Genetics
1/30
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
31 Terms
Describe the chemistry involved in DNA polymerization and complementarity.
DNA polymerization.
DNA polymerase adds dNTPs to the 3’ -OH of the growing strand; synthesis occurs 5’ → 3’.
Complementarity.
Bases par via hydrogen bonds: A-T, G-C
Describe the packaging of DNA in the human nucleus.
See image.
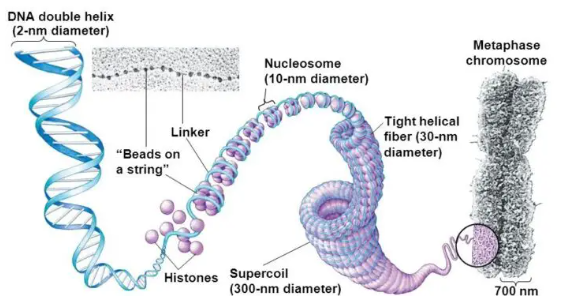
Describe the steps of DNA replication (initiation, elongation, proofreading).
Initiation:
Begins at origins of replication.
Helicase unwinds DNA → forms replication fork.
Single-strand binding proteins stabilize strands.
Primase synthesizes short RNA primers.
Elongation:
DNA polymerase adds nucleotides 5′ → 3′ from the primer.
Leading strand: continuous synthesis.
Lagging strand: discontinuous → Okazaki fragments.
DNA ligase joins fragments.
Proofreading:
DNA polymerase has 3′ → 5′ exonuclease activity.
Removes incorrectly paired nucleotides.
Ensures high replication fidelity.
Describe the phases of the cell cycle and the morphological characteristics that define each stage of cytokinesis/mitosis.
See image.
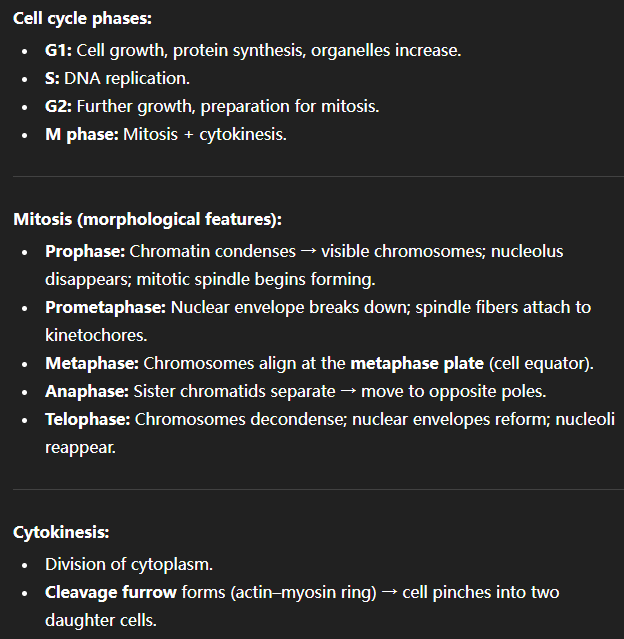
Explain the differences between heterochromatin and euchromatin
Heterochromatin: tightly packed, inactive.
Euchromatin: loosely packed, more accessible, allows access for RNA polymerase and transcription factors.
Describe the role of histones in DNA compaction.
Histones are the proteins which package DNA into chromosomes and play an important role in the structural organisation and regulation of DNA. Without histones, the unwound DNA in chromosomes would be very long.
