genetics exam 3
1/250
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
251 Terms
requirements of genetic material
must have complex information in stable form
must accurately reproduce and transit info
must be expressed to produce other molecules (must have the capacity to encode phenotypic traits)
must have the capacity to vary
johann friedrich miescher
discovered DNA (nuclein) in 1869
realization that the nucleus was the physical basis of heredity
isolated DNA from pus of wounds
mitosis and meiosis
chromosomes became focus of interest because of movements in what
protein theory
chromosomes found to be proteins and DNA
DNA is chemically simple
proteins are chemically complex
proteins and DNA
major components of chromatin
4 nitrogenous bases and 1 sugar
what is DNA made up of
proteins
in the original idea what was the most likely source of genetic material
being more chemically complex
frederick griffith
discovered principle of transformation in 1928
injected mice with different strains of bacterial pneumonia
some unknown component of dead virulent IIIS pneumonia cells “transformed” live nonvirulent IIR cells into live IIIS
(IIR- do not cause disease)
(IIIS- cause disease)
conclusion: a substance in the heat-killed virulent bacteria genetically transformed the type IIR bacteria into live, virulent type IIIS bacteria
avery, macleod, and mccarty
repeated griffiths work in test tubes cultures rather than live mice
found that purified IIS DNA from dead cells transforms live IIR into live IIIS
treating dead IIIS with DNase stops the transformation, but treating dead cells with protease or RNase does not
conclusion: because only DNase destroyed the transforming substance, the transforming substance is DNA
hershey and chase
used bacteriophage T2 to demonstrated that DNA, not protein, is the genetic material that viruses inject into bacteria to produce new viruses
labeled virus DNA with 32P- radioactive isotope of phosphate
labeled virus proteins with 35S- radioactive isotope of sulfur
labeled virus DNA with 32P
labeled virus protein with 35S
allowed labeled phages to attack separate batches of bacteria, then washed away leftover phages and “ghosts”
conclusion: found 32P inside bacteria and in progeny phages and found 35S only in ghosts
albrecht kossel
discovered 4 bases (A,C,G,T) in late 1800s
phoebus aaron
worked out the structure of the nucleotide
sugar, base, PO4
structure of the nucleotide
