GENE221 - Module 3 - The accessory genome – how mobile genetic elements shape bacterial evolution
1/56
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
57 Terms
Core genome
Located on chromosome(s)
Genes including essential genes conserved in all members of a species
Accessory genome
Located on plasmids, genomic islands, transposons, lysogenic phage
Non-essential genes of variable occurrence in a bacterial species
Acquired through horizontal transmission
Pangenome
Core genome + Accessory Genome
Replicon
Capable of autonomous replication i.e plasmids
What are the five things a plasmid must do?
Replicate
Keep host happy
Segregate
Keep host under control
Spread
How does a plasmid keep its host happy?
By limiting the amount of copy numbers, depending on the size of the plasmid. Larger plasmids take more resources to be replicated. The plasmid has an interest in keeping the host cell alive.
What is the general principle of plasmid replication?
Plasmid DNA replication controlled by plasmid encoded inhibitor that acts at oriV
As cell size increases, inhibitor concentration decreases and plasmid replication initiated
Replication results in further copies of inhibitor gene and more inhibitor which limits plasmid replication again
oriV
The origin of "vegetative" replication is where DNA replication begins. oriV encodes 2 different noncoding RNAs (RNA I, RNA II) and the DNA origin of replication
RNA II (ColE1 plasmid)
non-coding RNA (600nts) encoded by oriV, that binds to oriV and allows initiation of DNA replication by acting as a primer
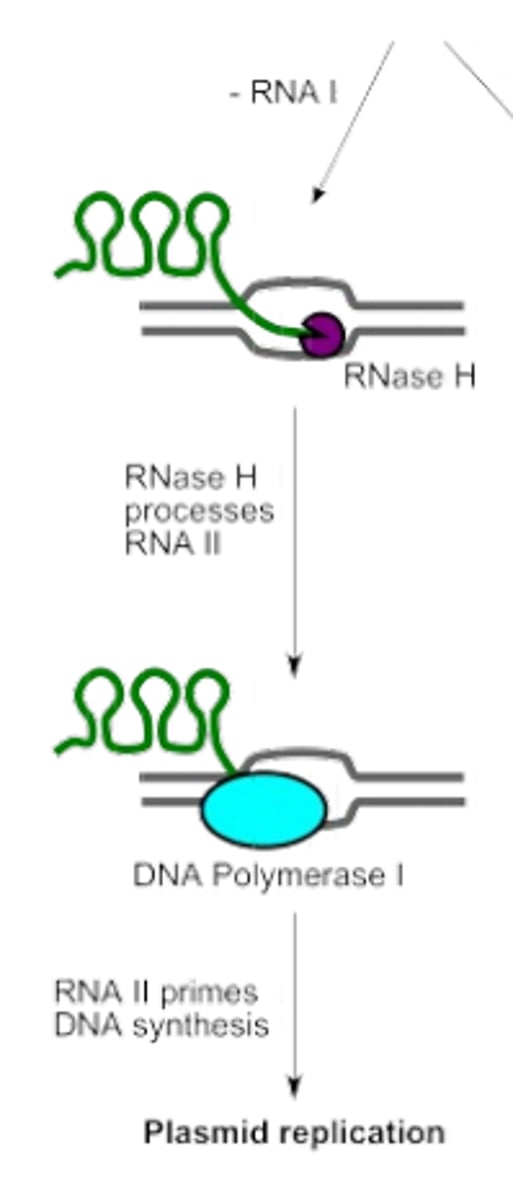
RNA I (ColE1 plasmid)
anti-sense non-coding RNA (108nts), that in the presence of rop protein complementarily binds to RNA II, and disables DNA replication.
rom/rop
inhibitor protein needed for copy number control
ColE1
High copy plasmid (15-20 copies inside the cell)
RepF1A
Low copy plasmid (1-2 copies inside the cell)
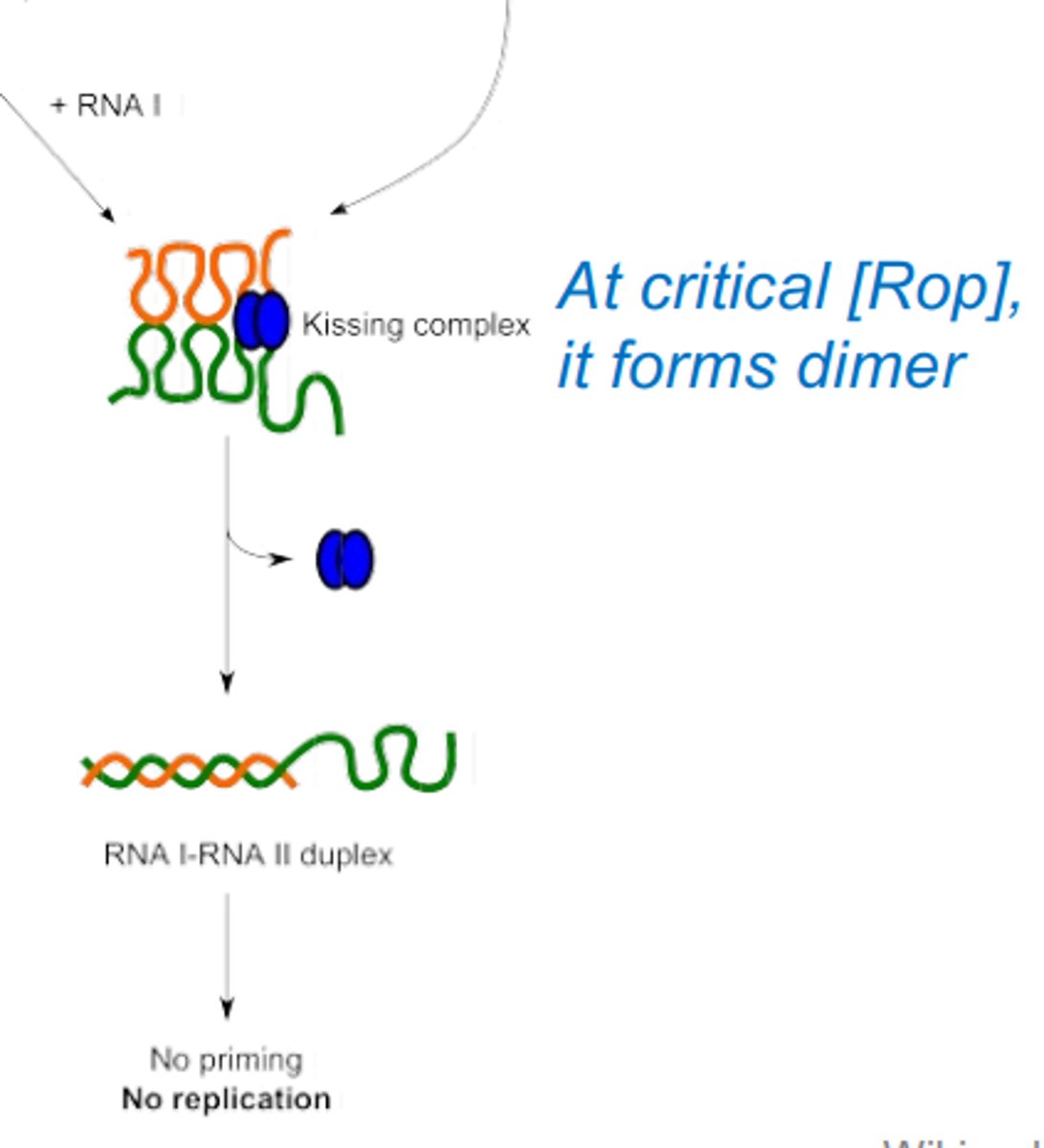
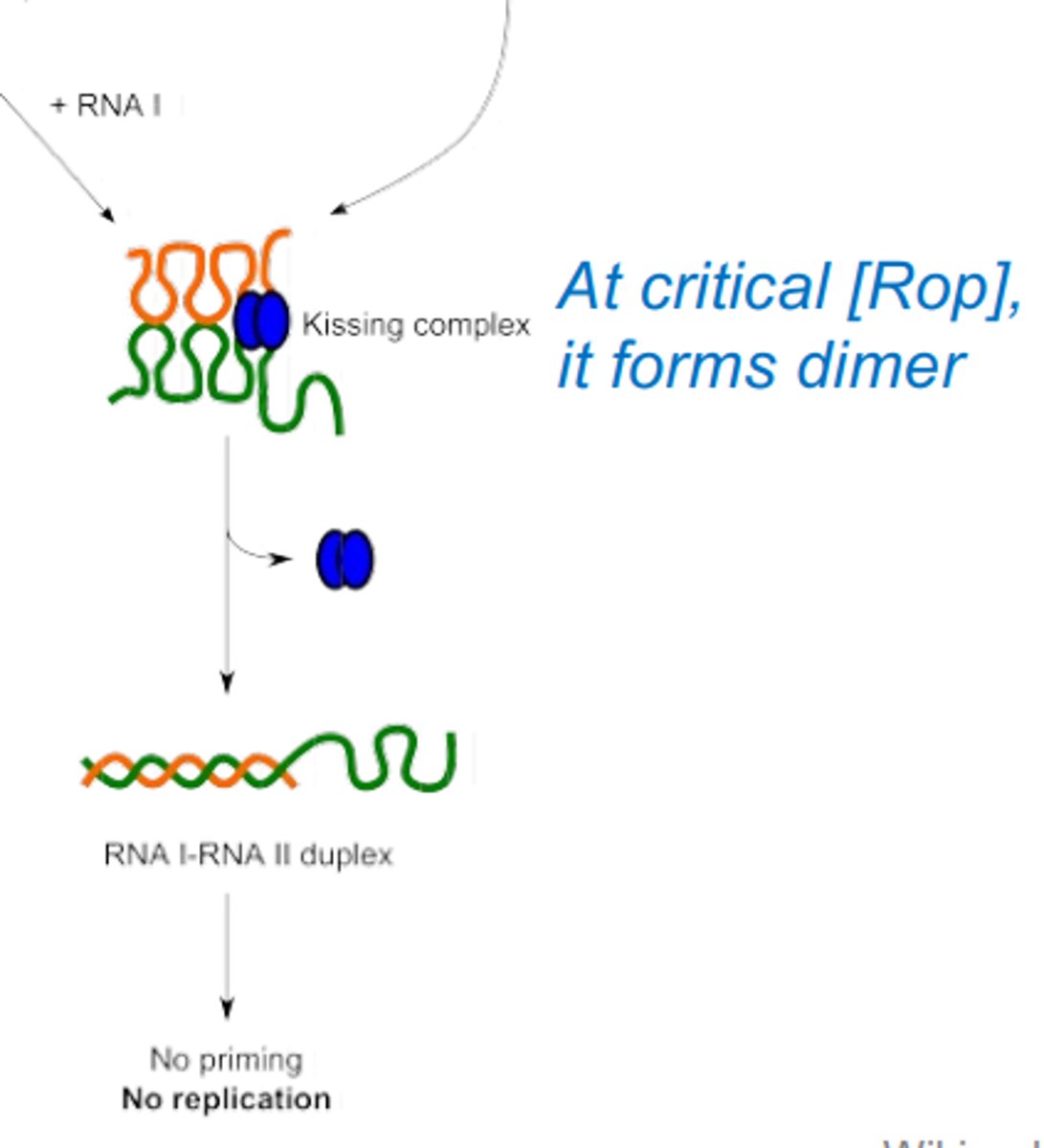
How does the rop protein inhibit replication of the ColE1 protein?