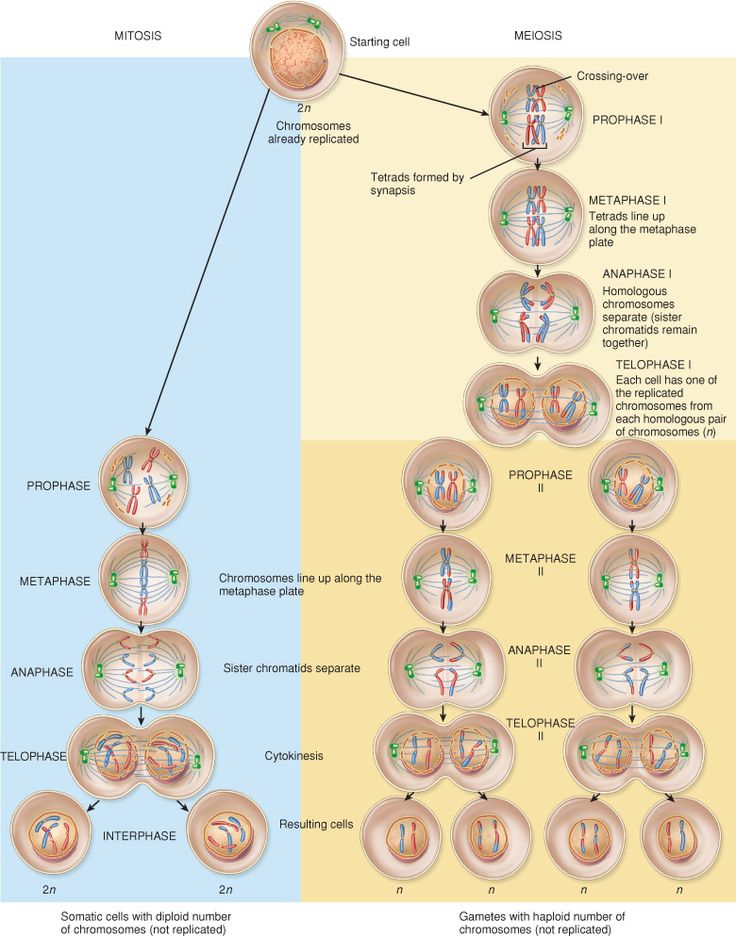
Compare and contrast the mechanism of mitosis and meiosis
Mitosis: repliaction division of normal (somatic cells) in their cell cycle.
Ends up with two daughter cells which are diploid cells
Meiosis: reduction division of sex cells (gametes)
Crossing over happens during prophase I, creates new combinations of alleles, ensuring genetic variation in gametes
Ends up with four daughter cells which are haploid cells
Describe gene expression and explain what effect this can have on a cells biology.
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product, and therefore dictates difference in cells.
Explain the terms phenotype and genotype.
Phenotype: the set of observable characteristics resulting from the interaction of its genotype with the environment; an organism’s observed properties.
Genotype: the part of the genetic makeup of a cell which determines one of its characteristics; an organism’s full hereditary information.
Understand and define the following terms: gene, allele, phenotype, genotype, locus.
Genes – Segments of DNA that code for proteins and determine traits (e.g., eye color, height).
Alleles – Different versions of a gene (e.g., the gene for eye colour has alleles for blue, brown, or green eyes).
Genotype = Your genetic makeup (the DNA you inherit).
Phenotype = Your physical traits (how your genes are expressed).
Locus: the specific, fixed physical location or "address" of a gene or DNA sequence on a chromosome.
Describe the following patterns of inheritance: recessive, dominant, codominant, X-linked recessive.
See notion page.
Calculate frequency of genotypes and phenotypes using Punnet squares
whatever…
Interpret family trees and draw conclusions regarding possible patterns of inheritance.
whatever…
Distinguish between simple/monogenic and complex disorders/traits.
Simple Mendelian inherited diseases
− One gene = one protein = (largely) one phenotype
− Ratios are easily defined within a given population - Dominant:Recessive (3:1)
− Easily able to identify family history and potential carriers and phenotype
Complex disorders
− Multiple interacting genes - more than one gene must be inherited to present with symptoms
− Caused by a combination of factors: e.g. genetic, environmental, lifestyle factors
E.g. cancer, Alzheimer's, asthma, MS, autoimmune diseases
− Lacks a clear-cut pattern of inheritance
− Can cause predisposition, but does not guarantee presentation.
The more complicated the mode of inheritance, the more genes, the more environmental component(s); the more difficult it is to identify the gene(s), provide diagnostic certainty & design/develop therapies to match.
Describe key RNA processing steps involved in translation.
Transcription: synthesis of RNA using DNA as a template. Occurs in the nucleus and is performed by a molecular complex that contains the enzyme RNA polymerase.
5’ GTP capping: addition of a G-P-P-P to protect the RNA from degradation and assist in transport across the nuclear membrane. Protects from 5’ exonucleases.
3’ Polyadenylation (Poly-A tail): Pre-mRNA tail is removed and addition of a 200-250 adenosine molecules are added to the 3’ end of the mRNA. Provides stability of the mRNA. Protects from 3’ exonucleases.
Splicing: Removal of introns and joining of exons by the spliceosome to create a single mature mRNA.
Alternate splicing: Pre-mRNA can be spliced into multiple different isoforms, leading to the creation of different proteins with distinct biological functions.
RNA nuclear export: after post transcriptional modification the mature mRNA is transported from the nucleus to the cytoplasm.
Describe the function on ribosomes in translation.