phoebus levene
proposed the tetranucleotide theory 1910
boring structure- A = C = G = T
they are all connected in a square like structure
erwin chargaff
has a rule
A=T and G=C but A+T does not equal G+C in 1948 (disproved the tetranucleotide theory)
rosalind franklin
used X-ray diffraction data to suggest helix form with with of 3.4 angstroms
OH in 2’
what makes RNA less stable than DNA
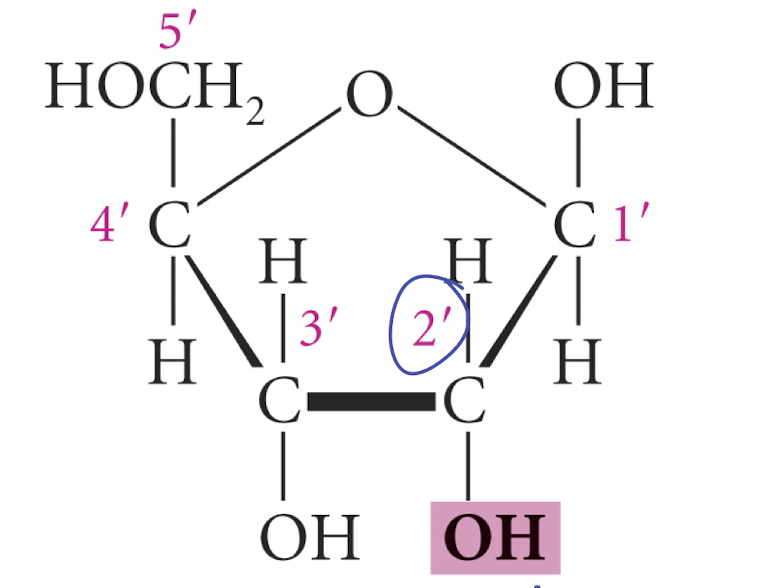
draw structure of ribose
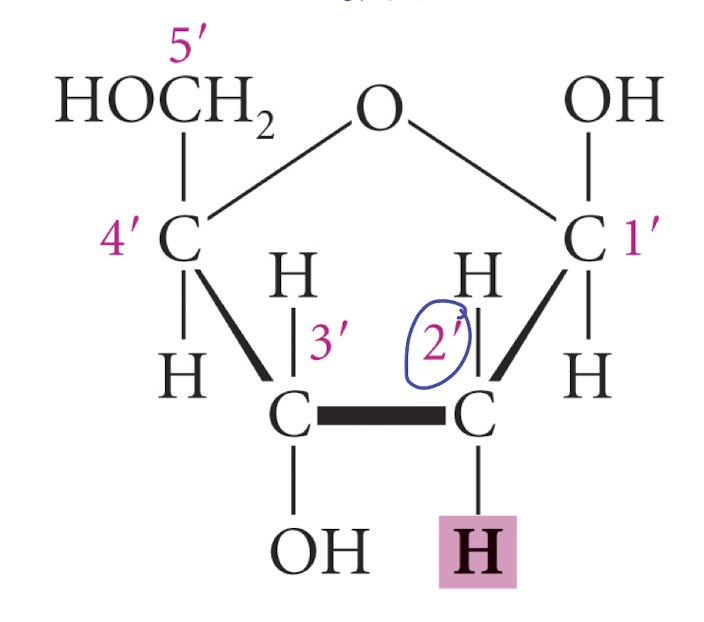
draw structure of deoxyribose
purine
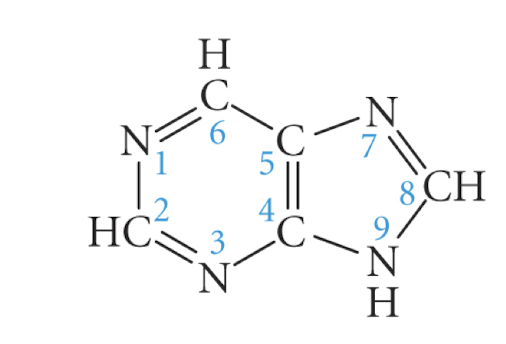
adenine
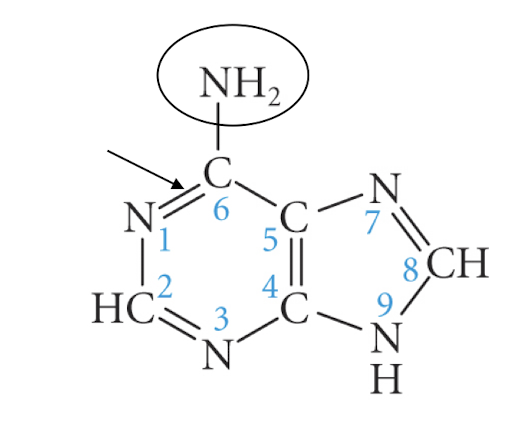
guanine
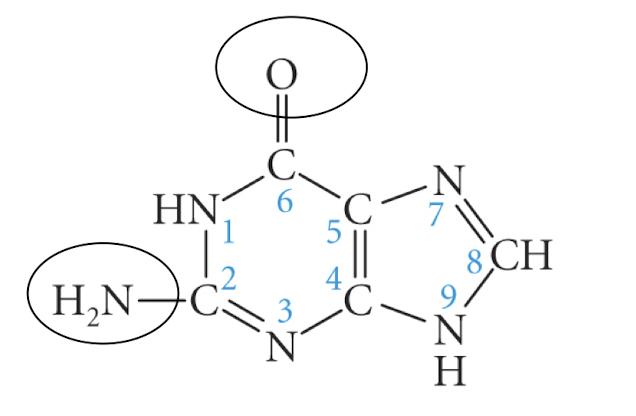
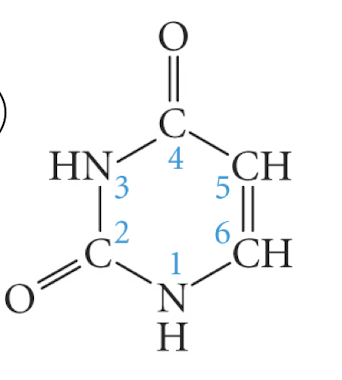
pyrimidine
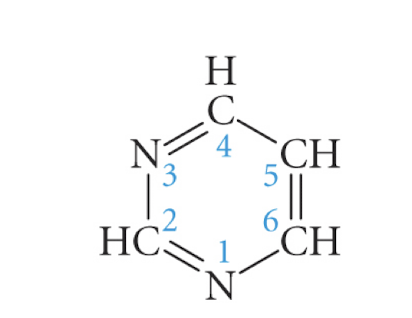
cytosine
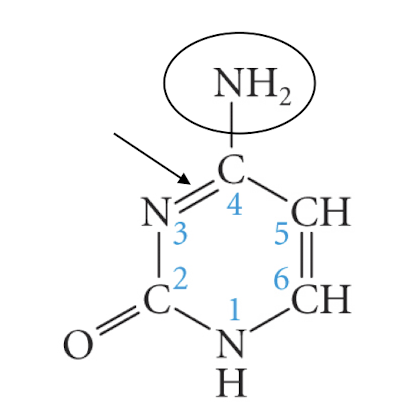
thymine
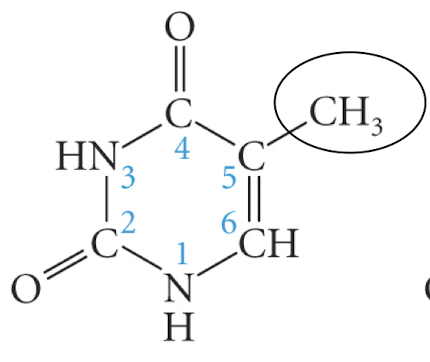
uracil
watson and crick
early information on structure of DNA
discovered the double helix model in 1953
antiparallel
antiparallel B form
form of DNA that is 10 bases per turn
obey chargaffs rules, bases on the inside, sugat PO4 on outside
features of double helix model
two sugar PO4 backbones with strong covalent bonds
base-pairing due to weak H bonding
G:C has 3 bonds
A:T has 2 bonds
distance between base pairs is even, about 3.4 A
major and minor grooves, right-handed twist
chains are polar, must go in opposite directions
CG
content of __ ___ in double stranded DNA is related to its stability
dsDNA molecules with higher GC content have correspondingly higher melting temperatures
more recent evidence suggests that this is due to base stacking interactions rather than the number of hydrogen bonds between base pairs
antiparallel
chains are polar and must go in opposite directions
polarity
given by 5’ and 3’ C,s of sugar
requirements of genetic material
must have complex information in stable form
must accurately reproduce and transmit info
must be expressed to produce other molecules (must have the capacity to encode phenotypic traits)
must have the capacity to vary
DH model
stable storage of complex info- bases can go in many orders (coding) but stay fixed once ordered
replicable- one chain determines other
expressible- linear code system can work for other polymers
changes in bases- (order or composition) create mutations that fuel variation
B-DNA
standard double helix
10 bases per turn
most stable configuration
most common
right handed helix
A-DNA
