PATH 381 - Module 5: Biomarker Evaluation and Acute Coronary Syndrome
1/69
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
70 Terms
NIH definition of a biomarker
characteristic that is objectively measured and evaluated as an indicator of healthy biological processes, pathogenic processes, or pharmacologic responses to therapeutic intervention
IPCS definition of a biomarker
any measurable substance/structure/process or its products that can influence/predict the incidence of an outcome/disease
WHO definition of a biomarker
- any measurement reflecting interaction between biological system and a poptential hazard (chemical, physical, biological)
- measured response may be functional and physiological, biochemical, or molecular
4 criteria of an ideal/clinically useful biomarker (and explain)
1. measureable and interpretive: specific, sensitive, quantifiable results/processes that can be interpreted
2. cost efficient and safe: (decision triad) fast, affordable, safe, meaningful results
3. consistent and accurate: reliably predict disease course and outcomes, measured using test performance metrics
(can act as surrogate marker)
4. applicable: effective for wide-scale use, safe and consistent across multiple patient populations
surrogate marker
biomarker that stands in for a real clinical outcome because the real outcome takes too long or is too hard to measure (eg. blood pressure in suspected stroke)
limitations of biomarkers (5 and explain)
1. cost:
- can be expensive, must fit budget and clinical need
2. analyte stability and storage:
- Some analytes degrade or change over time
- Poor storage = inaccurate results
3. standardization:
- different manufacturers use different methods
- no standardization = results won’t match across labs
4. measurement errors:
- test may lack accuracy or precision
- equipment/transport issues can = inaccurate levels
- even a “good biomarker” is useless if the measurement method is bad
5. confounding factors:
- biomarker levels vary by age, sex, ethnicity, weight, etc.
- need adjusted reference ranges
- if not accounted for → misinterpretation
3 steps in biomarker evaluation process (and what its for)
1. analytical validation: is the test accurate?
2. qualification: does the biomarker actually relate to the disease?
3. utilization: given the evidence, can it be used in practice?
step 1: analytical validation (5 specifications)
→ Must determine specifications of biomarker:
1. limit of detection
2. limit of quantification
3. reference value
4. cut-off conc
5. total imprecision at cut-off conc
- all specifications must be determined before biomarker data is relevant
step 2: qualification
→ look at research showing:
- biomarker levels change with disease
- biomarker is involved in the disease pathway
- treatments that change the biomarker also change real clinical outcomes
(Checks the strength of evidence linking biomarker ↔ disease)
step 3: utilization
→ Decide if:
- the validation and qualification evidence is enough
- the biomarker fits the intended use (e.g., diagnosis, monitoring, prognosis)
what are 3 sources of biomarker variation
- bias
- measurement errors
- confounding factors
stages of variability for biomarkers
1. pre-analytical: biological variation and sample collection
2. analytical (during measurements)
pre-analytical: biological variations and 3 ex
diff in health, disease, stress, dehydration, age, sex, diet, etc.
eg:
- random vs fasting glucose
- supine vs standing BP
- morning vs afternoon cortisol levels
pre-analytical: sample collection variation
issues with tube type, mislabelling, needle size, transport/storage temp
eg:
- hemolyzed sample (shear force→lysed RBC)
- biomarker instability at wrong temp
analytical variation
- during measurement
- interferences from drugs/endogenous compounts
- instrument performance changes
- errors in post-test calculations
eg. precision and accuracy problems (can get worse as more widely used → even in same lab, lot-to-lot variation = differences between batches)
how do labs minimize analytical variability?
total quality management framework
goal of evidence-based lab medicine and biomarker qualification
use high-quality evidence to decide whether a biomarker/lab test actually improves patient outcomes
5 principles of evidence-based lab medicine
1. asking the question
2. searching for evidence
3. appraising the evidence
4. applying the evidence
5. assessing the experience
evidence-based lab medicine: asking the question (and 6 common study themes)
identify the real unmet clinical needs/problems → common study themes:
1. pre-analytical issues (timing, sample type)
2. analytical/test performance (accuracy, imprecision)
3. diagnostic value (usefulness of screening
4. prognosis/treatment selection
5. cost-effectiveness
6. whether test should be used/discontinued
evidence-based lab medicine: searching for evidence (requires what?)
- use existing strong evidence (eg. systematic reviews, meta analysis) or develop new evidence
- evidence must reflect routine lab practice AND research settings
- to prove test improves outcomes, need intervenion/"test-and-act" study, often using an RCT = (test result + resulting clinical action).
evidence-based lab medicine: appraising the evidence (and 4 questions)
critically evaluate research/evidence to judge validity and usefulness → ask critical appraisal questions:
- is the question clear?
- what are the results?