At a critical concentration rop protein provides anti-sense regulation. Forms a kissing complex between RNA I and RNA II, by creating a dimer. This is an RNA duplex.
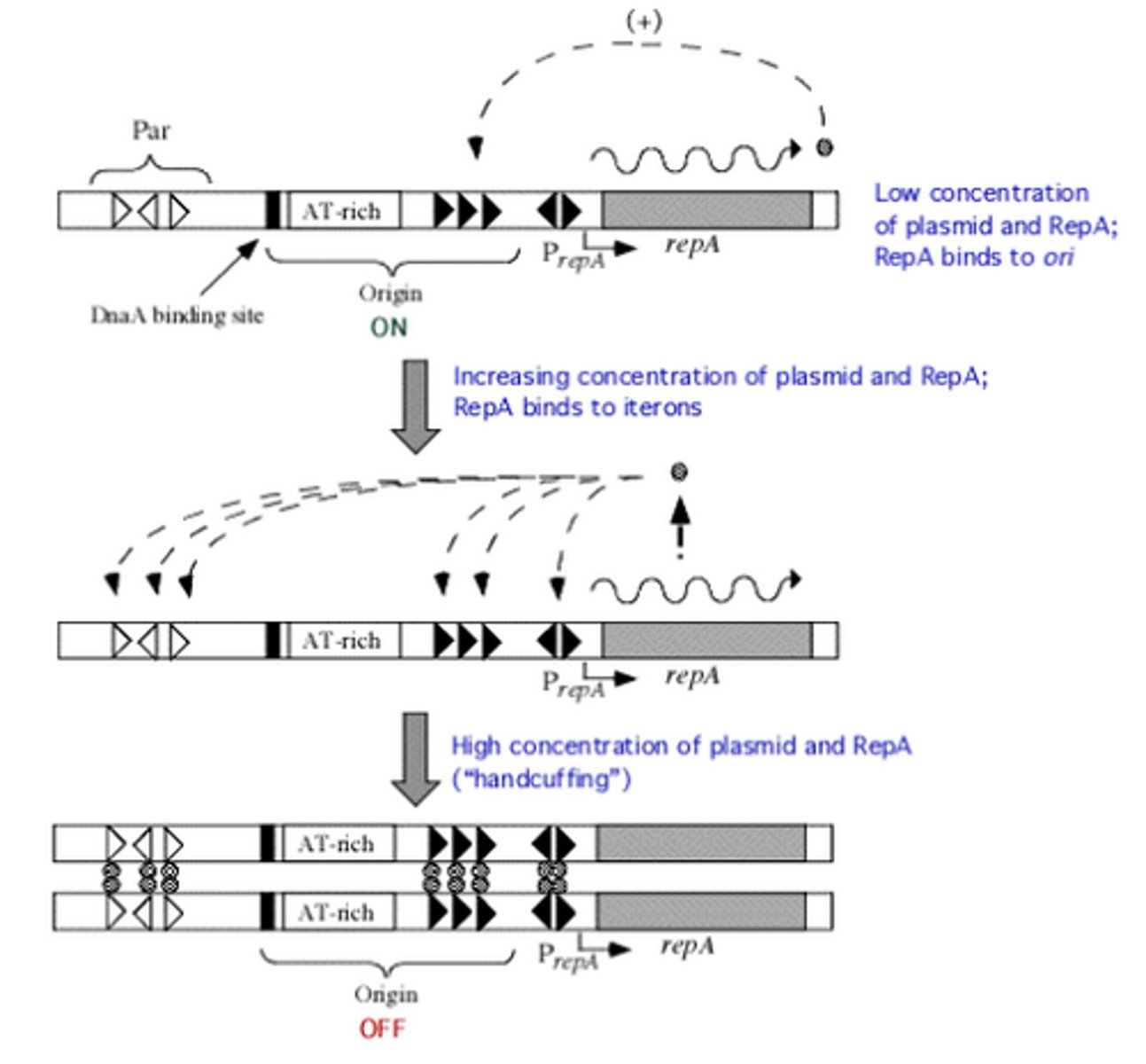
Iterons
repeated sequences in DNA
RepA
Protein that binds to iterons on OriV and allows for DNA replication. Causes increased expression of RepA. Then once RepA concentrations increase, binds to other iterons, and 'handcuffs' plasmids together, stopping replication.
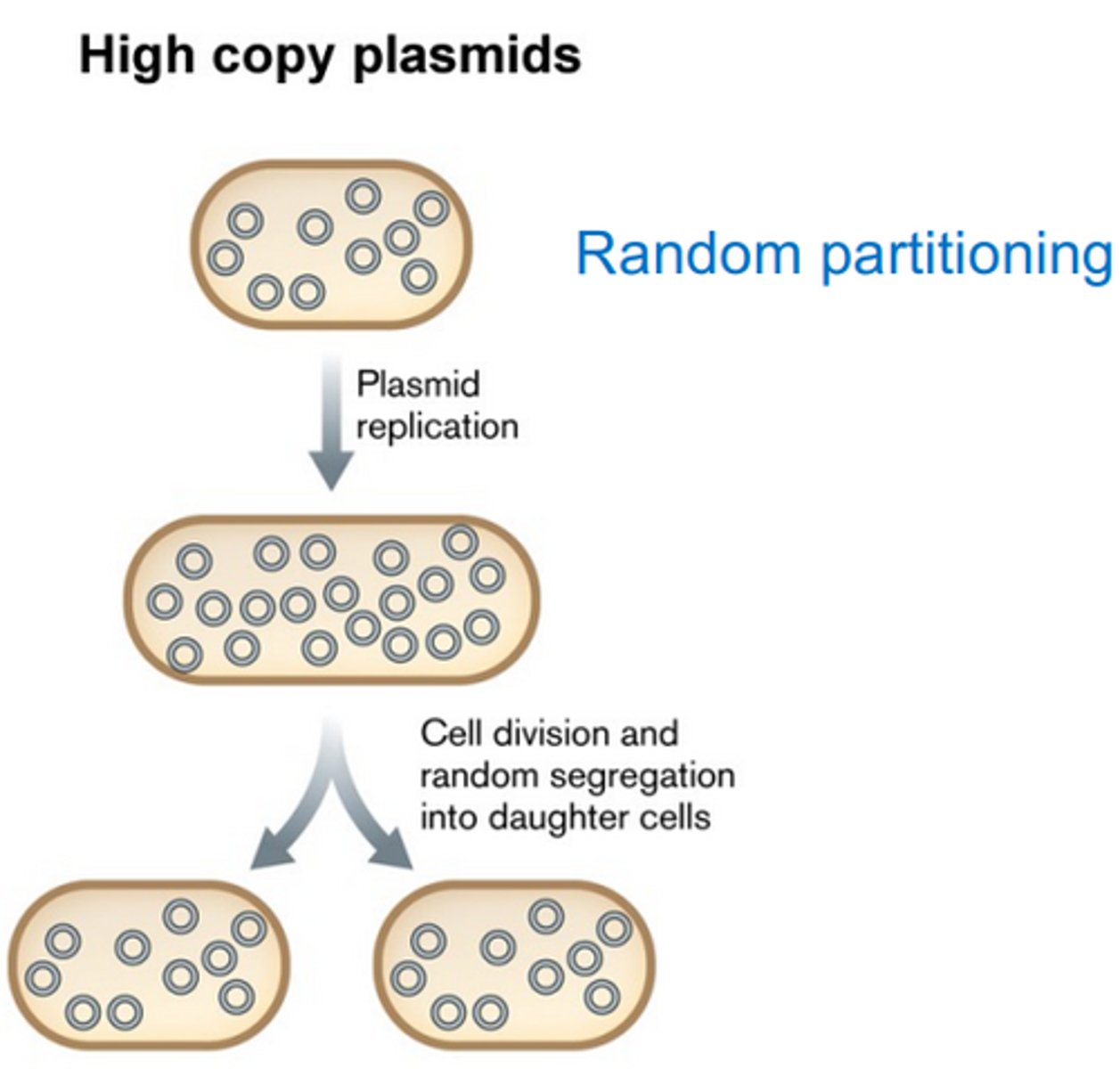
Partitioning in high copy plasmids
Done randomly, as long as copy number is higher than 15, there is a one in a million chance of a daughter cell to not contain the plasmid.