tight wound double helix
11 bases per turn
has been found in some DNA-protein complexes in spores and bacteria
right handed helix
Z-DNA
has left-handed twists
12 bases per turn
found in regions of heavy transcription
viruses
some of these use RNA as their genetic material
fraenkel conrat and singer
used tobacco to determine RNA carries genetic material in tabacco mosiac virus
degrade both types of TMV to yield RNA and coat proteins
mix RNA of one type with protein of the other to create hybrind viruses
infect tobacco with the hybrids
mRNA, rRNA, tRNA
large forms of RNA
mRNA
transmits info from DNA
3% of RNA
more of an intermediate
rRNA
translation machinery
majority of RNA in a cell
tRNA
interpreters of translation
inverted repeats
in RNA (sometimes DNA) can lead to self-pairing structures
hairpin structures
secondary structure
extensive structure of this in RNA is thought to serve important regulatory function
sometimes U pairs with G in RNA
primary structure
simply nucleotide sequence
secondary structure
DNA: double helix
RNA: stems and loops in RNA
methylation in prokaryotes
commonly methyl A or C often used to distinguish bacteria chromosome from foreign (viral) DNA
restriction enzyme defense
innate immunity
defense against its own restriction enzymes
methylation in eukaryotes
epigenetic phenomenon
methyl group added to base, usually C
changes how proteins interact with DNA
major eukaryotic gene regulation method
used to regulate gene expression (chromatin structure)
genomes of viruses
very small (1kb-340kb) nucleic acid
genetic material varies with species
DNA or RNA
single or double
Short genes (100-8000bp) without introns
genes can overlap
very little non-gene sequence
fred sanger
first sequenced viral genome
used 5386 bp circular ssDNA
found gene B lies entirely within gene A
retroviruses
are always RNA viruses that must convert their genome into DNA before integrating into the host genome as a provirus
include at least 3 genes
gag, pol, env
must be reverse transcribed into double stranded DNA, then infect
gag
gene in retroviruses that encodes viral capsid proteins
pol
gene in retroviruses that encodes the reverse transcriptase and integrase
env
gene in retroviruses that encodes glycoproteins that surround the capsid
virus chromosomes
chromosomes are packaged almost naked
no attached structural proteins
retroviruses have reverse transcriptase packaged with the viral RNA
retrovirus