- are the results valid? (internal validity)
- are the results relevent to patient/population? (external validity)
evidence-based lab medicine: applying the evidence (and 4 questions)
decide if evidence fits real clinical context, and if the question was right → question if evidence is relevant to problem:
- does test meet clinical needs?
- is the analytical performance good enough?
- does it meet standard of care?
- will using this test actually improve outcomes?
evidence-based lab medicine: assessing the experience (3)
- learning experience
- promotes applying the principles into daily practice = quality improvement
- after implementing test, evaluate if it actually improved outcomes
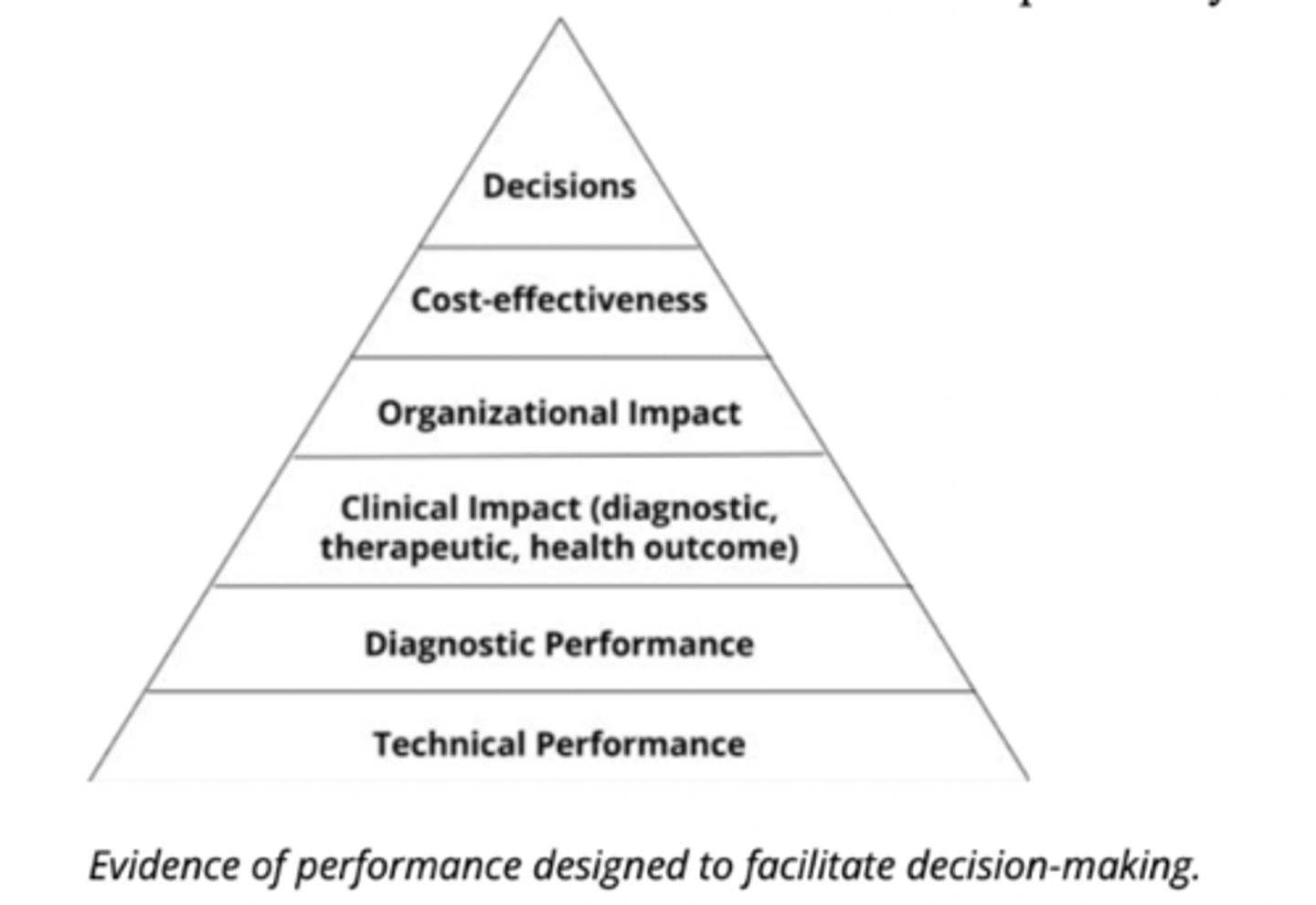
utilization/evidence of performance of a biomarker (5)
technical performance > diagnostic performance > clinical impact > organizational impact > cost-effectiveness > decisions
(must work on the lower level to work at any of the higher levels)
evaluation tools/test performance metrics (4)
1. NPV and PPV
2. Sensitivity and Specificity
3. Odds Ratio (OR)
4. Receiver Operating Characteristics (ROC)
PPV and NPV (what is it, equation, when does it inc/dec)
PPV:
- Probability a positive result is a true positive.
