miRNA and Gene Regulation
1/20
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
21 Terms
Recall from before…
Central Dogma: DNA → transcription → RNA → translation → protein
there are many layers of transcription regulation (e.g. PIC assembly, elongation control, transcription factors, etc.)
but gene regulation also occurs after mRNA is transcribed; post-transcriptional regulation
alternative splicing, mRNA stability (5’cap/A’tail), miRNA
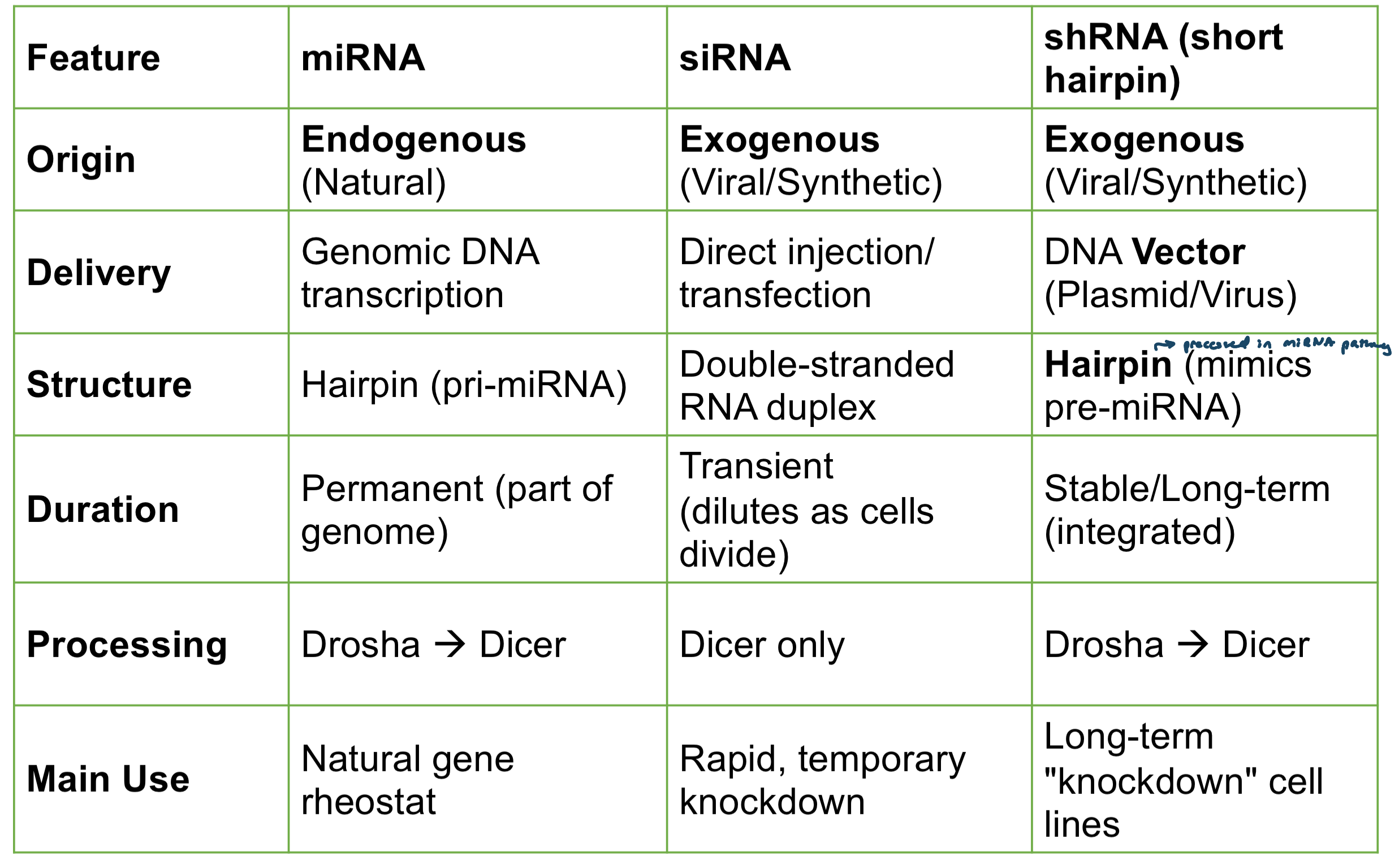
miRNA
short non-coding RNA molecules (~21-22 nts)
regulates mRNAs by RNA-RNA basepairing
results in translational repression and/or target dgradation
each miRNA can target one or several mRNAs
over 1/3 of protein-coding genes are regulated by miRNA