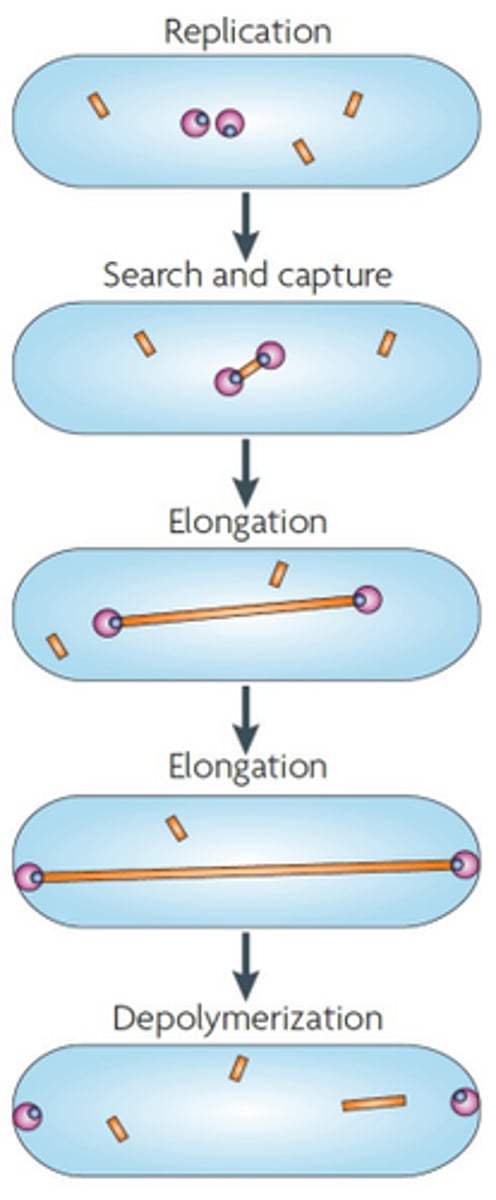
Partitioning in low copy plasmids
An actin-like protein named ParM attaches to ParR which is attached to parS, which is a centromere-like region in the plasmid. Then ParM polymerises, and lengthens, separating the plasmids to either pole of the cell, ensuring that when the cell divides, each daughter will have a plasmid copy. Once the plasmid reaches the poles, the ParM filament depolymerises.
Why do we call plasmids molecular parasites?
Because they typically do not confer any advantage to their host.
ParM
Protein involved in partitioning in low copy plasmids. Forms an actin-like filament via polymerisation. Binds to the ParR protein. Depolymerises once the plasmid reaches the pole of the cell. Yellow rectangle in diagram.
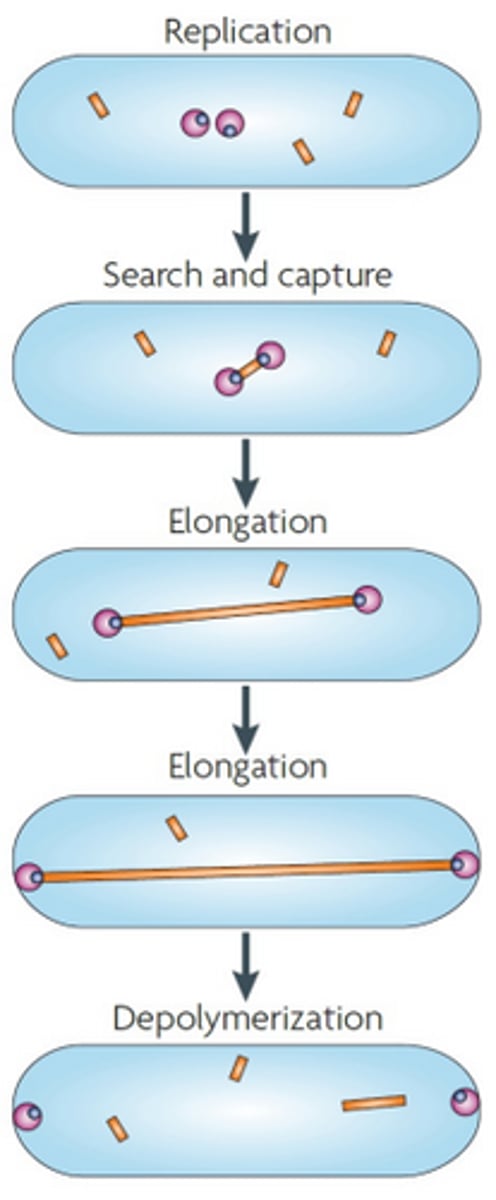
ParR
Protein involved in partitioning low copy plasmids. Binds the parS region on the low copy plasmid, and then is bound by ParM protein. Blue circle in diagram.
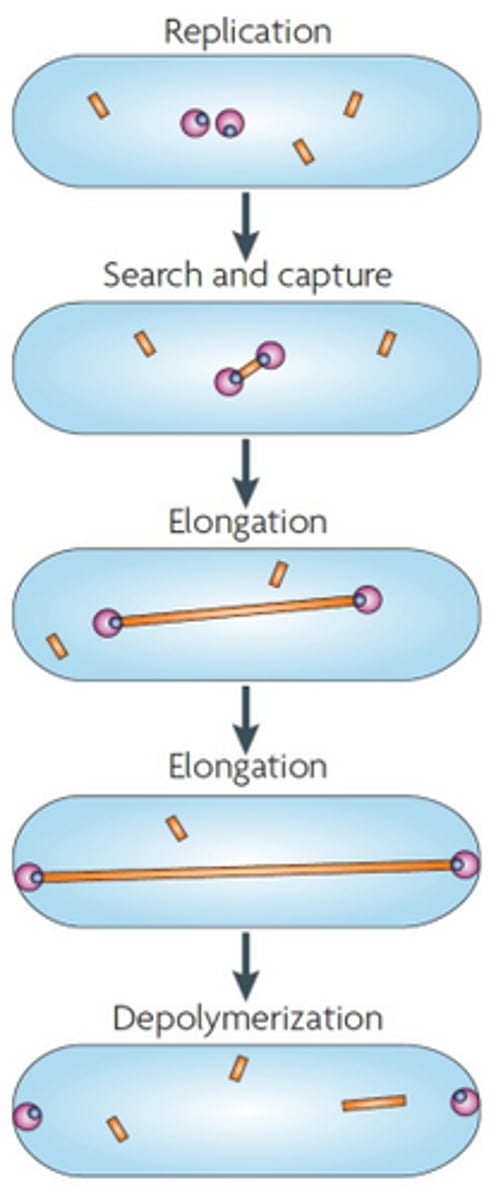
parS region
centromere-like region on the low copy plasmid that is bound by ParR.
Multimeric plasmid
Because plasmids have identical DNA sequences, they can undergo homologous recombination, and form larger multimeric plasmids.
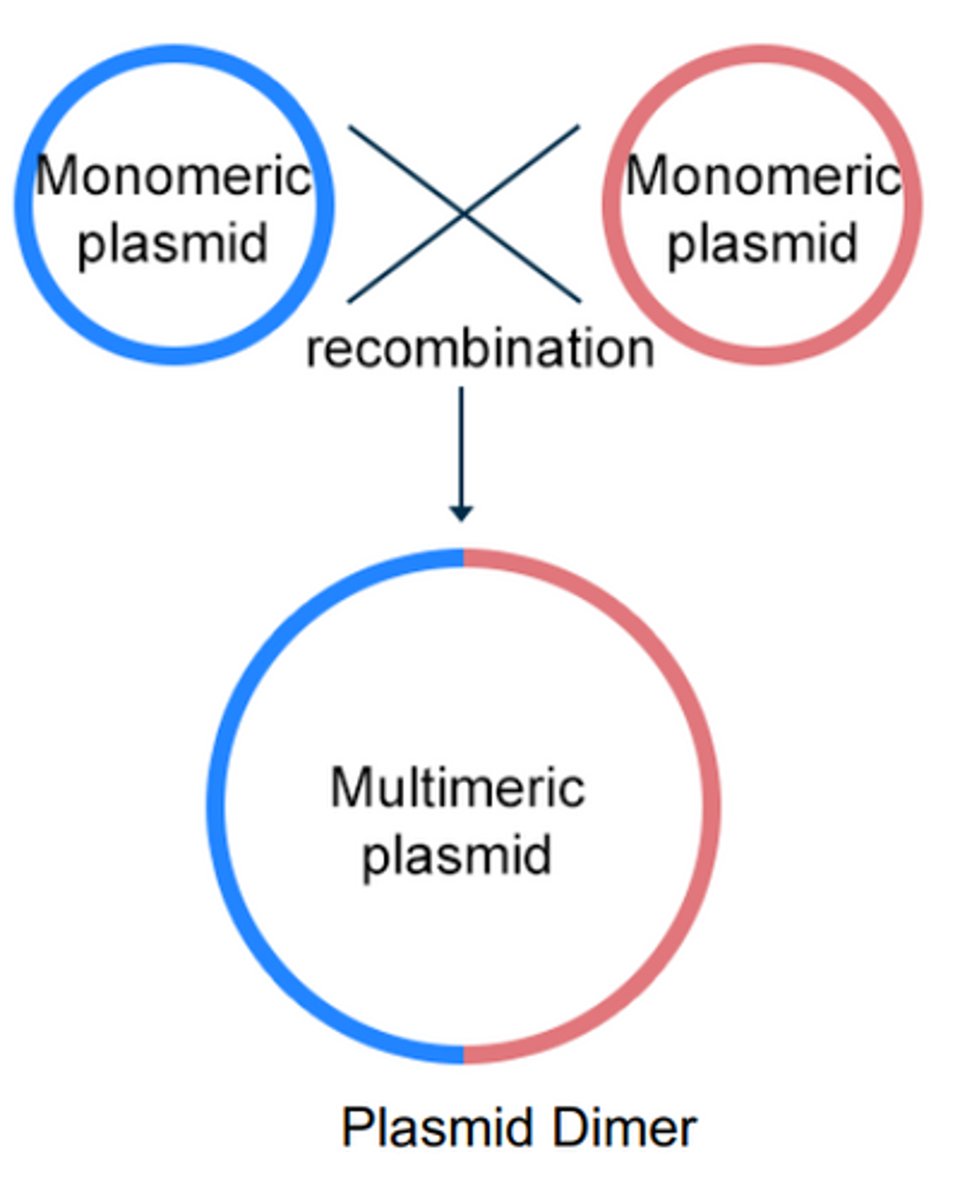
Problem with oligomerization for plasmids
Larger multimeric plasmid means the plasmid is more likely to be lost when the cell divides, i.e there will be one daughter cell with and one without.
Differences between F and ColE1 plasmid when resolving multimers
Both plasmids have site-specific recombination available, however the F plasmid encodes its own recombinase (ResD acts at fcr location), whereas ColE1 requires XerCD (acts at cer) which is encoded for by the host cell chromosome. ResD and XerCD act on the specific sites to resolve multimers back into monomers.