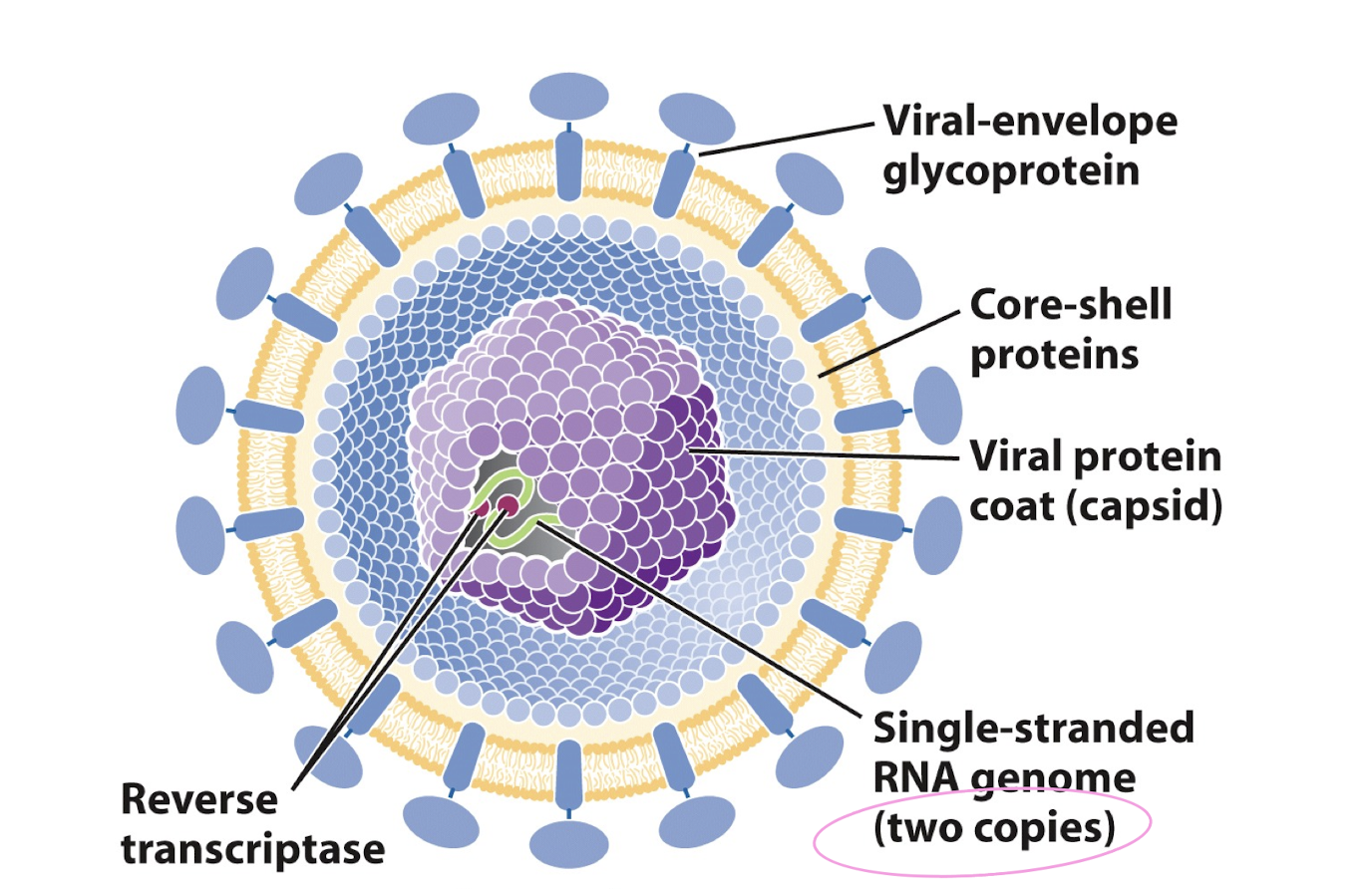
retrovirus infection
virus attaches to host cell at receptors in the membrane
the viral core enters the host cell
viral RNA uses reverse transcriptase to make complementary DNA, and viral RNA degrades
reverse transcriptase synthesizes the second DNA strand
the viral DNA enters the nucleus and is integrated into the host chromosome, forming a provirus
on activation, proviral DNA transcribes viral RNA, which is exported to the cytoplasm
in the cytoplasm, the viral RNA is translated
viral RNA, proteins, new capsids, and envelopes are assembled
an assembled virus buds from the cell membrane
once virus is in chromosome, never comes out
HTLV I
human retrovirus
leukemia after long latency
HTLV II
human retrovirus
possible leukemia
HIV1
human retrovirus
AIDS
derived from recombined SIV from red capped mangabey and spot nosed monkey that subsequently infected chimpanzees
HIV2
human retrovirus
AIDS
derived from SIV from sooty mangabeys
viral species
non-retroviral ssRNA genomes can be either positive or negative strand depending on the viral species
influenza (-)
common cold (+)
polio (+)
SARS (+)
hep C (+)
rotavirus
double stranded RNA viruses
gemini and parvo
single stranded DNA viruses
T2, T4, phage, HPV
single stranded DNA viruses
positive
RNA _____ strand
RNA genome directly codes for protein
viral genome can be used right away for protein synthesis
negative
RNA _____ strand
RNA genome is complimentary to the protein coding strand
have to make copy then can code for protein
influenza
rapid changes occur through genetic recombination
three main types: A, B, and C
most cases are A: divided into subtypes based upon expression of hemagglutinin (HA) and neuraminidase (NA)
H1N1
spanish flu
influenza pandemic
1918
50,000,000 dead
H2N2
asian flu
influenza pandemic
1957
2,000,000 dead
H3N2
hong kong flu
influenza pandemic
1968
1,000,000
H1N1
swine flu
influenza pandemic
2009
280,000
antigenic drift
small genetic changes resulting from mutations during viral replication
slow change
antigenic shift
reassortment of genetic material from different viruses
rapid change and lots of change
low
RNA has ____ fidelity
makes more mistakes
bacteria and archaea genomes
genomes that has double stranded DNA
usually one main circular chromosome
can have small supplemental chromosomes (plasmids)
bacteria and archaea plasmids
circular with range of sizes (3kb-400kb)
optional- not present in all individuals