many key biological processes are controlled by miRNA, such as development, cell differentiation and and diseases (from misregulation)
over 1500 miRNA have been uncovered in humans
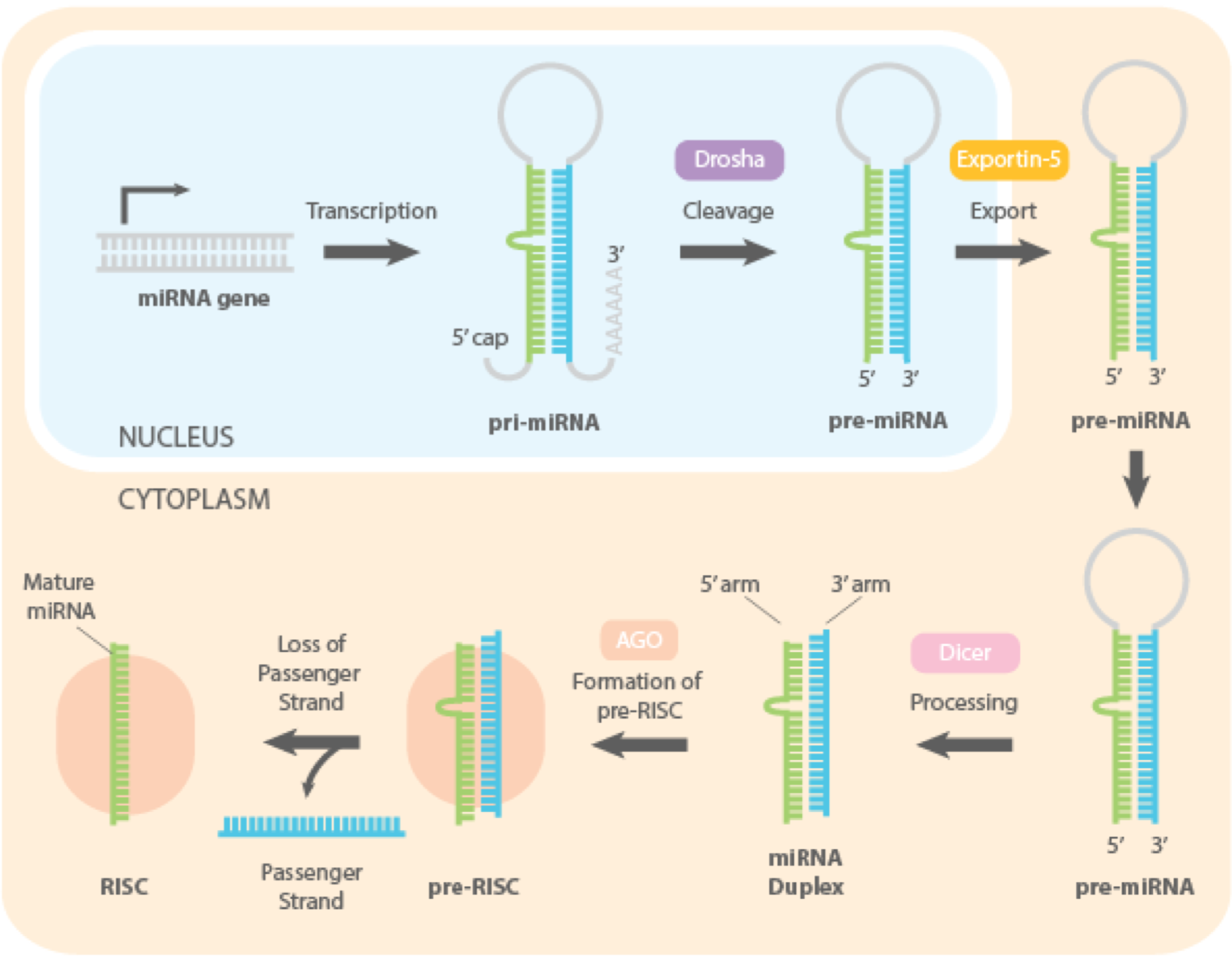
miRNA Processing: Step 1
Transcription
miRNAs are transcribed by RNA Pol II and therefore contain a 5’cap and 3’ poly(A) tail
a miRNA can be transcribed from its own gene or be part of an intron of another gene
following transcription, the pri-miRNA will form a hairpin with a loop
in some cases, multiple hairpins will be formed within one pri-miRNA
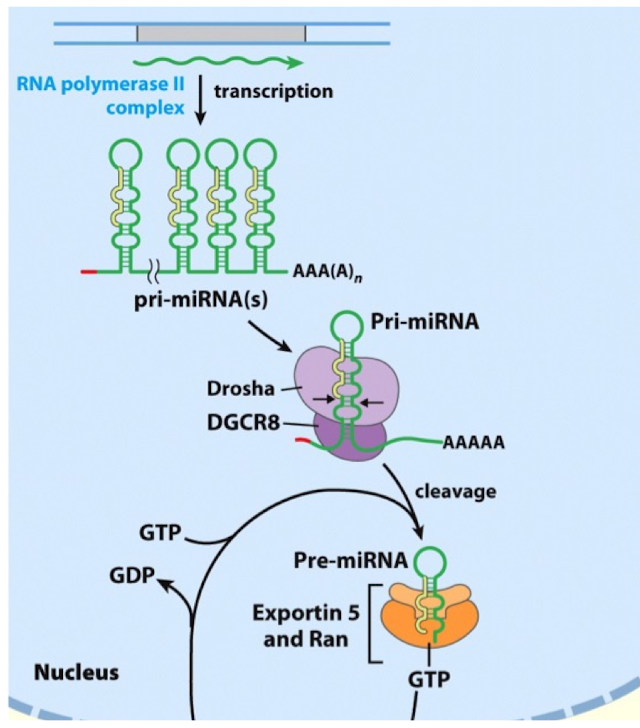
miRNA Processing: Step 2
Nuclear Processing
each hairpin is first processed by Drosha, an endoribonuclease in the nucleus that will introduce a first cleavage to form a ~70 mer pre-miRNA