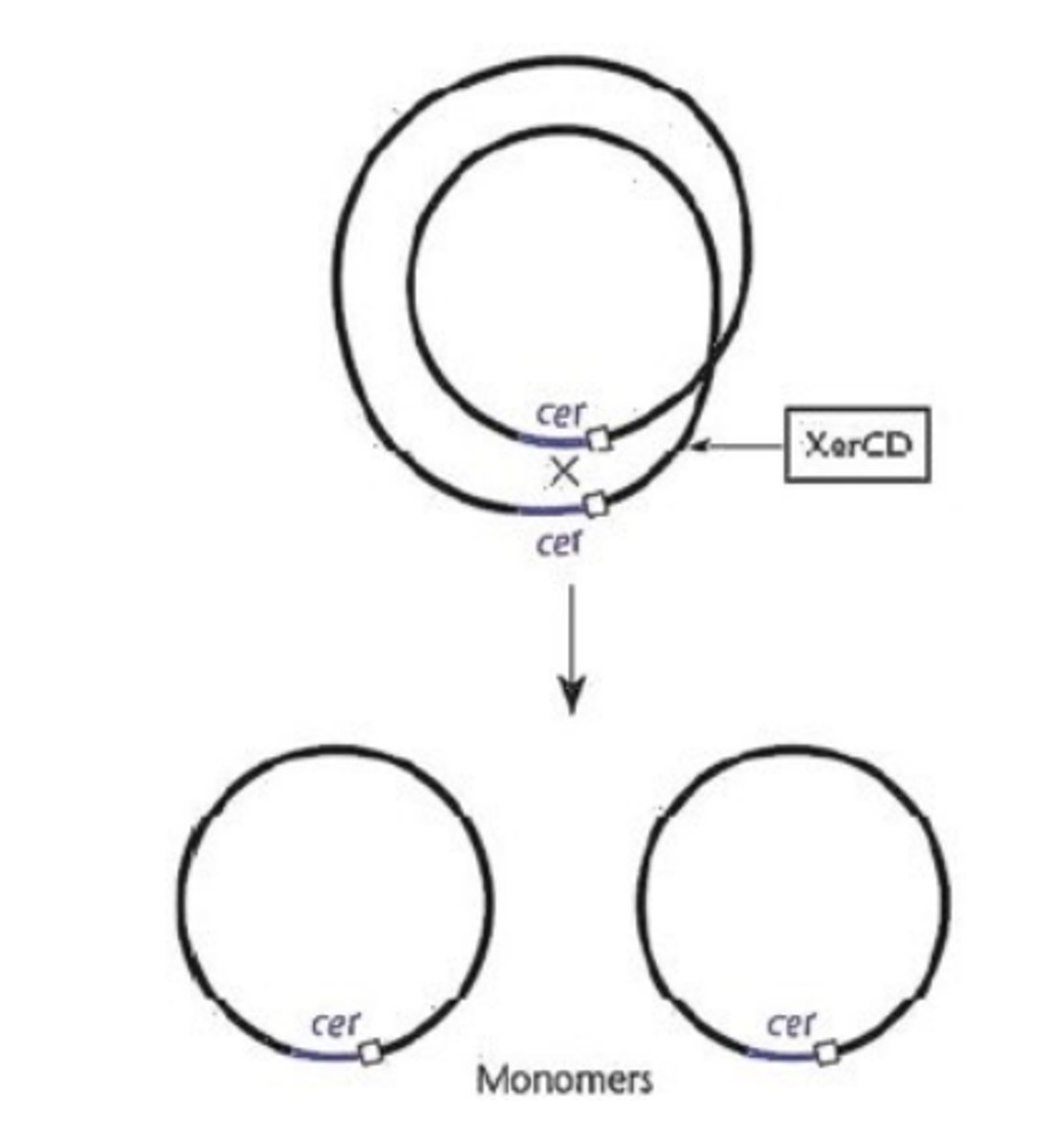
Why do host cells have recombinase enzymes (e.g XerCD in e.coli)?
Because when the chromosome is being replicated it can homologously recombine with its copy, which the cell would not want.
What is the problem that plasmids solve through post-segregational killing systems?
Cells without the plasmid have a fitness advantage, and would outcompete infected cells. Therefore the plasmids kill all cells that are not infected through post-seg killing systems.
Toxin-antitoxin system
Addiction system, a form of post-segregational killing. Plasmid encodes a stable toxin and an unstable antintoxin. When both are present the toxin is nullified. However if the cell somehow loses the plasmid, the toxin remains and the unstable antitoxin degrades, leading to cell death.
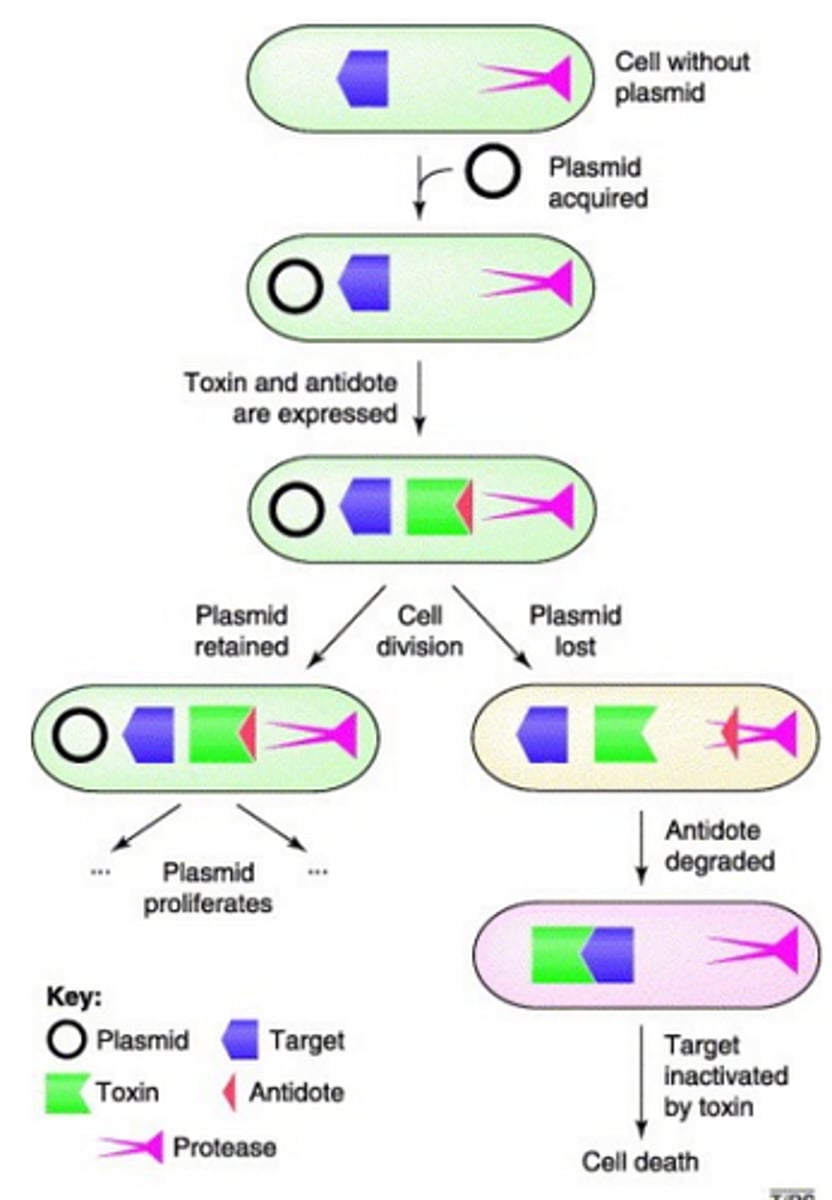
ColE1 plasmid post segregation killing system (genes and function)
Two genes, imm and ColE1. ColE1 gene synthesises ColicinE1 which kills bacteria, and imm encodes for immunity to ColicinE1.
What are the F plasmid post segregation killing systems (2)
hok/sok and ccdAB
hok gene
Host-killing. Encodes for killer peptide, leading to membrane depolarisation, the mRNA for the hok protein lasts for 20 minutes.
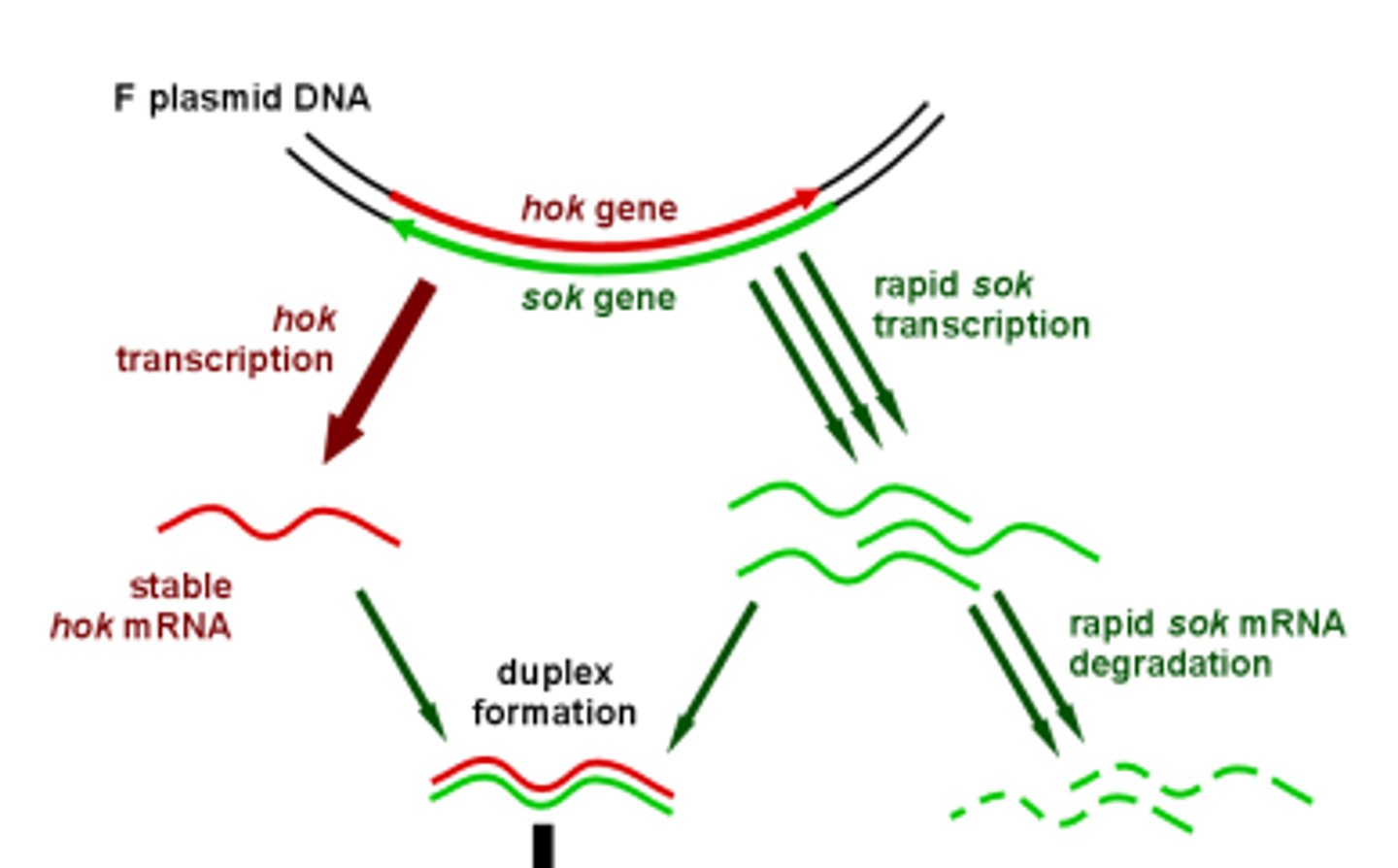
sok gene
Supressor of host killing. Antisense RNA of the hok/sok gene, and binds to the hok mRNA. Unstable, half life of about 5 minutes.