- PPV = TP / (TP + FP)
- Increases when disease prevalence ↑
NPV:
- Probability a negative result is a true negative.
- NPV = TN / (TN + FN)
- Decreases when disease prevalence ↑
Sensitivity and Specificity (what is it, equation)
Sensitivity:
- "True positive rate": Ability to correctly detect people with the disease
- Sens = TP / (TP + FN)
Specificity:
- "True negative rate": Ability to correctly identify people without the disease.
- Spec = TN / (TN + FP)
Odds Ratio (OR)
strength of association between exposure and event:
- OR < 1 → exposure lowers odds
- OR = 1 → no effect
- OR > 1 → exposure increases odds
LRs (equations, what it measures, how to interpret)
LR+: Sensitivity / (1 - Specificity)
- how does pos test impact probability of disease?
- larger = more likely
LR- : (1 - Sensitivity) / Specificity
- how does neg test impact probability of disease?
- smaller = less likely
why are LRs useful? (3)
- do NOT depend on prevalence (don't vary in diff pop)
- Can apply directly to one patient
- Turn test result into a probability
LRs and Baye's Theorum
- translates LRs into probability of disease
- post-test probability = pre-test probability × LR
why use ROC curves?
to visualize performance of biomarker at various cut-off settings:
- evaluate overall test performance
- decide the best cut-off point
- compare multiple biomarkers/tests
- understand how test accuracy changes depending on the threshold used
axes in ROC curve
Y axis: true positive rate (sensitivity)
X axis: false positive rate (1-specificity)
ROC curve: area under the curve (AUC)
- summarizes ROC curve into one number that represents how good the test is (higher AUC = better test at distinguishing)
AUC values (ideal, typical, poor)
ideal: 1.0 (100% distinguishablity)
typical: ~0.7 (~70% distinguishability)
poor: 0.5 (50% distinguishability = no better than flipping a coin)
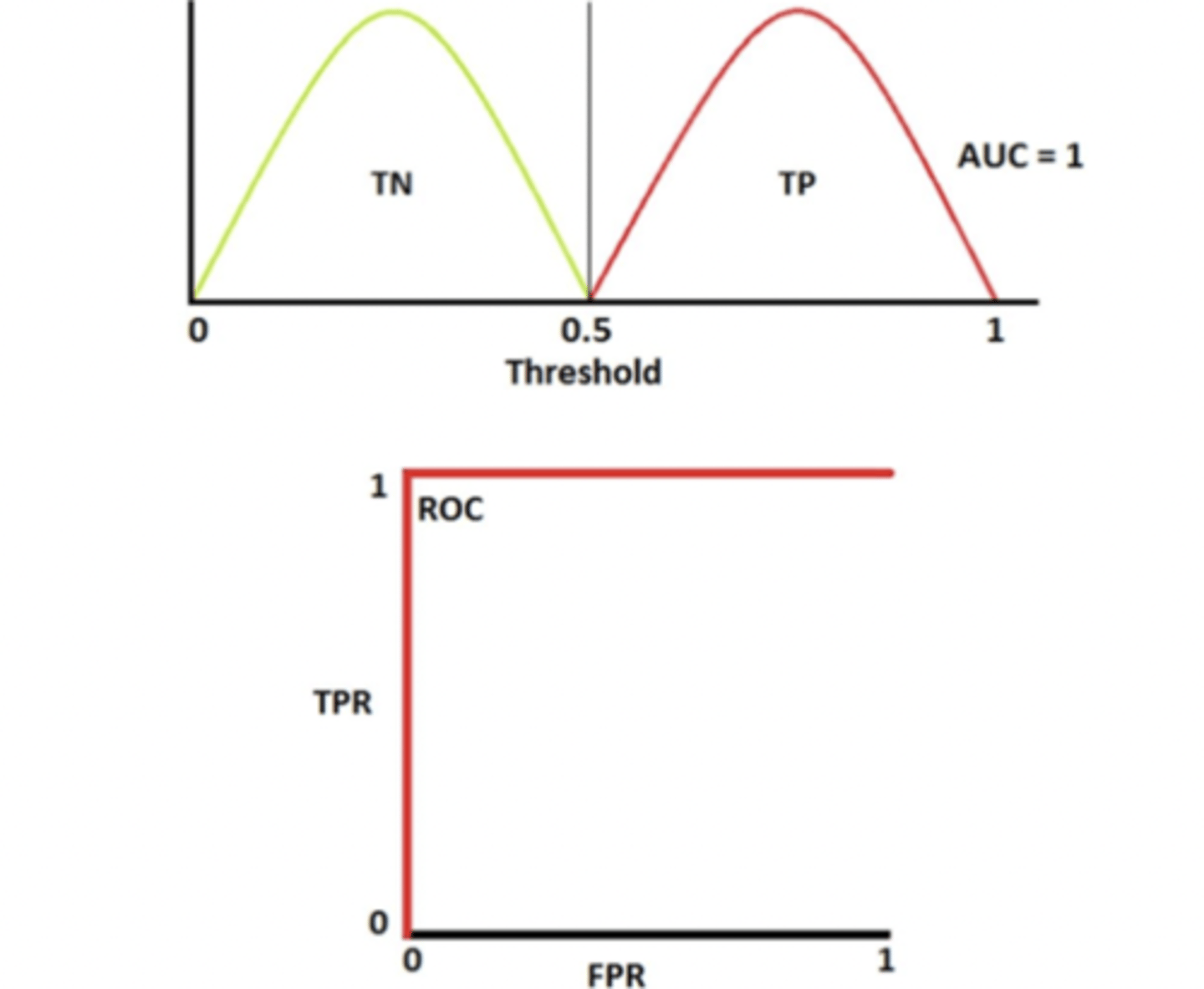
ideal biomarker ROC curve
- population distributions (disease vs non-disease) have no overlap
- AUC = 1.0 (100% chance distinguishability)
- ROC reaches top left corner (ie full area covered)
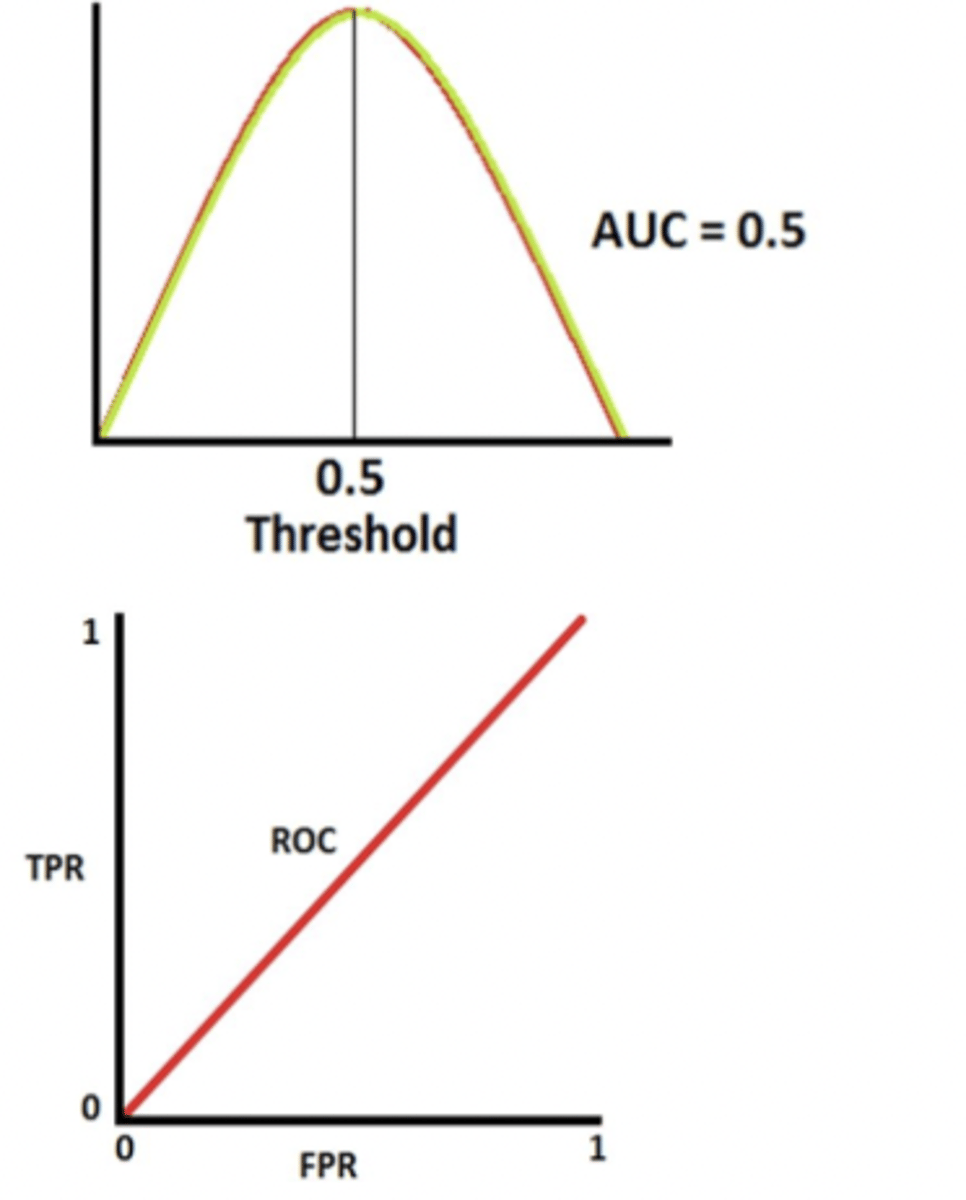
poor biomarker ROC curve
- complete overlap between disease and non-disease distrib.