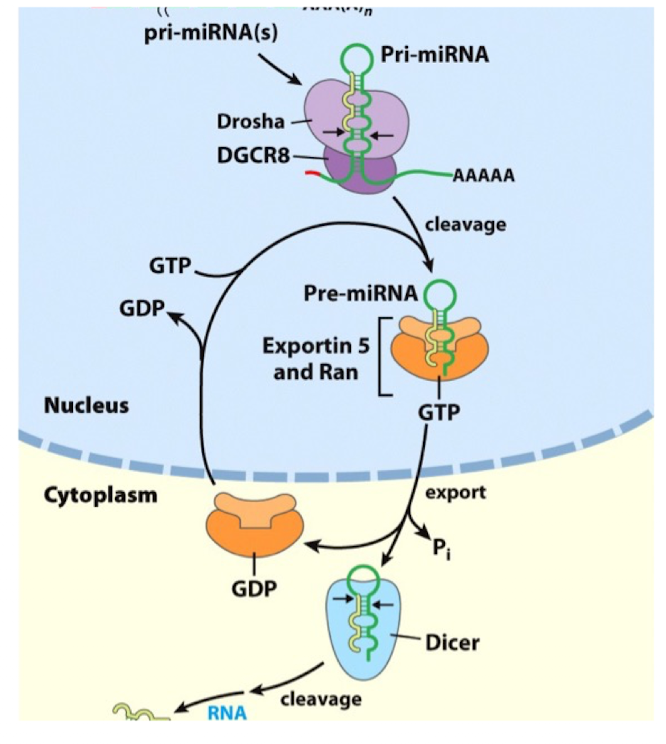
miRNA Processing: Step 3
Export
the nuclear export is mediated by Exportin-5 associated to Ran GTPase bound to GTP
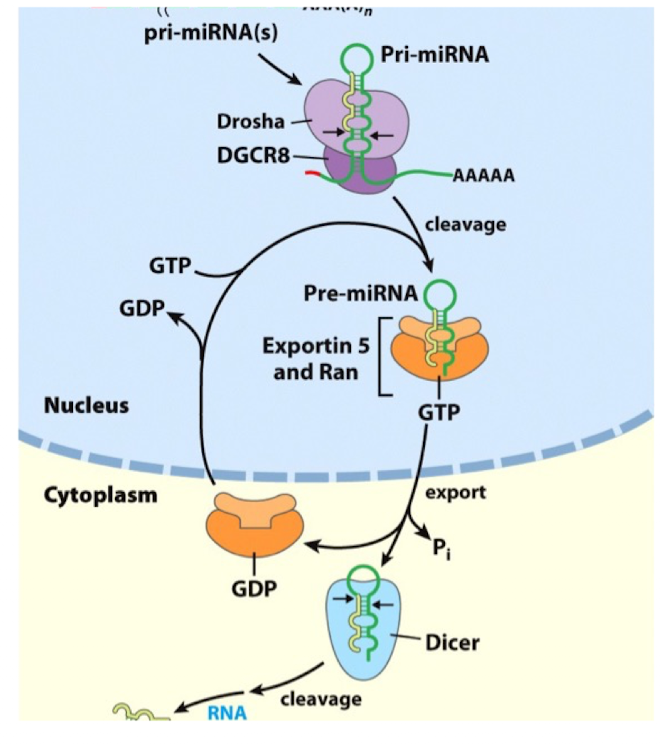
miRNA Processing: Step 4
Cytoplasmic Dicing
following export, the pre-miRNA is released to cytoplasm
the enzyme Dicer cleaves the loop of the hairpin, leaving a short double-stranded RNA duplex
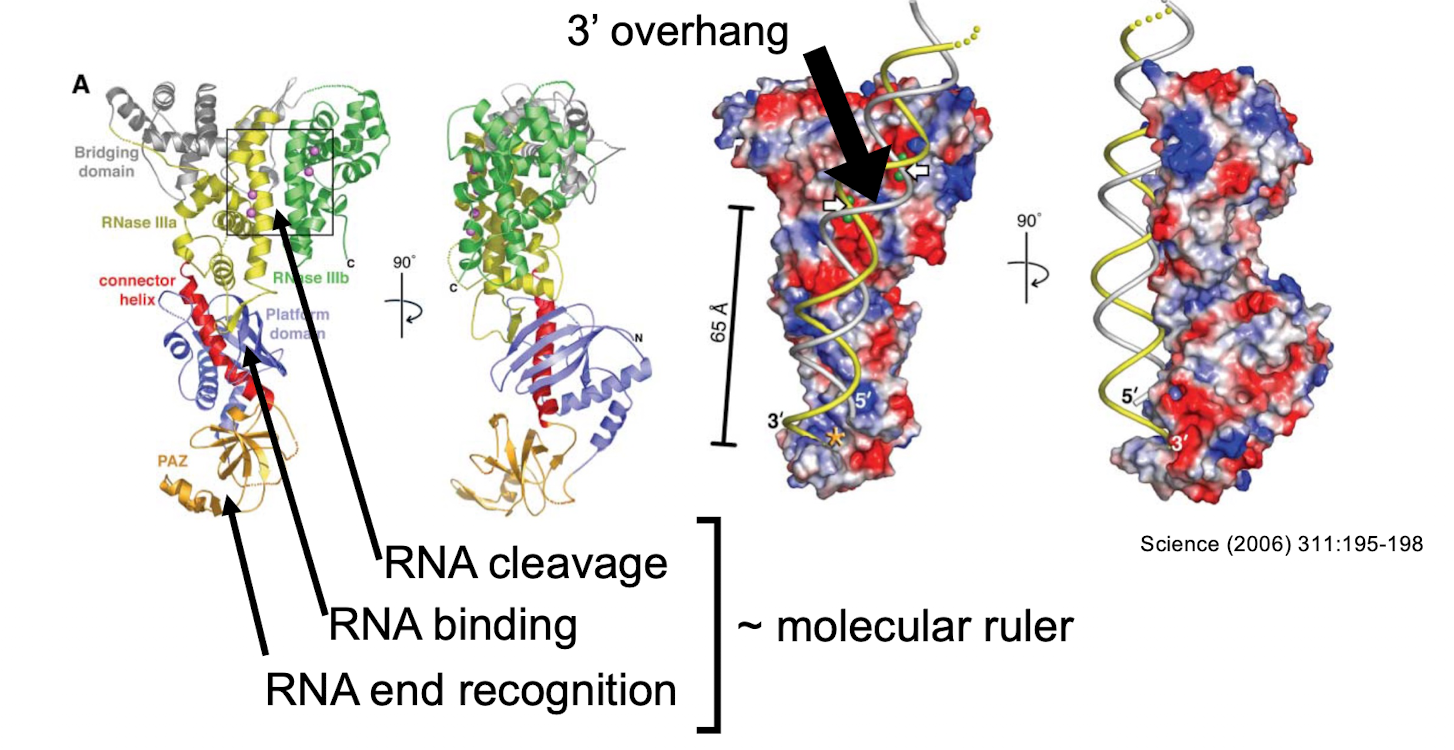
Dicer RNAse III
class III RNAse that specifically recognizes double stranded RNA molecules with a 3’ overhang
endonuclease active sites are placed 6.5nm from the RNA end recognition site (~21 nts long)
binding to the backbone ensures processing in a non specific manner
one enzyme can process multiple different pre-miRNA hairpins