Ribosomes are the site of protein synthesis, occurs in the cytoplasm.
mRNA passes through the ribosomes and each codon is matched up with a transfer RNA (tRNA) carrying an amino acid.
tRNA is removed and the amino acids are joined together into a protein one at the time.
Translation continues until a stop codon is encountered.
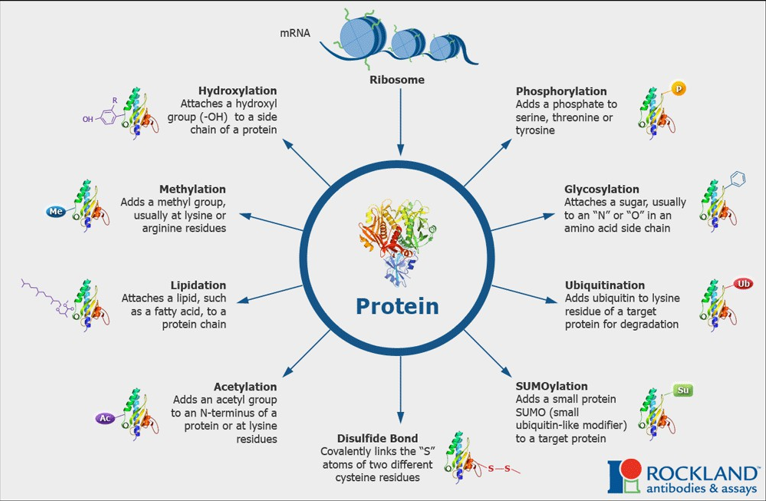
Provide examples of post-translational modifications of proteins.
Occurs in the endoplasmic reticulum and Golgi apparatus.
Hydroxylation: attaches a hydroxyl (-OH) to a side chain of a protein.
Methylation: adds a methyl group usually at lysine or arginine residues.
Lipidation: adds a lipid, such as a fatty acid, to a protein chain.
Acetylation: adds an acetyl group to an N-terminus of a protein or at lysine residues.
Disulfide bond: covalently links the “S” atoms of two different cysteine residues.
SUMOylation: adds a small protein SUMO (small ubiquitin-like modifer) to a target protein.
Ubiquitination: adds ubiquitin to lysine residue of a target protein for degradation.
Glycosylation: attaches a sugar, usually to an “N” or “O” in an amino acid side chain.
Phosphorylation: adds a phosphate to serine, theronine or tyrosine.
Give examples of technologies used to measure gene expression
Polymerase chain reaction (PCR): amply a specific segment of DNA between two regions of known sequence.
RNA sequencing: isolate RNA → fragment RNA into short segments → convert RNA fragments into cDNA → ligate sequencing adapters and amplify → perform NGS sequencing → map sequencing reads to the transcriptome/genome.
Single cell RNA sequencing: tissue dissociation leads to single cell suspension → single cell RNA sequencing → bioinformatics analysis
Spatial transcriptomics: maps gene expression (mRNA) directly within intact tissue sections, preserving the spatial, architectural, and cellular context
What are the implications of population genetics in disease?
Population genetics: study of how a population changes over time leading to a species evolving/mutating (mixture of Evolution (Darwin) and Hereditary (Mendel) models)
− Founded by Hardy and Weinberg
− Study of the frequency and interaction of alleles and genes in a population
− Medical genetics uses the principle of population genetics to identify genetic susceptibilities to common diseases and potential treatments
What are the factors that influence genetic variation and allelic frequency?
Factors influencing allele frequencies
1. Natural selection: alleles for more fitter organisms become selected
2. Sexual selection: alleles for more sexually attractive organisms become more frequent
3. Mutations: new alleles appear due to mistakes in DNA
4. Genetic drift: changes in allele frequency due to random chance
5. Gene flow: changes in allele frequency due to mixing of new populations
Outline the Hardy Weinberg equation.
Hardy-Weinberg Equilibrium
− Relationship between frequency of alleles at a genetic locus and the genotypes resulting from those alleles
− Used to predict gene frequencies with a given population
− Equation: p2 + 2pq + q2 = 1 or p + q = 1
p = dominant allele A
p2 = frequency of AA (dominant homozygous)
q = recessive allele a
q2 = frequency of aa (recessive homozygous)
2pq = frequency of Aa
− Limitations • Works under assumptions
The population is large and matings are random with respect to the locus in question (no-inbreeding or small populations)
Individuals of all genotypes must be able to reproduce (no genotype selection)
No random mutations (Dominant (A) becomes recessive (a) spontaneously)
Must be a dominant or recessive trait, Mendelian inheritance must be maintained
No significant immigration of individuals with allele frequencies very different from the endogenous population
Describe the deviations from the Hardy-Weinberg equilibrium.