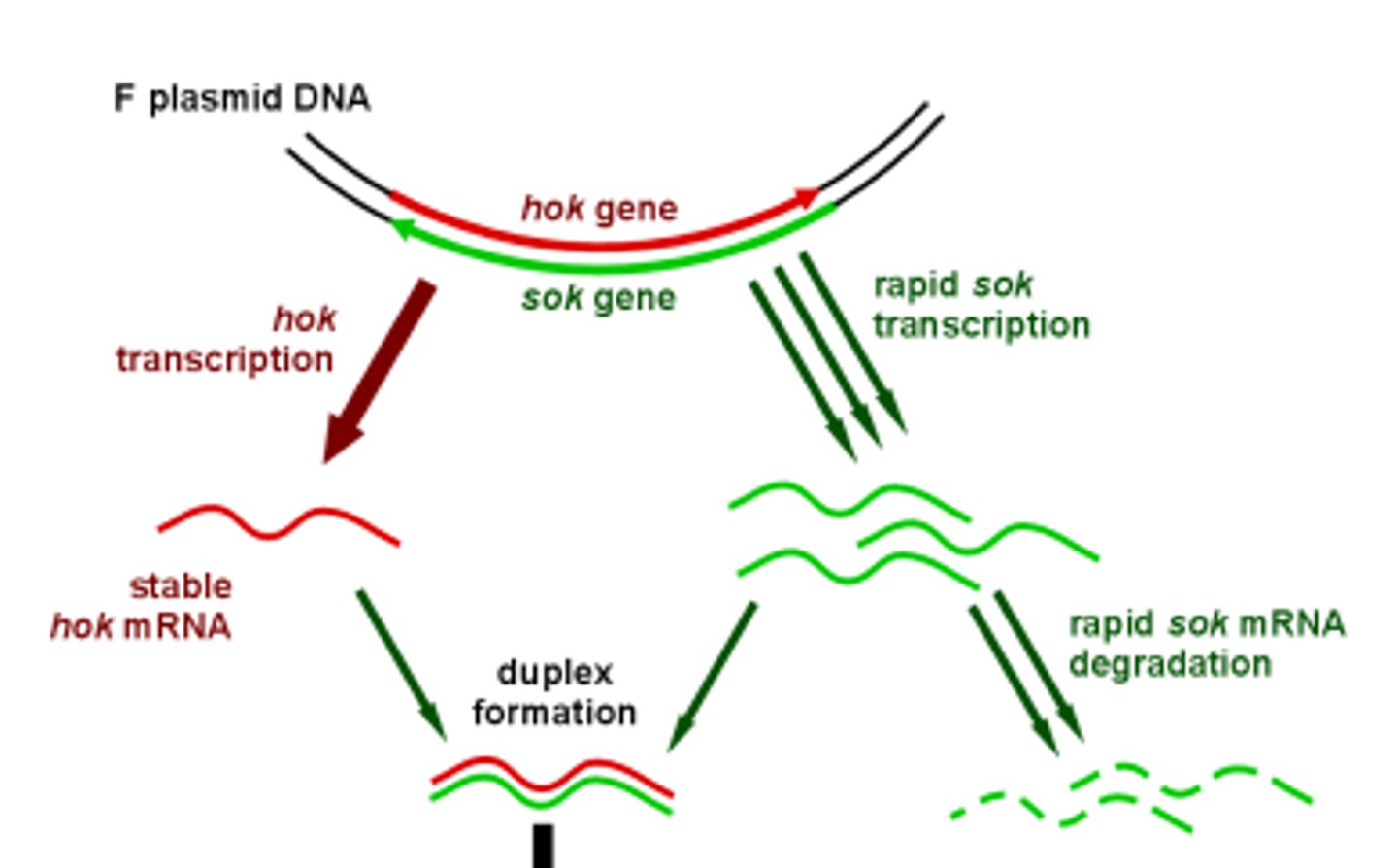
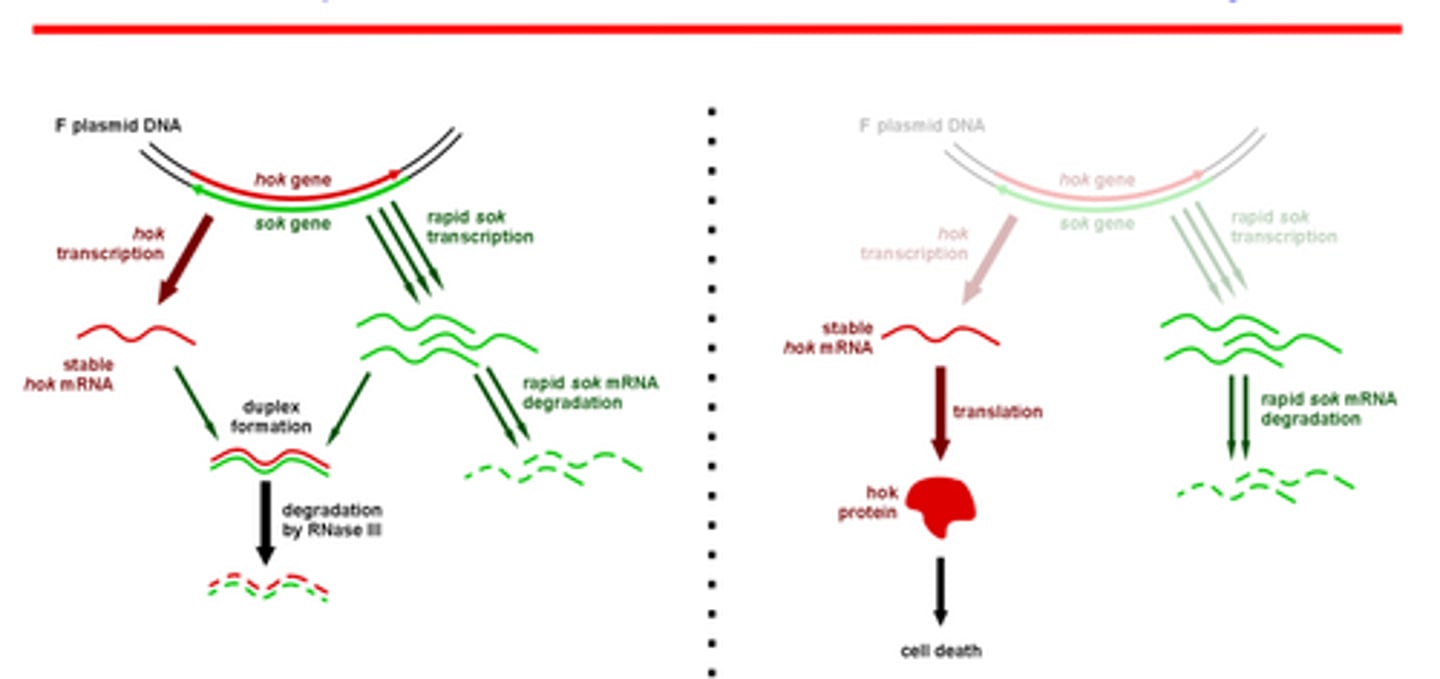
How does the F plasmid use hok/sok to kill cells without the F plasmid?
The hok mRNA is bound by the sok RNA and forms a duplex which is then degraded by Rnase III. Because hok is more stable than sok, if the host loses the plasmid, the sok RNA will degrade, and allow remaining hok mRNA to be translated into hok killer protein.
tra genes
Encode the mating pore and DNA mobilisation functions for conjugative transfer in the F plasmid.
mob gene
Relaxase that nicks DNA at the oriT of the ColE1 plasmid, allowing for mobilisation.
How does ColE1 transfer by conjugation?
It needs to use machinery produced by conjugal plasmids like the F plasmid to then mobilise and become conjugated.
Lysogen
Host cell with bacteriophage DNA integrated into its chromosome.
Temperate bacteriophages
Phages that can operate between two lifestyles, lysogenic or lytic.
Lambda phage integration
Phage uses attP site on the phage DNA and attB site on the bacterial chromosome, and then uses λ integrase to combine the prophage and the chromosome. the attP and attB sites become attL and attR.