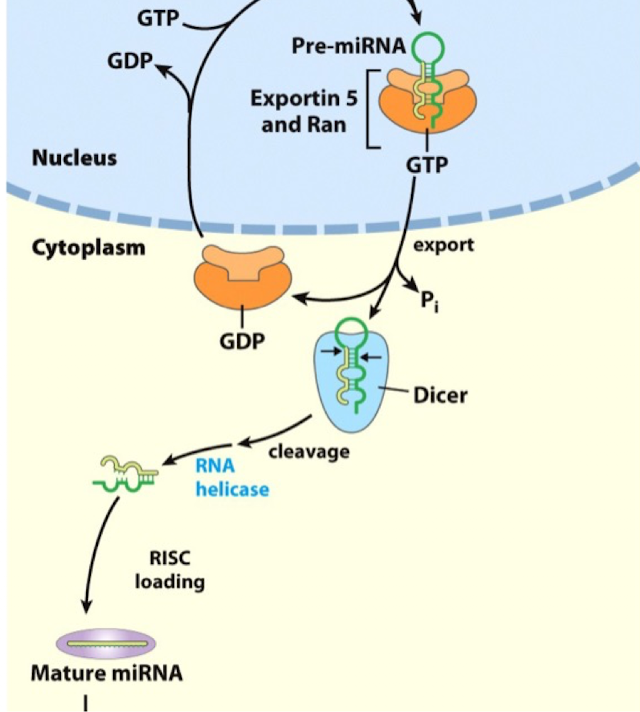
miRNA Targeting: Step 1
following the formation of the short 22 nts duplex, the passenger strand is evicted by a helicase
the complement strand is usually kept and loaded into the RISC complex
miRNA Targeting: Step 2
mature miRNA is loaded on RIS (RNA induced silencing complex; a multisubunit protein complex)
miRNA Targeting: Step 3
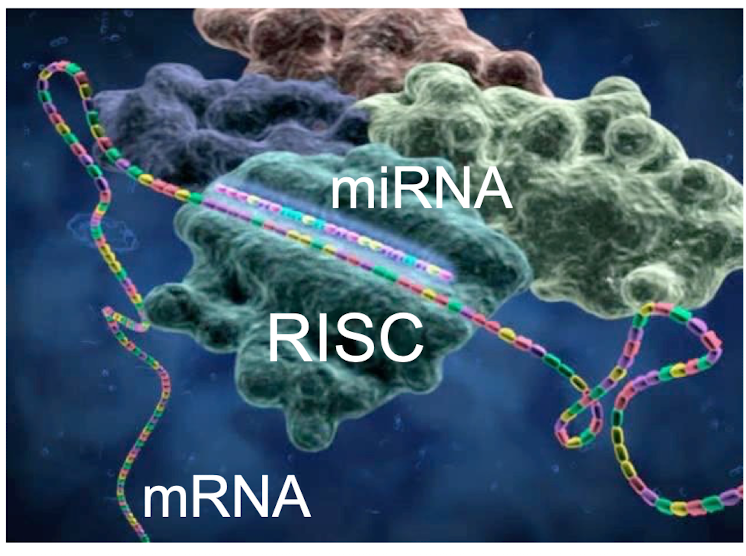
with the help of RISC, the miRNA will then hybridize with the targeted mRNA (often at 3’UTR)
a seed sequence (conserved 7-mer) in the 5’ region of the miRNA typically aligns with the complementary regions
if the sequence has partial complementary with mRNA, the mRNA will remain stable but translation will be inhibited (in most cases)