Deviations from H-W Equilibrium
Allele frequencies for inherited traits will remain stable unless populations undergo selection, migration, mutation or assortative (non- random) mating.
Deviations from the Hardy-Weinberg equilibrium give us an indication of the reason for a genetic evolution (change) within a given population.
Explain and provide an example of selective advantage and negative selection.
Negative selection - alleles lost by selection are replaced by recurrent new mutations in order to maintain the observed disease incidence
− In an infinitely large population, a recessive allele (a) will approach 0 but never reach it
− E.g. Duchenne's Muscular Dystrophy
Recessive x-linked
Mutation in dystrophin gene - breakdown of muscle cells, mutation is lethal, unable to reproduce
Largest protein on x chromosome - vulnerable to mutation
‘Spontaneous mutations’ occur in one-third of DMD cases
This is true for all x-linked diseases, where the genetic phenotype is lethal
Selection advantage - particular trait becomes advantageous to reproduction (adaptive trait) as it provides resistance to other
− Balanced polymorphism/Heterozygous advantage
Selective pressure for Ss genotype (resistance + no development of sickle cell), as both SS and ss are subjected to selection by malaria and anaemia respectively
− E.g. Sickle cell anaemia
Autosomal recessive
Causes misfolding of haemoglobin - sickle cells cannot pass through small blood vessels, transport of oxygen is massively reduced
Malaria parasite cannot reproduce in sickle cells - resistance to this protozoan infection
Describe the inheritance of mitochondrial genome
Only maternal mitochondrial inheritance occurs.
Most mitochondria is in the tail of the sperm. After fertilisation the tail of the sperm is directed towards degradation. Remaining sperm mitochondria are lost by chance due to exponential cell division during embryonic development. Highly debated if any mtDNA is transferred to offspring from the father. Incidences of paternal leakage of mtDNA (Schwatz and Vissing (2002) – muscle tissue).
Describe the role of mitochondrial genes and elaborate on why mitochondrial mutations are mostly associated with reduction in energy demands
Role of Mitochondrial Genes
The human mitochondrial genome (mtDNA) is small, typically consisting of 16,569 base pairs that encode 37 essential genes. These genes are indispensable for mitochondrial function:
Protein Coding (13 genes): These provide instructions for making subunits of the enzyme complexes (I, III, IV, and V) involved in oxidative phosphorylation (OXPHOS).
Translation Machinery (24 genes): These encode 22 transfer RNAs (tRNAs) and 2 ribosomal RNAs (rRNAs) necessary for assembling mitochondrial proteins within the mitochondrion itself.
These genes are essential for converting energy from food into usable ATP. The 13 encoded proteins are part of the respiratory chain, which pumps protons across the inner mitochondrial membrane, establishing a gradient that drives ATP synthase.
Why Mutations Reduce Energy Production
Mitochondrial mutations cause a decline in energy output through several key mechanisms, often leading to a "bioenergetic deficit":
Disruption of Oxidative Phosphorylation (OXPHOS): Mutations in mtDNA alter the proteins that constitute the respiratory complexes (I, III, IV, V), making them non-functional. For example, a mutation in the MT-ATP6 gene, which is essential for the ATP synthase complex (complex V), reduces the mitochondria's ability to convert ADP into ATP.
Oxidative phosphorylation is the final, high-yield stage of cellular respiration in eukaryotes, occurring in the inner mitochondrial membrane. It generates ATP by transferring electrons from NADH and through an electron transport chain (ETC) to oxygen, creating an electrochemical proton gradient that drives ATP synthase. This process generates approximately 90% of cellular ATP.
Impaired Protein Synthesis: Mutations in tRNA or rRNA genes slow down or disrupt the synthesis of mitochondrial proteins, weakening the entire ETC.
Elevated Reactive Oxygen Species (ROS): Mutant mitochondria often produce higher levels of harmful molecules called ROS. These molecules can damage the remaining mitochondrial DNA and proteins, creating a vicious cycle of further degradation and diminished energy output.
Heteroplasmy and Threshold Effect: Cells contain many mitochondria, and a "mixture" of mutated and normal DNA (heteroplasmy) can exist. If the percentage of mutant DNA exceeds a specific threshold (often 60–90%), the cell can no longer compensate, resulting in a severe decrease in energy production.