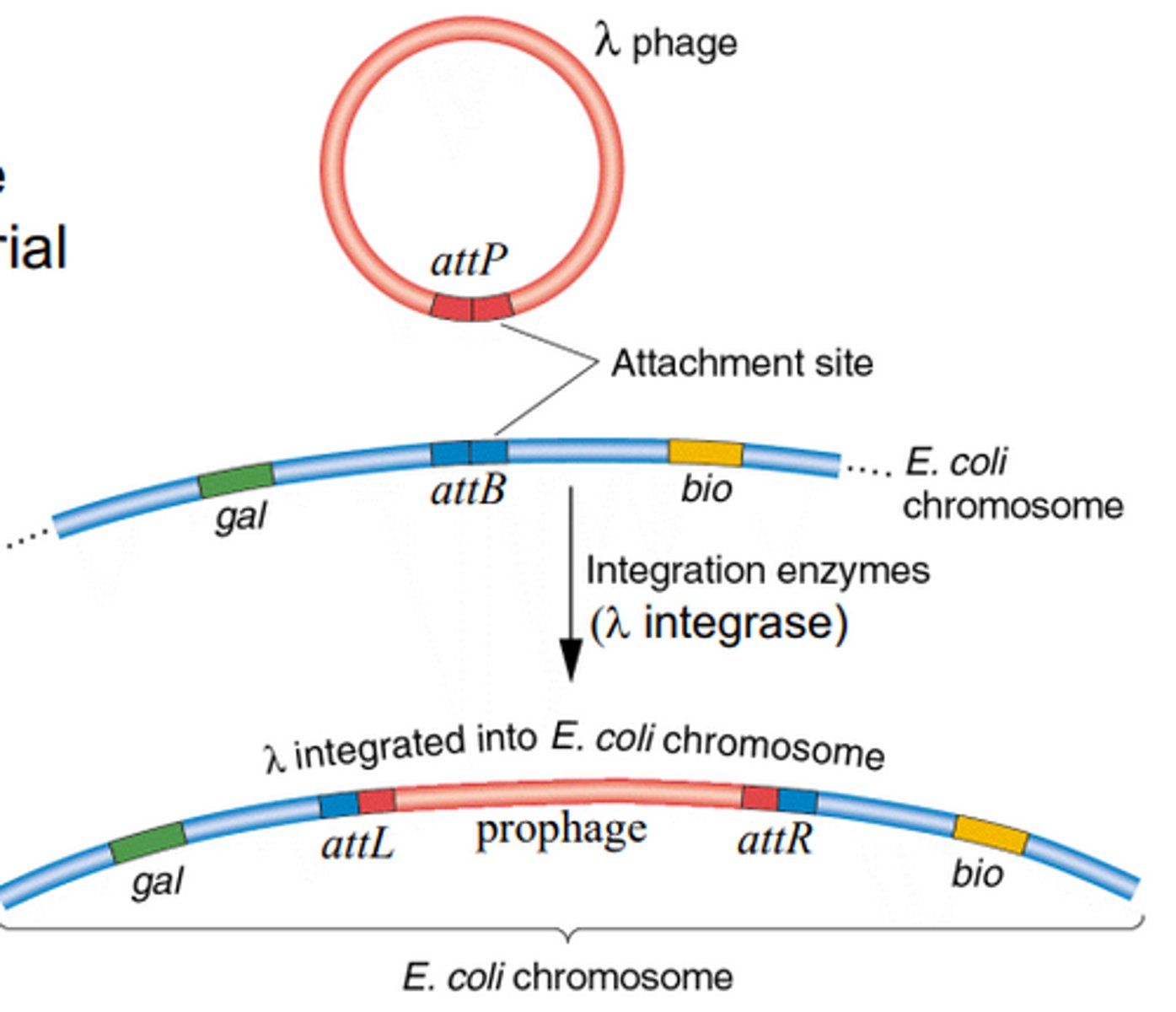
attP
Attach to the prophage. Used by λ integrase to attach prophage to attB
attB
attach to the bacterial (chromosome). used by λ integrase to attach prophage to chromosome.
λ integrase
enzyme used to connect attP and attB to integrate lambda phage into the bacterial chromosome inside the lysogenic cycle.
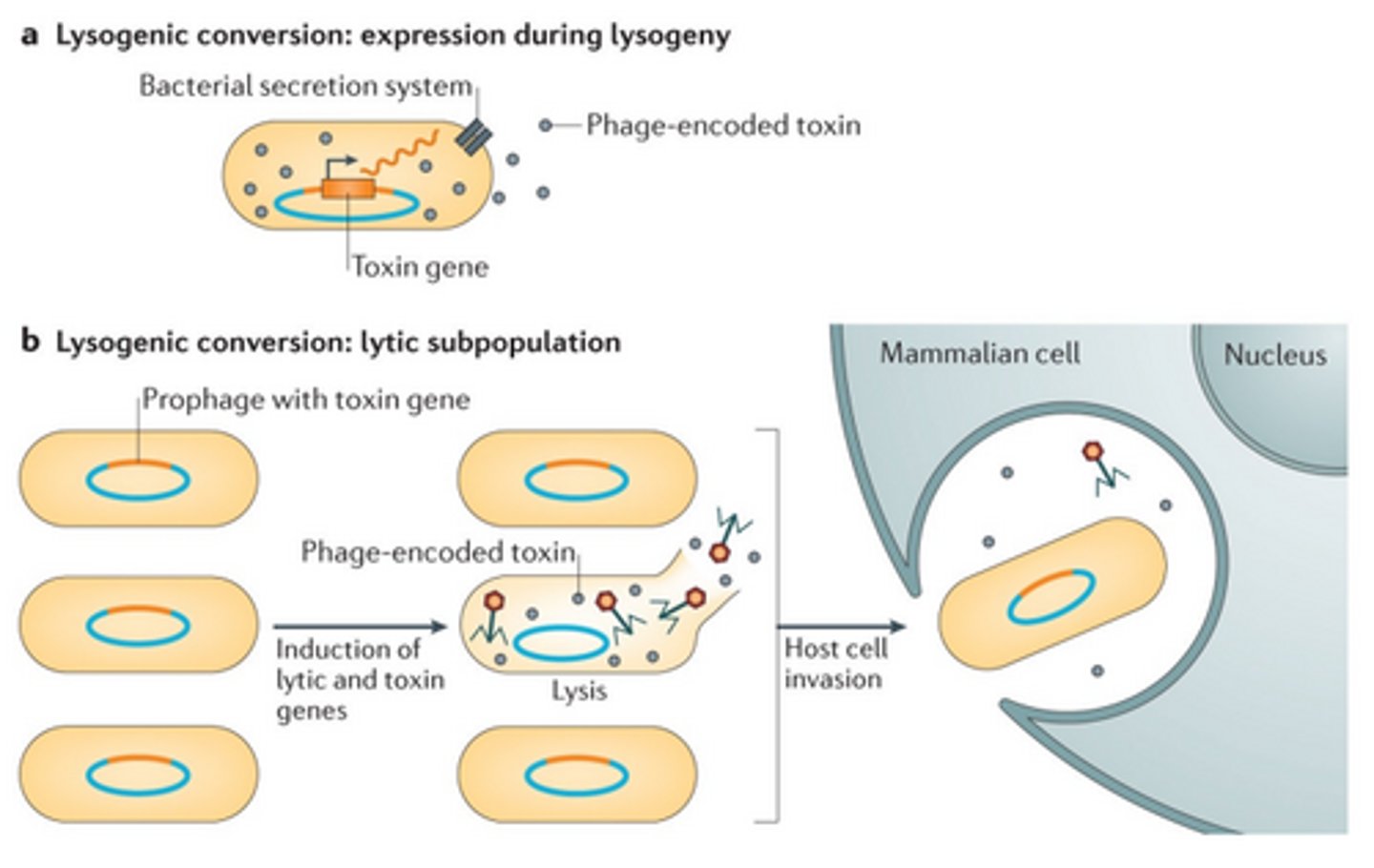
Lysogenic conversion
Genes from the inserted prophage become expressed in the host cell, during lysogeny.
When can prophages inside the bacterial chromosome secrete toxins and why?
Toxin genes can be expressed passively, and then secreted out of the host cell via the bacterial secretion system (salmonella), or otherwise the toxic gene is only expressed when the cell enters the lytic cycle. This means that the lysed cell dies, but it allows the toxin to work for the other bacterial cells in order to infect mammalian cells.
Virulence
Degree of pathogenicity of strains
Transposases
Enzymes that allow transposons to move
Transposition
Movement of the transposon genetic element
Insertion Sequence (IS)
Encodes only the protein needed for transposition (transposase). The ends of all known IS elemetns show inverted terminal repeats.