if the sequence has near perfect complementarity, the miRNA will induce mRNA cleavage degradation
miRNA Targeting Figure
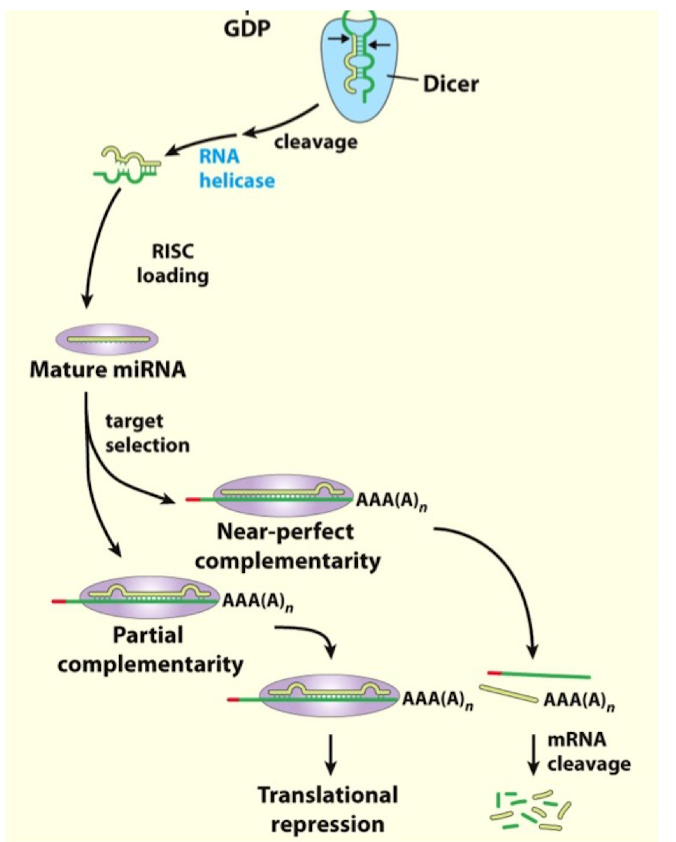
How does miRNA Regulate mRNA Translation
can repress translation at the initiation stage by blocking the cap recognition site or stage of recruiting 60s ribosome subunit
can block mRNA circularization
ribosome dropoff
degradation by cap removal and/or tail removal
30% of human proteins is regulated by miRNA
RNAi technology
scientists can co-opt miRNA tech to perform targeted knock down of specific mRNAs using the base-pairing strategies (RNA interference)
RNAi Technology Steps
protein of interest
design or order siRNA/shRNA
targets mRNA