Degradation of Mitochondria: Dysfunctional mitochondria are sometimes removed via autophagy (mitophagy), reducing the total cellular capacity for ATP production.
Describe mitochondrial inheritance and its role in mitochondrial disease.
Mitochondrial inheritance involves inheritance via the mtDNA and the mitochondrial genome which codes for proteins.
− Mitochondrial inheritance passes through maternal lines
− Mitochondrial genes are mostly associated with energy production (OxPhos)
− Mitochondrial mutations are often associated with the central nervous system due to the large energy demands
Reactive oxygen species (ROS) ▪ Number of adverse effects - bind lipids, lipid peroxidation (breakdown of cell walls, cell death), mix in DNA (increase mutation rates) ▪ Role to move proteins into intermembrane space (I-IV) ▪ Increase of ROS - build-up of ROS through donation of protons, superoxides - can interact with biology in number of ways (mostly damage DNA) ▪ Number of reactive molecules and free radicals derives from molecular oxygen
Biochemical process in close proximity to genetic material - mitochondrial genome more susceptible ▪ Theory of ageing - more DNA mutations, RNA doing mutation is error-prone • Mutations combatted by having multiple copies of genome
Comparison to nuclear DNA
− 0.0005% of genetic information
− 1 small circular DNA
− Multiple copies
− 2-10 genomes per mitochondria (multiple mt per cell; 100-10000 genomes per cell)
− 93% coding region - no room for error
Explain the terms penetrance, variable expressivity and heteroplasmy
Penetrance: the probability that a given phenotype will appear given that the phenotype related genotype is present. − Penetrance (P) = (Phenotype/Genotype) * 100
Variable expressivity: the variations in a phenotype among individuals carrying a particular genotype (the range or intensity of the trait in question).
Heteroplasmy: the mixed lineages of mtDNA which can cause highly variable expression of a disease (dependent on the defective copy number).
Describe the structure and number of chromosomes in a human cell.
Every cell normally contains 23 pairs of chromosomes, for a total of 46. 22 pairs are called autosomes and look the same in both males and females. The 23rd pair, the sex chromosomes, differ between males XY and females XX. In nucleus of each cell, the DNA molecule is packaged into thread-like structures called chromosomes. Each chromosome is made up of DNA tightly coiled many times around proteins called histones that support its structure.
Diploid: cell that contains 2 sets of homologous chromosomes
Haploid: cell that contains on a single set of genes
Explain the terms aneuploidy, trisomy and euploidy.
Large-scale mutation resulting in abnormal number of chromosomes, or structurally abnormal chromosomes – arisen from errors during gamete formation.
Numerical abnormality:
Aneuploidy: when gametes are forming, can end up with additional or missing chromosomes. - The chromosomes altered give rise to genetic disorders:
- Turner’s syndrome: only 1 X chromosome (so only X instead of XX or XY; called monosomy)
short stature, amenorrhea, lack of secondary sexual development
always female
- Down syndrome: extra chromosome 21, so 3 in total (XXX or trisomy); hypotonia, characteristic facial features, developmental delay
- Patau syndrome: trisomy 13; cleft lip + palate, severe CNS anomaly, polydactyly
- Klinefelter Syndrome (47, XXY); extra X chromosome o Males only; hypogonadism and reduced fertility, increased stature
Trisomy 21 – down syndrome
- (90-95%) A duplication of chromosome 21 (either from sperm or egg)
- Mosaic trisomy 21 (2-3%): not all cells contain duplication of chr.21
- Translocation trisomy (4-5%): part of chr.21 translocates to another chromosome (usually 13,14,15, 22)
- Considered the only viable autosomal trisomy
Because the number of protein-coding sequences for chr21 is the smallest of any human chromosome (~220 genes) [except Y-chromosome]
Euploidy: State of having a cell or organism with an integral multiple of the monoploid number (e.g. in humans, n = 23 (monoploid), 2n = 46 (diploid) are euploidy)
Describe the mechanism of chromosomal translocation, deletions, duplications, inversions and insertions, and provide a clinical example where chromosomal rearrangement occurs.
see learning obj…
Elaborate on how a chromosomal rearrangement can lead to a normal individual (increasing genetic diversity) or cause a disease/disorder.
Depending on the rearrangement type, location & amount affects the presentation of that mutation (some don’t present symptomology, some present mild, some present severe). Mutations that occur (de novo or otherwise) can actually gain function (duplication) when expanded, and therefore contribute to evolutionary mutations/changes. However other instances (such as deletion) can result in the loss of function of that protein (disease/disorders arise).
Everyone has chromosomal abnormalities.