- ROC curve is a diagonal line (y = x)
- AUC = 0.5 (50% of distinguishability)
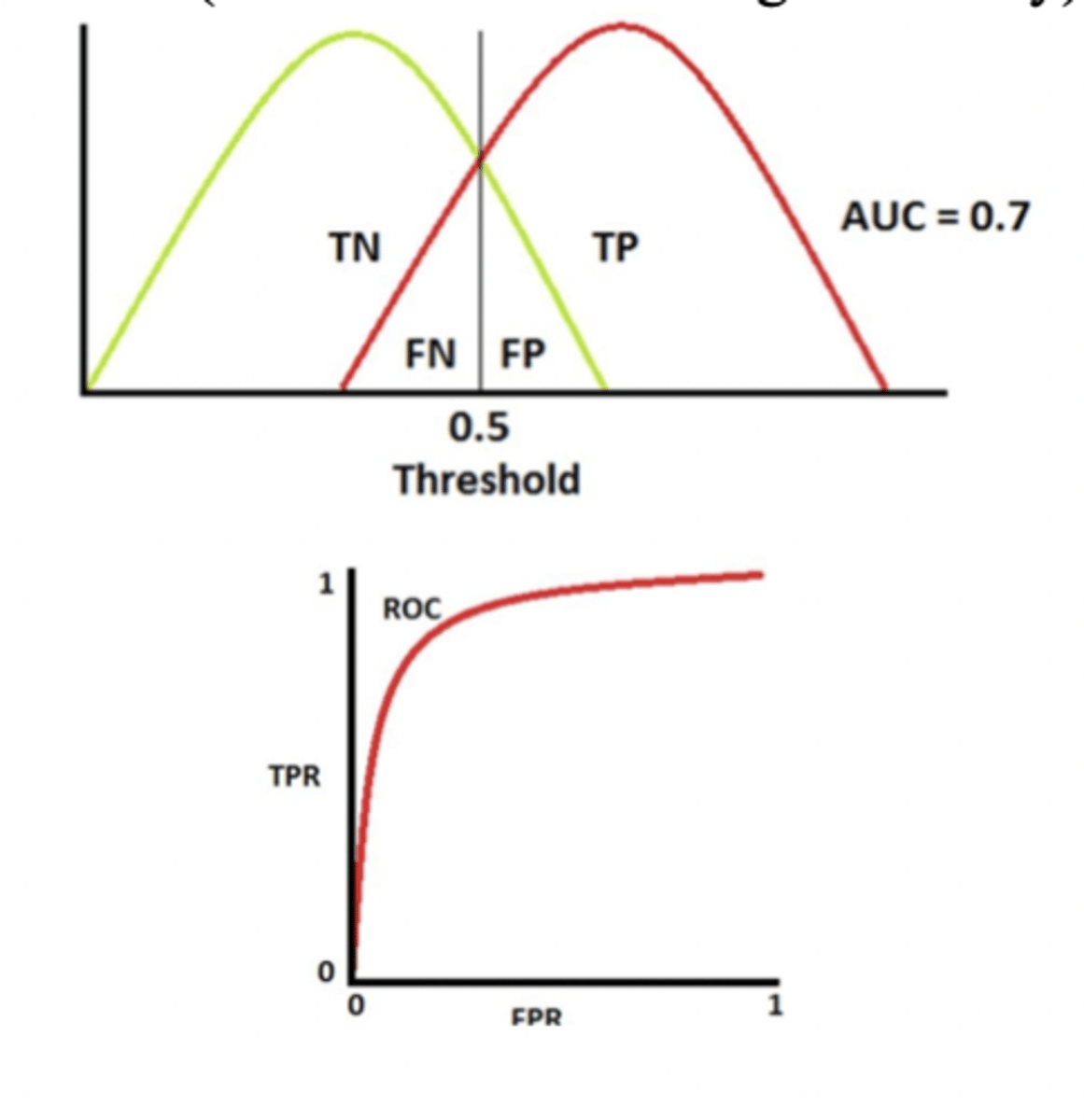
typical biomarker ROC curve
- some overlap between groups (realistic scenario)
- ROC curve goes upward but not perfectly
- AUC ~0.7 (moderate-to-good discrimination)
How to create an ROC curve (6 steps)
1. choose possible cut-off
2. calculate spec and sensitivity at that cut-off
3. plot sensitivity vs false positive rate (1-specificity)
4. repeat for may different cut-offs
5. connect points = make curve
6. measure AUC to summarize test performance
why are randomized control trials (RCTs) needed (3)
- biomarker may look good in research but not acc improve patient outcomes
- to prove biomarker is clinically useful, must compare to current gold standard
- RCTs are the only way to whether changes in biomarker actually cause better outcomes
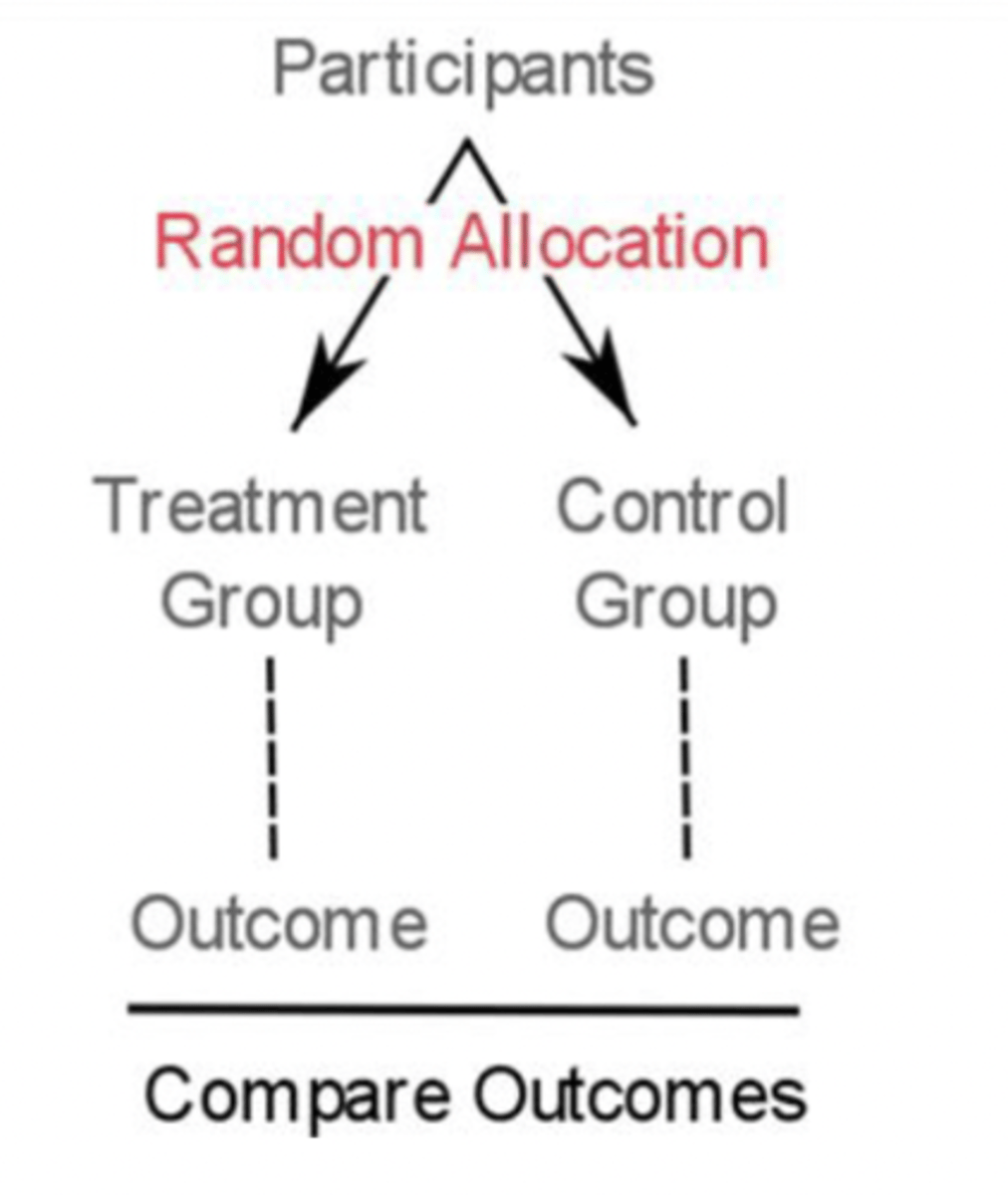
RCT overview (3)
- patients randomly assigned to treatment (new biomarker) or control (current gold standard)
- may be blinded or double-blinded (prevents bias)
- outcomes compared → tells you if the new biomarker actually improves patient care
Phases of RCTs and explain
I: 'first in human'
- first patient group, examines potential use using earlier work
II: 'first in patient'
- randomized, controlled testing (use earlier work/phase I)
- validating biomarker in real clinical pop
- identifies promising approaches to be used for phase III
III: 'multi-site trials'
- large, randomized, can be placebo-controlled/uncontrolled, blinded
- confirms biomarker performance across labs and populations
- needed before widespread use
designs for cancer biomarker trials
1. target/enrichment: only biomarker+ (or -) patients
2. allcomers: everyone regardless of biomarker
3. adaptive: trial adjusts as it goes
target/enrichment designs (3)
- only include patients with (or without) the biomarker of interest.