siRNA is already double-stranded; shRNA are incorporated into plasmid to be delivered into the cell
shRNA has to be transcribed and processed first in order to do its function, but is more stable for longer silencing
transfect cells
verify that the mRNA or protein of interest is downregulated
study protein of interest
Relationship between mRNA and Protein is not 1:1 DATA (Was hinted at to be on exam!)
Total RNA vs protein in human brain samples
y-axis: protein produced per amount of mRNA
There is a positive correlation (r = 0.56) → more mRNA generally = more protein
However, r ≠ 1, so mRNA levels do not perfectly predict protein levels
Interpretation of points:
On the line of best fit: expected protein from mRNA level
Below the line: less protein than expected → post-transcriptional repression (reduced translation)
Above the line: more protein than expected → enhanced translation
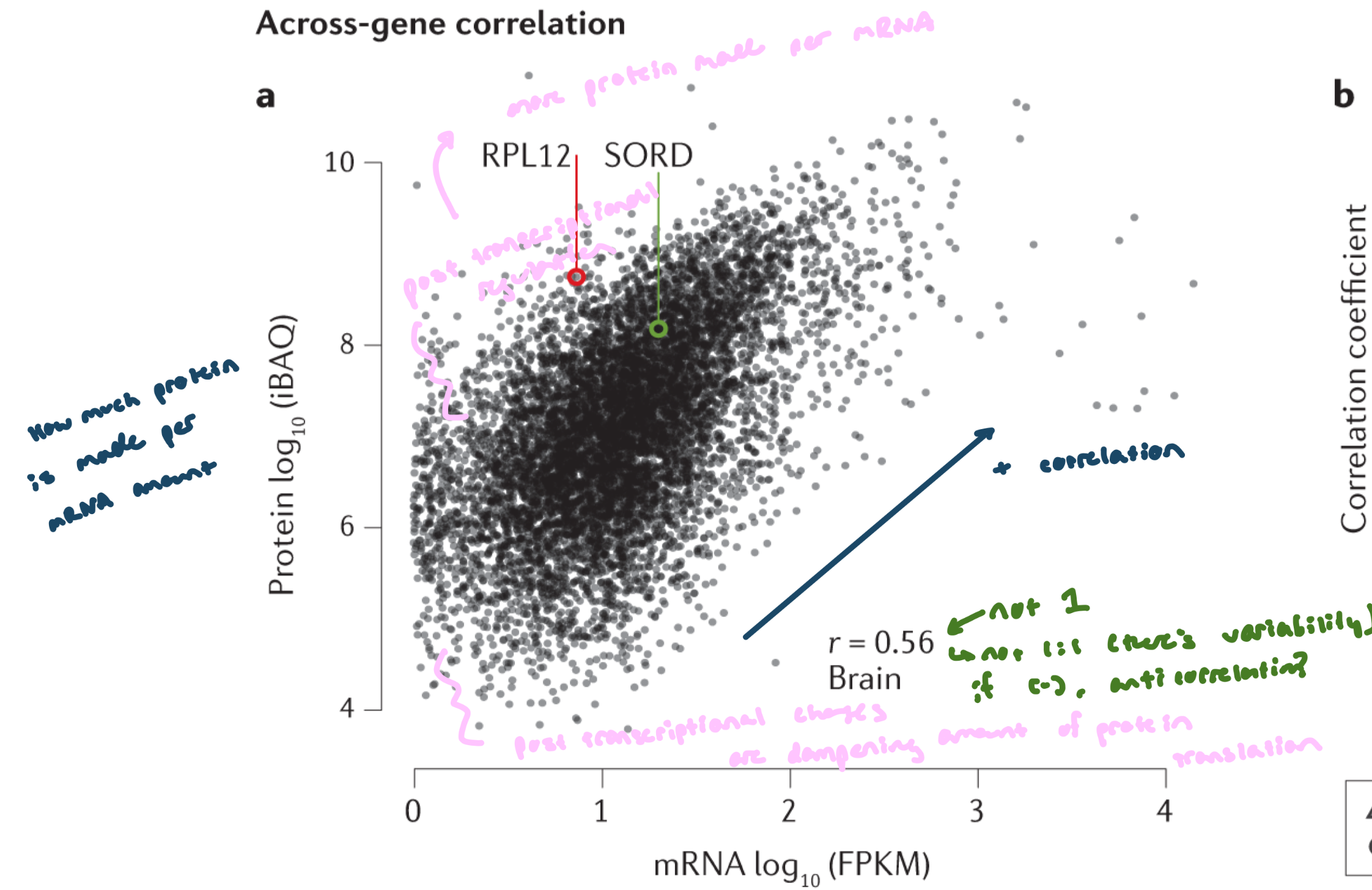
Relationship between mRNA and Protein is not 1:1 DATA
Correlation between RNA and Protein across diff studies in diff species
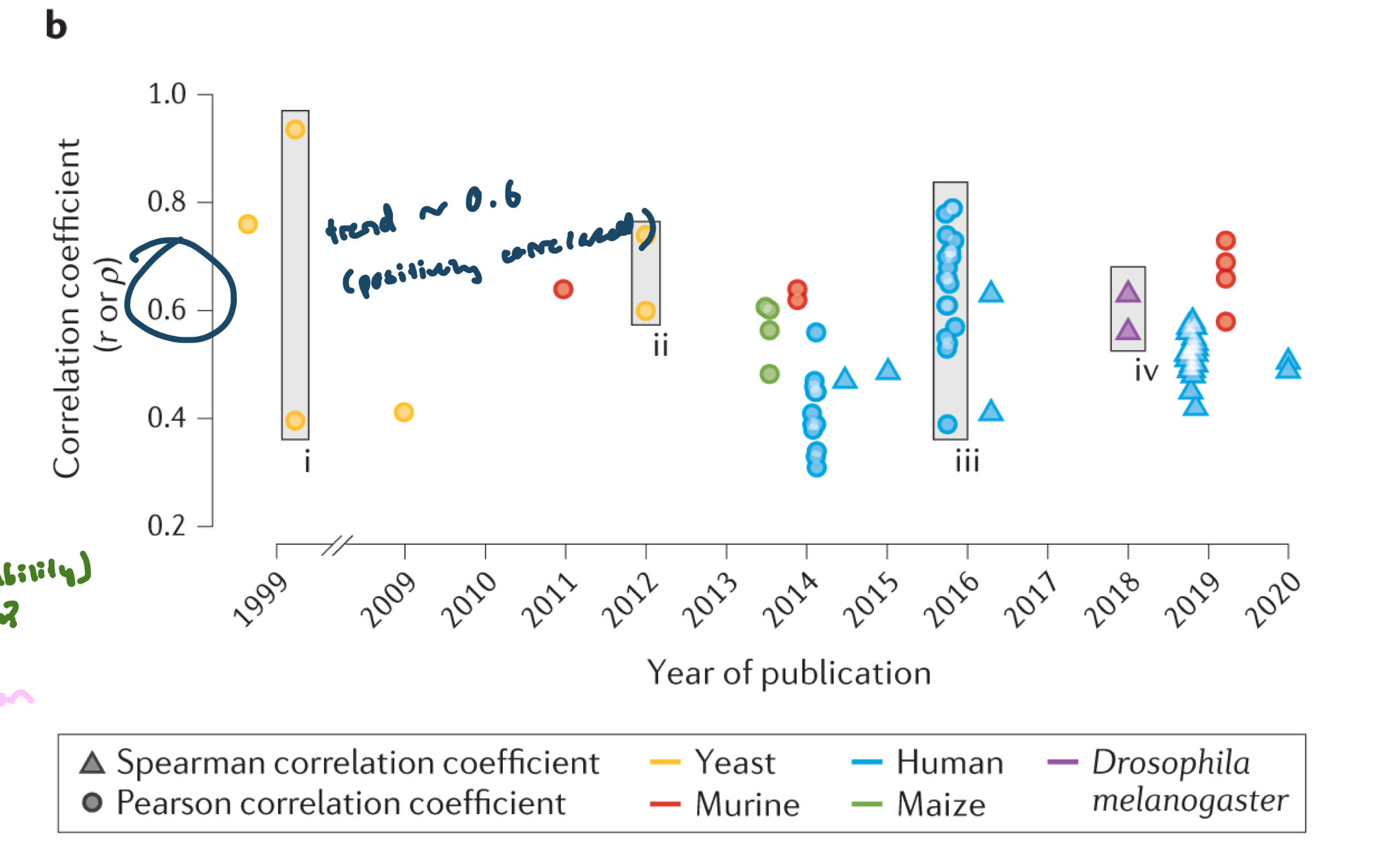
Relationship between mRNA and protein is gene specific
Each dot = one tissue (same gene across different tissues)
Correlation (R) shows how well mRNA levels predict protein levels
Key interpretations:
Low R value: weak mRNA–protein relationship; greater contribution from post-transcriptional regulation
High R value: strong mRNA–protein relationship; gene expression mainly controlled at the transcriptional level
Examples:
RPL12: low correlation → strong post-transcriptional regulation