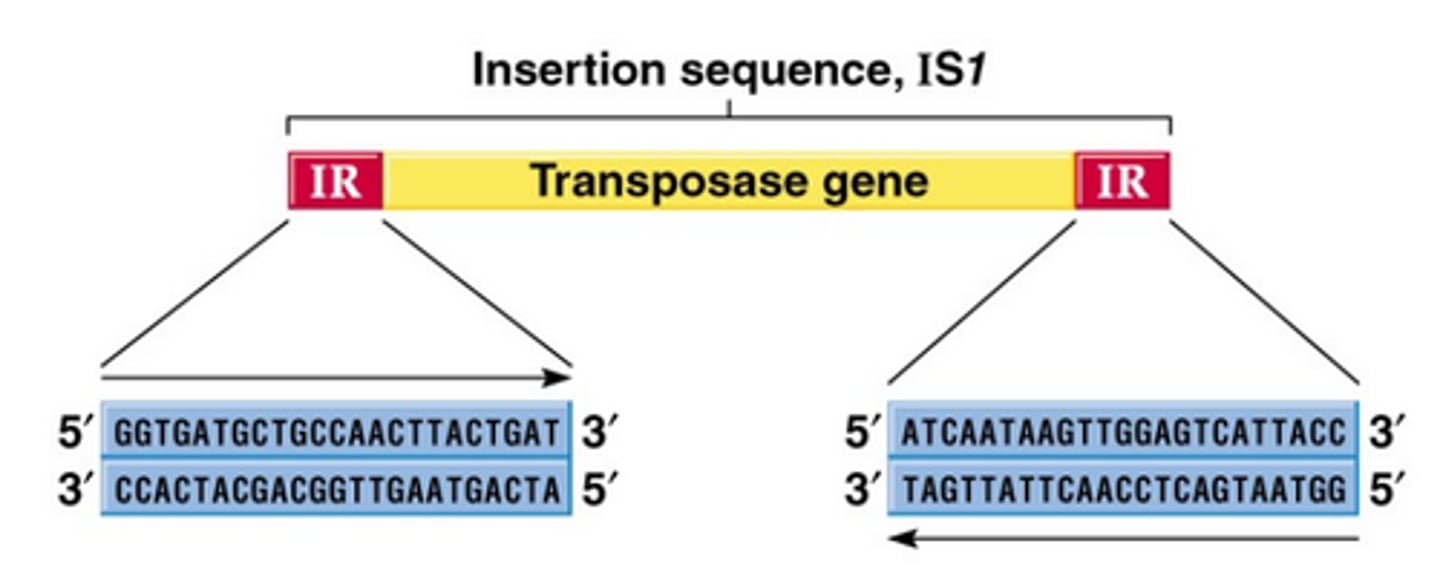
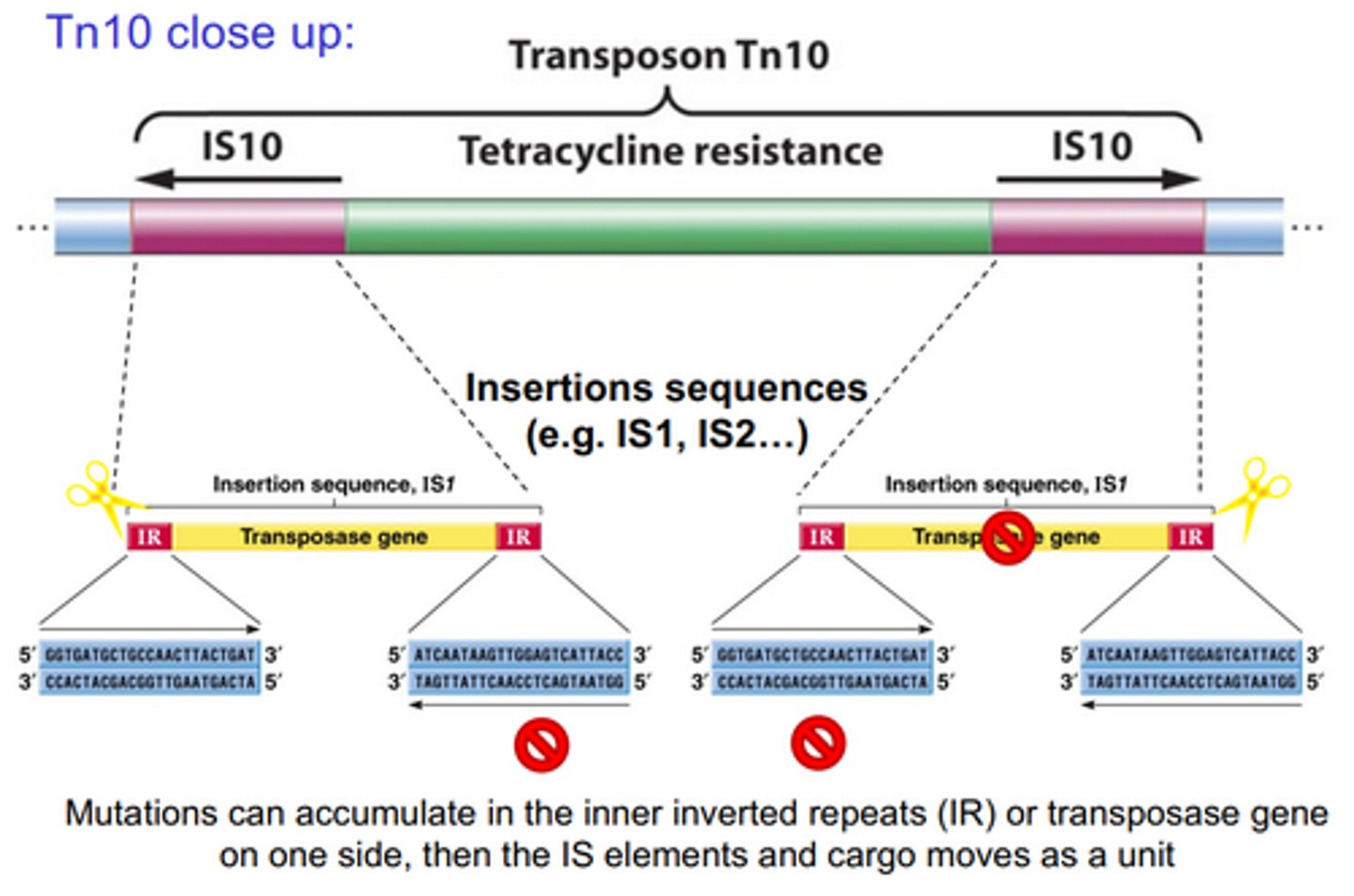
Composite transpoosons
Two IS elements flanking a central region carrying other genes. IS elements supply the transposase and ITR recognition signals. Often only one IS makes the active transposase.
How do composite transposons that excise chromosomal DNA form? e.g antibacterial genes
The IS elements on either side of the central region have IR sequences that tell transposase where to cut. The innermost IR sequences can be mutated over time, and then the transposase will cut only twice for two IS elements, and the entire composite transposon will be excised. Often the transposase genes are also mutated, but as long as one copy is functional, the composite transposon can be excised.
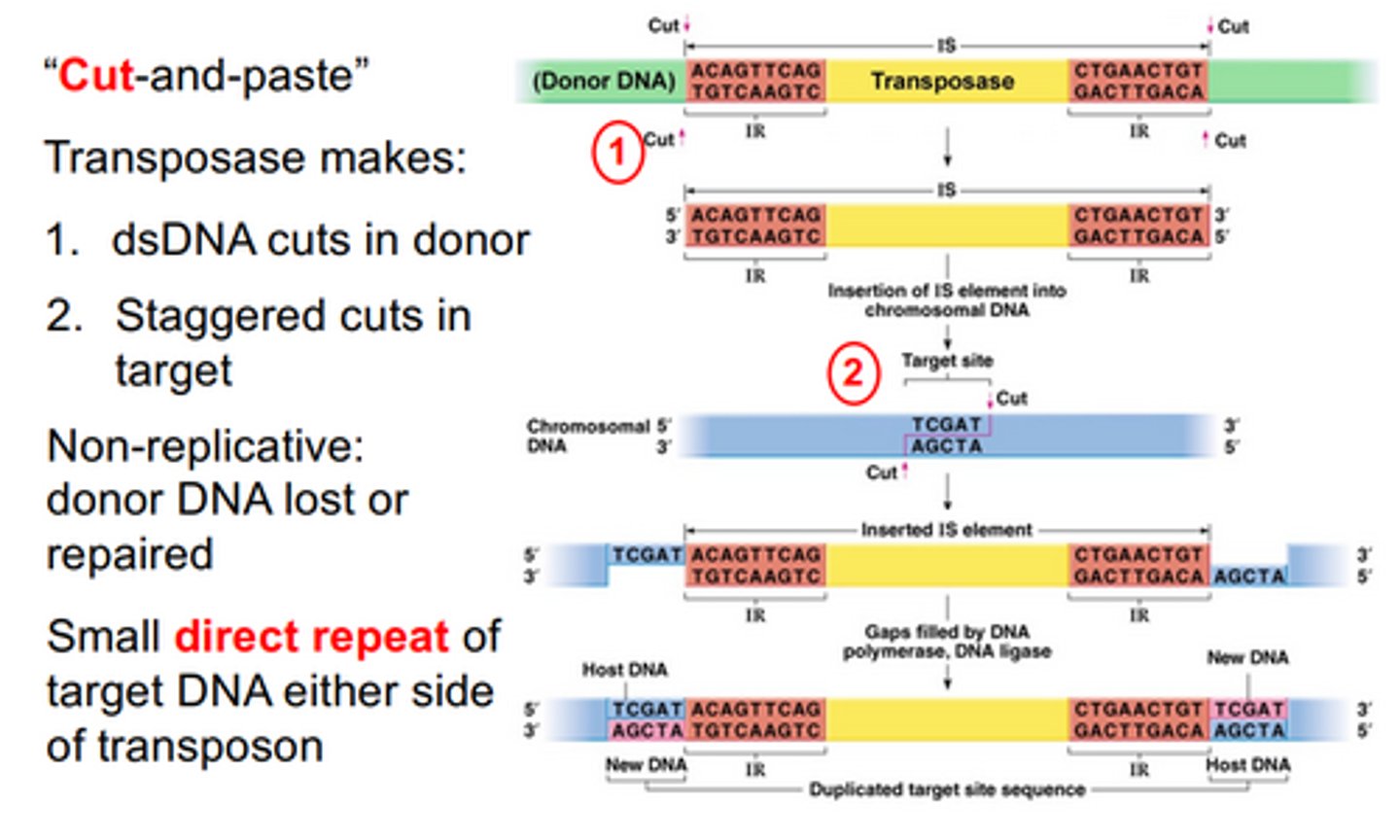
cut-and-paste
Transposase makes dsDNA cut in the donor, to excise IS. Then it makes staggered cuts in the target. This is non-replicative, meaning the donor loses DNA (if composite transposon), or has to repair where the IS was excised. Then the IS is inserted between the staggered cuts, and the host then repairs the cuts and replaces lost DNA. This forms a direct repeat of the target DNA on either side of the transposon.
Direct Orientation
Transposon elements like IRs have to line up in the same direction in order to be homologously recombined, either for replicon fusion (Hfr) or for deletions (F').