encode a few supplemental genes
often multiple copies per cell
replicate separately from main chromosome
optional under ideal conditions- take work by cell to be maintained so if not beneficial they are usually lost
main chromosome in bacteria and archaea
single, usually circular
defined sizes range from 0.2Mb- 8.7Mb
moderate number of genes: 800-5000
short genes without introns
very little non-gene sequence
packaging problem
if stretched out straight, the E coli genome of 4.6 × 106 would be 1000X the length of the cell
has to package genome
supercoiling
represents a way to conserve space in circular DNA
the ends must not be free to rotate
DNA in cells tends to be negatively supercoiled
accomplished by topoisomerase
topoisomerases
accomplishes supercoiling (create kinks in molecule so they take up less space)
cut one or both strands
prokaryotic main chromosome
negative supercoiling
most common supercoiling
easier to unwind strands during replication and transcription
supercoiled DNA uses less space than relaxed DNA
second level packaging
tight coiling based on proteins and RNAs
basic proteins bind to acidic DNA
short RNA “twist ties” hold loops
positive charged proteins to negative charged DNA
organelle genome
in eukaryotes (chloroplasts and mitochondria)
small circular double stranded chromosome
short genes (1000-5000bp) with few introns
few genes
very little non-gene sequence
multiple copies per cell, maternal inheritance
mitochondria and chloroplasts
contain DNA
encodes some polypeptides used by the organelle, rRNA, and some tRNAs
protein function stays in these 2 organelle
encoding is in chromosomal DNA
endosymbiotic theory
proposes that mitochondria and chloroplasts were once free-living bacteria
both organelles are similar to eubacteria, and DNA sequences found within them are also similar to eubacteria
present day cell
approximately 1 billion to 1.5 billion years ago, an anaerobic eukaryotic cell engulfed an aerobic eubacterial call through endocytosis
the aerobic endosymbiont evolved into mitochondria
likewise, endocytosis of a photosynthesizing eubacterium
led to the evolution of modern eukaryotic cells with mitochondria and chloroplasts
evidence of endosymbiotic theory
mitochondria and chloroplasts have double membrane
mitochondria and chloroplasts have their own DNA with a circular genome
mitochondria and chloroplasts have their own ribosomes which exhibit sensitivity to antibiotics directed at prokaryotic protein synthesis