SORD: positive/high correlation → mainly transcriptional control
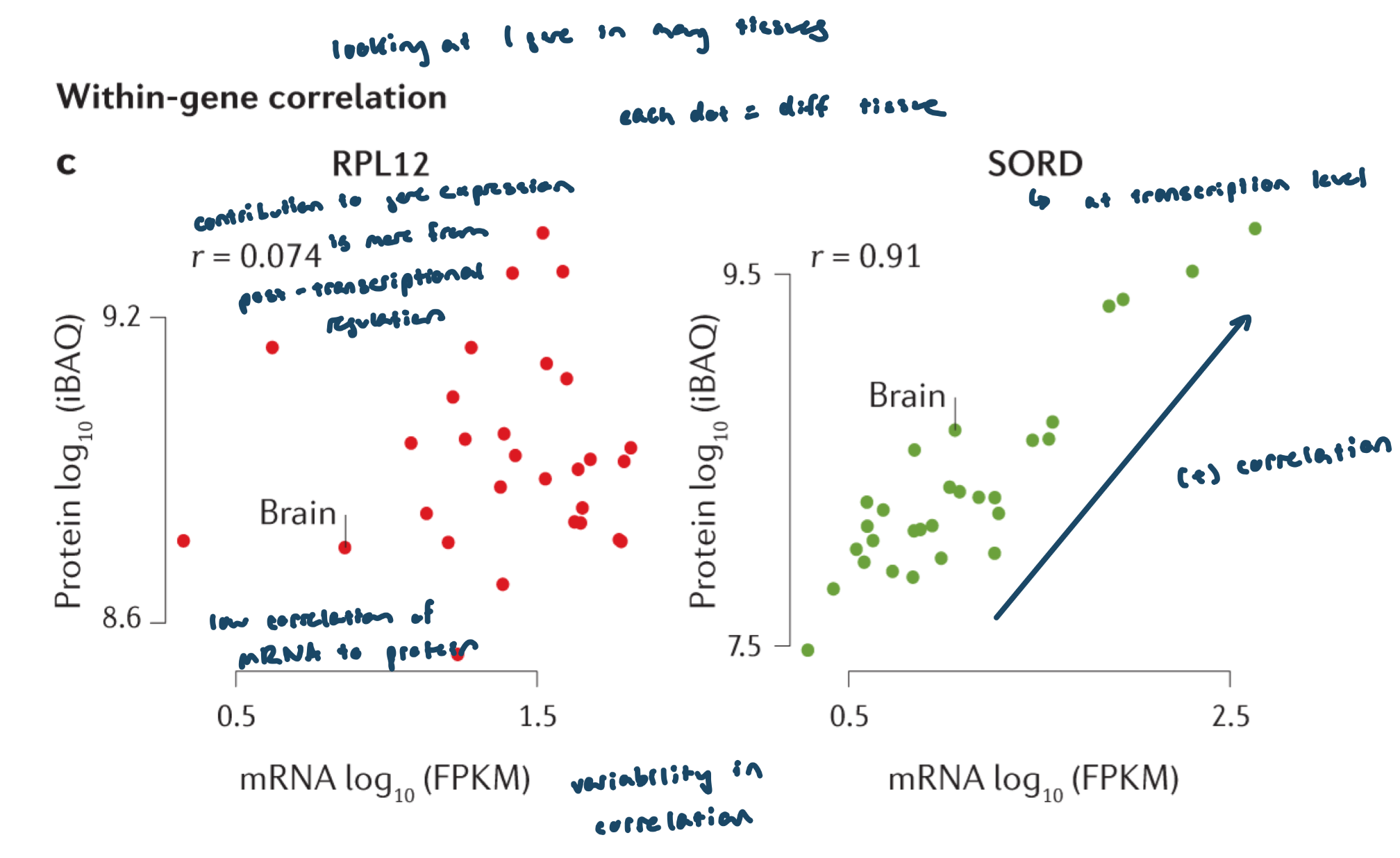
Relationship between mRNA and protein is gene specific
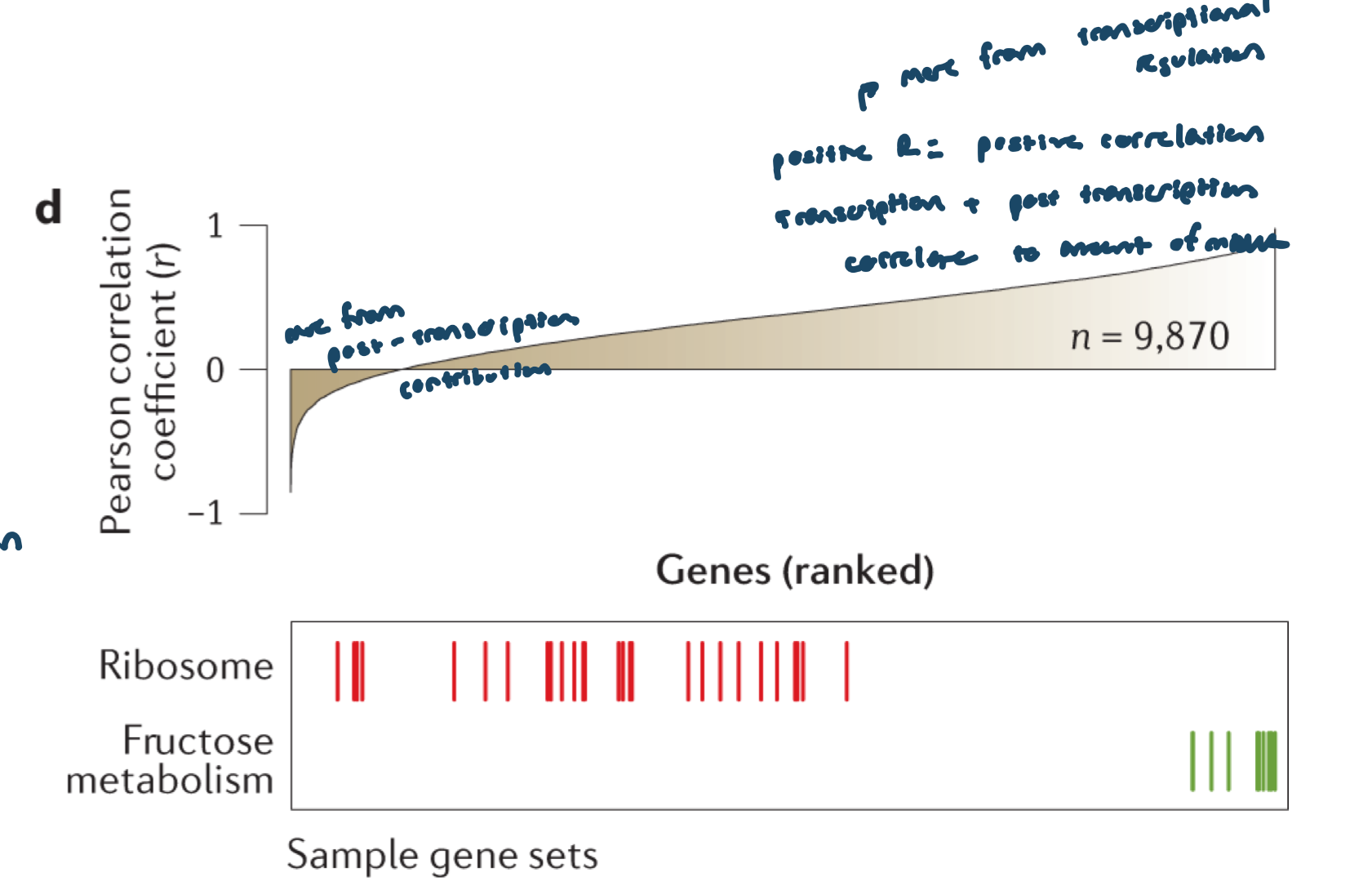
correlation for all genes across tissues; certain gene classes have higher RNA:protein correlation than others
miRNA Targeting: Thermodynamic assymmetry rule
the strand whose 5’end is less stable (more AU pairs, fewer GC pairs) is preferentially selected as the guide strand (easier to denature)
wy? the Ago protein RISC subunit anchors the 5’ phosphate of the guide strand
if the 5’ end of a strand is loose due to weak AU bonding, it is much easier for the Ago protein to grab that end and begin unwinding the duplex
miRNA vs siRNA vs shRNA