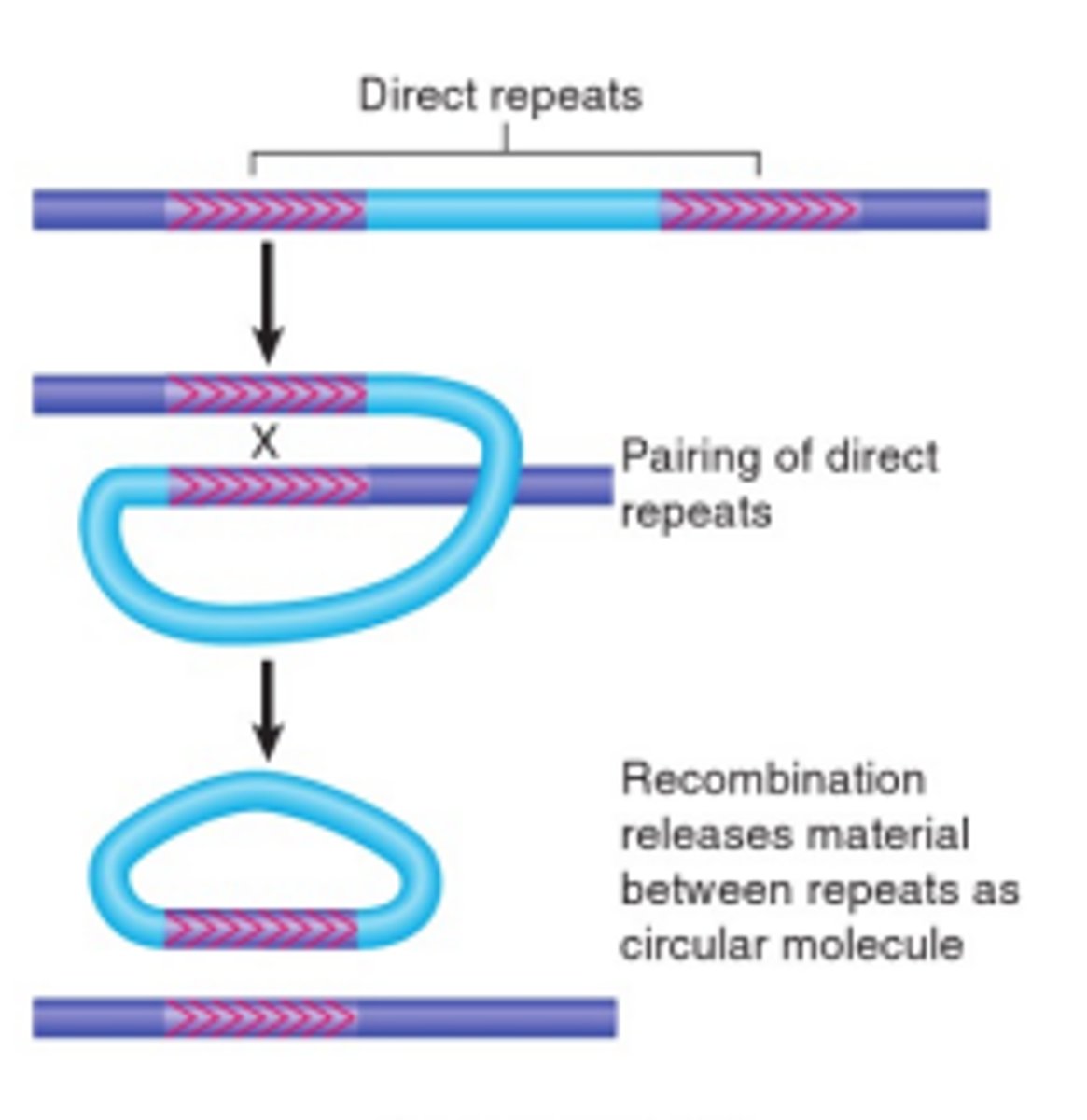
What direction do genes that are high copy number (e.g ribosomal genes) run?
Typically in the direction of DNA replication. Otherwise DNA polymerase and RNA polymerase would often run into each other.
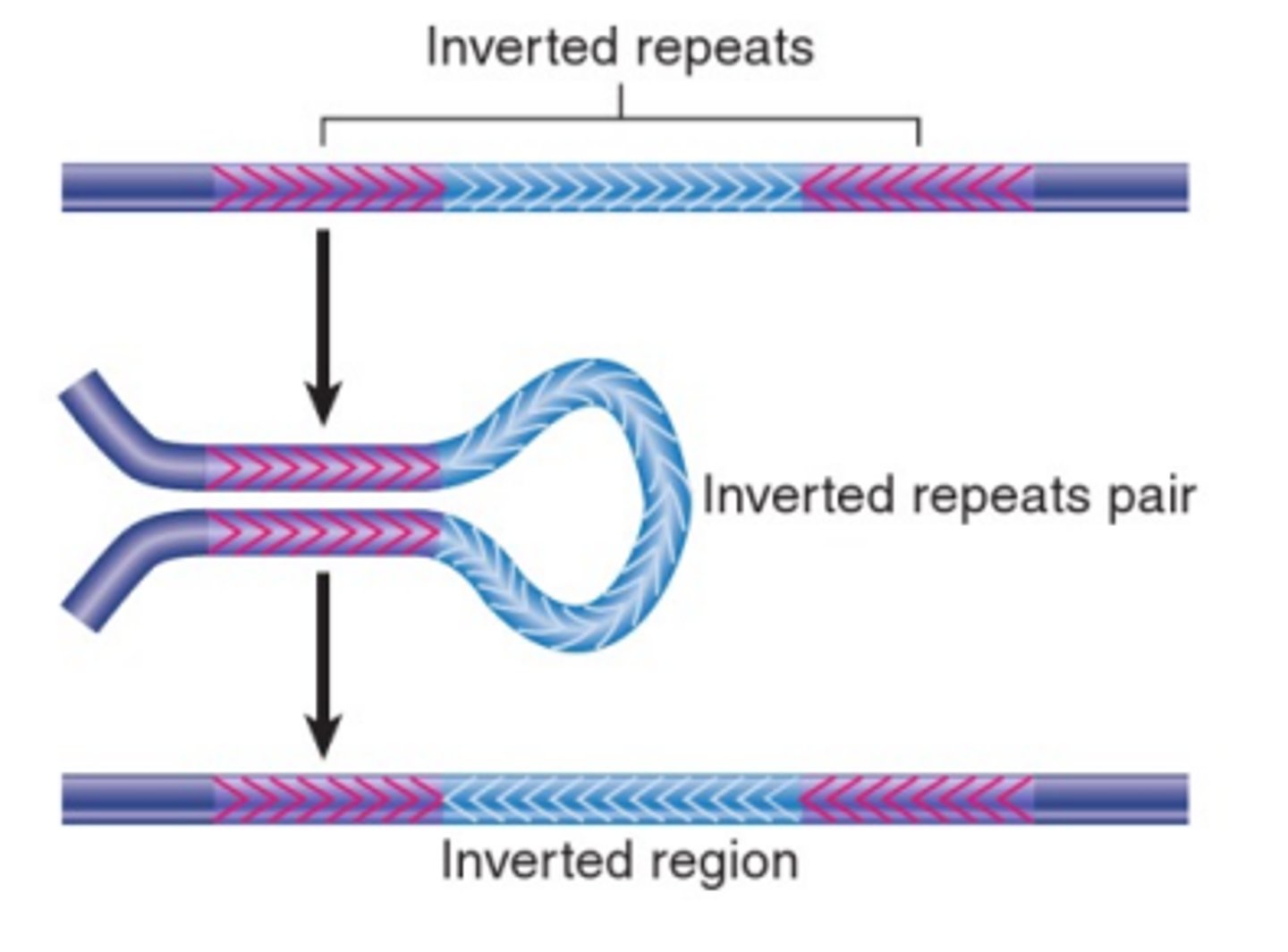
How do inversions in DNA occur?
Transposons that are inverted can only recombine homologously if the DNA between them is inverted. This creates DNA that runs in the opposite direction.
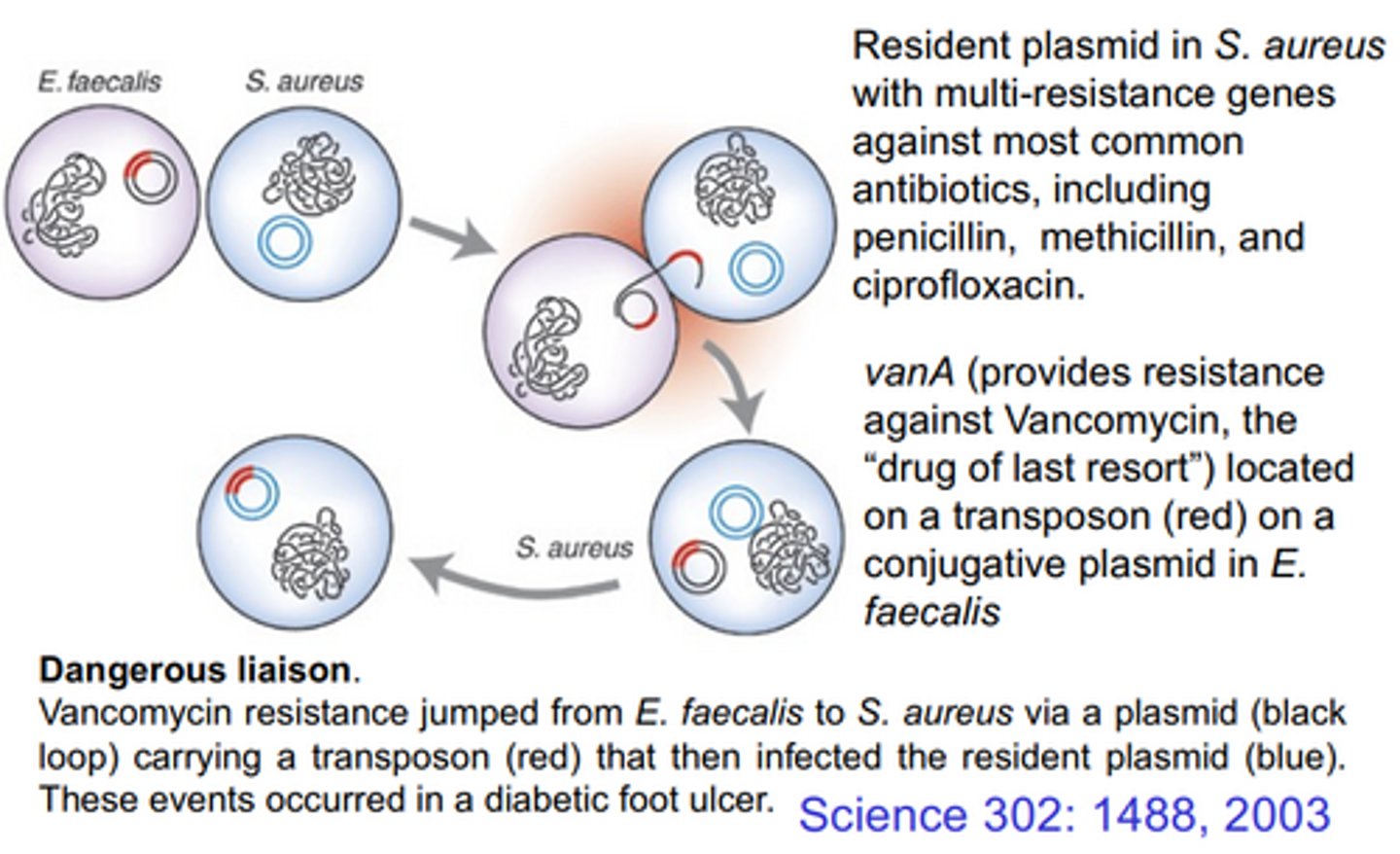
MRSA
Methicillin Resistant Staph. aureus
Vancomycin
The antibot of last resort
How did vanomycin resistance evolve in MRSA?
SA already had its own antibot resistant plasmid, but then it received a plasmid from e.faecalis containing vanA through conjugation. This vanA was on a transposon, and hopped from the new plasmid to the resident plasmid, creating a MR and VR plasmid.