Global gene regulation and expression in bacteria
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
Bacterial gene
1000 bases
Absence of introns
95% of the sequence codes for proteins
Grouped in operons
Operons
Segment of DNA containing genes that operate under the signal of the same promoter. One mRNA → several proteins
SIgma factors
Interact with an RNA polymerase (RNAP) core enzyme to forme the RNAP holoenzyme and direct the complex to promoter regions
Facilitates the opening of the double strand DNA
Start of the transcription
Dissociation and release of the sigma factor
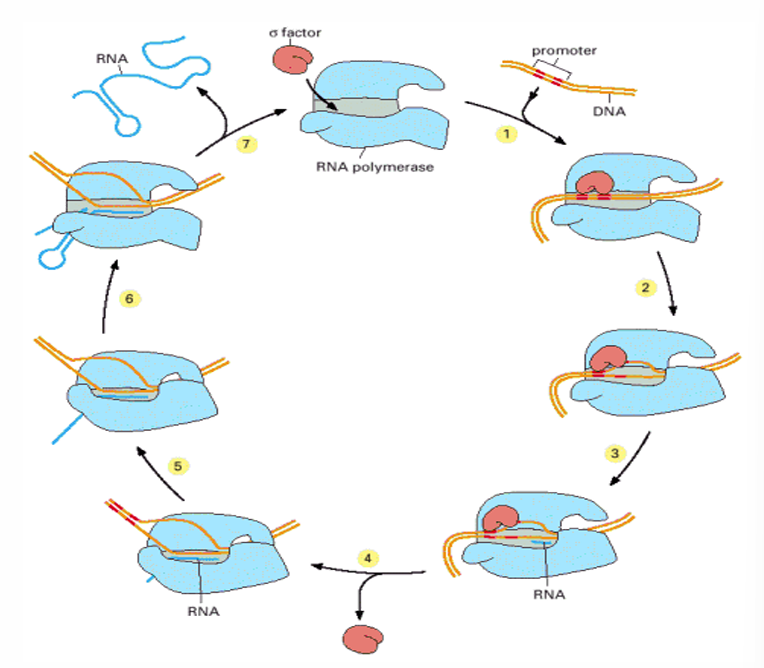
Initiation
Plasmid recognition
Elongation
Addition of nucleotides to the new DNA strand
Termination
Transcript sequences that encode and RNA hairpin and terminal uridine-rich segment
Termination by Rho enzyme (ATP dependent RNA translocase) → releases RNA by forcing uncharacterized structural changes in elongation complex
Activity of DNA translocase Mfd and ATP
Constitutive genes
Genes continuously expressed in the cell
Regulated genes
Only expressed under certain physiological conditions or in response to stimuli
Regulation by alternative sigma factors
Activation of transcription genes under stress conditions
Recruit core enzyme after binding to specific sequences different than the sigma factor (sigH)
Regulation by transcription factors
Proteins that bind to the DNA in cis of the promoter or to the polymerase
Regulators Helice-Turn-Helice (HTH)
Bind to DNA via alpha helices
2 segments of 20 amino acids separated by 4 residues
In dimers
Regulators ribbon helice helice (RHH)
Antiparallel β-strands (ribbons) framed within an α helical scaffold
Use the anti-parallel β-sheet to recognize specific nucleotide sequences and α-helices to anchor the β-sheet in the DNA major groove
Positive regulation
Direct contact with the C-terminal domain of the α subunit
Negative regulation
Repression by steric encumbrance
Repression by a loop in DNA
Lactose operon
The lactose operon is a set of genes in bacteria responsible for the uptake and metabolism of lactose. It consists of structural genes like lacZ, lacY, and lacA, which encode specific proteins involved in lactose utilization
Riboswitch regulations
Regulatory segment of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the proteins encoded by the mRNA. No regulatory protein is needed to mediate the interaction
Regulations by Hfq proteins
Hfq is an RNA-binding protein
Facilitates the short and imperfect base-pairing interactions of regulatory small RNAs (sRNAs) with trans-encoded target mRNAs
Permit or suppress the protein synthesis
Post-translational regulations
Control the activity, the localisation, the stability and the function of proteins after their synthesis
Proteolytic cleavage: inactive precursors that must be cleaved to be active
Adding a chemical group: phosphorylation, acetylation, methylation
Formation of a disulfide bridge
Transcription factors in two-component systems
Used to perceive and adapt to environmental changes (osmolarity, chemotaxis, nutritional deficiency, antibiotic resistance, and virulence)
Histidine kinase: sensor, responsible for detecting external stimuli and activate by phosphorylation HTH
Regulatory response protein (HTH), which binds to the promoter regions of target genes to induce their expression