’ direction.
DNA polymerase I goes back and replaces primer with DNA nucleotides and Ligase seals the games between Leading Strand fragments.
Leading Strand:
is made continuously.
Lagging strand:
is made in pieces called Okazaki Fragments. Ligase seals the gaps between them to make it one continuous strand.
Termination of Replication
Telomerase extends the end of the chromosome by adding extra repetitive DNA to the 3’ overhang (this prevents chromosomes from shrinking after each replication. Then Primase and DNA Polymerase III can fill the complementary strand.
DNA Replication Summary
Helicase unzips both strands of DNA, Topoisomerase prevents unzipped strands from twisting, Primase sets primers, DNA Polymerase III builds new DNA by adding nucleotides to primers, DNA Polymerase I replaces primers with nucleotides, Ligase seals Okazaki Fragments, and Telomerase
Exergonic Reactions
Spontaneous, with a negative deltaG.
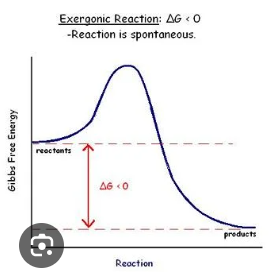
Endergonic Reactions
Non-Spontaneous, with a positive deltaG.