- used when believe only a specific subgroup will benefit
- aim to understand safety, tolerability, clinical benefit in specific subgroup
allcomer designs and when are they used
Include all eligible patients regardless of biomarker status → Used when:
- Evidence for the biomarker is unclear
- Biomarker prevalence is high (>50%)
- No cut-off is established yet
- Biomarker takes too long to measure
adaptive designs
- multiple biomarkers tested under one big protocol
- trial adjusts as it runs (more patients moved to promising subgroups, and weak subgroups dropped)
acute coronary syndrome (ACS)
- umbrella term for conditions where blood flow to heart decreases
- includes myocardial infarction (MI) and unstable angina
- responsible for ~1/3 deaths in ppl >35
Unstable Angina
ischemic symptoms without elevations in biomarkers (cTn) or ECG changes
myocardial infarction (MI)
- 'heart attack'
- when heart doesn't get enough oxygen due to blocked blood flow (blocked coronary artery)
- term used when evidence of myocardial necrosis (cell death) due to acute ischemia (reduced blood flow)
- either STEMI or NSTEMI
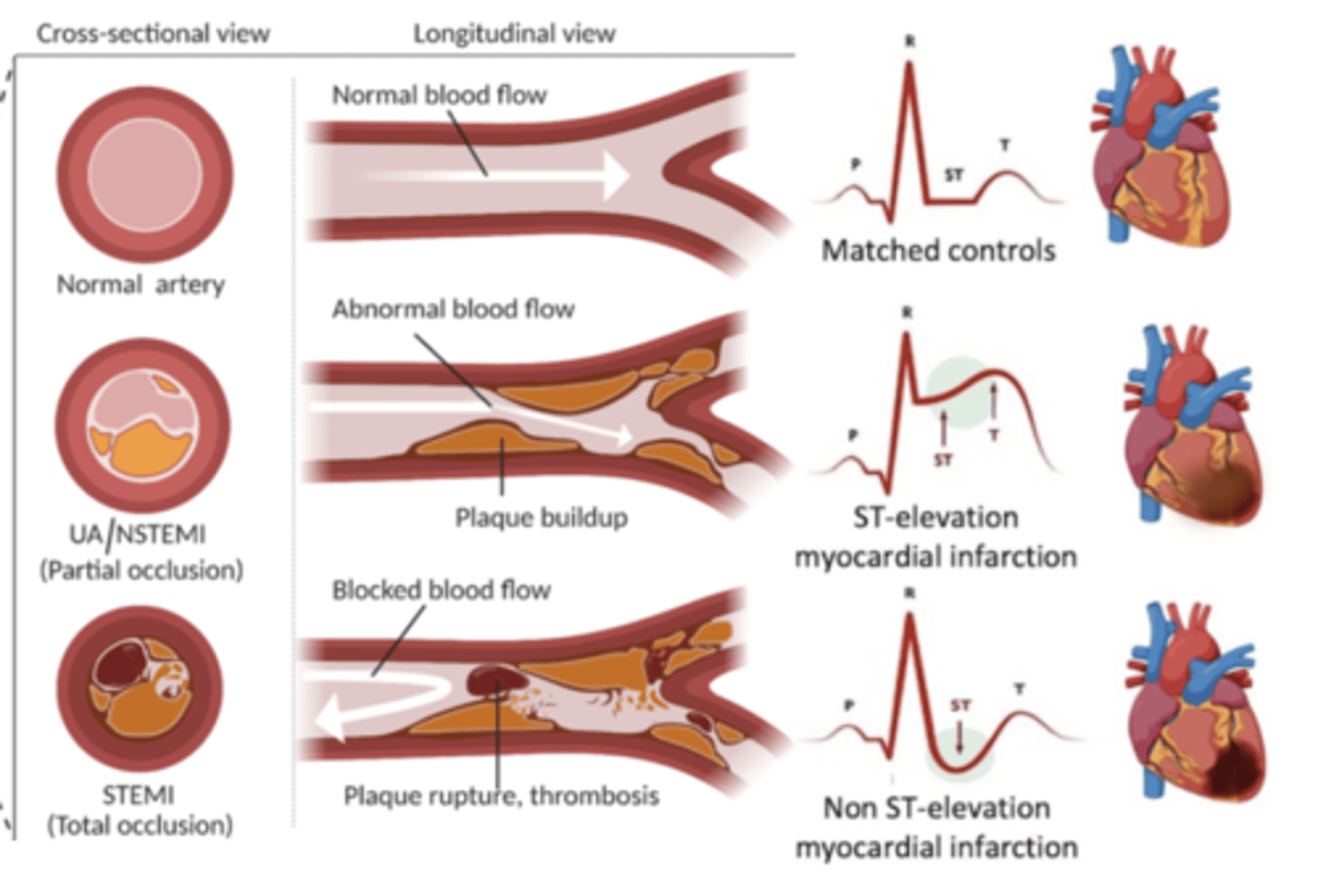
types of MI
STEMI (ST elevation MI):
- ST elevation on ECG
- total occlusion/blockage of coronary artery
NSTEMI (non ST elevation MI):
- no ST elevation on ECG
- partial occlusion/blockage of coronary artey
- troponin (cTn) elevated
criteria for diagnosing MI (3)
1. acute myocardial injury with clinical evidence of myocardial ischemica
2. detection of rise/fall in cTn
3. at least one clinical sign of acute ischemia
clinical signs of acute ischemia (5)
- typical ischemic symptoms (chest pain, jaw/arm pain, dyspnea, nausea, fainting)
- imaging evidence of new loss of myocardiam or new wall motion
- pathological Q waves
- coronary thrombus (blood clot) on angiography/autopsy
- ECG changes indicative of ischemia
pathophysiology of ACS
usually following plaque rupture, formation of thrombus, occlusion of vessel → decreased bloodflow to part of heart → ischemia and then infarction
general qualities of an ideal biomarker (5)
- accurate diagnosis/prediction
- fast, affordable, meaningful results
- value beyond existing tests
- easy to measure
- can act as a surrogate marker
specific qualities for an MI biomarker (6)
- accurately distinguish MI from other causes
- high sensitivity and specitivity for MI
- clear separation between MI and non-MI levels
- useful in staging, prognosis, intervention
- direct link to pathophysiology of MI
- can predict occurrence
History of ACS Biomarkers
1950s: AST, LDH
1960s: CK, discovery of troponins
1970s: cTnT, cTnI, CKMB, myoglobin released early after MI
1980s: CKMB 'mass' immunoassay = more sensitive