currently, several single cell eukaryotes harbor symbiotic bacteria
mitochondria and chloroplasts DNA sequences are more closely related to eubacterial genes than eukaryotic genes
nuclear genome
part of eukaryotic chromosome:
has multiple linear double stranded DNA chromosomes
total size extremely variable
many genes that are spaced apart
elaborate condensation around proteins
usually by histone proteins
size
____ of eukaryotic chromosome
variable within organism
variable between closely related species
centromere
______ of eukaryotic chromosome
special sequences in DNA bind special proteins
essential for segregation of chromosomes in mitosis or meiosis
can be point or regional
no universal specific sequence requirement but tend to be very A-T rich
contains a special variant histone CenH3 that replaces H3
made up of heterochromatin
euchromatin
less condensed chromatin
located on chromosome arms
unique types of sequences
many genes present
replicated throughout the S phase
transcription occurs often
crossing over is common
heterochromatin
more condensed chromatin
located at centromeres, telomeres, and other specific places
repeated sequences
few genes present
replicated in late S phase
infrequent transcription
crossing over is uncommon
centromere
special sequences in DNA bind special kineticore proteins
point of attachment for spindle fibers
fibers bind to proteins, not DNA itself
position varies widely between chromosomes
telocentric, acrocentric, metacentric
without one there is no mechanism for a piece of DNA to be retained in the nucleus after cell division
acentric pieces of DNA are lost during cell division
telomeres
caps that stabilize ends of linear chromosomes
made up of heterochromatin
necessary to prevent degradation
special repetitive sequences
same sequence on every chromosome in genome
very similar between species
mammals: TTAGGG
tetrahymena: TTGGGG
G rich
shelterin
multisubunit protein complex that protect the ends of telomeres
banding patterns
characteristics of chromosomes used to identify
numbering system for human chromosomes
relates to chromosome composition
euchromatin
heterochromatin
euchromatin
light staining, decondenses, actively transcribed
heterochromatin
dark staining, stays condensed, not transcribed
constitutive heterochromatin
is always condensed
facultative heterochromatin
can convert to euchromatin depending on conditions