1990s: rapid CKMB mass assays, 3rd-gen cTnT assay
2000s-2010s: high-sensitivity troponin assays
rise and fall curves for biomarkers
- help compare biomarkers
- myoglobin, CK, CKMB rise fast but low specificity
- troponin has slightly later response but much more specific → best overall marker
myoglobin (timing, pros, cons)
Timing: rises quickly (1-3h), falls early
Pros: high sensitivity, good for early detection and ruling out MI
Cons: low specificity (also elevated in skel muscle damage → rhabdomyolosis)
creatine kinsase (CK)
- older biomarker
- not specific
CKMB (timing, pros, cons)
cardiac-associated isoform of CK
Timing: rises 4-6h after MI, stays elevated for 24-48h
Pros: detect early refraction, rapid, cost-efficient, better specificity than CK/myoglobin
Cons: much less specific than cTn, can still rise with skel muscle injury
cardiac troponin (cTnI, cTnT) - what is it, pros, cons
proteins released from necrotic myocytes when irreversible damage occurs
Pros:
- highest sensitivity and specificity for MI
- prognostic indicators
- can detect recent MI up to 2 weeks after onset
Cons:
- slightly later rise (but outweighed by high accuracy)
what does detection of cTn mean? what does it not mean?
- indicates and quantifies cardiomyocyte damage and injury
- does not indicate underlying mechanisms, or ischemic/nonischemic causes
- does not automatically mean ACS (just detects damage)
why is cTn preferred?
- cTnI and cTnT highly specific and sensitive biomarkers of myocardial injury because unique to the heart
- remain as intact proteins and degradation products
- can detect MI for up to 2 weeks after onset
what is considered elevated cTn
defined as value that exceeds 99th percentile of normal reference populations
when are troponin levels measured
first presentation (within 6h)
AND
6-12 hours after pain onset (bc cTn release is delayed)
what is the next step if cTn levels are inconsistant with clinical symptoms?
CKMB testing may be used to assess differences
3 challenges of MI biomarkers
1. biotin use
2. muscle damage
3. chronic kidney disease
MI biomarkers and biotin use
- many immunoassays (including cTn) use streptavidin beads
- high biotin intake (hair/skin supplements) → excess biotin in blood → disrupts assay binding → falsely low/high troponin results depending on the assay design
MI biomarkers and muscle damage
- CK and myoglobin not specific
- trauma → muscle damage → elevated CK/myoglobin (even without MI)
- lowers specificity, can falsely suggest MI
MI biomarkers and CKD and how can you distinguish?
patients with CKD often have elevated cTnT (bc reduced clearence, mild myocardial stress)
key distinguisher: NO rise and fall in cTnT due to CKD